Abstract
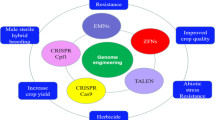
This chapter considers the genome editing technologies that have been utilized for breeding horticultural plants. Many examples of the successful application of genome editing technologies including ZFN, TALEN, and especially CRISPR/Cas systems in improving diverse characteristics of horticultural plants are mentioned and discussed. Based on the literature review, CRISPR/Cas technology has proved its potential in altering many genes of interest in horticultural plants including fruits, vegetables, and ornamental plants for improving agronomically important traits and attributes such as growth rate, seed size, flowering time, flower color, storage time, resistance to biotic stresses, tolerance to abiotic stresses, herbicide tolerance, metabolism, fruit color, fruit ripening, and so forth. This advanced technology paves the way for more favorable and precise manipulation of plant genomes to improve crop performance.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
1 Introduction
Horticulture goes back to ancient times as an important sector within agriculture and has improved a lot during human civilization. Horticultural crops are typically known as vegetables, fruits, and ornamental plants which are planted and harvested for food, medical, and cosmetic purposes [1]. Nevertheless, horticultural crop cultivation continues to face a variety of challenges, such as serious environmental concerns, the spread of viruses and pests, climate change, and increasing population growth [2]. Conventional breeding, molecular markers, and genetic modification have been used for improving traits in horticultural plants. However, faster development of better varieties requires more precise and safe techniques such as genome editing systems.
2 Non-CRISPR/Cas Genome Editing Systems and Their Applications in Horticultural Plants
Before the discovery of CRISPR/Cas, other genome editing techniques were used to modify traits in plants, but not so frequently in horticultural crops. Strictly speaking, there is no report on the application of meganucleases [3], the first developed genome editing tool, in horticultural crops. Later, after the rise of Zinc Finger Nuclease (ZFN) technology, two groups applied ZFN in the functional analysis and characterization of genes in apple and fig [4] as well as tomato [5] plants. Thereafter, TALEN-based gene editing technology was developed and effectively employed in some economically important horticultural crops such as rapeseed (Brassica oleracea) [6], potato (Solanum tuberosum) [7,8,9,10,11,12] and tomato (Solanum lycopersicum) [13, 14]. The first instance of genome editing in a horticulture crop was accomplished in 2013 using TALEN in Brassica oleracea [6]. Later, in potato, TALEN was used for silencing the vacuolar invertase gene to improve the processing and cold storage of potato tubers [12].
3 CRISPR/Cas Systems in Horticultural Plants
The CRISPR/Cas genome editing technology was developed almost two years later than the discovery of TALEN proteins [15]. The CRISPR/Cas technology is expected to open up novel opportunities for the improvement of horticultural plants. This technology has already been used widely in horticultural plant breeding projects (see Table 14.1). For example, CRISPR/Cas9 was employed in the genome editing of sweet orange and tomato hairy roots [16, 17]. Several works have endeavored to enhance floricultural characteristics in ornamental plants, such as flower size, shape, color, aroma, shelf life, stress tolerance, etc. Through knocking down the Argonaute7, CRISPR/Cas9 was exploited to develop the first targeted mutation in the needle-leaf of tomato [18]. Since then, some research have been published on the potential uses of CRISPR/Cas in improving fruit quality, plant architecture, and shelf life as well as protecting plants from biotic and abiotic stresses [19]. The technique is now being exploited for a variety of fruits and vegetable crops, including watermelon, mustard, tomato, potato, and cabbage, etc. Table 14.1 presents a wide variety of genome-edited horticultural plants including vegetables, ornamentals, and fruit crops for different types of traits. Until now, just a handful number of teams have utilized CRISPR/Cas-mediated genome-editing in ornamental plants, because it is challenging to edit genomes of the plants without any genome sequence information. Thus, more whole-genome sequencing data should be generated. In addition, for the improvement of polyploid plant species, such as tetraploid roses and hexaploid chrysanthemum plants, a very effective editing platform has yet to be developed. Eleven research papers concerning genetic manipulation through CRISPR/Cas in ornamental plants have been collected from previously published research articles (Table 14.1). Specialized application of CRISPR/Cas such as base-editing has been reported for tomato [20]; however, any research generating mutations in horticultural plants other than tomato have not been reported using either the base-editing or prime-editing approaches yet.
4 Making Horticultural Plants More Tolerant to Abiotic and Biotic Stresses
Pathogens including bacteria, fungi, and viruses can cause a variety of diseases in plants. This hinders plant growth and development, which can result in significant losses and hence increase agricultural production costs. Nevertheless, plant tolerance to biotic stresses can be significantly improved through the utilization of the CRISPR/Cas technology.
Two separate strategies are utilized to develop virus-resistant plants: editing the virus genome and modifying the plant genes responsible for susceptibility to viruses. Viruses typically employ the transcription and translation machinery found in the host plant. Using CRISPR/Cas tools, sensitivity (S) genes may have their expression altered to protect plants from viruses, for example by knocking off translation initiation factors. For example, producing bananas with endogenous banana streak virus resistance was made possible by the CRISPR/Cas9 technique [25] through introucing mutation in the integrative viral components that inhibited the transcription and translation of viral proteins in banana trees. An example of modifying plant genes to introduce more resistance is apple for which CRISPR/Cas9 RNPs were introduced into protoplasts to target the DIPM-1, -2, and -4 genes which are fire blight resistance inhibitors [26]. The benefit of this transient expression was revealed through a decrease in undesirable mutations. The MdDIPM-4 gene was also knocked out in apple plants by the other researchers. Surprisingly, the FLP/FRT recombination system was used to eliminate the T-DNA harboring the expression cassettes for CRISPR/Cas9 from the genome of the transformed plants [27]. Also Citrus (Citrus sinensis orange and C. paradisi grapefruit) mutants were created by genome editing showing significant tolerance to Xanthomonas causing citrus canker [28, 29]. Citrus plants have the CsLOB1 gene, which makes them susceptible to Xanthomonas citri subsp. citri [30]. This gene’s promoter region contains elements necessary for the pathogenicity factor PthA4 of the bacterium to interact, which promotes the emergence of disease symptoms [31]. The PthA4 factor’s binding sites were edited using CRISPR/Cas9, which reduced the bacteria’s capacity to infect Citrus sinensis [29]. The Wanjincheng orange variety’s CsLOB1 gene’s promoter region was altered using some vector constructs. The rate of identified mutations varies between 11.5% and 64.7% according to the structure. As a result, four canker-resistant citrus mutant lines were selected. Significant plant resistance was obtained when a complete deletion happened in the promoter region of CsLOB1 where the PthA4 effector binds. Both Cas9 and Cpf1 have been used in similar investigations [32]. The CRISPR/Cas9-mediated editing of the CsWRKY22 gene, which codes for a different transcription factor, was another strategy to improve the bacterial canker resistance in Wanjincheng orange [33]. Genome editing also helped in the development of mutant bananas containing the DMR6 gene that are resistant to the Xanthomonas bacteria-caused banana wilt [34]. Many plant diseases are brought on by fungal infections. The introduction of CRISPR/Cas9 technology has brought novel promises for breeding crops resistant to a variety of fungal diseases by altering pathogen-sensitivity genes. It is well known that plants’ sensitivity genes make it easier for pathogens to invade and infect them. For example, CRISPR/Cas9 has made possible to knock out the MLO-7 gene, which negatively regulates resistance to powdery mildew pathogen, Erysiphe necator, in grapevine [26]. RNPs were used to deliver sgRNA to plants, and the mutagenesis frequency was reported quite low (0.1–6.9%). The RNP-mediated editing approach has been developed in recent studies [35]. Three MLO gene mutations caused grapevine plants to be 77% less sensitive to powdery mildew [36]. Moreover, by the deletion of the WRKY52 gene, a jasmonic acid pathway’s negative regulator, highly resistant grapevine plants to Botrytis cinerea have been produced [37, 38]. A noteworthy number of sgRNAs targeting various sequences within the first exon of the WRKY52 gene, were generated, and it was discovered that mutations in two alleles of the gene were more protective against the pathogen than those in only one allele. It is helpful to employ genome editing to explain how certain genes contribute to bring resistance to a disease. For instance, through knocking out the pathogenesis-related protein 4b gene (VvPR4b), grapevines became less resistant to Plasmopara viticola [39]. The VvPR4b gene produces the chitinase II-like protein which is required to stop hyphae development in P. viticola.
The fungal pathogen Botryosphaeria dothidea infects apple plants, causing serious damage. Increased response to this pathogen has been observed in apple calli after CNGC2 gene knockout [40]. Salicylic acid levels were found to be increasing at the same time as the expression of the gene encoding PR protein was still being inhibited. The gene CNGC2 was selected for targeted mutagenesis, but this was not the right alternative because mutations in this gene can have negative influences, like significantly reduced fertility. In another research, transient expression of the CRISPR/Cas9 in Theobroma cacao, the cacao plant, resulted in more resistant embryos and leaves against Phythophtora tropicalis [41]. The TcNPR3 gene was selected as the candidate for modification because it is an inhibitor of the defense mechanism. These findings support the outlook of cacao breeding for resistance to P. tropicalis.
It has been indicated that a Clpsk1 gene mutation makes watermelon plants more resistant to the fungus Fusarium oxysporum f. sp. niveum [42]. With CRISPR/Cas9 technology, the alteration of the genes controlling the pathogen-sensitivity in plants seems to be an efficient and fast method to make resistant plants against viral, bacterial and fungal diseases.
A few studies report that horticultural plants had been improved for abiotic stress tolerance using genome editing. As an example, the watermelon acetolactate synthase (ClALS) gene knocked out by CRISPR/Cas9, promoted the development of watermelons with herbicide tolerance [43]. The ALS gene was employed as an indicator for CRISPR/Cas9-mediated base editing, resulting in the development of herbicide (chlorosulfuron) tolerant pear trees (Pyrus communis L.) [44]. Similar CsALS gene editing in orange (Carrizo citrange) led to the mutant varieties that were tolerant to the herbicide imazapyr [45].
Therefore, editing the genomes of horticultural plants by CRISPR/Cas9 can be advantageous for developing crops resistant to a variety of (a)biotic stresses. Nevertheless, it seems essential to obtain stable mutants and completely study how genome editing influences traits and metabolic processes in plants.
5 Editing of Phenotypic Characteristics of Horticultural Plants
There are reports that genome editing can alter a variety of plant characteristics, including plant growth and morphology, fruit maturation time, fruit color, metabolism, and shelf life. The semi-dwarf banana plant (Musa acuminata “Gros Michel”) was produced through CRISPR/Cas9-mediated editing of the MaGA20ox2 genes regulating gibberellin biosynthesis [46]. The mutant plants displayed less growth than the wild-type plants, however thicker and deep-green leaves. The modified plants’ cells were structurally different compared to the wild-type plants. The findings of these works are crucial for the screening of dwarf banana genotypes because tall trees are susceptible to strong winds, which causes significant harvest losses. In grapevine plants, the targeted mutagenesis of the VvCCD8 gene resulted in more branches per shoot than in wild-type plants [47]. Plant phytohormones known as strigolactones prevent the development of axillary buds. The CRISPR/Cas9 editing system helped to functionally characterize the VvCCD8 gene’s role in the regulation of shoot branching. It is then planned to look into additional mechanisms for controlling the morphology and development of shoots in grapevines. In strawberry, it appeared feasible to produce white colored fruits using CRISPR/Cas9. The RAP (reduced anthocyanins in petioles) gene, which encodes a glutathione S-transferase enzyme, was targeted using CRISPR/Cas9 gene editing system [48]. This enzyme binds anthocyanins allowing them to move from the cytosol to the vacuole. It may be possible to produce strawberry types with popular white berries by editing the RAP gene.
CRISPR/Cas9-mediated genome editing is also considered as an efficient method for enhancing the nutritional value of fruits by editing their genomes. By mutating the lycopene epsilon-cyclase (LCYε) locus, it was possible to produce bananas with higher levels of β-carotene [49]. The amount of β-carotene rose six-fold in the fruit pulp of the mutant lines, whereas the α-carotene and lutein levels considerably diminished. Limited efforts have been made to apply CRISPR/Cas9 to create plants with improved phenotypes in red raspberry (Rubus idaeus L.). The flavone 3-hydrolase (F3′H), one of the important enzymes in flavonoid biosynthetic pathway, was knocked out in an investigation [50]. The gene MYB-16-like, functioning as a prickle formation regulator in raspberries, was another candidate for editing [51]. However, in both examples, it proved challenging for researchers to develop new seedlings from the generated raspberry calli.
Watermelon (Citrullus lanatus) seeds with mutations in the ClBG1 (β-glucosidase) gene had smaller seeds but better emerging due to decreased abscisic acid levels [52]. This gene can influence seed size and germination, which is a crucial characteristic of watermelon breeding. In another research, the role of the genes involved in fruit ripening such as 1-aminocyclopropane-1-carboxylate oxidase 1 (MaACO1) was evaluated and its sequences altered using CRISPR/Cas9 to find a way to extend the shelf life of bananas [53]. The obtained mutants developed smaller fruits with a slower maturity (60 days in comparison with 21 days for control bananas), which improved the storage capacity. The modified banana fruits additionally contained a higher vitamin C content. The CTR1-like, ROS1, and CmNAC-NOR genes in the melon (Cucumis melo var. cantalupensis) were knocked out employing CRISPR/Cas9, resulting in the development of melons with prolonged maturation and a long shelf life [54,55,56]. Thus, the reviewed studies demonstrated the sustainability of utilizing CRISPR/Cas9 to modify numerous characteristics in horticultural plants, for instance to improve fruit color and flavor, alter ripening and storage times, and alter growth-related traits.
6 Modification of the Flowering Period and Lifetime
Some studies indicate that horticultural plants can have their flowering time, floral life span, and flower shape and color altered by employing the CRISPR/Cas9. The function of numerous genes in the formation of flowers and fruits has been successfully identified in both wild and cultivated strawberry plants using the CRISPR/Cas9 technology. Since auxins are essential to the development of strawberries, FveARF8 and FveTAA1 genes were modified resulting in homozygous FveARF8 mutant plants with larger size and faster growth compared to the control plants [57]. Furthermore, mutations in FaTM6 and FveSEP3 strawberry genes resulted in parthenocarpy and an inaccurate fruit phenotype, as well as abnormal petals, anthers, and pollen grains [58, 59]. Thus, it has been demonstrated how these genes contribute to the growth of strawberry flowers and berries.
The flowering mechanisms of fruit plants can be manipulated with the use of genome editing. Growth traits have been manipulated in tomato to alter fruit development and flowering patterns. The transcriptional cofactors encoding genes such as BLADE-ON-PETIOLE (BOP) in tomato, can influence the inflorescences structure, and gene editing can be used to reduce the number of flowers per inflorescence by editing SlBOP genes [60]. Rapid flowering and early harvest are induced by CRISPR/Cas9-mediated targeted mutagenesis in the flowering inhibitor SELF-PRUNING 5G [61]. Editing the coding sequences of SlDML2 [62], SlORRM4 [63], the RIN locus [64], or the cis-regulatory domain of SlCLV3 [65] also affects the fruit development and ripening. The characteristics of the wild tomato relative (Solanum pimpinellifolium) were significantly altered by multiplex targeting of many genes crucial for tomato domestication, resulting in mutants that resembled domesticated tomatoes [66, 67]. Apple and pear mutant plants were obtained via knocking out of the TFL1 flowering repressor gene [68]. Only a few pear mutant plants (9% of all) but many apple mutant tree lines (93%) showed early flowering. Genome editing was used to clarify the significance of the AcCen4, AcCen, and SyGl genes role in delaying flowering in kiwifruit plants (Actinidia chinensis) [69, 70]. So there is a chance to get horticultural plants that flower earlier, which would decrease the period it takes for them to yield fruit. Likewise, in blueberry (Vaccinium corymbosum L.) plants, gene editing was performed on the gene CENTRORADIALIS (CEN) [71]. Based on the evidence for TFL1/CEN-like genes in kiwifruit, pear, and apple, the researchers hypothesized that the knocking out of this gene would result in early flowering [68,69,70]. However, attempts to use CEN gene editing to influence blueberry plant flowering were ineffective. Moreover, compared to the control plants, mutant plants grew far more slowly. Additional examination are yet to be done to explain the dwarf phenotype resulted in these mutant seedlings of CEN-edited blueberry plants [71].
Genes that control aging and corollas color in petunias, gentian, lily, ipomoea, chrysanthemum, and orchids have been studied by some researchers using genome editing [40, 72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89]. For instance, plants of the petunia cultivar “Mirage Rose” have been modified targeting the PhACO1 gene, which regulates the biosynthesis of the hormone ethylene [86]. As a result, petunia plants with less ethylene synthesis and flowers that lasted longer were developed. In Japanese morning glory (Ipomoea nil, “Violet”) plants, the EPH1 gene, a regulator of petal aging, was also knocked out to decrease flower wilting [76]. The problem of altering the color of flower corollas in ornamentals has been the center of many investigations. The genes carotenoid cleavage dioxygenase 4 (CCD4) and dihydroflavonol-4-reductase (DFR) were knocked out in Ipomoea nil plants [84, 85], which allowed for the modification of flower color. One of the key enzymes responsible for the production of flavonoids, flavone 3-hydrolase (F3′H), was knocked out by another study [74]. As a result, Torenia fournieri’s flower color converted from bright blue to white. In another research, the essential enzyme of carotenoid synthesis, PDS, was mutated, resulting in the creation of mutant Lilium pumilum and L. longiflorum with chimeric morphologies and different flower colors [87]. CRISPR/Cas9 gene editing has only been used in a few orchid species [73, 83]. Hence, it has been demonstrated that it is possible to modify horticultural plants’ flowering time, flower color, and flowering period using the CRISPR/Cas9 technology.
7 Conclusion
CRISPR/Cas9 technology has so far demonstrated its efficacy to modify horticultural plant genomes. The genomes of these plants have been edited to regulate the period of fruit ripening and flowering, to increase plant resistance to biotic and tolerance to abiotic stresses, and to enhance the characteristics related to plant growth and fruit flavor. Better editing and transcription-activation techniques as well as improved enzymes for more effective genome editing are now available for the development of novel types of horticultural plants.
References
Ravichandra, N., Ravichandra, N.: Horticulture and its role in the national economies. Hortic. Nematol. 1–3 (2014)
Mishra, R., Joshi, R.K., Zhao, K.: Genome editing in rice: recent advances, challenges, and future implications. Front. Plant Sci. 9, 1361 (2018)
Voytas, D.F.: Plant genome engineering with sequence-specific nucleases. Annu. Rev. Plant Biol. 64, 327–350 (2013)
Peer, R., Rivlin, G., Golobovitch, S., Lapidot, M., Gal-On, A., Vainstein, A., Tzfira, T., Flaishman, M.A.: Targeted mutagenesis using zinc-finger nucleases in perennial fruit trees. Planta. 241, 941–951 (2015)
Hilioti, Z., Ganopoulos, I., Ajith, S., Bossis, I., Tsaftaris, A.: A novel arrangement of zinc finger nuclease system for in vivo targeted genome engineering: the tomato LEC1-LIKE4 gene case. Plant Cell Rep. 35, 2241–2255 (2016)
Sun, Z., Li, N., Huang, G., Xu, J., Pan, Y., Wang, Z., Tang, Q., Song, M., Wang, X.: Site-specific gene targeting using transcription activator-like effector (TALE)-based nuclease in Brassica oleracea. J. Integr. Plant Biol. 55, 1092–1103 (2013)
Sawai, S., Ohyama, K., Yasumoto, S., Seki, H., Sakuma, T., Yamamoto, T., Takebayashi, Y., Kojima, M., Sakakibara, H., Aoki, T., Muranaka, T., Saito, K., Umemoto, N.: Sterol side chain reductase 2 is a key enzyme in the biosynthesis of cholesterol, the common precursor of toxic steroidal glycoalkaloids in potato. Plant Cell. 26, 3763–3774 (2014)
Ma, J., Xiang, H., Donnelly, D.J., Meng, F.-R., Xu, H., Durnford, D., Li, X.-Q.: Genome editing in potato plants by agrobacterium-mediated transient expression of transcription activator-like effector nucleases. Plant Biotechnol. Rep. 11, 249–258 (2017)
Forsyth, A., Weeks, T., Richael, C., Duan, H.: Transcription activator-like effector nucleases (TALEN)-mediated targeted DNA insertion in potato plants. Front. Plant Sci. 7, 1572 (2016)
Butler, N.M., Baltes, N.J., Voytas, D.F., Douches, D.S.: Geminivirus-mediated genome editing in potato (Solanum tuberosum L.) using sequence-specific nucleases. Front. Plant Sci. 7, 1045 (2016)
Nicolia, A., Proux-Wéra, E., Åhman, I., Onkokesung, N., Andersson, M., Andreasson, E., Zhu, L.-H.: Targeted gene mutation in Tetraploid potato through transient TALEN expression in protoplasts. J. Biotechnol. 204, 17–24 (2015)
Clasen, B.M., Stoddard, T.J., Luo, S., Demorest, Z.L., Li, J., Cedrone, F., Tibebu, R., Davison, S., Ray, E.E., Daulhac, A.: Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotechnol. J. 14, 169–176 (2016)
Lor, V.S., Starker, C.G., Voytas, D.F., Weiss, D., Olszewski, N.E.: Targeted mutagenesis of the tomato Procera gene using transcription activator-like effector nucleases. Plant Physiol. 166, 1288–1291 (2014)
Čermák, T., Baltes, N.J., Čegan, R., Zhang, Y., Voytas, D.F.: High-frequency, precise modification of the tomato genome. Genome Biol. 16, 1–15 (2015)
Malzahn, A., Lowder, L., Qi, Y.: Plant genome editing with TALEN and CRISPR. Cell Biosci. 7, 1–18 (2017)
Ron, M., Kajala, K., Pauluzzi, G., Wang, D., Reynoso, M.A., Zumstein, K., Garcha, J., Winte, S., Masson, H., Inagaki, S.: Hairy root transformation using agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol. 166, 455–469 (2014)
Jia, H., Wang, N.: Targeted genome editing of sweet orange using Cas9/SGRNA. PLoS One. 9, e93806 (2014)
Brooks, C., Nekrasov, V., Lippman, Z.B., Van Eck, J.: Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 166, 1292–1297 (2014)
Kulus, D.: Genetic resources and selected conservation methods of tomato. J. Appl. Bot. Food Qual. 91, 135–144 (2018)
Chen, L., Yang, D., Zhang, Y., Wu, L., Zhang, Y., Ye, L., Pan, C., He, Y., Huang, L., Ruan, Y.-L., Lu, G.: Evidence for a specific and critical role of mitogen-activated protein kinase 20 in uni-to-binucleate transition of microgametogenesis in tomato. New Phytol. 219, 176–194 (2018)
Wan, L., Wang, Z., Tang, M., Hong, D., Sun, Y., Ren, J., Zhang, N., Zeng, H.: CRISPR-Cas9 gene editing for fruit and vegetable crops: strategies and prospects. Horticulturae. 7, 193 (2021)
Rukavtsova, E.B., Zakharchenko, N.S., Lebedev, V.G., Shestibratov, K.A.: CRISPR-Cas genome editing for horticultural crops improvement: advantages and prospects. Horticulturae. 9, 38 (2023)
Xiong, J.-S., Ding, J., Li, Y.: Genome-editing technologies and their potential application in horticultural crop breeding. Hortic. Res. 2, 15019 (2015)
Xu, J., Hua, K., Lang, Z.: Genome editing for horticultural crop improvement. Hortic. Res. 6 (2019)
Tripathi, J.N., Ntui, V.O., Ron, M., Muiruri, S.K., Britt, A., Tripathi, L.: CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun. Biol. 2, 46 (2019)
Malnoy, M., Viola, R., Jung, M.-H., Koo, O.-J., Kim, S., Kim, J.-S., Velasco, R., Nagamangala Kanchiswamy, C.: DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 7, 1904 (2016)
Pompili, V., Dalla Costa, L., Piazza, S., Pindo, M., Malnoy, M.: Reduced fire blight susceptibility in apple cultivars using a high-efficiency CRISPR/Cas9-FLP/FRT-based gene editing system. Plant Biotechnol. J. 18, 845–858 (2020)
Jia, H., Omar, A.A., Orbović, V., Wang, N.: Biallelic editing of the LOB1 promoter via CRISPR/Cas9 creates canker-resistant ‘Duncan’ grapefruit. Phytopathology. 112, 308–314 (2022)
Peng, A., Chen, S., Lei, T., Xu, L., He, Y., Wu, L., Yao, L., Zou, X.: Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 15, 1509–1519 (2017)
Gottwald, T.R., Graham, J.H., Schubert, T.S.: Citrus canker: the pathogen and its impact. Plant Health Prog. 3, 15 (2002)
Hu, Y., Zhang, J., Jia, H., Sosso, D., Li, T., Frommer, W.B., Yang, B., White, F.F., Wang, N., Jones, J.B.: Lateral organ boundaries 1 is a disease susceptibility gene for citrus bacterial canker disease. Proc. Natl. Acad. Sci. 111, E521–E529 (2014)
Jia, H., Orbović, V., Wang, N.: CRISPR-LbCas12a-mediated modification of citrus. Plant Biotechnol. J. 17, 1928–1937 (2019)
Wang, L., Chen, S., Peng, A., Xie, Z., He, Y., Zou, X.: CRISPR/Cas9-mediated editing of CsWRKY22 reduces susceptibility to Xanthomonas citri subsp. Citri in Wanjincheng orange (Citrus sinensis (L.) Osbeck). Plant Biotechnol. Rep. 13, 501–510 (2019)
Tripathi, J.N., Ntui, V.O., Shah, T., Tripathi, L.: CRISPR/Cas9-mediated editing of DMR6 orthologue in banana (Musa spp.) confers enhanced resistance to bacterial disease. Plant Biotechnol. J. 19, 1291 (2021)
Osakabe, Y., Liang, Z., Ren, C., Nishitani, C., Osakabe, K., Wada, M., Komori, S., Malnoy, M., Velasco, R., Poli, M.: CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat. Protoc. 13, 2844–2863 (2018)
Pessina, S., Lenzi, L., Perazzolli, M., Campa, M., Dalla Costa, L., Urso, S., Valè, G., Salamini, F., Velasco, R., Malnoy, M.: Knockdown of MLO genes reduces susceptibility to powdery mildew in grapevine. Hortic. Res. 3 (2016)
Leisen, T., Bietz, F., Werner, J., Wegner, A., Schaffrath, U., Scheuring, D., Willmund, F., Mosbach, A., Scalliet, G., Hahn, M.: CRISPR/Cas with ribonucleoprotein complexes and transiently selected telomere vectors allows highly efficient marker-free and multiple genome editing in Botrytis cinerea. PLoS Pathog. 16, e1008326 (2020)
Wang, X., Tu, M., Wang, D., Liu, J., Li, Y., Li, Z., Wang, Y., Wang, X.: CRISPR/cas9-mediated efficient targeted mutagenesis in Grape in the first generation. Plant Biotechnol. J. 16, 844–855 (2018)
Wan, D.-Y., Guo, Y., Cheng, Y., Hu, Y., Xiao, S., Wang, Y., Wen, Y.-Q.: CRISPR/cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Hortic. Res. 7 (2020)
Zhou, H., Bai, S., Wang, N., Sun, X., Zhang, Y., Zhu, J., Dong, C.: CRISPR/Cas9-mediated mutagenesis of MdCNGC2 in Apple Callus and VIGS-mediated silencing of MdCNGC2 in fruits improve resistance to Botryosphaeria dothidea. Front. Plant Sci. 11, 575477 (2020)
Fister, A.S., Landherr, L., Maximova, S.N., Guiltinan, M.J.: Transient expression of CRISPR/Cas9 machinery targeting TcNPR3 enhances defense response in Theobroma cacao. Front. Plant Sci. 9, 268 (2018)
Zhang, M., Liu, Q., Yang, X., Xu, J., Liu, G., Yao, X., Ren, R., Xu, J., Lou, L.: CRISPR/Cas9-mediated mutagenesis of clpsk1 in watermelon to confer resistance to Fusarium oxysporum f. sp. niveum. Plant Cell Rep. 39, 589–595 (2020)
Tian, S., Jiang, L., Cui, X., Zhang, J., Guo, S., Li, M., Zhang, H., Ren, Y., Gong, G., Zong, M.: Engineering herbicide-resistant watermelon variety through CRISPR/cas9-mediated base-editing. Plant Cell Rep. 37, 1353–1356 (2018)
Malabarba, J., Chevreau, E., Dousset, N., Veillet, F., Moizan, J., Vergne, E.: New strategies to overcome present CRISPR/Cas9 limitations in apple and pear: efficient dechimerization and base editing. Int. J. Mol. Sci. 22, 319 (2020)
Alquézar, B., Bennici, S., Carmona, L., Gentile, A., Peña, L.: Generation of transfer-DNA-free base-edited citrus plants. Front. Plant Sci. 13 (2022)
Shao, X., Wu, S., Dou, T., Zhu, H., Hu, C., Huo, H., He, W., Deng, G., Sheng, O., Bi, F.: Using CRISPR/Cas9 genome editing system to create MaGA20ox2 gene-modified semi-dwarf Banana. Plant Biotechnol. J. 18, 17 (2020)
Ren, C., Guo, Y., Kong, J., Lecourieux, F., Dai, Z., Li, S., Liang, Z.: Knockout of VvCCD8 gene in grapevine affects shoot branching. BMC Plant Biol. 20, 1–8 (2020)
Gao, Q., Luo, H., Li, Y., Liu, Z., Kang, C.: Genetic modulation of RAP alters fruit coloration in both wild and cultivated strawberry. Plant Biotechnol. J. 18, 1550–1561 (2020)
Kaur, N., Alok, A., Kumar, P., Kaur, N., Awasthi, P., Chaturvedi, S., Pandey, P., Pandey, A., Pandey, A.K., Tiwari, S.: CRISPR/Cas9 directed editing of lycopene epsilon-cyclase modulates metabolic flux for β-carotene biosynthesis in Banana fruit. Metab. Eng. 59, 76–86 (2020)
Miller, S.: Gene Editing of Red Raspberry (Rubus Idaeus L.) with CRISPR/Cas9 Knocking out F3′H. Norwegian University of Life Sciences, Ås (2019)
Khadgi, A.: Uncovering Genetic and Transcriptomic Regulation of Prickle Development in Red Raspberry. Cornell University (2020)
Wang, Y., Wang, J., Guo, S., Tian, S., Zhang, J., Ren, Y., Li, M., Gong, G., Zhang, H., Xu, Y.: CRISPR/Cas9-mediated mutagenesis of ClBG1 decreased seed size and promoted seed germination in Watermelon. Hortic. Res. 8 (2021)
Hu, C., Sheng, O., Deng, G., He, W., Dong, T., Yang, Q., Dou, T., Li, C., Gao, H., Liu, S.: CRISPR/Cas9-mediated genome editing of MaACO1 (aminocyclopropane-1-carboxylate oxidase 1) promotes the shelf life of Banana fruit. Plant Biotechnol. J. 19, 654 (2021)
Giordano, A., Santo Domingo, M., Quadrana, L., Pujol, M., Martín-Hernández, A.M., Garcia-Mas, J.: CRISPR/cas9 gene editing uncovers the roles of constitutive triple response 1 and repressor of silencing 1 in melon fruit ripening and epigenetic regulation. J. Exp. Bot. 73, 4022–4033 (2022)
Hooghvorst, I., López-Cristoffanini, C., Nogués, S.: Efficient knockout of phytoene desaturase gene using CRISPR/Cas9 in melon. Sci. Rep. 9, 1–7 (2019)
Liu, B., Santo Domingo, M., Mayobre, C., Martín-Hernández, A.M., Pujol, M., Garcia-Mas, J.: Knock-out of CmNAC-NOR affects melon climacteric fruit ripening. Front. Plant Sci. 13 (2022)
Zhou, J., Wang, G., Liu, Z.: Efficient genome editing of wild strawberry genes, vector development and validation. Plant Biotechnol. J. 16, 1868–1877 (2018)
Martín-Pizarro, C., Triviño, J.C., Posé, D.: Functional analysis of the TM6 MADS-box gene in the octoploid strawberry by CRISPR/Cas9-directed mutagenesis. J. Exp. Bot. 70, 885–895 (2019)
Pi, M., Hu, S., Cheng, L., Zhong, R., Cai, Z., Liu, Z., Yao, J.-L., Kang, C.: The MADS-box gene FveSEP3 plays essential roles in flower organogenesis and fruit development in woodland strawberry. Hortic. Res. 8 (2021)
Xu, C., Park, S.J., Van Eck, J., Lippman, Z.B.: Control of inflorescence architecture in tomato by BTB/POZ transcriptional regulators. Genes Dev. 30, 2048–2061 (2016)
Soyk, S., Müller, N.A., Park, S.J., Schmalenbach, I., Jiang, K., Hayama, R., Zhang, L., Van Eck, J., Jiménez-Gómez, J.M., Lippman, Z.B.: Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in Tomato. Nat. Genet. 49, 162–168 (2017)
Lang, Z., Wang, Y., Tang, K., Tang, D., Datsenka, T., Cheng, J., Zhang, Y., Handa, A.K., Zhu, J.-K.: Critical roles of DNA demethylation in the activation of ripening-induced genes and inhibition of ripening-repressed genes in Tomato fruit. Proc. Natl. Acad. Sci. 114, E4511–E4519 (2017)
Yang, Y., Zhu, G., Li, R., Yan, S., Fu, D., Zhu, B., Tian, H., Luo, Y., Zhu, H.: The RNA editing factor SLORRM4 is required for normal fruit ripening in Tomato. Plant Physiol. 175, 1690–1702 (2017)
Ito, Y., Nishizawa-Yokoi, A., Endo, M., Mikami, M., Shima, Y., Nakamura, N., Kotake-Nara, E., Kawasaki, S., Toki, S.: Re-evaluation of the rin mutation and the role of RIN in the induction of tomato ripening. Nat. Plant. 3, 866–874 (2017)
Rodríguez-Leal, D., Lemmon, Z.H., Man, J., Bartlett, M.E., Lippman, Z.B.: Engineering quantitative trait variation for crop improvement by genome editing. Cell. 171, 470–480. e478 (2017)
Li, T., Yang, X., Yu, Y., Si, X., Zhai, X., Zhang, H., Dong, W., Gao, C., Xu, C.: Domestication of wild tomato is accelerated by genome editing. Nat. Biotechnol. 36, 1160–1163 (2018)
Zsögön, A., Čermák, T., Naves, E.R., Notini, M.M., Edel, K.H., Weinl, S., Freschi, L., Voytas, D.F., Kudla, J., Peres, L.E.P.: De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 36, 1211–1216 (2018)
Charrier, A., Vergne, E., Dousset, N., Richer, A., Petiteau, A., Chevreau, E.: Efficient targeted mutagenesis in apple and first time edition of pear using the CRISPR-cas9 system. Front. Plant Sci. 10(Feb) (2019)
Varkonyi-Gasic, E., Wang, T., Cooney, J., Jeon, S., Voogd, C., Douglas, M.J., Pilkington, S.M., Akagi, T., Allan, A.C.: Shy girl, a kiwifruit suppressor of feminization, restricts gynoecium development via regulation of cytokinin metabolism and signalling. New Phytol. 230(May), 1461–1475 (2021)
Varkonyi-Gasic, E., Wang, T., Voogd, C., Jeon, S., Drummond, R.S.M., Gleave, A.P., Allan, A.C.: Mutagenesis of kiwifruit CENTRORADIALIS-like genes transforms a climbing woody perennial with long juvenility and axillary flowering into a compact plant with rapid terminal flowering. Plant Biotechnol. J. 17, 869–880 (2019)
Omori, M., Yamane, H., Osakabe, K., Osakabe, Y., Tao, R.: Targeted mutagenesis of CENTRORADIALIS using CRISPR/Cas9 system through the improvement of genetic transformation efficiency of tetraploid highbush blueberry. J. Hortic. Sci. Biotechnol. 96, 153–161 (2021)
Kishi-Kaboshi, M., Aida, R., Sasaki, K.: Generation of gene-edited Chrysanthemum morifolium using multicopy transgenes as targets and markers. Plant Cell Physiol. 58, 216–226 (2017)
Kui, L., Chen, H., Zhang, W., He, S., Xiong, Z., Zhang, Y., Yan, L., Zhong, C., He, F., Chen, J., Zeng, P., Zhang, G., Yang, S., Dong, Y., Wang, W., Cai, J.: Building a genetic manipulation tool box for Orchid biology: identification of constitutive promoters and application of CRISPR/Cas9 in the Orchid, Dendrobium officinale. Front. Plant Sci. 7, 2036 (2016)
Nishihara, M., Higuchi, A., Watanabe, A., Tasaki, K.: Application of the CRISPR/Cas9 system for modification of flower color in Torenia fournieri. BMC Plant Biol. 18, 331 (2018)
Nitarska, D., Boehm, R., Debener, T., Lucaciu, R.C., Halbwirth, H.: First genome edited poinsettias: targeted mutagenesis of flavonoid 3′-hydroxylase using CRISPR/Cas9 results in a colour shift. Plant Cell Tissue Organ Cult. 147, 49–60 (2021)
Shibuya, K., Watanabe, K., Ono, M.: CRISPR/cas9-mediated mutagenesis of the EPHEMERAL1 locus that regulates petal senescence in Japanese morning glory. Plant Physiol. Biochem. 131, 53–57 (2018)
Su, S., Xiao, W., Guo, W., Yao, X., Xiao, J., Ye, Z., Wang, N., Jiao, K., Lei, M., Peng, Q., Hu, X., Huang, X., Luo, D.: The CYCLOIDEA-RADIALIS module regulates petal shape and pigmentation, leading to bilateral corolla symmetry in Torenia fournieri (Linderniaceae). New Phytol. 215, 1582–1593 (2017)
Subburaj, S., Chung, S.J., Lee, C., Ryu, S.-M., Kim, D.H., Kim, J.-S., Bae, S., Lee, G.-J.: Site-directed mutagenesis in petunia × hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep. 35, 1535–1544 (2016)
Sun, L., Kao, T.H.: CRISPR/cas9-mediated knockout of PiSSK1 reveals essential role of S-locus F-box protein-containing SCF complexes in recognition of non-self S-RNases during cross-compatible pollination in self-incompatible Petunia inflata. Plant Reprod. 31, 129–143 (2018)
Takahashi, S., Yoshida, C., Takahashi, H., Nishihara, M.: Isolation and functional analysis of EPHEMERAL1-LIKE (EPH1L) genes involved in flower senescence in cultivated Japanese gentians. Int. J. Mol. Sci. 23, 5608 (2022)
Tasaki, K., Higuchi, A., Watanabe, A., Sasaki, N., Nishihara, M.: Effects of knocking out three anthocyanin modification genes on the blue pigmentation of gentian flowers. Sci. Rep. 9, 15831 (2019)
Tasaki, K., Yoshida, M., Nakajima, M., Higuchi, A., Watanabe, A., Nishihara, M.: Molecular characterization of an anthocyanin-related glutathione S-transferase gene in Japanese gentian with the CRISPR/Cas9 system. BMC Plant Biol. 20, 370 (2020)
Tong, C.G., Wu, F.H., Yuan, Y.H., Chen, Y.R., Lin, C.S.: High-efficiency CRISPR/Cas-based editing of Phalaenopsis or chid mads genes. Plant Biotechnol. J. 18, 889–891 (2020)
Watanabe, K., Kobayashi, A., Endo, M., Sage-Ono, K., Toki, S., Ono, M.: CRISPR/cas9-mediated mutagenesis of the dihydroflavonol-4-reductase-B (DFR-B) locus in the Japanese morning glory Ipomoea (Pharbitis) nil. Sci. Rep. 7, 10028 (2017)
Watanabe, K., Oda-Yamamizo, C., Sage-Ono, K., Ohmiya, A., Ono, M.: Alteration of flower colour in ipomoea nil through CRISPR/Cas9-mediated mutagenesis of carotenoid cleavage dioxygenase 4. Transgenic Res. 27, 25–38 (2018)
Xu, J., Kang, B.-C., Naing, A.H., Bae, S.-J., Kim, J.-S., Kim, H., Kim, C.K.: CRISPR/Cas9-mediated editing of 1-aminocyclopropane-1-carboxylate oxidase1 enhances petunia flower longevity. Plant Biotechnol. J. 18, 287–297 (2020)
Yan, R., Wang, Z., Ren, Y., Li, H., Liu, N., Sun, H.: Establishment of efficient genetic transformation systems and application of CRISPR/Cas9 genome editing technology in Lilium pumilum dc. Fisch. and Lilium longiflorum White Heaven. Int. J. Mol. Sci. 20, 2920 (2019)
Zhang, B., Xu, X., Huang, R., Yang, S., Li, M., Guo, Y.: CRISPR/Cas9-mediated targeted mutation reveals a role for AN4 rather than DPL in regulating venation formation in the corolla tube of Petunia Hybrida. Hortic. Res. 8 (2021)
Zhang, B., Yang, X., Yang, C., Li, M., Guo, Y.: Exploiting the CRISPR/Cas9 system for targeted genome mutagenesis in Petunia. Sci. Rep. 6, 20315 (2016)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2024 The Author(s)
About this chapter
Cite this chapter
Mahna, N., Nayeri, S. (2024). Genome Editing in Horticultural Plants: Present Applications and Future Perspective. In: Ricroch, A., Eriksson, D., Miladinović, D., Sweet, J., Van Laere, K., Woźniak-Gientka, E. (eds) A Roadmap for Plant Genome Editing . Springer, Cham. https://doi.org/10.1007/978-3-031-46150-7_14
Download citation
DOI: https://doi.org/10.1007/978-3-031-46150-7_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-46149-1
Online ISBN: 978-3-031-46150-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)