Abstract
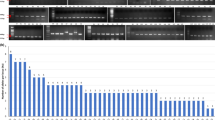
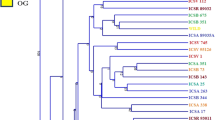
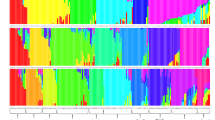
Genetic structure of 142 parent lines of sorghum [Sorghum bicolor (L.) Moench] was analyzed using model-based approach based on SSR markers. Forty-one selected from 103 SSR markers were used to analyze the parent lines, which generated 189 alleles revealed by each marker ranging from 2 to 11 with an average of 4.6 per marker. The polymorphic information content (PIC) value was 0.543 with a range of 0.089 to 0.850. All the parent lines were assigned to 7 subgroups, named Kafir, Kaoliang, Feterita, Shallu, Hegari, Milo and Durra. Parent lines without clear pedigree record were clustered into their corresponding groups, and genetic components of each line were estimated by Q-values. Information of this study would be useful for breeders to conclude their genetic background and select appropriate parents for germplasm improvement and hybrid breeding, and thus improve the efficiency of breeding programs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ali, M.L., Rajewski, J.F., Baenziger, P.S., Gill, K.S., Eskridge, K.M., Dweikat, I. 2008. Assessment of genetic diversity and relationship among a collection of US sweet sorghum germplasm by SSR markers. Mol. Breed. 21:497–509.
Anderson, J.A., Churchill, G.A., Autrique, J.E., Sorrellls, M.E., Tanksley, S.D. 1993. Optimizing parental selection for genetic linkage maps. Genome 36:181–186.
Barro-Kondombo, C., Sagnard, F., Chantereau, J., Deu, M., vom Brocke, K., Durand, P., Gozé, E., Zongo, J.D. 2010. Genetic structure among sorghum landraces as revealed by morphological variation and microsatellite markers in three agroclimatic regions of Burkina Faso. Theor. Appl. Genet. 120:1511–1523.
Chen, Q.S. 2005. Comparative analysis of specialty index and genetic similarity on soybean germplasm with resistance to Cercospora sojina Hara. China Biotech. 25 (suppl.):155–158. (in Chinese)
Choukan, R., Hossainzadeh, A., Ghannadha, M.R., Warburton, M.L., Talei, A.R., Mohammadi, S.A. 2006. Use of SSR data to determine relationships and potential heterotic groupings within medium to late maturing Iranian maize inbred lines. Field Crops Res. 95:212–222.
Doyle, J.F., Doyle, J.L. 1990. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Focus 12:13–15.
Evanno, G., Reganut, S., Goudet, J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecology 14:2611–2620.
Falush, D., Stephens, M., Pritchard, J.K. 2003. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164:1567–1587.
Kong, L., Dong, L., Hart, G.E. 2000. Characteristics, linkage-map positions, and allelic differentiation of Sorghum bicolor (L.) (Moench) DNA simple-sequence repeats (SSRs). Theor. Appl. Genet. 101:438–448.
Li, H., Ruan, C.J., da Silva, J.A.T. 2009. Identification and genetic relationship based on ISSR analysis in a germplasm collection of sea buckthorn (Hippophae L.) from China and other countries. Scientia Hortic. 123:263–271.
Mutegi, E., Sagnard, F., Semagn, K., Deu, M., Muraya, M., Kanyenji, B., de Villiers, S., Kiambi, D., Herselman, L., Labuschagne, M. 2011. Genetic structure and relationships within and between cultivated and wild sorghum (Sorghum bicolor (L.) Moench) in Kenya as revealed by microsatellite markers. Theor. Appl. Genet. 122:989–1004.
Panaud, O., Chen, X., McCouch, S. R. 1996. Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol. General Genet. 252:597–607.
Poehlman, J.M. 1986. Breeding Field Crops. Van Nostrand Reinhold Company, New York, USA, pp. 522–527.
Reif, J.C., Melchinger, A.E., Xia, X.C., Warburton, M.L., Hoisington, D.A., Vasal, S.K., Beck, D., Bohn, M., Frisch, M. 2003. Use of SSRs for establishing heterotic groups in subtropical maize. Theor. Appl. Genet. 107:947–057.
Shehzad, T., Okuizumi, H., Kawase, M., Okuno, K. 2009. Development of SSR-based sorghum (Sorghum bicolor (L.) Moench) diversity research set of germplasm and its evaluation by morphological traits. Genet. Res. Crop Evol. 56:809–827.
Simko, I., Eujayl, I., van Hintum, T.J.L. 2012. Empirical evaluation of DArT, SNP, and SSR marker-systems for genotyping, clustering, and assigning sugar beet hybrid varieties into populations. Plant Sci. 184:54–62.
Wang, M.L., Zhu, C., Barkey, N.A., Chen, Z., Erpelding, J.E., Murray, S.C., Tuinstra, M.R., Tesso, T., Pederson, G.A., Yu, J. 2009. Genetic diversity and population structure analysis of accessions in the US historic sweet sorghum collection. Theor. Appl. Genet. 120:13–23.
Wu, C.L., Zhang, Q.Q., Dong, B.X., Li, S.F., Zhang, C.Q. 2010. Analysis of genetic structure and relationships of maize inbred lines in China. Acta Agron. Sin. 36:1820–1831.
Yu, X., Bai, G., Luo, N., Chen, Z., Liu, S., Liu, J., Warnke, S. E., Jiang, Y. 2011. Association of simple sequence repeat (SSR) markers with submergence tolerance in diverse populations of perennial ryegrass. Plant Sci. 180:391–398.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Anioł
Electronic Supplementary Material (ESM)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wang, L.M., Jiao, S.J., Jiang, Y.X. et al. Genetic Structure Analysis of Sorghum Parent Lines Based on SSR Markers. CEREAL RESEARCH COMMUNICATIONS 41, 359–365 (2013). https://doi.org/10.1556/CRC.2013.0018
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/CRC.2013.0018