Abstract
Background
Entomopathogenic nematodes (EPNs) belonging to the Heterorhabditis spp. harbour symbiotically associated bacteria which are toxic to a wide range of insect pests. Isolation, purification, characterization and mass multiplication of such bacteria will be a promising strategy in the management of the pests. This study was carried out to isolate the EPN from different locations, isolate and purify the bacterial colonies, characterize the bacteria through morphological and molecular strategies and to test the efficacy of different bacteria in the control of polyphagous Tetranychus truncatus Ehara mites.
Results
EPNs were isolated from soil samples at 11 localities of Kerala State, India, and used to infect the Galleria mellonella L. larvae. Bacteria associated with the haemolymph of the infected larvae were isolated, which on NBTA medium have produced circular to irregular, entire, opaque and smooth colonies. Sequence characterization of the 16S rRNA revealed nine isolates namely: one symbiotic bacterium Photorhabdus luminescens, two Pseudomonas aeruginosa, five Ochrobactrum sp. and one Stenotrophomonas maltophilia. Phylogenetic analysis using the sequences has further confirmed the bacterial identity. Evaluation of the cell suspension (CS) and cell-free supernatant (CFS) of P. luminescens, P. aeruginosa and Ochrobactrum sp. for their adulticidal and ovicidal efficiencies on T. truncatus had identified significant adulticidal effects by P. luminescens, followed by P. aeruginosa. After 96 h of treatment, P. luminescens at 108 cells/ml resulted in a significantly higher mortality rate of adult mites (64.00 and 60.67%, respectively, for CFS and CS), compared to that resulted by P. aeruginosa (38.67 and 33.33%).
Conclusions
Results of this study showed that P. luminescens associated with the EPN Heterorhabditis spp. is a promising biocontrol agent for T. truncatus.
Similar content being viewed by others
Background
Entomopathogenic nematodes (EPNs) belonging to the families Steinernematidae and Heterorhabditidae are obligate insect parasites which can be used as potential biocontrol agents against important soil dwelling pests (Lacey and Georgis 2012). The bacteria symbiotically associated with EPNs are toxic to a wide range of insect pests (Guo et al. 1999). Nematodes protect the bacteria from the host’s immune system and help in their transportation by forming a special vesicle in the infective L3 juvenile stage, as in case of Steinernematidae, or retain throughout the intestine as in case of Heterorhabditidae. The bacteria help in maintaining ideal conditions for the reproduction of the nematodes by releasing some of the nutrients. They also release antimicrobial substances, which prevent the growth of other bacteria (Boemare et al. 1996).
Photorhabdus luminescens is the main bacterial species symbiotically associated with the EPN Heterorhabditis (Boemare et al. 1996). Pathogenicity of this bacterium against insect pests has been well documented by earlier scientists (Bowen et al. 1998). Additionally, few studies report the pathogenicity of this bacterium to mite species (Kulkarni et al. 2017). However, many other bacteria are also associated with EPNs (Babic et al. 2000). Entomopathogenicity of some of these bacteria associated with Heterorhabditis spp. has also been reported (Salgado-Morales et al. 2019).
Spider mites (Tetranychidae) are serious sucking pests of many agricultural and horticultural crops. Among the spider mites, Tetranychus truncatus Ehara (Acari: Tetranychidae) is a major species infesting economically important crops of Kerala State, India (Bennur et al. 2015). In this study, bacteria associated with the EPNs Heterorhabditis spp. were evaluated under laboratory conditions for their bio-efficacy against T. truncatus.
Methods
Collection of soil samples
Soil samples were collected randomly from 11 localities of Kerala State, India. From each locality, 500 g soil was collected from 15 cm depth, sealed and labelled in polybags and brought to the laboratory.
Isolation of Heterorhabditis from the soil
Following the baiting technique (Bedding and Akhurst 1975), EPNs were isolated from the soil samples. Larvae of greater wax moth, Galleria mellonella L. (Lepidoptera: Pyralidae) obtained from the culture maintained by All India Network Project on Agricultural Acarology (AINPAA), Kerala Agricultural University, India, were used for the isolation and maintenance of EPNs. Four to five healthy last-instar larvae of G. mellonella were released separately into plastic containers of 250 ml size, containing the soil samples collected from different locations. The soil samples were moistened before transferring to the plastic container to maintain adequate moisture content. Mouth of the container was covered with muslin cloth, placed upside down and stored at room temperature. Infected larvae which turned to brick red colour and died within 24–48 h, indicating infection by the EPNs were picked and washed thoroughly with distilled water and transferred to White’s trap (White 1927) and kept in dark at 28 °C. For the multiplication and maintenance of EPNs, infective juveniles (IJs) released from the cadavers along the sides of the Petri plate were collected, stored in vials and used as stock for infecting the Galleria larvae. To get infected, four to five larvae were released into Petri plate lined with filter paper, moistened with IJ suspension (1 ml), incubated at 28 °C in a BOD incubator and observed for mortality. The dead cadavers were used for the isolation of EPNs-associated bacteria.
Isolation of bacteria from the EPNs
Bacteria were isolated from the EPNs infected cadavers and inoculated on NBTA plates. Within 24–48 h of death, the cadavers were collected and surface sterilized with 70% ethanol under aseptic conditions. The cadavers were dissected with a sterile blade, and a drop of haemolymph was streaked on Petri plates containing 10–20 ml of NBTA media (peptone 5 g, beef extract 3 g, NaCl 5 g, bromothymol blue dye 0.025 g, 2,3,5 triphenyl-tetrazolium chloride (TTC) 0.04 g, agar 15 g and distilled water 1000 ml) using a sterile inoculation loop (Woodring and Kaya 1988). Same medium was used for isolation, purification and maintenance of the bacteria. The Petri plates were then sealed tightly by using parafilm and kept in BOD incubator for 24–48 h at the optimum temperature of 28 °C and dark conditions and observed for colony proliferation. Individual pure colonies were selected, streaked on NBTA plates and subcultured continuously to get pure colonies of uniform size and morphology.
Identification of the bacteria
Bacteria associated with EPNs were studied for their cultural characters and 16S rRNA sequences. The colony characters such as shape, colour, edge, margin, elevation and surface as well as change in the colour of media, were recorded. The isolates were also subjected to Gram’s reaction.
For molecular characterization, 16S rRNA gene was amplified and sequenced through colony PCR on fresh cultures of 24–48 h. Reaction mixture (50 μl) was prepared by adding EmeraldAmp GT PCR Master mix (25 μl), water (23 μl), forward and reverse primers (1 μl each) and bacterial culture was added by picking a single colony with tooth pick. Universal primer sequences (27f/1492r) 5′ AGAGTTTGATCCTGGCTCAG 3′ (forward) and 5′ ACGGCTACCTTGTTACGACTT 3′ (reverse) were used (Mulla et al. 2017). Thermal program followed had initial denaturation at 94 °C for 4 min, followed by 35 cycles consisting of denaturation at 94 °C for 30 s, annealing at 55.7 °C for 45 s and extension at 72 °C for 45 s and the final extension at 72 °C for 8 min (Veriti, Thermo Fisher Scientific). Products were electrophoresed (1.4% agarose gel) and sequenced. Contigs of 16S rRNA gene were analysed for homology and phylogenetic relations along with 25 accessions of Ochrobactrum spp., seven accessions of Photorhabdus luminescens, 10 sequences of P. aeruginosa and five accessions of Stenotrophomonas spp. retrieved from GenBank. Phylogenetic tree was constructed by neighbour joining method with 500 bootstrap replications.
Bioassay of EPN-associated bacteria against Tetranychus truncatus
Bacterial cells as well as cell-free supernatant (CFS) of five bacterial isolates (one isolate of P. luminescens, two isolates of P. aeruginosa and two isolates of Ochrobactrum) were evaluated independently against the eggs and adults of T. truncatus at 104, 105, 106, 107 and 108 cells/ml concentrations. The Pseudomonas and Photorhabdus isolates were selected because of their high pathogenicity against some of the insect pests (Salgado-Morales et al. 2019). Since Ochrobactrum spp. is reported to be pathogenic (Fu and Liu 2019) and non-pathogenic (Babic et al. 2000) to insects, its isolates were also included. The emerging multidrug-resistant opportunistic pathogen Stenotrophomonas (Brooke 2012) was excluded from the study. Experiments were laid out in completely randomized design with 26 treatments including control and three replications per treatment.
For the ovicidal assay, ten gravid females of T. truncatus were transferred on the leaf bit of mulberry (5 × 5 cm2 each), placed in a Petri plate lined with wet cotton and allowed for laying eggs. After 24 h, the mites were removed and 25 eggs were retained per leaf bit. For adulticidal assay, three leaf bits with 25 gravid female mites were used for each treatment. Sterile liquid broth of NBT (150 ml) was taken in a conical flask and a loop full of fresh bacterial culture was added aseptically, sealed and maintained in an orbital shaker at 200 rpm for 24–48 h and used as mother culture. Serial dilution, plating and colony counting method were followed to assess the initial concentration of the mother culture. Based on the initial concentration, 104, 105, 106, 107 and 108 cell/ml were prepared by serial dilution. Ten millilitre each of these dilutions was centrifuged at 4000 rpm for 20 min for the preparation of cell suspension and cell-free supernatant and sprayed separately on leaf bits with egg and adult mites, using a hand atomizer (2 ml/leaf bit). Per cent mortality was recorded by observing under a stereo binocular microscope (LEICA EZ4 HD) at 24, 48, 72 and 96 h of spraying.
Results
Eleven isolates of EPNs Heterorhabditis spp. were isolated from the soil samples by baiting method. The infective juvenile suspension prepared from the EPNs isolates (Fig. 1A) was used for infecting healthy Galleria larvae (Fig. 1B–D). Bacteria were isolated from the infected cadavers (Fig. 1E) and cultured on NBTA media. When haemolymph from the infected cadavers of G. mellonella was streaked on NBTA plates, bacterial colonies developed within 24–48 h. Eleven bacteria were isolated from the EPN-infected Galleria cadavers and coded based on the locality from where soil was collected for EPN isolation (Table 1).
A Infective juvenile suspension of nematodes collected in the White’s trap. B Healthy Galleria larva. C Entomopathogenic nematode infected cadaver. D Nematodes inside the host body. E Colonies produced by bacteria on NBTA media. F Change in media colour due to pigmentation by CF1 isolate. G Gram-stained bacterial cells (100X magnification)
Cultural characterization of the bacteria
The colony characteristics of the isolates KL1 and KT1 were similar on NBTA medium (Table 2), with circular shape, irregular edges, opaque, flat and smooth surface. Similarly, the isolates MP1, MT1, FR1, EKM1, HI1, HQ1 and HS1 were similar, but differed from the isolates, KL1 and KT1 in colour of the colony. The colour varied from red (FR1, MT1, HI1) to pinkish red (HS1, MP1, EKM1, HQ1). Elevation of the colonies varied from flat (KL1. KT1, FR1, EKM1), low convex (MP1, HS1) to raised (MT1, CF1, HI1, HQ1, WH1). Isolates WH1 and CF1 were similar, producing dark red colonies with white margin and the media turned from yellowish to blue (Fig. 1F). Circular to irregular, opaque, raised, smooth colonies were produced. Under 100X oil immersion microscope, the cells appeared red in colour (Fig. 1G) and were identified as Gram’s negative. However, the shape of the cell varied among the isolates from short to long rods.
Molecular characterization
Using the universal primers, 16S rRNA gene from eleven isolates was amplified (Fig. 2) and sequenced. Out of the eleven isolates, only nine isolates were identified, as the sequencing information for the remaining two isolates (WH1 and MT1) was incomplete even after repeated sequencing. Of the nine isolates identified, isolates KL1 and KT1 were identified as Pseudomonas aeruginosa, while the isolate HQ1 was identified as Stenotrophomonas maltophilia (Table 3). The isolate CF1 showed maximum sequence homology with Photorhabdus luminescens, while the remaining isolates MP1, FR1, EKM1, HI1 and HS1 belonged to Ochrobactrum (FR1—O. pseudogrignonensis, HI1—O. anthropi, EKM1, MP1 and HS1—undescribed species).
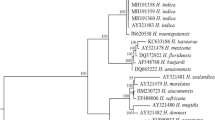
The phylogenetic analysis of 16S rRNA gene from the nine isolates along with 47 related accessions from the GenBank has resulted in a phylogenetic tree with two clusters (Fig. 3). Cluster A had subcluster A1 accommodating all sequences of Ochrobactrum and A2 accommodating the accessions of Stenotrophomonas. Cluster B had subcluster B1 accommodating Photorhabdus luminescens accessions and B2 accommodating P. aeruginosa accessions.
Bioassay of the EPN-associated bacteria against Tetranychus truncatus
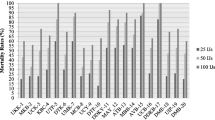
Five bacterial isolates (CF1—P. luminescens, KL1—P. aeruginosa strain 1, KT1—P. aeruginosa strain 2, HI1—Ochrobactrum sp. strain 1 and FR1—Ochrobactrum sp. strain 2) were evaluated for their biocontrol efficacy. Though the isolates had non-significant ovicidal action (Fig. 4), substantial adulticidal effects were recorded (Fig. 5, Additional file 1: Tables S1 and S2). The highest egg mortality of 30.67% was observed at 96 h, in the treatment with 108 cells/ml of P. luminescens. In adulticidal treatments, no mortality was observed up to 24 h but after 48 h, mortality rate had increased. The P. luminescens isolate was significantly superior over other isolates in both CS and CFS forms. At 96 h, treatment with 108 cells/ml of P. luminescens recorded significantly higher mortality of 60.67%. The same treatment recorded 64.00% mortality in its CFS form. This was followed by the other treatments of P. luminescens at 107 cells/ml (52.00% in CS and 56.00% in CFS), 105 cells/ml (50.67% in CS and 53.33% in CFS), 106 cells/ml (49.33% in CS and 52.00% in CFS) and 104 cells/ml (49.33% in CS and 48.00% in CFS).
Among treatments of P. aeruginosa, the highest mortality of 38.67% in CFS and 33.33% in CS was recorded in the treatment with 108 cells/ml of strain 2 (KT1 isolate). No significant mortality rate was observed in any of the treatments of Ochrobactrum spp.
Discussion
In this study, nine bacterial isolates were isolated from the haemolymph of EPNs (Heterorhabditis spp.) infected G. mellonella cadaver. This included one species of symbiotic bacteria, P. luminescens and eight non-symbiotic/associated bacteria namely two isolates of P. aeruginosa, five isolates of Ochrobactrum and one isolate of Stenotrophomonas maltophilia.
Photorhabdus is the predominant genus of entomopathogenic bacteria, which is a natural symbiont of the EPN, Heterorhabditis spp. (Boemare et al. 1993). It is Gram negative and bioluminescent. The bacterium supports growth of the nematode in the association, but is a deadly pathogen to most of the insect pests (Clarke 2008). They reside in the gut of the nematodes and get released into the insect haemocoel soon after the EPN infects the insect and kill them within 24–48 h. Other than the symbionts, few associated bacteria also reside within the system of Heterorhabditis. Bacterial species such as Alcaligenes aquatilis, Alcaligenes faecalis, Enterococcus mundtii, Pseudomonas protegens, Serratia nematodiphila, Serratia marcescens and Stenotrophomonas maltophilia were previously isolated from the Heterorhabditis infected Galleria cadaver (Ruiu et al. 2017).
Ochrobactrum is a Gram-negative opportunistic bacterium (Brucellaceae) which mostly occurs singly. This bacterium is related to Brucella, Phyllobacterium, Rhizobium and Agrobacterium. Occurrence of natural dixenic association between the bacterial symbiont P. luminescens and the bacterium Ochrobactrum sp. in Heterorhabditis species was reported by Babic et al. (2000). In this study, five isolates of Ochrobactrum were found in association with the EPN-infected Galleria. Akhurst (1982) reported the antibiotic properties of the Xenorhabdus sp. (now Photorhabdus) which inhibits the growth of other microorganisms. However, Aujoulat et al. (2019) observed the ability of Ochrobactrum sp. to grow well even in the presence of P. luminescens due of its resistance to the antibiotics produced by the Photorhabdus.
Pseudomonas includes disease-causing opportunistic bacteria. P. aeruginosa is a free living, Gram-negative bacterium having extensive metabolic diversity. The Stenotrophomonas spp., formerly isolated as P. maltophilia, is an emerging multidrug-resistant opportunistic pathogen. Ruiu et al. (2017) isolated S. maltophilia from the Heterorhabditis infected Galleria cadaver.
In the phylogenetic tree, the sequences of Ochrobactrum spp. and Stenotrophomonas spp. formed two subclades indicating the divergence of these genera from a common ancestry. In clade 2, Photorhabdus and Pseudomonas subclustered, indicating the evolutionary relationship of the two species. Velasco et al. (1998) reported the similarity of O. anthropi and O. intermedium with Photorhabdus luminescens subsp. akhurstii. All the subclades have further branched into subclusters within the species, indicating significant inter-specific diversity.
Efficacy of the symbiotic bacterium P. luminescens against different insect pests was reported by several workers (Kumar et al. 2014). However, studies on the efficacy of this bacterium on mites are limited. Due to the virulent properties and the ability to infect a wide range of insect hosts, it is a promising candidate for agricultural use as a mass-produced biological control agent (Gerdes et al. 2015). Kumar et al. (2014) studied the bio-efficacy of broth cultures of different isolates of P. luminescens against some sucking pests (Aphis gossypii and Tetranychus macfarlanei) and defoliators (Plutella xylostella and Spodoptera litura). The isolate Z-8–1 was found to be effective against A. gossypii after 24 h with 100% nymphal mortality and the isolate Z-3–1 was effective against T. macfarlanei with 100% mortality after 36 h. None of the isolates were pathogenic to defoliators.
In this study, adult mortality rate was higher by P. luminescens treatment, compared to P. aeruginosa and Ochrobactrum. At the highest concentration of 108 cells/ml, P. luminescens has resulted in 60.67% mortality. Mortality was negligible with the other isolates. This clearly showed that P. luminescens is more effective against T. truncatus than the other bacteria. The EPN-associated bacteria: Ochrobactrum as well as P. aeruginosa were also reported to possess insecticidal activity against few pests.
Salgado-Morales et al. (2019) assessed the pathogenicity of P. luminescens HIM3 and P. aeruginosa NA04 isolated from Heterorhabditis indica against G. mellonella and few other insect pests. Both the isolates were found to be highly virulent to G. mellonella with nearly complete mortality after 24 h. P. luminescens was virulent to both of the insect hosts but the pathogenicity of P. aeruginosa varied with the hosts. They also noticed that a high concentration of P. aeruginosa was required for infecting Tenebrio molitor. P. aeruginosa recorded only (10%) mortality of Diatraea magnifactella Dyar after 36 h of treatment, but mortality was complete with P. luminescens. In the present study, mortality of T. truncatus was less when treated with P. aeruginosa compared to P. luminescens. The bacterium Ochrobactrum tritici isolated from the EPN Oscheius chongmingensis was reported to cause 93.33% mortality of G. mellonella (Fu and Liu 2019). But Ochrobactrum isolates did not exhibit appreciable activity against T. truncatus in this study.
Conclusions
This study identified a potential isolate of P. luminescens with significant adulticidal action against T. truncatus. Techniques for mass-multiplying P. luminescens are to be standardized for utilization of this bacterial isolate in mite pest management.
Availability of data and materials
Additional data associated with this study can be accessed from the Master’s thesis submitted by AMN. The bacterial cultures and the pure lines of mites are maintained at Kerala Agricultural University, India.
Abbreviations
- BOD:
-
Biological oxygen demand
- CFS:
-
Cell-free supernatant
- CS:
-
Cell suspension
- EPN:
-
Entomopathogenic nematodes
References
Akhurst RJ (1982) Antibiotic activity of Xenorhabdus spp., bacteria symbiotically associated with insect pathogenic nematodes of the families Heterorhabditidae and Steinernematidae. Microbiology 128(12):3061–3065. https://doi.org/10.1099/00221287-128-12-3061
Aujoulat F, Pages S, Masnou A, Emboule L, Teyssier C, Marchandin H, Gaudriault S, Givaudan A, Jumas-Bilak E (2019) The population structure of Ochrobactrum isolated from entomopathogenic nematodes indicates interactions with the symbiotic system. Infect Genet Evol 70:131–139. https://doi.org/10.1016/j.meegid.2019.02.016
Babic I, Fischer-LeSaux M, Giraud E, Boemare N (2000) Occurrence of natural dixenic associations between the symbiont Photorhabdus luminescens and bacteria related to Ochrobactrum spp. in tropical entomopathogenic Heterorhabditis spp. (Nematoda, Rhabditida). Microbiology 146(3):709–718. https://doi.org/10.1099/00221287-146-3-709
Bedding RA, Akhurst RJ (1975) A simple technique for the detection of insect parasitic rhabditid nematodes in soil. Nematologica 21:109–110
Bennur S, Abida PS, Valsala PA, Mathew D, Bhaskar H (2015) DNA barcoding of spider mites (Prostigmata: Tetranychidae) in vegetables using COI and ITS2 marker. Genome 58(5):195. https://doi.org/10.1139/gen-2015-0087
Boemare NE, Akhurst RJ, Mourant RG (1993) DNA relatedness between Xenorhabdus sp. (Enterobacteriaceae) symbiotic bacteria of entomopathogenic nematodes, and a proposal to transfer Xenorhabdus luminescens to new genus, Photorhabdus. Int J Syst Evol Microbiol 43(2):249–255. https://doi.org/10.1099/00207713-43-2-249
Boemare N, Laumond C, Mauleon H (1996) The entomopathogenic nematode-bacterium complex: biology, life cycle and vertebrate safety. Biocontrol Sci Technol 6(3):333–346. https://doi.org/10.1080/09583159631316
Bowen D, Rocheleau TA, Blackburn M, Andreev O, Golubeva E, Bhartia R (1998) Insecticidal toxins from the bacterium Photorhabdus luminescens. Science 280(5372):2129–2132. https://doi.org/10.1126/science.280.5372.2129
Brooke JS (2012) Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clin Microbiol Rev 25(1):2–41. https://doi.org/10.1128/CMR.00019-11
Clarke DJ (2008) Photorhabdus: a model for the analysis of pathogenicity and mutualism. Cell Microbiol 10(11):2159–2167. https://doi.org/10.1111/j.1462-5822.2008.01209.x
Fu JR, Liu QZ (2019) Evaluation and entomopathogenicity of gut bacteria associated with dauer juveniles of Oscheius chongmingensis (Nematoda: Rhabditidae). Microbiol Open 8(9):e00823. https://doi.org/10.1002/mbo3.823
Gerdes E, Upadhyay D, Mandjiny S, Bullard-Dillard R, Storms M, Menefee M, Holmesi LD (2015) Photorhabdus luminescens: virulent properties and agricultural applications. J Agric for 3(5):171–177. https://doi.org/10.11648/j.ajaf.20150305.12
Guo L, Fatig RO, Orr GL, Schafer GW, Strickland JA, Sukhapinda K, Woodsworth AT, Petell JK (1999) Photorhabdus luminescens W-14 insecticidal activity consists of at least two similar but distinct proteins. Purification and characterization of Toxin A and Toxin B. J Biol Chem 274:9836–9842. https://doi.org/10.1074/jbc.274.14.9836
Kulkarni RA, Prabhuraj A, Ashoka J, Hanchinal SG, Hiregoudar S (2017) Generation and evaluation of nanoparticles of supernatant of Photorhabdus luminescens (Thomas and Poinar) against mite and aphid pests of cotton for enhanced efficacy. Curr Sci 112(11):2312–2316. https://doi.org/10.18520/cs/v112/i11/2312-2316
Kumar KV, Vendan KT, Nagaraj SB (2014) Isolation and characterization of entomopathogenic symbiotic bacterium, Photorhabdus luminescens of Heterorhabditis indica from soils of five agro climatic zones of Karnataka. Biosci Biotechnol Res Asia 11(1):129–139. https://doi.org/10.13005/bbra/1244
Lacey LA, Georgis R (2012) Entomopathogenic nematodes for control of insect pests above and below ground with comments on commercial production. J Nematol 44(2):218–225
Mulla SR, Joshi S, Das J (2017) Isolation, cloning and expression of insecticidal-protein-encoding gene tcdA from Photorhabdus luminescens in Escherichia coli. Int J Curr Microbiol Appl Sci 6(9):1718–1724. https://doi.org/10.20546/ijcmas.2017.609.212
Ruiu L, Virdis B, Mura ME, Floris I, Satta A, Tarasco E (2017) Oral insecticidal activity of new bacterial isolates against insects in two orders. Biocontrol Sci Technol 27(7):886–902. https://doi.org/10.1080/09583157.2017.1355964
Salgado-Morales R, Martínez-Ocampo F, Obregon-Barboza V, Vilchis-Martínez K, Jimenez-Perez A, Dantan-Gonzalez E (2019) Assessing the pathogenicity of two bacteria isolated from the entomopathogenic nematode Heterorhabditis indica against Galleria mellonella and some pest insects. Insects 10(3):83. https://doi.org/10.3390/insects10030083
Velasco J, Romero C, Lopez-Goni I, Leiva J, Díaz R, Moriyón I (1998) Evaluation of the relatedness of Brucella spp. and Ochrobactrum anthropi and description of Ochrobactrum intermedium sp. nov., a new species with a closer relationship to Brucella spp. Int J Syst Evol Microbiol 48(3):759–768. https://doi.org/10.1099/00207713-48-3-759
White GF (1927) A method for obtaining infective nematode larvae from cultures. Science 66(1709):302–303. https://doi.org/10.1126/science.66.1709.302-b
Woodring JL, Kaya HK (1988) Steinernematid and heterorhabditid nematodes: A handbook of biology and techniques. Southern Cooperative Series Bulletin 331, Arkansas Agricultural Experiment Station, Fayetteville, AR
Acknowledgements
Not applicable.
Funding
This work was financially supported by Department of Biotechnology (DBT), Ministry of Science and Technology, Govt. of India (DBT-HRD/CPBMB/CoH/2019).
Author information
Authors and Affiliations
Contributions
HB conceived the research hypothesis, supervised the research progress, performed the morphological characterization of EPN and mites, reared and maintained the mites and revised the manuscript. AMN performed the experiments and wrote the draft manuscript. DM designed the experiments for the molecular characterization of the bacterial isolates, performed the bioinformatics analyses and revised the manuscript. GD designed the experiments for the isolation, purification and morphological characterization of bacteria. SMR designed sequence characterization experiments and managed the funding. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable, because our manuscript reports neither studies involving human participants nor human data or human tissue.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
Adulticidal effect of cell suspension (CS) of bacterial isolates against Tetranychus truncatus. Table S2. Adulticidal effect of cell-free supernatant (CFS) of bacterial isolates against Tetranychus truncatus
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ashwini, M.N., Bhaskar, H., Mathew, D. et al. Isolation and evaluation of bacteria associated with entomopathogenic nematode Heterorhabditis spp. against the spider mite, Tetranychus truncatus Ehara (Acari: Tetranychidae). Egypt J Biol Pest Control 32, 87 (2022). https://doi.org/10.1186/s41938-022-00586-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s41938-022-00586-8