Abstract
Electrochemically active bacteria (EAB) receive considerable attention in sustainable biotechnology, since they are essential components in microbial fuel cells (MFCs) that are able to generate electricity from biomass wastes. EAB are also expected to be applied to the production of valued chemicals in microbial electrosynthesis systems (MESs) with the supply of electric energy from electrodes. It is, therefore, important to deepen our understanding of EAB in terms of their physiology, genetics and genomics. Knowledge obtained in these studies will facilitate the engineering of EAB for developing more efficient biotechnology processes. In this article, we summarize current knowledge on Shewanella oneidensis MR-1, a representative EAB extensively studied in the laboratory. Studies have shown that catabolic activities of S. oneidensis MR-1 are well tuned for efficiently conserving energy under varied growth conditions, e.g., different electrode potentials, which would, however, in some cases, hamper its application to biotechnology processes. We suggest that understanding of molecular mechanisms underlying environmental sensing and catabolic regulation in EAB facilitates their biotechnological applications.
Similar content being viewed by others
Introduction
Electrochemically active bacteria (EAB) are capable of electric interactions with extracellularly located redox-active materials, such as electrodes (Chang et al. 2006; Sydow et al. 2014). Owing to their abilities to catalyze various intracellular catabolic reactions with the aid of electrodes, EAB serves as essential components in bioelectrochemical systems (BESs), including microbial fuel cells (MFCs) and microbial electrosynthesis systems (MESs) (Schröder et al. 2015). MFCs are devices that use EAB as catalysts for the oxidative conversion of fuels, including organic compounds, into electricity (Logan et al. 2006; Watanabe 2008), and are regarded as a promising technology for energy-saving wastewater treatment (Lefebvre et al. 2011; Yoshizawa et al. 2014). MESs are engineered systems in which EAB catalyze reductive reactions for producing valued chemicals using electricity as an energy source (Nevin et al. 2010; Rabaey and Rozendal 2010). The application of EAB in biotechnology, therefore, offers the potential to develop new technologies for producing energy and chemicals from sustainable resources, and extensive studies have been conducted to deepen our understanding on the physiology, genetics, and genomics of EAB towards their practical applications (for reviews, refer to Fredrickson et al. 2008; Lovley 2012; Sydow et al. 2014; Kouzuma et al. 2015, 2018).
Although many EAB have been isolated and characterized in the laboratory (Doyle and Marsili 2015; Kokko et al. 2016; Ueoka et al. 2018), our knowledge on molecular mechanisms underlying electrochemical interactions of EAB with electrodes has mostly been gained in studies using model EAB, including Shewanella oneidensis MR-1 and Geobacter sulfurreducens PCA (Sydow et al. 2014). These bacteria were originally isolated as dissimilatory metal-reducing bacteria that can respire using insoluble metal oxides as electron acceptors (Myers and Nealson 1988; Caccavo et al. 1994), and later studies have shown that they can also transfer electrons to electrodes and generate current in MFCs without adding electron-shuttle compounds (Kim et al. 1999; Bond and Lovley 2003). It has been discovered that these EAB intrinsically possess extracellular electron transfer (EET) pathways, consisting mainly of periplasmic and membrane-bound cytochromes, which serve as conduits between intracellular catabolic reactions and extracellular conductive materials (e.g., electrodes) (Shi et al. 2016). Comparative evaluation of activities of different EAB in same BESs is limited (Newton et al. 2009; Kato et al. 2013).
Shewanella oneidensis MR-1 is a facultative anaerobe belonging to the Gammaproteobacteria (Venkateswaran et al. 1999) and is one of the most extensively studied EAB due to its annotated genome sequence (Heidelberg et al. 2002) and ease of cultivation and genetic manipulation (Sydow et al. 2014). Genetic and biochemical studies have revealed that MR-1 has well-developed respiratory networks for efficiently discharging electrons to various organic and inorganic electron acceptors, e.g., oxygen, fumarate, nitrate, thiosulfate, trimethylamine N-oxide [TMAO], dimethyl sulfoxide [DMSO], and solid-phase electron acceptors, such as metal oxides and electrodes (Fredrickson et al. 2008; Sturm et al. 2015). Since MR-1 is also able to accept electrons from a low potential electrode through the EET pathway (Ross et al. 2011), this strain is expected to serve as a host for synthesizing valued chemicals in MES. In addition, studies have indicated that MR-1 has well-developed environment-sensing and regulatory systems (Kouzuma et al. 2015) that are used to modulate catabolic pathways and biofilm structures in response to environmental stimuli, including shifts in electrode potentials (Hirose et al. 2018; Kitayama et al. 2017). The abilities to sense and respond to electrode potentials are considered to be particularly important features of this bacterium, since underlying mechanisms can be used for the electrochemical control of bacterial metabolism in BESs. For these reasons, MR-1 is suitable as a model for investigating how EAB interact with electrodes and regulate their metabolism in BESs.
In this article, we summarize current knowledge on S. oneidensis MR-1 with particular focuses on its catabolic and electron-transport pathways, sensing and regulatory systems and biofilm structures. We also describe the metabolic engineering of this strain for future biotechnological applications. By summarizing results of these studies, we suggest that molecular-level understanding of this model bacterium provides information indispensable for engineering EAB for future use in sustainable biotechnology processes. The aim of this article is not to summarize current knowledge on MFCs and other BESs, for which readers are referred to other articles (e.g., Santoro et al. 2017; Li et al. 2018b; Kouzuma et al. 2018).
Catabolic and electron-transport pathways
The level of electric current generated by S. oneidensis MR-1 in BES is dependent on its catabolic activity. It has been known that MR-1 is able to utilize low molecular-weight organic compounds as substrates for the current generation, including lactate, pyruvate, formate, and N-acetylglucosamine (NAG) (Kouzuma et al. 2015). These substrates are completely oxidized to CO2 via the tricarboxylic acid (TCA) cycle under aerobic conditions, while they are partially oxidized to form acetate as the major end product under anaerobic conditions, including electricity-generating conditions (Kouzuma et al. 2015). Catabolic electron-transport pathways in MR-1 have extensively been studied (Fig. 1), and this chapter summarizes current knowledge on these pathways with focuses on their unique features.
Catabolic pathways in S. oneidensis MR-1. Growth substrates and end metabolites are highlighted with boxes, while missing steps are indicated with gray dashed arrows. *Enzymes whose gene expression is regulated by the Arc system (Gao et al. 2008); †enzymes whose gene expression is regulated by the cAMP/CRP system (Charania et al. 2009). NAG: N-acetylglucosamine; NAG-6P: N-acetylglucosamine-6-phosphate; KDG-6P: 2-keto-3-deoxygluconate 6-phosphate; PEP: phosphoenolpyruvate; Q: quinone; QH2: quinol; NagP: N-acetylglucosamine transporter; NagK: N-acetylglucosamine kinase; NagA: N-acetylglucosamine-6-phosphate deacetylase; NagB: glucosamine/fructose-6-phosphate aminotransferase; Fbp: fructose-1,6-bisphosphatase; Fba: fructose-bisphosphate aldolase; TpiA: triosephosphate isomerase; Pgi: glucose-6-phosphate isomerase; Zwf: glucose-6-phosphate 1-dehydrogenase; Edd: phosphogluconate dehydratase; Eda: 2-dehydro-3-deoxyphosphogluconate aldolase/(4S)-4-hydroxy-2-oxoglutarate aldolase; PpsA: pyruvate, water dikinase; PckA: phosphoenolpyruvate carboxykinase; Pdh: pyruvate dehydrogenase; GltA: citrate synthase; Acn: aconitate hydratase; Icd: isocitrate dehydrogenase; SucAB: 2-oxoglutarate dehydrogenase; SucCD: succinyl-CoA synthetase; Sdh: succinate dehydrogenase/fumarate reductase; Fum: fumarate reductase; Mdh: malate dehydrogenase; AceB: malate synthase; AceA: isocitrate lyase; Pta: phosphate acetyltransferase; AckA: acetate kinase; Acs: acetyl-CoA synthetase; Fdh: formate dehydrogenase; Dld: d-lactate dehydrogenase, quinone dependent; Lld: l-lactate dehydrogenase; LdhA: d-lactate dehydrogenase, NAD dependent; Glf: glucose uniporter; GalP: galactose:H+ symporter; Glk: glucokinase; Xks1: xylulokinase; Xyl2: xylitol dehydrogenase; Xyl1: NAD(P)H-dependent xylose reductase
MR-1 is unable to utilize glucose, the major starting material in many biotechnology processes (Naik et al. 2010), as a catabolic substrate due to the lack of functional glucose transporters (Nakagawa et al. 2015). The Embden–Meyerhof–Parnas (EMP) and Entner–Doudoroff (ED) pathways are the two known glucose catabolic pathways in organisms, while the ATP yield per mole of glucose in the EMP pathway (2 ATP) is higher than that in the ED pathway (1 ATP), indicating that the ED pathway is a low energy-yield process (Aristidou and Penttilä 2000). Genome analysis of MR-1 has revealed that 6-phosphofructokinase, an essential enzyme in the EMP pathway, is not present (Rodionov et al. 2011). In contrast, all genes necessary for reconstructing the ED pathway are identified in the MR-1 genome, and NAG is catabolized via the ED pathway. NAG is an intermediate metabolite in the chitin-degradation pathway in aquatic bacteria, and most strains in the genus Shewanella are able to grow on and generate current from NAG (Yang et al. 2006; Zhao et al. 2010). In addition, MR-1 has a gene (SO_4085) for chitinase A, a chitin-degradation enzyme, and generates current, albeit at a low level, with chitin as the sole substrate (Li et al. 2017b).
NagP (SO_3503) is a NAG permease, a transporter protein categorized into neither the phosphoenolpyruvate carbohydrate phosphotransferase system nor the ABC transporter (Yang et al. 2006). For transporting 1 mol of NAG, NagP consumes 1 mol of ATP and produces 1 mol of fructose-6P that is further catabolized in the ED pathway. Interestingly, it has been documented that aerobic cultivation of MR-1 in glucose-rich media promoted laboratory evolution for the acquisition of glucose-utilizing capacity (Howard et al. 2012). The whole genome analysis identified a mutation in transcriptional repressor NagR (SO_3516) in the evolved strain (Chubiz and Marx 2017). NagR regulates the expression of nagP and nagK, a gene encoding a NAG kinase (SO_3507), and the study has shown that the deletion of nagR promotes the expression of nagP and nagK and the catabolism of glucose. In addition, several studies employed bioengineering approaches (genetic supplementation of MR-1 with genes for glucose transporter and glucokinase) to construct glucose-utilizing strains from MR-1 (Choi et al. 2014; Nakagawa et al. 2015).
Xylose is abundantly present in nature as a major component of the plant cell wall (Scheller 2017), and some bacteria are able to catabolize xylose. The MR-1 genome encodes genes for xylose reductase (SO_0900) and xylulokinase (SO_4230) (Sekar et al. 2016), while a gene for xylose transporter is not present. Adaptive evolution is a powerful tool to confer a missing function on a bacterial strain, and a xylose-utilizing mutant was obtained from MR-1 after incubation in media containing a high concentration of xylose (Sekar et al. 2016). Whole genome sequencing revealed a single nucleotide mutation in a gene encoding unknown membrane protein (SO_1396), which resulted in the substitution of a glutamate residue to histidine and conferred the activity to bind to and transport xylose (Sekar et al. 2016). Li et al. (2017a) exploited a synthetic-biology approach (the introduction of genes for xylose facilitator, d-xylose reductase, xylitol dehydrogenase and d-xylulokinase from Candida intermedia and Scheffersomyces stipites) to construct an MR-1 derivative able to generate current form xylose.
Pyruvate is a major product of glycolysis and utilized in diverse catabolic and anabolic pathways, including gluconeogenesis, pyruvate fermentation, amino-acid biosynthesis and the TCA cycle. Among known enzymes for pyruvate oxidation, MR-1 has a pyruvate dehydrogenase complex (PDH, comprised of AceEF and LpdA, SO_0424 to SO_0426) and pyruvate formate lyase (Pfl, SO_2912). The PDH complex is composed of pyruvate dehydrogenase, dehydrolipoate acyltransferase and dihydrolipoate dehydrogenase and catalyzes NAD-dependent oxidative decarboxylation of pyruvate to form acetyl-CoA, CO2 and NADH mostly in association with aerobic respiration (Pinchuk et al. 2011). In contrast, Pfl catalyzes the oxidation of pyruvate to form acetyl-CoA and formate and is a key enzyme in diverse anaerobes (Knappe et al. 1984). Carbon-flux and deletion-mutant analyses showed that oxygen inactivates the Pfl-dependent pathway and facilitates the NAD-dependent oxidation of pyruvate to CO2 and acetyl-CoA (Tang et al. 2007; Pinchuk et al. 2011). PflA, a Pfl-activating enzyme encoded by SO_2913 is also known to be oxygen sensitive (Tang et al. 2007; Crain and Broderick 2014).
Formate is a key intermediate metabolite in anaerobic microbiota, including those in MFCs (Luo et al. 2016). In MR-1, formate transporter FocA (SO_2911) transfers formate from cytoplasm to periplasm, followed by the oxidation of formate by membrane-bound three-subunit formate dehydrogenases (FDHs), composed of a formate-catalytic α subunit, iron-sulfur β subunit and cytochrome b-containing γ subunit. The genome of MR-1 encodes three FDH genes, fdnGHI (SO_0101 to SO_0103), fdhABC1 (SO_4509 to SO_4511) and fdhABC2 (SO_4513 to SO_4515). An FDH complex oxidizes formate to CO2 at the periplasmic space with membrane quinones as electron acceptors and generates the proton motive force (PMF) (Kane et al. 2016). Although there is no well-established idea on how MR-1 differentially uses these FDHs, it has been known that Fdn does not work in minimal media supplemented with NAG and fumarate (Kane et al. 2016). Fdn in MR-1 is a homologue of nitrate-inducible formate dehydrogenase FDH-N in Escherichia coli (Jormakka et al. 2002), suggesting that nitrate reduction would induce the expression of fdnGHI. MR-1 may also have an NAD-dependent formate dehydrogenase (SO_3922), while it has been reported that this gene does not significantly affect catabolism in MR-1 (Li et al. 2018a). However, it has also been described that the introduction of NAD-dependent FDH in Moraxella sp. increases the current density in MFC (Mordkovich et al. 2013).
Diverse anaerobic bacteria have hydrogenases that catalyze reversible oxidation of hydrogen to proton (Wang et al. 2003; Das and Veziroglu 2008; Drake et al. 2008). In addition, it is known that catalytic and electron-transfer subunits of formate-hydrogen lyase (FHL), hydrogenase and formate dehydrogenase are linked to each other, enhancing the efficiency of electron transfer between hydrogen production and formate oxidation (Redwood et al. 2008; Mcdowall et al. 2014). Several reports have described that MR-1 utilizes hydrogen as an electron donor under fumarate, azo-dye and solid-metal reducing conditions and as an electron sink in the absence of electron acceptors (Liu et al. 2002; Meshulam-Simon et al. 2007; Le Laz et al. 2014). In MR-1, hydrogen production and consumption are catalyzed by two types of hydrogenase, namely, [Fe/Ni] hydrogenase (Hya, SO_2089 to SO_2099) and [Fe/Fe] hydrogenase (Hyd, SO_3920 to SO_3926), and these enzymes contribute to the maintenance of intracellular redox states and the conservation of energy coupled to the reducing-power recycling (Kreuzer et al. 2014). MR-1 is considered to differentially utilize these hydrogenases depending on the oxygen partial pressure (Kreuzer et al. 2014); Hya seems to be able to produce hydrogen under microaerobic and anaerobic conditions, while Hyd specifically and dominantly work under strict anaerobic conditions. On the other hand, hydrogen consumption is mainly catalyzed by Hya (Meshulam-Simon et al. 2007). In BESs, hydrogenases are considered to play important roles in discharging intracellular reducing equivalents as hydrogen and ingesting hydrogen as an energy source (Croese et al. 2014). It is known that hydrogen is produced at the surface of cathodic electrodes and serves as the reducing power for carbon fixation by autotrophs (Aulenta et al. 2008; Nevin et al. 2010). MR-1 is also able to utilize molecular hydrogen generated at a cathode, which promotes current generation and growth in BESs (Hirose et al. unpublished results).
Under electron-acceptor limitation, pyruvate is reduced to lactate by NADH-dependent fermentative lactate dehydrogenase (LDH) encoded by ldhA (SO_0968) (Pinchuk et al. 2011). Analyses of a glucose-utilizing derivative of MR-1 indicated that LdhA is involved in the lactate fermentation in glucose-amended media (Nakagawa et al. 2015). In contrast, for the oxidation of lactate, MR-1 utilizes stereospecific LDH, including l-LDH (LldEFG, SO_1520 to SO_1518) and d-LDH (Dld, SO_1521) (Pinchuk et al. 2009). Dld is an FAD-dependent bidirectional LDH that transfers electrons to the membrane quinone pool. This enzyme is co-expressed with a putative lactate permease encoded by lldP (SO_1522) (Kasai et al. 2017), indicating that the D-lactate uptake is tightly linked to the consumption by Dld. Lld is a three-subunit enzyme whose expression is regulated by transcriptional regulator LlpR (SO_3460) that is activated by l-lactate. In contrast, Dld is regulated by a cyclic AMP (cAMP)-dependent global regulatory system (Kasai et al. 2017). This may be because d-lactate is the major product of lactate fermentation in many fermentative bacteria (Brutinel and Gralnick 2012) and more commonly supplied to MR-1 than l-lactate.
Acetyl-CoA can be incorporated into the TCA cycle. In MR-1, the TCA cycle, which includes NADH-dependent redox enzymes, isocitrate dehydrogenase (icd, SO_1538), 2-oxoglutarate dehydrogenase (sucAB, SO_1930 to SO_1931), malate dehydrogenase (mdh, SO_0770), functions only under aerobic conditions. Transcriptomics, proteomics and metabolomics analyses showed that the level of 2-oxoglutarate dehydrogenase is significantly decreased under anaerobic conditions, resulting in an incomplete cycle (Tang et al. 2007; Grobbler et al. 2015; Hirose et al. 2018). The TCA cycle of MR-1 has a branching node at 2-oxoglutarate that is converted to malate and succinate by AceAB (SO_1483 to SO_1484) without producing NADH under reductive conditions (Kouzuma et al. 2015). The TCA cycle in MR-1 is, therefore, unable to oxidize acetyl-CoA to form CO2 under anaerobic conditions, resulting in the accumulation of acetate as the major end product (Kouzuma et al. 2015). In anaerobic cultures of MR-1, acetyl-CoA is mainly catalyzed by Pta (SO_2916) and AckA (SO_2915) to form acetate via acetyl phosphate with the production of ATP by the substrate-level phosphorylation (Hunt et al. 2010). In contrast, other EAB, including Geobacter spp. and Shewanella algae, are able to oxidize acetate to CO2 coupled to metal respiration, since the complete TCA cycle is operational in these bacteria even under anaerobic conditions (Bond and Lovley 2003; Szeinbaum et al. 2017).
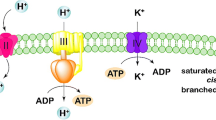
Reducing equivalents (NADH and quinols) generated in the catabolic pathways are transferred to respiratory electron acceptors via NADH dehydrogenases and electron-transfer networks (Fig. 2). MR-1 has three different types of NADH dehydrogenases (NDH); type1 NDH (Nuo, SO_1009 to SO_1021) that translocates two protons per electron, type II NDH (Ndh, SO_3517) that does not translocate proton, and two type III NDH (Nqr1, SO_0902 to SO_0907, and Nqr2, SO_1103 to SO_1108) that translocates one Na+ per electron. Most freshwater bacteria are known to possess proton-translocating NDHs, while marine bacteria possess Na+-translocating NDHs; on the other hand, bacteria that have three different types of NDHs are scarce (Melo et al. 2004). Although limited knowledge is available concerning the respective roles of these NDHs in MR-1, it has been reported that Nuo and Nqr2 preferentially work under aerobic conditions (Duhl et al. 2018). A recent study has also found that Nuo is preferentially utilized at high electrode potentials in BESs (Hirose et al. 2018).
Electron-transport pathways in S. oneidensis MR-1. Electron acceptors are marked with boxes. *Enzymes whose gene expression is regulated by the Arc system (Gao et al. 2008); †enzymes whose gene expression is regulated by the cAMP/CRP system (Charania et al. 2009). OM: outer membrane; IM: inner membrane; TMAO: trimethylamine N-oxide; TMA: trimethylamine; DMSO: dimethyl sulfoxide; DMS: dimethyl sulfide; Q: quinone; QH2: quinol; CytC oxidase: cytochrome c oxidase; TorA: trimethylamine-N-oxide reductase; TorC: trimethylamine-N-oxide reductase cytochrome c subunit; PsrABC: polysulfide reductase; SirACD: sulfite reductase; Hya: hydrogenase; LDH: lactate dehydrogenase; NDH: NADH dehydrogenase; FDH: formate dehydrogenase; CymA: membrane anchored tetraheme cytochrome c; NrfA: nitrite reductase; FccA: fumarate reductase flavoprotein subunit; NapA: periplasmic nitrate reductase; NapB: cytochrome c-type protein; DmsE: periplasmic decaheme cytochrome c; DmsA: anaerobic dimethyl sulfoxide reductase subunit A; DmsB: anaerobic dimethyl sulfoxide reductase subunit B; DmsF: extracellular dimethyl sulfoxide/manganese oxide reductase outer membrane translocase; OmcA: outer-membrane decaheme cytochrome c; MtrC: outer-membrane decaheme cytochrome c; MtrB: outer-membrane protein; MtrA: outer-membrane periplasmic decaheme cytochrome c component
MR-1 is able to utilize various organic and inorganic electron acceptors (e.g., oxygen, fumarate, nitrate, thiosulfate, TMAO, DMSO, solid metal oxides and electrodes) (Fredrickson et al. 2008; Sturm et al. 2015), and reconstructed electron-transfer networks are shown in Fig. 2. Membrane quinones (ubiquinones and menaquinones) are reduced by membrane-bound dehydrogenases, including NDHs, LDHs, and FDHs, and activities of these dehydrogenases are known to be affected by external redox states (Hirose et al. 2018). For instance, in the presence of low-potential electrodes (e.g., below 0 V vs. standard hydrogen electrode, SHE), electrons are mainly supplied to quinones by LDHs and FDHs, while NDH (Nuo) also contributes to the reduction of quinones in the presence of high-potential electrodes (e.g., above + 0.4 V) (Hirose et al. 2018).
Electrons in quinols are transferred to periplasmic electron-transport proteins (small c-type cytochromes, such as STC [SO_2727] and FccA [SO_0970]) via cytoplasmic membrane-bound cytochromes, including CymA (SO_4591), and subsequently transferred to terminal reductases in the periplasm and those bound to the cytoplasmic membrane (Marritt et al. 2012). MR-1 has a variety of terminal reductases, including fumarate reductase (FccA, SO_0970), nitrate reductase (Nap, SO_0845 to SO_0849), nitrite reductase (NrfA, SO_3980), TMAO reductase (SO_1232 to SO_1234), DMSO reductase (Dms, SO_1427 to SO_1432), cytochrome c oxidases (aa3 type [SO_4606 to SO_4609] and cbb3 type [SO_2361 to SO_2364]) and bd-type cytochrome oxidase [SO_3285 to SO_3286]). In addition, MR-1 has outer-membrane cytochrome complexes (mtrBAC/omcA [SO_1776 to SO_1779] and mtrFED [SO_1780 to SO_1782]) that transfer electrons to extracellular solid electron acceptors (the EET pathway), such as metal oxides and electrodes in BESs (Bretschger et al. 2007; Newton et al. 2009). These terminal reductases are considered to compete against each other on electrons, and it has been demonstrated that a double-knockout mutant of genes for cytochrome c oxidases (ΔSO_2364/SO_4607) generates more current in MFCs than the wild-type MR-1 (Kouzuma et al. 2012a). For efficient transfer of electrons to external solid electron acceptors, MR-1 secrets electron-shuttle compounds, such as water-soluble quinones (Newman and Kolter 2000) and riboflavin (Marsili et al. 2008).
As outlined above, extensive studies have been performed on enzymes and electron-transport proteins in S. oneidensis MR-1, and information thus obtained would serves as the basis for metabolic engineering of this strain for producing valued chemicals.
Sensing and regulatory systems
Studies have also reported sensing and regulatory systems that function in S. oneidensis MR-1. It has been known that, in MR-1, the expression of genes coding for catabolic and electron-transport pathways is mostly controlled by global transcriptional regulators, such as an anoxic redox-control two-component system (Arc-TCS) and a cyclic AMP (cAMP) receptor protein (CRP). In this section, we focus on the roles of these global regulators in transcriptional regulation of the catabolic and electron-transport pathways; those regulated by Arc-TCS and CRP are highlighted in Figs. 1 and 2.
Arc-TCS is extensively studied sensing systems that are known to regulate the expression of genes involved in the TCA cycle in enterobacteria, including E. coli (Georgellis et al. 2001). Arc-TCS in E. coli consists of sensor histidine kinase ArcB and response regulator ArcA. Among them, ArcB recognizes the redox state of ubiquinone and menaquinone in the cytoplasmic membrane and modifies its kinase activity (Georgellis et al. 2001). The kinase activity of ArcB is activated under reducing conditions, and ArcA is thus phosphorylated for regulating the expression of target genes. For instance, phosphorylated ArcA represses the expression of genes involved in the aerobic metabolism, such as the TCA cycle, and activates the expression of genes related to the anaerobic metabolism, such as acid fermentation (Iuchi and Lin 1988; Gunsalus and Park 1994).
In S. oneidensis MR-1, Arc-TCS is involved in regulating a variety of cellular functions. Different from Arc-TCS in E. coli, however, that in MR-1 is comprised of three components, namely, sensor histidine kinase ArcS (SO_0577), phosphotransferase HptA (SO_1327) and response regulator ArcA (SO_3988) (Lassak et al. 2010). ArcS, localized in the cytoplasmic membrane, is a complex protein that possesses a periplasmic calcium channel, chemotaxis receptor domain (CaChe-domain), histidine kinase domain, two transmembrane domains, two cytoplasmic Per Arnt Sim (PAS)-sensing domains, and two receiver domains (Lassak et al. 2013). ArcS senses the redox states of ubiquinone and/or menaquinone in the cytoplasmic membrane (Lassak et al. 2010; Hirose et al. 2018), and it has been suggested that the kinase activity of ArcS is activated, when cysteine residues in ArcS retain free thiol groups under reductive conditions (Lassak et al.2010, 2013). HptA functions as phosphotransferase and transfers a phosphoryl group between ArcS and ArcA (Lassak et al. 2010, 2013). ArcA, a response regulator, has an N-terminal receiver domain of a phosphoryl group and a C-terminal DNA-binding domain (Lassak et al. 2013). Phosphorylated-ArcA binds to transcriptional regulatory regions of its target genes and regulates their transcription (Gao et al. 2008). Target genes of Arc-TCS in MR-1 are different from those in E. coli (Gao et al. 2008; Lassak et al. 2010, 2013); transcription and proteome analyses have reported that expression levels of several anaerobic respiratory genes, such as nap, fccA, dms, mtrC, omcA and cymA, are decreased in an arcA-deletion mutant under anaerobic conditions (Gao et al. 2008; Yuan et al. 2012). However, it is likely that ArcA indirectly regulates the expression of omcA and mtrC, since ArcA-binding consensus sequences are not present in upstream regions of these genes (Gao et al. 2008). A recent study has reported that Arc-TCS in MR-1 senses electrode potentials via membrane quinones and directly or indirectly regulates the expression of many genes involved in energy conservation, e.g., PDH, Nuo NDH and ATP synthase (Hirose et al. 2018). Consequently, in the presence of a high-potential (e.g., + 0.4 V and above) electrode, MR-1 preferentially utilizes the NADH-dependent pathway rather than the formate-dependent pathway for intracellular electron transfer, resulting in the generation of a large PMF in the presence of a large electromotive force (Hirose et al. 2018).
CRP is also a well-known global transcriptional regulator that is distributed among diverse organisms, including bacteria. CRP activates or represses the transcription of target genes, when it binds to cAMP, a second messenger of particular importance (Botsford and Harman 1992; Shimada et al. 2011; Gancedo 2013). It has been suggested that the activity of CRP depends on intracellular levels of cAMP that is synthesized by adenylate cyclase (Cya) and degraded by cAMP phosphodiesterase (Cpd). The CRP regulatory system in E. coli has extensively been studied and is known to control the expression of genes involved in catabolic pathways, such as glycolysis (Botsford and Harman 1992; Hollands et al. 2007). In E. coli, the intracellular cAMP level is regulated by CyaA that synthesizes cAMP from ATP in the cytoplasm (Botsford and Harman 1992). In the absence of glucose, the catalytic activity of CyaA is activated, and the intracellular cAMP concentration is increased, resulting in the activation of CRP-dependent expression of β-galactosidase, a regulatory circuit known as catabolite repression (Botsford and Harman 1992).
Different from E. coli, S. oneidensis MR-1 uses CRP for regulating the expression of many genes for enzymes involved in the anaerobic respiration, such as fumarate, nitrate and nitrite reductases and the EET pathway (Saffarini et al. 2003; Charania et al. 2009; Kasai et al. 2015). It has been reported that a crp-deletion mutant exhibits the deficiency in anaerobic respiration with DMSO, fumarate, nitrate and ferric iron (Saffarini et al. 2003; Charania et al. 2009). The expression of genes encoding the Mtr proteins (OmcA, MtrC, MtrA and MtrB) that constitute the EET pathway in MR-1 is directly regulated by CRP and upregulated in response to the increase in the intracellular cAMP concentration (Kasai et al. 2015). Recently, it has also been found that the transcription of dld and lldP encoding d-lactate dehydrogenase and lactate permease, respectively, is also regulated by CRP (Kasai et al. 2017). The CRP-dependent coordinated expression of catabolic and respiratory pathways is considered to be beneficial for MR-1 to survive in nutrient-limited natural environments.
Based on the genome sequence, MR-1 is considered to possess three putative Cya (CyaA, CyaB and CyaC) (Fredrickson et al. 2008). A study has shown that, similar to the crp-deletion mutant, a cyaA/cyaC double-deletion mutant is unable to utilize several terminal electron acceptors, including DMSO, fumarate and ferric iron (Saffarini et al. 2003; Charania et al. 2009), and it has been considered that CyaA and CyaC are responsible for synthesizing cAMP and activating CRP under anaerobic conditions. A cyaC single-deletion mutant exhibited a partial deficiency in anaerobic respiration, whereas the loss of cyaA did not substantially affect the anaerobic growth (Charania et al. 2009). Additionally, several genes involved in the anaerobic respiration (e.g., dms and mtr) and flagellum biosynthesis are shown to be down-regulated in the cyaC-deletion mutant, suggesting that CyaC plays the central role in the activation of anaerobic respiration in MR-1 (Charania et al. 2009). Several studies have also investigated roles of the cAMP/CRP regulatory system in MR-1 by constructing deletion mutants for CpdA, the major cAMP phosphodiesterase in this organism. Yin et al. (2016) have reported that a cpdA-deletion mutant exhibits impaired aerobic growth due to reduced expression of cytochrome bd and cbb3 oxidase genes. CpdA in MR-1 is also considered to be involved in the regulation of amino acid metabolism in a cAMP-independent manner (Kasai et al. 2018). On the other hand, little is known about upper signaling cascades that regulate the activities of CyaC and CpdA, and further study is, therefore, necessary to understand the whole picture of the cAMP/CRP-regulatory system in MR-1.
In addition to Arc-TCS and CRP, electron transport regulator (EtrA) and ferric uptake regulator (Fur) appear to affect the expression of genes involved in the anaerobic respiration. EtrA is an analogue of fumarate nitrate reduction regulator (Fnr) in E. coli and directly responds to environmental oxygen levels using a 4Fe-4S cluster (Crack et al. 2004; Moore et al. 2006). It has been reported that the expression of anaerobic respiratory genes, including mtr, cymA, nap, fccA and dms, in MR-1 are decreased in an etrA-deletion mutant (Cruz-García et al. 2011). Furthermore, EtrA appears to regulate the expression of catabolic pathways, such as gluconeogenesis and the TCA cycle (Cruz-García et al. 2011).
Fur is also known to affect the expression of the mtr genes. Fur senses the intracellular iron level and regulates the expression of many genes involved in iron homeostasis and uptake (Griggs and Konisky 1989; Andrews et al. 2003). In E. coli, Fur acts as an iron-responsive repressor and represses the transcription of target genes by binding to a specific sequence in promoter regions of these genes under iron-rich conditions (de Lorenzo et al. 1987). In MR-1, a study has demonstrated that the transcription level of the mtr genes is decreased by iron depletion (Yang et al. 2009). In addition, it has also been reported that the transcription level of the mtr genes is decreased in a deletion mutant of a siderophore biosynthesis gene (SO_3030) under MnO2-reducing conditions (Kouzuma et al. 2012b). These findings suggest that the intracellular iron level is important for the expression of the mtr genes. Furthermore, the expression level of the mtr gene is decreased in a fur-deletion mutant, and a Fur-binding site is predicted in the upstream region of omcA (Wan et al. 2004). However, another study has also shown that the expression of these genes is responsive to iron, but not to Fur (Yang et al. 2008).
Taken together, it is shown that Arc and CRP play central roles in the regulation of the catabolic and electron-transport pathways in MR-1, whereas the expression of these genes is also considered to be under the control of complex regulatory networks. For instance, it has been reported that the transcription of ArcA and EtrA is influenced by one another. (Gao et al. 2010). Despite the importance of these global regulators, however, knowledge is limited as to how these regulators are controlled by upper signaling cascades. Further investigation is, therefore, needed to understand the whole regulatory networks that control the expression of catabolic and electron-transport pathways in MR-1.
Biofilm formation and electron transfer
Shewanella oneidensis MR-1 is well known for its ability to transfer electrons to extracellular solid electron acceptors, such as iron oxide and anodes (Kouzuma et al. 2015), and it has been shown that MR-1 forms biofilm on solid electron acceptors. This bacterium also forms biofilm on a cathode in MES (Borole et al. 2011), where MR-1 receives electrons. Since the formation of biofilm is considered to facilitate electric interactions of MR-1 with solid electron acceptors, it is important to understand factors regulating the structure, composition and activity of electrochemically active biofilm.
The structure of biofilm formed by S. oneidensis MR-1 has been investigated under various conditions. Under aerobic conditions, MR-1 forms biofilms with mushroom-like structures typically found for many bacteria, such as E. coli and Pseudomonas aeruginosa (Thormann et al. 2004). Using flow cells in combination with confocal laser-scanning microscopes (CLSMs), processes of biofilm formation have been observed under the flow of an aerobic medium, showing that, in the initial phase, MR-1 cells attach to solid surfaces and form micro-colonies (Thormann et al. 2004). MR-1 subsequently secretes extracellular polymeric substances (EPS) and form thick and uneven biofilms with mushroom-like protrusions (Thormann et al. 2004; Kitayama et al. 2017). When the flow of the medium is subsequently stopped, MR-1 cells rapidly detach from solid surfaces, and biofilm structures are collapsed. In this process, the decrease in oxygen concentrations in biofilms is considered to be the cue that is sensed via several global regulators, e.g., EtrA, Arc and CRP (Thormann et al. 2005).
Biofilm formed by MR-1 under anaerobic conditions has been investigated in BES during current generation. Bretschger et al. (2007) observed MR-1 cells attached onto an anode of a batch-mode MFC using a scanning electron microscope; unfortunately, however, the biofilm structure observed in that study is considered to be far different from intact ones, since it was observed under a vacuum condition after rigorous washing and drying. Recent studies have reported the development of optically accessible electrochemical flow cells (EFCs) and used them to observe electrode-attached cells at initial phases of biofilm formation (Stoeckl et al. 2016; Lu et al. 2017). Kitayama et al. (2017) also developed optically accessible EFCs and used them for observing the development and maturation of intact biofilms formed by green fluorescence protein (GFP)-expressing MR-1 on graphite electrodes under active current generation and medium-flow conditions. It has been reported by Kitayama et al. (2017) that, different from lumpy biofilms formed under aerobic conditions, mature current-generating biofilms formed on an electrode poised at + 0.4 V (vs. SHE) are flat, even, dense and thin (~ 10 µm in thickness) (Fig. 3). Interestingly, the structure of mature biofilm formed at an electrode potential of 0 V (vs. SHE) is entirely different from that at + 0.4 V, and it is rough and uneven, suggesting that the strength of electric interaction determines biofilm structure (Fig. 3). In addition, Kitayama et al. (2017) have found that, compared to biofilms formed under aerobic and 0 V conditions, + 0.4 V biofilm has unique electrochemical properties (as analyzed by cyclic voltammetry) and contains low levels of extracellular polysaccharides. It has also been reported that the structure and electrochemical property of current-generating biofilms formed under flow conditions are different from those under static conditions (Kitayama et al. 2017). These results illustrate the dynamic feature of electrochemically active biofilm formed by MR-1 that is tuned in response to changes in environmental conditions, including medium flow, electron acceptors and electrode potentials.
Structures of live biofilms formed on graphite plates under aerobic (a) and electricity-generating (b, c) conditions by GFP-expressing S. oneidensis MR-1 in the presence of medium flow. For current generation, potentials of graphite-plate electrodes were poised at 0 V (b) and + 0.4 V (c) vs. Ag/AgCl reference electrodes. Top images are XZ projections, while bottom images are XY projections
In electrochemically active biofilms, MR-1 transfers electrons to electrodes via direct and mediated electron transfer (DET and MET, respectively) mechanisms (Choi et al. 2018). In the case of DET, MR-1 cells directly attach onto electrode surfaces using outer membrane c-type cytochrome, such as OmcA and MtrC (Shi et al. 2007). A study has shown that MR-1 produces nanowire-like extensions of its outer membrane containing OmcA and MtrCAB for long-distance DET (Pirbadian et al. 2014). Such membrane extensions are associated with membrane vesicles that are known to be produced by diverse Gram-negative bacteria (Zhou et al. 1998; Gorby et al. 2008; Schwechheimer and Kuehn 2015). It has been reported that MR-1 produces membrane vesicles under electron acceptor-limited (anaerobic) conditions, and they are assembled into filamentous structures to form membrane extensions (Pirbadian et al. 2014). Membrane extensions and membrane vesicles are considered to contribute to electron transfer to distant solid electron acceptors. MR-1 is also able to transfer electrons to solid electron acceptors via the MET mechanism with secreted electron-mediator compounds, such as riboflavin (Marsili et al. 2008). This mechanism is considered to work in mature electrochemically active biofilm on electrodes particularly for transferring electrons from top-layer cells to electrodes (Xiao et al. 2017; Choi et al. 2018). A recent report has described that EPSs help retain riboflavin in biofilms (Xiao et al. 2017). These observations suggest that the biofilm structure is important for MR-1 to efficiently transfer electrons to solid electron acceptors, such as electrodes.
It is also important to understand how S. oneidensis MR-1 regulates the formation of electrochemically active biofilms. It has been known that 3′5′-cyclic diguanosine monophosphate (c-di-GMP) is a second messenger that is involved in the regulation of biofilm formation in diverse bacteria (Jenal et al. 2017). Studies on several bacteria have also shown that c-di-GMP is also used for regulating a variety of cellular processes, including virulence formation, flagellum synthesis, and cell detachment (Hengge et al. 2016). In bacteria, c-di-GMP is synthesized by diguanylate cyclase (DGC) with the GGDEF domain (Paul et al. 2004), while it is degraded by phosphodiesterase (PDE) with the EAL and/or HD-GYP domains (Christen et al. 2005; Ryan et al. 2006). DGC synthesizes c-di-GMP from two GTPs (Paul et al. 2004). PDEs are classified into two types based on the catalyst domains. PDE with EAL degrades c-di-GMP to pGpG (Christen et al. 2005), and pGpG is subsequently degraded to GMP by oligoribonuclease (Orr et al. 2015). On the other hand, PDE with the EAL and HD-GYP domains directory degrades c-di-GMP to GMP (Ryan et al. 2006). When the intracellular c-di-GMP concentration is increased, cells start to form biofilm and repress motility (Thormann et al. 2006). In the KEGG database, S. oneidensis MR-1 has 43 DGCs and 23 PDEs, including bifunctional proteins (http://www.genome.jp/kegg-bin/show_organism?org=son). Thormann et al. (2006) have also reported that MxdA (SO_4180), one of DGCs, promotes biofilm formation, while Chao et al. (2013) have shown that PdeB (SO_0437), a bifunctional protein with the GGDEF and EAL domains, represses biofilm formation by degrading c-di-GMP. These results suggest that DGCs and PDEs are important for biofilm formation by MR-1, whereas functions of other proteins remain unclear.
Another study has shown that MxdB, a glycosyltransferase known as an extracellular polymeric saccharides production protein, promotes EPS production and biofilm formation under aerobic conditions in flow cells (Thormann et al. 2006). In addition, adhesive proteins are necessary for bacterial cells to attach onto solid surfaces and form biofilms. AggA (SO_4320) is an agglutination protein in MR-1 and an aggA-deletion mutant markedly is deficient in biofilm formation (de Vriendt et al. 2005). Zhou et al. (2015) have also found that biofilm promoting-factor proteins (BpfA, BpfD and BpfG; SO_4317, SO_4322 and SO_4323) are key proteins for cell attachment and biofilm formation; in particular, the disruption of bpfA results in almost complete loss of biofilm-forming ability. BpfA is regulated by flagella master regulator FlrA that responds to the intracellular c-di-GMP concentration in S. putrefaciens (Cheng et al. 2017). These adhesive and agglutination proteins are thus identified as key factors for biofilm formation.
It has also been reported that type IV pili play essential roles in the attachment of cells to solid surfaces (Thormann et al. 2004). In that study, deletion mutants for the pilT and mshA genes (SO_3351 and SO_4105, respectively), which are known to be involved in the twitching motility and pili biosynthesis, respectively, were deficient in the early attachment to solid surfaces (Thormann et al. 2004). In contrast, a disruption mutant for FlrA, a master regulator of flagellum biosynthesis, is shown to exhibit promoted biofilm formation (Cheng et al. 2017). In relation to this finding, it is interesting that an FlrA homologue, FleQ, in P. aeruginosa is regulated in a c-di-GMP-dependent manner (Hickman and Harwood 2008). Finally, it has been documented that extracellular DNA (eDNA) serves as a structural component in all stages of the formation of aerobic biofilms under static and hydrodynamic conditions (Gödeke et al. 2011). In addition, that study has also shown that eDNA is released by cell lysis mediated by the activation of prophages in the genome of MR-1 (Gödeke et al. 2011). It would be interesting to examine if eDNA is also involved in electrochemical activities of MR-1 biofilms.
Metabolic engineering in combination with electrochemistry
Metabolic engineering of EAB, e.g., S. oneidensis MR-1, has been examined mainly for the purpose of producing valued chemicals in MES with the aid of electric energy (Kracke et al. 2018). On the other hand, since electricity generation in MFCs has mostly been attempted in association with the treatment of organic wastes, naturally occurring microbiomes rather than pure cultures of EAB have been exploited in MFCs. Limited studies have, therefore, been conducted for metabolic engineering of EAB used for electricity generation (Choi et al. 2014; Nakagawa et al. 2015; Li et al. 2017a).
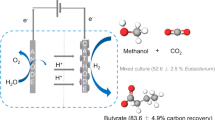
In some studies, MES is definitively used to denote process, in which carbon dioxide is fixed to produce organic compounds with electricity as the reducing power (Kracke et al. 2018), while processes, in which electricity is supplied for increasing the efficiency of bio-production of chemicals from organic feedstock, are termed electro-fermentation (EF) (Moscoviz et al. 2016). Since S. oneidensis MR-1 is incapable of CO2 fixation, studies have exploited this EAB for EF processes with electrodes either as electron acceptors or donors (Kracke et al. 2018).
Several studies have shown that electrodes (anodes) can be used to recover excess electrons generated in fermentation pathways, thereby improving fermentation efficiencies (termed unbalanced fermentation) (Liu et al. 2013). S. oneidensis MR-1 is unable to grow fermentatively in the absence of electron acceptors, and it has been suggested that an excessively reduced quinone pool exerts inhibitory effects on the catabolic pathway (Kane et al. 2016). This idea is considered to be related to the recent finding that Arc-TCS senses the redox state of membrane quinones for regulating the expression of key catabolic enzymes, including Nuo (Hirose et al. 2018); the down-regulation of Nuo may result in the accumulation of NADH and stagnation of fermentative catabolic pathways. Flynn et al. (2010) have demonstrated that the amount of ethanol produced from glycerol by an engineered MR-1 can be increased in BES, in which electrons generated by glycerol oxidation are discharged to an electrode. Anode-assisted EF was also examined for the production of acetoin from lactate by an engineered MR-1, in which codon-optimized genes for acetolactate synthase and acetolactate decarboxylase in Bacillus subtilis were introduced (Bursac et al. 2017). These studies show that electric interactions with anodes facilitate the discharge of intracellular excess electrons, thereby enabling fermentative production of target compounds by engineered MR-1 derivatives.
Studies have also attempted to utilize cathodic electron flows for MES (Kracke et al. 2018), while most of them have produced acetate from CO2 using non-engineered acetogens, and only limited studies have exploited metabolic engineering in combination with MES (Torella et al. 2015). It should also be noted that the electron transfer from cathodes to acetogens is considered to be mediated by hydrogen, but not by direct electron transfer. In contrast, MR-1 is also able to receive electrons from a cathode, and it has been demonstrated that the EET pathway is necessary for this electron transfer (Ross et al. 2011). This is not surprising, since potential windows of EET components substantially overlap with each other (Firer-Sherwood et al. 2008), and electrons are thus transferred from a low-potential electrode to membrane quinones via the EET pathway. A recent study has also shown that MR-1 is able to generate PMF by transferring electrons supplied from an electrode to oxygen as a respiratory electron acceptor, resulting in the production of ATP and NADH (Rowe et al. 2018). Several studies have examined engineered MR-1 for bioproduction in the presence of cathodes (Jeon et al. 2015, 2018). For instance, it has been demonstrated that isobutanol is produced by engineered MR-1 with NAG, pyruvate and lactate as mixed substrates under microaerobic conditions (approximately 10 mg l−1 within 48 h), and the production was stimulated in BES by supplying electrons to the EAB (Jeon et al. 2015). This study suggests that the cathodic electron supply would be a possible option for enhancing a yield of production in an EF process.
Despite laboratory demonstrations for anode- and cathode-assisted EF processes, it should be noted that production yields and rates reported in these EF studies are much lower than those for conventional fermentative processes, and further studies are, therefore, needed for EF processes to be practically used for bioproduction. As summarized elsewhere (Kracke et al. 2018), production rates reported for EF processes are roughly less than one-tenth compared to those for conventional fermentation processes. We suggest that one reason for low production rates in the EF process would be unfavorable regulation of catabolic pathways in EAB. For instance, it has been reported that some important catabolic enzymes, e.g., Nuo, PDH and ATPase, in MR-1 are down-regulated under reduced conditions and in the presence of a low-potential electrode (Hirose et al. 2018). It is likely that MR-1 has acquired such regulatory systems to efficiently utilize limited nutrients and survive in the natural environment, a situation substantially different from that in a nutrient-rich fermentation tank. It is, therefore, suggested that the engineering of sensing and regulatory systems is also necessary for achieving a high production yield with EAB.
In addition, studies have proposed genetic approaches to improve electrochemical activities of EAB (Kouzuma et al. 2010; Zheng et al. 2015; Yu et al. 2018). For instance, Yu et al. (2018) have shown that a knockout mutant of Pseudomonas aeruginosa for a gene encoding sigma factor RpoS exhibits improved electrochemical activity and biofilm formation. Such approaches may also be applied to S. oneidensis MR-1 for improving its electrochemical activity. Besides, Kouzuma et al. (2010) have shown that the EET activity of MR-1 can be enhanced by manipulating cell-surface properties, in which the disruption of a gene involved in the cell surface polysaccharide biosynthesis results in hydrophobic cell surface and enhanced EET activity. These approaches can be used in association with metabolic engineering to construct an engineered EAB with high production efficiencies.
Conclusions
Based on extensive knowledge on catabolic and regulatory systems, studies have attempted to engineer metabolic pathways in MR-1 for its application to electrode-assisted production of valued chemicals. Although successful laboratory examples have been reported, production efficiencies reported in these studies are much lower than those reported for conventional fermentation processes (Kracke et al. 2018). This would be attributable to the facts that MR-1 was isolated from lake sediment, and its metabolic pathways are well tuned for surviving in nutrient-limited natural environments (Hirose et al. 2018). It is, therefore, suggested that, to improve production efficiencies, further studies are necessary for deepening our understanding of sensing and regulatory systems in MR-1 and developing an engineering platform for controlling the expression of desired metabolic pathways. In addition, since the biofilm structure is the key for efficient electron transfer between cells and an electrode, we need to further our understanding on molecular mechanisms for biofilm development and develop methodological bases for engineering biofilms of EAB. These studies will open up novel biotechnology for the production of valued chemicals that are impossible to be produced by the conventional biotechnology.
Abbreviations
- EAB:
-
electrochemically active bacteria
- BES:
-
bioelectrochemical system
- MFC:
-
microbial fuel cell
- MES:
-
microbial electrosynthesis system
- EF:
-
electro-fermentation
- EET:
-
extracellular electron transfer
- NAG:
-
N-acetylglucosamine
- TCA:
-
tricarboxylic acid
- EMP:
-
Embden–Meyerhof–Parnas
- ED:
-
Entner–Doudoroff
- NAG-6P:
-
N-acetylglucosamine-6-phosphate
- KDG-6P:
-
2-keto-3-deoxygluconate 6-phosphate
- PEP:
-
phosphoenolpyruvate
- Q:
-
quinone
- QH2:
-
quinol
- NagP:
-
N-acetylglucosamine transporter
- NagK:
-
N-acetylglucosamine kinase
- NagA:
-
N-acetylglucosamine-6-phosphate deacetylase
- NagB:
-
glucosamine/fructose-6-phosphate aminotransferase
- Fbp:
-
fructose-1,6-bisphosphatase
- Fba:
-
fructose-bisphosphate aldolase
- TpiA:
-
triosephosphate isomerase
- Pgi:
-
glucose-6-phosphate isomerase
- Zwf:
-
glucose-6-phosphate 1-dehydrogenase
- Edd:
-
phosphogluconate dehydratase
- Eda:
-
2-dehydro-3-deoxyphosphogluconate aldolase/(4S)-4-hydroxy-2-oxoglutarate aldolase
- PpsA:
-
pyruvate, water dikinase
- PckA:
-
phosphoenolpyruvate carboxykinase
- Pdh:
-
pyruvate dehydrogenase
- GltA:
-
citrate synthase
- Acn:
-
aconitate hydratase
- Icd:
-
isocitrate dehydrogenase
- SucAB:
-
2-oxoglutarate dehydrogenase
- SucCD:
-
succinyl-CoA synthetase
- Sdh:
-
succinate dehydrogenase/fumarate reductase
- Fum:
-
fumarate reductase
- Mdh:
-
malate dehydrogenase
- AceB:
-
malate synthase
- AceA:
-
isocitrate lyase
- Pta:
-
phosphate acetyltransferase
- AckA:
-
acetate kinase
- Acs:
-
acetyl-CoA synthetase
- Dld:
-
d-lactate dehydrogenase, quinone dependent
- Lld:
-
l-lactate dehydrogenase
- LdhA:
-
d-lactate dehydrogenase, NAD dependent
- Glf:
-
glucose uniporter
- GalP:
-
galactose:H+ symporter
- Glk:
-
glucokinase
- Xks1:
-
xylulokinase
- Xyl2:
-
xylitol dehydrogenase
- Xyl1:
-
NAD(P)H-dependent xylose reductase
- OM:
-
outer membrane
- IM:
-
inner membrane
- TMAO:
-
trimethylamine N-oxide
- TMA:
-
trimethylamine
- DMSO:
-
dimethyl sulfoxide
- DMS:
-
dimethyl sulfide
- CytC oxidase:
-
cytochrome c oxidase
- TorA:
-
trimethylamine-N-oxide reductase
- TorC:
-
trimethylamine-N-oxide reductase cytochrome c subunit
- PsrABC:
-
polysulfide reductase
- SirACD:
-
sulfite reductase
- Hya:
-
hydrogenase
- LDH:
-
lactate dehydrogenase
- NDH:
-
NADH dehydrogenase
- FDH:
-
formate dehydrogenase
References
Andrews SC, Robinson AK, Rodríguez-Quiñones F (2003) Bacterial iron homeostasis. FEMS Microbiol Rev 27:215–237
Aristidou A, Penttilä M (2000) Metabolic engineering applications to renewable resource utilization. Cur Opin Biotechnol 11:187–198
Aulenta F, Reale P, Catervi A, Panero S, Reale P, Rossetti S (2008) Kinetics of trichloroethene dechlorination and methane formation by a mixed anaerobic culture in a bio-electrochemical system. Electrochim Acta 53:5300–5305
Bond DR, Lovley DR (2003) Electricity production by Geobacter sulfurreducens attached to electrodes. Appl Environ Microbiol 69:1548–1555
Borole AP, Reguera G, Ringeisen B, Wang ZW, Feng Y, Kim BH (2011) Electroactive biofilms: current status and future research needs. Energy Environ Sci 4:4813–4834
Botsford JL, Harman JG (1992) Cyclic AMP in prokaryotes. Microbiol Rev 56:100–122
Bretschger O, Obraztsova A, Sturm CA, Chang IS, Gorby YA, Reed SB, Culley DE, Reardon CL, Barua S, Romine MF, Zhou J, Beliaev AS, Bouhenni R, Saffarini D, Mansfeld F, Kim BH, Fredrickson JK, Nealson KH (2007) Current production and metal oxide reduction by Shewanella oneidensis MR-1wild type and mutants. Appl Environ Microbiol 73:7003–7012
Brutinel ED, Gralnick JA (2012) Preferential utilization of d-Lactate by Shewanella oneidensis. Appl Environ Microbiol 78:8474–8476
Bursac T, Gralnick JA, Gescher J (2017) Acetoin production via unbalanced fermentation in Shewanella oneidensis. Biotechnol Bioeng 114:1283–1289
Caccavo F, Lonergan DJ, Lovley DR, Davis M, Stolz JF, McInerney MJ (1994) Geobacter sulfurreducens sp. nov., a hydrogen- and acetate-oxidizing dissimilatory metal-reducing microorganism. Appl Environ Microbiol 60:3752–3759
Chang IS, Moon H, Bretschger O, Jang JK, Park HI, Nealson KH, Kim BH (2006) Electrochemically active bacteria (EAB) and mediator-less microbial fuel cells. J Microbiol Biotechnol 16:163–177
Chao L, Rakshe S, Leff M, Spormann AM (2013) PdeB, a cyclic di-GMP-specific phosphodiesterase that regulates Shewanella oneidensis MR-1 motility and biofilm formation. J Bacteriol 195:3827–3833
Charania MA, Brockman KL, Zhang Y, Banerjee A, Pinchuk GE, Fredrickson JK, Beliaev AS, Saffarini DA (2009) Involvement of a membrane-bound class III adenylate cyclase in regulation of anaerobic respiration in Shewanella oneidensis MR-1. J Bacteriol 191:4298–4306
Cheng YY, Wu C, Wu JY, Jia HL, Wang MY, Wang HY, Zou SM, Sun RR, Jia R, Xiao YZ (2017) FlrA represses transcription of the biofilm-associated bpfA operon in Shewanella putrefaciens. Appl Environ Microbiol 83:e02410–16
Choi D, Lee SB, Kim S, Min B, Choi IG, Chang IS (2014) Metabolically engineered glucose-utilizing Shewanella strains under anaerobic conditions. Bioresour Technol 154:59–66
Choi S, Kim B, Chang IS (2018) Tracking of Shewanella oneidensis MR-1 biofilm formation of a microbial electrochemical system via differential pulse voltammetry. Bioresour Technol 254:357–361
Christen M, Christen B, Folcher M, Schauerte A, Jenal U (2005) Identification and characterization of a cyclic di-GMP-specific phosphodiesterase and its allosteric control by GTP. J Biol Chem 280:30829–30837
Chubiz LM, Marx CJ (2017) Growth trade-offs accompany the emergence of glycolytic metabolism in Shewanella oneidensis MR-1. J Bacteriol 199:e00827-16
Crack J, Green J, Thomson AJ (2004) Mechanism of oxygen sensing by the bacterial transcription factor fumarate–nitrate reduction (FNR). J Biol Chem 279:9278–9286
Crain AV, Broderick JB (2014) Pyruvate formate-lyase and its activation by pyruvate formate-lyase activating enzyme. J Biol Chem 289:5723–5729
Croese E, Jeremiasse AW, Marshall IP, Spormann AM, Euverink GJ, Geelhoed JS, Stams AJ, Plugge CM (2014) Influence of setup and carbon source on the bacterial community of biocathodes in microbial electrolysis cells. Enzyme Microb Technol 61:67–75
Cruz-García C, Murray AE, Rodrigues JL, Gralnick JA, McCue LA, Romine MF, Löffler FE, Tiedje JM (2011) Fnr (EtrA) acts as a fine-tuning regulator of anaerobic metabolism in Shewanella oneidensis MR-1. BMC Microbiol 11:64
Das D, Veziroglu TN (2008) Advances in biological hydrogen production processes. Int J Hydrog Energy 33:6046–6057
De Lorenzo V, Wee S, Herrero M, Neilands JB (1987) Operator sequences of the aerobactin operon of plasmid ColV-K30 binding the ferric uptake regulation (fur) repressor. J Bacteriol 169:2624–2630
De Vriendt K, Theunissen S, Carpentier W, De Smet L, Devreese B, Van Beeumen J (2005) Proteomics of Shewanella oneidensis MR-1 biofilm reveals differentially expressed proteins, including AggA and RibB. Proteomics 5:1308–1316
Doyle LE, Marsili E (2015) Methods for enrichment of novel electrochemically-active microorganisms. Bioresour Technol 195:273–282
Drake HL, Gößner AS, Daniel SL (2008) Old acetogens, new light. Ann N Y Acad Sci 1125:100–128
Duhl KL, Tefft NM, TerAvest MA (2018) Shewanella oneidensis MR-1 utilizes both sodium- and proton-pumping NADH dehydrogenases during aerobic growth. Appl Environ Microbiol 84:e00415-18
Firer-Sherwood M, Pulcu GS, Elliott SJ (2008) Electrochemical interrogations of the Mtr cytochromes from Shewanella: opening a potential window. J Biol Inorg Chem 13:849–854
Flynn JM, Ross DE, Hunt KA, Bond DR, Gralnick JA (2010) Enabling unbalanced fermentations by using engineered electrode-interfaced bacteria. MBio 1:e00190-10
Fredrickson JK, Romine MF, Beliaev AS, Auchtung JM, Driscoll ME, Gardner TS, Nealson KH, Osterman AL, Pinchuk G, Reed JL, Rodionov DA, Rodrigues JL, Saffarini DA, Serres MH, Spormann AM, Zhulin IB, Tiedje JM (2008) Towards environmental systems biology of Shewanella. Nat Rev Microbiol 6:592–603
Gancedo JM (2013) Biological roles of cAMP: variations on a theme in the different kingdoms of life. Biol Rev Camb Philos Soc 88:645–668
Gao H, Wang X, Yang ZK, Palzkill T, Zhou J (2008) Probing regulon of ArcA in Shewanella oneidensis MR-1 by integrated genomic analyses. BMC Genom 9:42
Gao H, Wang X, Yang ZK, Chen J, Liang Y, Chen H, Palzkill T, Zhou J (2010) Physiological roles of ArcA, Crp, and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. PLoS ONE 5:e15295
Georgellis D, Kwon O, Lin EC (2001) Quinones as the redox signal for the arc two-component system of bacteria. Science 292:2314–2316
Gödeke J, Paul K, Lassak J, Thormann KM (2011) Phage-induced lysis enhances biofilm formation in Shewanella oneidensis MR-1. ISME J 5:613–626
Gorby Y, Mclean J, Korenevsky A, Rosso K, El-Naggar MY, Beveridge TJ (2008) Redox-reactive membrane vesicles produced by Shewanella. Geobiology 6:232–241
Griggs DW, Konisky J (1989) Mechanism for iron-regulated transcription of the Escherichia coli cir gene: metal-dependent binding of Fur protein to the promoters. J Bacteriol 171:1048–1054
Grobbler C, Virdis B, Nouwens A, Harnisch F, Rabaey K, Bond PL (2015) Use of SWATH mass spectrometry for quantitative proteomic investigation of Shewanella oneidensis MR-1 biofilms grown on graphite cloth electrodes. Syst Appl Microbiol 38:135–139
Gunsalus RP, Park SJ (1994) Aerobic-anaerobic gene regulation in Escherichia coli: control by the ArcAB and Fnr regulons. Res Microbiol 145:437–450
Heidelberg JF, Paulsen IT, Nelson KE, Gaidos EJ, Nelson WC, Read TD, Eisen JA, Seshadri R, Ward N, Methe B, Clayton RA, Meyer T, Tsapin A, Scott J, Beanan M, Brinkac L, Daugherty S, DeBoy RT, Dodson RJ, Durkin AS, Haft DH, Kolonay JF, Madupu R, Peterson JD, Umayam LA, White O, Wolf AM, Vamathevan J, Weidman J, Impraim M, Lee K, Berry K, Lee C, Mueller J, Khouri H, Gill J, Utterback TR, McDonald LA, Feldblyum TV, Smith HO, Venter JC, Nealson KH, Fraser CM (2002) Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat Biotechnol 20:1118–1123
Hengge R, Gründling A, Jenal U, Ryan R, Yildiz F (2016) Bacterial signal transduction by cyclic di-GMP and other nucleotide second messengers. J Bacteriol 198:15–26
Hickman JW, Harwood CS (2008) Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol Microbiol 69:376–389
Hirose A, Kasai T, Aoki M, Umemura T, Watanabe K, Kouzuma A (2018) Electrochemically active bacteria sense electrode potentials for regulating catabolic pathways. Nat Commun 9:1083
Hollands K, Busby SJW, Lloyd GS (2007) New targets for the cyclic AMP receptor protein in the Escherichia coli K-12 genome. FEMS Microbiol Lett 274:89–94
Howard EC, Hamdan LJ, Lizewski SE, Ringeisen BR (2012) High frequency of glucose-utilizing mutants in Shewanella oneidensis MR-1. FEMS Microbiol Lett 327:9–14
Hunt KA, Flynn JM, Naranjo B, Shikhare ID, Gralnick JA (2010) Substrate-level phosphorylation is the primary source of energy conservation during anaerobic respiration of Shewanella oneidensis strain MR-1. J Bacteriol 192:3345–3351
Iuchi S, Lin EC (1988) arcA, a global regulatory gene in Escherichia coli mediating repression of enzymes in aerobic pathways. Proc Natl Acad Sci USA 85:1888–1892
Jenal U, Reinders A, Lori C (2017) Cyclic di-GMP: second messenger extraordinaire. Nat Rev Microbiol 15:271–284
Jeon JM, Park H, Seo HM, Kim JH, Bhatia SK, Sathiyanarayanan G, Song HS, Park SH, Choi KY, Sang BI, Yang YH (2015) Isobutanol production from an engineered Shewanella oneidensis MR-1. Bioprocess Biosyst Eng 38:2147–2154
Jeon JM, Song HS, Lee DG, Hong JW, Hong YG, Moon YM, Bhatia SK, Yoon JJ, Kim W, Yang YH (2018) Butyrate-based n-butanol production from an engineered Shewanella oneidensis MR-1. Bioprocess Biosyst Eng 41:1195–1204
Jormakka M, Törnroth S, Byrne B, Iwata S (2002) Molecular basis of proton motive force generation: structure of formate dehydrogenase-N. Science 295:1863–1868
Kane AL, Brutinel ED, Joo H, Maysonet R, VanDrisse CM, Kotloski NJ, Gralnick JA (2016) Formate metabolism in Shewanella oneidensis generates proton motive force and prevents growth without an electron acceptor. J Bacteriol 198:1337–1346
Kasai T, Kouzuma A, Nojiri H, Watanabe K (2015) Transcriptional mechanisms for differential expression of outer membrane cytochrome genes omcA and mtrC in Shewanella oneidensis MR-1. BMC Microbiol 15:68
Kasai T, Kouzuma A, Watanabe K (2017) CRP regulates d-lactate oxidation in Shewanella oneidensis MR-1. Front Microbiol 8:869
Kasai T, Kouzuma A, Watanabe K (2018) CpdA is involved in amino acid metabolism in Shewanella oneidensis MR-1. Biosci Biotechnol Biochem 82:166–172
Kato S, Hashimoto K, Watanabe K (2013) Iron-oxide minerals affect extracellular electron-transfer paths of Geobacter spp. Microbes Environ 28:141–148
Kim BH, Kim HJ, Hyun MS, Park DH (1999) Direct electrode reaction of Fe(III)-reducing bacterium, Shewanella putrefaciens. J Microbiol Biotechnol 9:127–131
Kitayama M, Koga R, Kasai T, Kouzuma A, Watanabe K (2017) Structures, compositions, and activities of live Shewanella biofilms formed on graphite electrodes in electrochemical flow cells. Appl Environ Microbiol 83:166–172
Knappe J, Neugebauer FA, Blaschkowski HP, Gänzler M (1984) Post-translational activation introduces a free radical into pyruvate formate-lyase. Proc Natl Acad Sci USA 81:1332–1335
Kokko ME, Mäkinen AE, Puhakka JA (2016) Anaerobes in bioelectrochemical systems. Adv Biochem Eng Biotechnol 156:263–292
Kouzuma A, Meng XY, Kimura N, Hashimoto K, Watanabe K (2010) Disruption of the putative cell surface polysaccharide biosynthesis gene SO3177 in Shewanella oneidensis MR-1 enhances adhesion to electrodes and current generation in microbial fuel cells. Appl Environ Microbiol 76:4151–4157
Kouzuma A, Hashimoto K, Watanabe K (2012a) Influences of aerobic respiration on current generation by Shewanella oneidensis MR-1 in single-chamber microbial fuel cells. Biosci Biotechnol Biochem 76:270–275
Kouzuma A, Hashimoto K, Watanabe K (2012b) Roles of siderophore in manganese-oxide reduction by Shewanella oneidensis MR-1. FEMS Microbiol Lett 326:91–98
Kouzuma A, Kasai T, Hirose A, Watanabe K (2015) Catabolic and regulatory systems in Shewanella oneidensis MR-1 involved in electricity generation in microbial fuel cells. Front Microbiol 6:609
Kouzuma A, Ishii S, Watanabe K (2018) Metagenomic insights into the ecology and physiology of microbes in bioelectrochemical systems. Bioresour Technol 255:302–307
Kracke F, Lai B, Yu S, Krömer JO (2018) Balancing cellular redox metabolism in microbial electrosynthesis and electro fermentation—a chance for metabolic engineering. Metab Eng 45:109–120
Kreuzer HW, Hill EA, Moran JJ, Bartholomew RA, Yang H, Hegg EL (2014) Contributions of the [NiFe]- and [FeFe]-hydrogenase to H2 production in Shewanella oneidensis MR-1 as revealed by isotope ratio analysis of evolved H2. FEMS Microbiol Lett 352:18–24
Lassak J, Henche A-L, Binnenkade L, Thormann KM (2010) ArcS, the cognate sensor kinase in an atypical Arc system of Shewanella oneidensis MR-1. Appl Environ Microbiol 76:3263–3274
Lassak J, Bubendorfer S, Thormann KM (2013) Domain analysis of ArcS, the hybrid sensor kinase of the Shewanella oneidensis MR-1 Arc two-component system, reveals functional differentiation of its two receiver domains. J Bacteriol 195:482–492
Le Laz S, Kpebe A, Lorquin J, Brugna M, Rousset M (2014) H2-dependent azoreduction by Shewanella oneidensis MR-1: involvement of secreted flavins and both [Ni–Fe] and [Fe–Fe] hydrogenases. Appl Microbiol Biotechnol 98:2699–2707
Lefebvre O, Uzabiaga A, Chang IS, Kim BH, Ng HY (2011) Microbial fuel cells for energy self-sufficient domestic wastewater treatment-a review and discussion from energetic consideration. Appl Microbiol Biotechnol 89:259–270
Li F, Li Y, Sun L, Li X, Yin C, An X, Chen X, Tian Y, Song H (2017a) Engineering Shewanella oneidensis enables xylose-fed microbial fuel cell. Biotechnol Biofuels 10:1–10
Li SW, Zeng RJ, Sheng GP (2017b) An excellent anaerobic respiration mode for chitin degradation by Shewanella oneidensis MR-1 in microbial fuel cells. Biochem Eng J 118:20–24
Li F, Li Y, Sun L, Chen X, An X, Yin C, Cao Y, Wu H, Song H (2018a) Modular engineering intracellular NADH regeneration boosts extracellular electron transfer of Shewanella oneidensis MR-1. ACS Synth Biol 7:885–895
Li M, Zhou M, Tian X, Tan C, McDaniel CT, Hassett DJ, Gu T (2018b) Microbial fuel cell (MFC) power performance improvement through enhanced microbial electrogenicity. Biotechnol Adv 36:1316–1327
Liu C, Gorby YA, Zachara JM, Fredrickson JK, Brown CF (2002) Reduction kinetics of Fe(III), Co(III), U(VI), Cr(VI), and Tc(VII) in cultures of dissimilatory metal-reducing bacteria. Biotechnol Bioeng 80:637–649
Liu CG, Xue C, Lin YH, Bai FW (2013) Redox potential control and applications in microaerobic and anaerobic fermentations. Biotechnol Adv 31:257–265
Logan BE, Hamelers B, Rozendal R, Schröder U, Keller J, Freguia S, Aelterman P, Verstraete W, Rabaey K (2006) Microbial fuel cells: methodology and technology. Environ Sci Technol 40:5181–5192
Lovley DR (2012) Electromicrobiology. Annu Rev Microbiol 66:391–409
Lu M, Chan S, Babanova S, Bretschger O (2017) Effect of oxygen on the per-cell extracellular electron transfer rate of Shewanella oneidensis MR-1 explored in bioelectrochemical systems. Biotechnol Bioeng 11:96–105
Luo S, Guo W, Nealson KH, Feng X, He Z (2016) 13C pathway analysis for the role of formate in electricity generation by Shewanella oneidensis MR-1 using lactate in microbial fuel cells. Sci Rep 6:1–8
Marritt SJ, McMillan DG, Shi L, Fredrickson JK, Zachara JM, Richardson DJ, Jeuken LJ, Butt JN (2012) The roles of CymA in support of the respiratory flexibility of Shewanella oneidensis MR-1. Biochem Soc Trans 40:1217–1221
Marsili E, Baron DB, Shikhare ID, Coursolle D, Gralnick JA, Bond DR (2008) Shewanella secretes flavins that mediate extracellular electron transfer. Proc Natl Acad Sci USA 105:3968–3973
McDowall JS, Murphy BJ, Haumann M, Palmer T, Armstrong FA, Sargent F (2014) Bacterial formate hydrogenlyase complex. Proc Natl Acad Sci USA 111:3948–3956
Melo AMP, Bandeiras TM, Teixeira M (2004) New insights into type II NAD(P)H: quinone oxidoreductases. Microbiol Mol Biol Rev 68:603–616
Meshulam-Simon G, Behrens S, Choo AD, Spormann AM (2007) Hydrogen metabolism in Shewanella oneidensis MR-1. Appl Environ Microbiol 73:1153–1165
Moore LJ, Mettert EL, Kiley PJ (2006) Regulation of FNR dimerization by subunit charge repulsion. J Biol Chem 281:33268–33275
Mordkovich NN, Voeĭkova TA, Novikova TA, Smirnov IA, Il’in VK, Soldatov PE, Tiurin-Kuz’min AI, Smolenskaia TS, Veĭko VP, Shakulov RS, Debabov VG (2013) Influence of NAD-dependent formate dehydrogenase on anaerobic respiration of Shewanella oneidensis MR-1. Mikrobiologiia 82:395–401
Moscoviz R, Toledo-Alarcón J, Trably E, Bernet N (2016) Electro-fermentation: how to drive fermentation using electrochemical systems. Trends Biotechnol 34:856–865
Myers CR, Nealson KH (1988) Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science 240:1319–1321
Naik SN, Goud VV, Rout PK, Dalai AK (2010) Production of first and second generation biofuels: a comprehensive review. Renew Sustain Energy Rev 14:578–597
Nakagawa G, Kouzuma A, Hirose A, Kasai T, Yoshida G, Watanabe K (2015) Metabolic characteristics of a glucose-utilizing Shewanella oneidensis strain grown under electrode-respiring conditions. PLoS ONE 10:e0138813
Nevin KP, Woodard TL, Franks AE, Summers ZM, Lovley DR (2010) Microbial electrosynthesis: feeding microbes electricity to convert carbon dioxide and water to multicarbon extracellular organic compounds. MBio 1:1–4
Newman DK, Kolter R (2000) A role for excreted quinones in extracellular electron transfer. Nature 405:94–97
Newton GJ, Mori S, Nakamura R, Hashimoto K, Watanabe K (2009) Analyses of current-generating mechanisms of Shewanella loihica PV-4 and Shewanella oneidensis MR-1 in microbial fuel cells. Appl Environ Microbiol 75:7674–7681
Orr MW, Donaldson GP, Severin GB, Wang J, Sintim HO, Waters CM, Lee VT (2015) Oligoribonuclease is the primary degradative enzyme for pGpG in Pseudomonas aeruginosa that is required for cyclic-di-GMP turnover. Proc Natl Acad Sci USA 112:5048–5057
Paul R, Weiser S, Amiot NC, Chan C, Schirmer T, Giese B, Jenal U (2004) Cell cycle-dependent dynamic localization of a bacterial response regulator with a novel di-guanylate cyclase output domain. Genes Dev 18:715–727
Pinchuk GE, Rodionov DA, Yang C, Li X, Osterman AL, Dervyn E, Geydebrekht OV, Reed SB, Romine MF, Collart FR, Scott JH, Fredrickson JK, Beliaev AS (2009) Genomic reconstruction of Shewanella oneidensis MR-1 metabolism reveals a previously uncharacterized machinery for lactate utilization. Proc Natl Acad Sci USA 106:2874–2879
Pinchuk GE, Geydebrekht OV, Hill EA, Reed JL, Konopka AE, Beliaev AS, Fredrickson JK (2011) Pyruvate and lactate metabolism by Shewanella oneidensis MR-1 under fermentation, oxygen limitation, and fumarate respiration conditions. Appl Environ Microbiol 77:8234–8240
Pirbadian S, Barchinger SE, Leung KM, Byun HS, Jangir Y, Bouhenni RA, Reed SB, Romine MF, Saffarini DA, Shi L, Gorby YA, Golbeck JH, El-Naggar MY (2014) Shewanella oneidensis MR-1 nanowires are outer membrane and periplasmic extensions of the extracellular electron transport components. Proc Natl Acad Sci USA 111:12883–12888
Rabaey K, Rozendal RA (2010) Microbial electrosynthesis—revisiting the electrical route for microbial production. Nat Rev Microbiol 8:706–716
Redwood MD, Mikheenko IP, Sargent F, Macaskie LE (2008) Dissecting the roles of Escherichia coli hydrogenases in biohydrogen production. FEMS Microbiol Lett 278:48–55
Rodionov DA, Novichkov PS, Stavrovskaya ED, Rodionova IA, Li X, Kazanov MD, Ravcheev DA, Gerasimova AV, Kazakov AE, Kovaleva GY, Permina EA, Laikova ON, Overbeek R, Romine MF, Fredrickson JK, Arkin AP, Dubchak I, Osterman AL, Gelfand MS (2011) Comparative genomic reconstruction of transcriptional networks controlling central metabolism in the Shewanella genus. BMC Genom 12(Suppl 1):S3
Ross DE, Flynn JM, Baron DB, Gralnick JA, Bond DR (2011) Towards electrosynthesis in Shewanella: energetics of reversing the mtr pathway for reductive metabolism. PLoS ONE 6:e16649
Rowe AR, Rajeev P, Jain A, Pirbadian S, Okamoto A, Gralnick JA, El-Naggar MY, Nealson KH (2018) Tracking electron uptake from a cathode into Shewanella cells: implications for energy acquisition from solid-substrate electron donors. MBio 9:1–19
Ryan RP, Fouhy Y, Lucey JF, Crossman LC, Spiro S, He YW, Zhang LH, Heeb S, Cámara M, Williams P, Dow JM (2006) Cell-cell signaling in Xanthomonas campestris involves an HD-GYP domain protein that functions in cyclic di-GMP turnover. Proc Natl Acad Sci USA 103:6712–6717
Saffarini DA, Schultz R, Beliaev A (2003) Involvement of cyclic AMP (cAMP) and cAMP receptor protein in anaerobic respiration of Shewanella oneidensis. J Bacteriol 185:3668–3671
Santoro C, Arbizzani C, Erable B, Ieropoulos I (2017) Microbial fuel cells: from fundamentals to applications. A review. J Power Sources 356:225–244
Scheller HV (2017) Plant cell wall: never too much acetate. Nat Plants 3:17024
Schröder U, Harnisch F, Angenent LT (2015) Microbial electrochemistry and technology: terminology and classification. Energy Environ Sci 8:513–519
Schwechheimer C, Kuehn MJ (2015) Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat Rev Microbiol 13:605–619
Sekar R, Shin HD, DiChristina TJ (2016) Activation of an otherwise silent xylose metabolic pathway in Shewanella oneidensis. Appl Environ Microbiol 82:3996–4005
Shi L, Squier TC, Zachara JM, Fredrickson JK (2007) Respiration of metal (hydr)oxides by Shewanella and Geobacter: a key role for multihaem c-type cytochromes. Mol Microbiol 65:12–20
Shi L, Dong H, Reguera G, Beyenal H, Lu A, Liu J, Yu HQ, Fredrickson JK (2016) Extracellular electron transfer mechanisms between microorganisms and minerals. Nat Rev Microbiol 14:651–662
Shimada T, Fujita N, Yamamoto K, Ishihama A (2011) Novel roles of cAMP receptor protein (CRP) in regulation of transport and metabolism of carbon sources. PLoS One 6:e20081
Stoeckl M, Schlegel C, Sydow A, Holtmann D, Ulber R, Mangold KM (2016) Membrane separated flow cell for parallelized electrochemical impedance spectroscopy and confocal laser scanning microscopy to characterize electro-active microorganisms. Electrochim Acta 220:444–452
Sturm G, Richter K, Doetsch A, Heide H, Louro RO, Gescher J (2015) A dynamic periplasmic electron transfer network enables respiratory flexibility beyond a thermodynamic regulatory regime. ISME J 9:1802–1811
Sydow A, Krieg T, Mayer F, Schrader J, Holtmann D (2014) Electroactive bacteria—molecular mechanisms and genetic tools. Appl Microbiol Biotechnol 98:8481–8895
Szeinbaum N, Lin H, Brandes JA, Taillefert M, Glass JB, DiChristina TJ (2017) Microbial manganese(III) reduction fuelled by anaerobic acetate oxidation. Environ Microbiol 19:3475–3486
Tang YJ, Hwang JS, Wemmer DE, Keasling JD (2007) Shewanella oneidensis MR-1 fluxome under various oxygen conditions. Appl Environ Microbiol 73:718–729
Thormann KM, Saville RM, Shukla S, Pelletier DA, Spormann AM (2004) Initial phases of biofilm formation in Shewanella oneidensis MR-1. J Bacteriol 186:8096–8104
Thormann KM, Saville RM, Shukla S, Spormann AM (2005) Induction of rapid detachment in Shewanella oneidensis MR-1 biofilms. J Bacteriol 187:1014–1021
Thormann KM, Duttler S, Saville RM, Hyodo M, Shukla S, Hayakawa Y, Spormann AM (2006) Control of formation and cellular detachment from Shewanella oneidensis MR-1 biofilms by cyclic di-GMP. J Bacteriol 188:2681–2691
Torella JP, Gagliardi CJ, Chen JS, Bediako DK, Colón B, Way JC, Silver PA, Nocera DG (2015) Efficient solar-to-fuels production from a hybrid microbial water-splitting catalyst system. Proc Natl Acad Sci USA 112:2337–2342
Ueoka N, Kouzuma A, Watanabe K (2018) Electrode plate-culture methods for colony isolation of exoelectrogens from anode microbiomes. Bioelectrochemistry 124:1–6
Venkateswaran K, Moser DP, Dollhopf ME, Lies DP, Saffarini DA, MacGregor BJ, Ringelberg DB, White DC, Nishijima M, Sano H, Burghardt J, Stackebrandt E, Nealson KH (1999) Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. Int J Syst Bacteriol 49:705–724
Wan XF, Verberkmoes NC, McCue LA, Stanek D, Connelly H, Hauser LJ, Wu L, Liu X, Yan T, Leaphart A, Hettich RL, Zhou J, Thompson DK (2004) Transcriptomic and proteomic characterization of the Fur modulon in the metal-reducing bacterium Shewanella oneidensis. J Bacteriol 186:8385–8400
Wang CC, Chang CW, Chu CP, Lee DJ, Chang BV, Liao CS (2003) Producing hydrogen from wastewater sludge by Clostridium bifermentans. J Biotechnol 102:83–92
Watanabe K (2008) Recent developments in microbial fuel cell technologies for sustainable bioenergy. J Biosci Bioeng 106:528–536
Xiao Y, Zhang E, Zhang J, Dai Y, Yang Z, Christensen HEM, Ulstrup J, Zhao F (2017) Extracellular polymeric substances are transient media for microbial extracellular electron transfer. Sci Adv 3:e1700623
Yang C, Rodionov DA, Li X, Laikova ON, Gelfand MS, Zagnitko OP, Romine MF, Obraztsova AY, Nealson KH, Osterman AL (2006) Comparative genomics and experimental characterization of N-acetylglucosamine utilization pathway of Shewanella oneidensis. J Biol Chem 281:29872–29885
Yang Y, Harris DP, Luo F, Wu L, Parsons AB, Palumbo AV, Zhou J (2008) Characterization of the Shewanella oneidensis Fur gene: roles in iron and acid tolerance response. BMC Genom 9:S11
Yang Y, Harris DP, Luo F, Xiong W, Joachimiak M, Wu L, Dehal P, Jacobsen J, Yang Z, Palumbo AV, Arkin AP, Zhou J (2009) Snapshot of iron response in Shewanella oneidensis by gene network reconstruction. BMC Genom 10:131
Yin J, Meng Q, Fu H, Gao H (2016) Reduced expression of cytochrome oxidases largely explains cAMP inhibition of aerobic growth in Shewanella oneidensis. Sci Rep 6:24449
Yoshizawa T, Miyahara M, Kouzuma A, Watanabe K (2014) Conversion of activated-sludge reactors to microbial fuel cells for wastewater treatment coupled to electricity generation. J Biosci Bioeng 118:533–539
Yu YY, Fang Z, Gao L, Song H, Yang L, Mao B, Yong YC (2018) Engineering of bacterial electrochemical activity with global regulator manipulation. Electrochem Commun 86:117–120
Yuan J, Wei B, Lipton MS, Gao H (2012) Impact of ArcA loss in Shewanella oneidensis revealed by comparative proteomics under aerobic and anaerobic conditions. Proteomics 12:1957–1969
Zhao JS, Deng Y, Manno D, Hawari J (2010) Shewanella spp. genomic evolution for a cold marine lifestyle and in situ explosive biodegradation. PLoS ONE 5:e9109
Zheng T, Xu YS, Yong XY, Li B, Yin D, Cheng QW, Yuan HR, Yong YC (2015) Endogenously enhanced biosurfactant production promotes electricity generation from microbial fuel cells. Bioresour Technol 197:416–421
Zhou L, Srisatjaluk R, Justus DE, Doyle RJ (1998) On the origin of membrane vesicles in gram-negative bacteria. FEMS Microbiol Lett 163:223–228
Zhou G, Yuan J, Gao H (2015) Regulation of biofilm formation by BpfA, BpfD, and BpfG in Shewanella oneidensis. Front Microbiol 6:790
Authors’ contributions
AH, TK, RK, YS and AK wrote parts of the manuscript, and KW edited the manuscript. All authors read and approved the final manuscript.
Acknowledgements
We thank Nanako Amano for technical assistance.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
All data generated or analyzed during this study are included in this article.
Consent for publication
Author have approved to submit this manuscript to Bioresources and Bioprocessing.
Funding
This work was supported by JSPS KAKENHI Grant Numbers: 15H01753, 16J08653, 17J05454 and 18K05399.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hirose, A., Kasai, T., Koga, R. et al. Understanding and engineering electrochemically active bacteria for sustainable biotechnology. Bioresour. Bioprocess. 6, 10 (2019). https://doi.org/10.1186/s40643-019-0245-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40643-019-0245-9