Abstract
Objective
Many sections of the health care system are facing a major challenge making infectious disease problematic to treat; antimicrobial resistance (AMR). Identification and surveillance of the resistome have been highlighted as one of the strategies to overcome the problem. This study aimed to screen for AMR genes in an oral microbiota, a complex microbial system continuously exposed to antimicrobial agents commonly used in dental practice.
Materials and methods
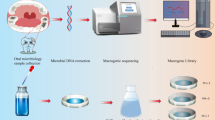
As a significant part of the oral microbiome cannot be conventionally cultured, a functional metagenomic approach was chosen. The human oral metagenomic DNA was extracted from saliva samples collected from 50 healthy volunteers in Norway. The oral metagenomic library was then constructed by ligating partially digested oral metagenome into pSMART BAC vector and introducing into Escherichia coli. The library was screened against antimicrobials in dental practices. All resistant clones were selected and analyzed.
Results
Screening of the oral metagenomic library against different antimicrobials detected multiple clones with resistance against chlorhexidine, triclosan, erythromycin, tetracycline, and sodium hypochlorite. Bioinformatic analysis revealed both already known resistance genes, including msr, mef(A), tetAB(46), and fabK, and genes that were not previously described to confer resistance, including recA and accB conferring resistance to sodium hypochlorite and chlorhexidine, respectively.
Conclusion
Multiple clones conferring resistance to antimicrobials commonly used in dental practices were detected, containing known and novel resistant genes by functional-based metagenomics. There is a need for more studies to increase our knowledge in the field.
Similar content being viewed by others
Introduction
Antimicrobial agents have saved uncountable numbers of lives for decades since the discovery of Penicillin; however, with a worldwide increase of antimicrobial resistance, infectious diseases currently have become more challenging to be treated. All uses of antimicrobials apply selective pressure to bacteria to evolve and develop antimicrobial resistance [1,2,3,4]. Discovery of resistance genes recovered from ancient samples showed that they were significantly similar to the modern resistance variants, suggesting antimicrobial resistance as an old natural phenomenon [5,6,7], but have recently become a problem possibly due to the selective pressures that accelerated the spreading of resistance genes through horizontal gene transfer [8,9,10]. Identification and surveillance of the resistome are, therefore, essential in the battle against antimicrobial resistance, as they will improve our understanding of resistance genes in each setting which can be used to design effective strategies to limit the spreading between organisms and environments [11,12,13].
The human oral cavity is a complex microbial system [14, 15], housing a selection of bacteria with more than 700 bacterial species [16,17,18]. It consists of several small ecosystems with unique environments such as keratinized and non-keratinized mucosa, the tongue, saliva, tonsils, teeth, and subgingival pockets together making up the oral microbiome [16, 19]. The species in the oral microbiome vary, from facultative aerobes to strict anaerobes. They are continuously exposed to antimicrobial agents from external products such as oral hygiene products as toothpaste, mouth rinse, agents used in dental treatment and food, and is therefore likely to develop antimicrobial resistance. Relevant examples of antimicrobials used in dental practices and dental hygiene products are chlorhexidine used in antimicrobial mouth rinses post-operative of surgical procedures [20], and for gingivitis and periodontitis patients who are unable to maintain adequate mechanical hygiene [21], sodium hypochlorite used as an irrigation agent during root canal treatment [22], sodium benzoate used in various toothpastes, cetyltrimethylammonium bromide (CTAB) found in throat lozenges and topical gels and conventional antibiotics for patients with risk factors pre-operative of surgical procedures. Studies have shown that the oral microbiome contains resistance genes against various antimicrobials agents such as β-lactams, tetracycline, tigecycline, amoxicillin, gentamicin, CTAB, erythromycin and cetylpyridinium chloride [1, 23,24,25,26,27,28].
Of the more than 700 oral bacterial species, one-third of them are not cultured in the laboratory yet due to difficult and unknown proper conditions for growth [29, 30], which has created challenges for characterizing the resistome in the oral microbiome. Functional metagenomics is a culture-independent approach, which relies on phenotypes of resistance genes, rather than the sequences of the resistance genes as in PCR and microarray [31,32,33]. It is, therefore, a method with the potential to discover completely novel resistance genes [34,35,36,37], without culturing bacteria. It involves cloning of metagenomic DNA into a vector, introducing into a surrogate bacterial host, and screening for clones of phenotypes of interest, such as resistance traits. Several novel resistance genes were identified from the oral metagenome through a functional metagenomic approach, such as tetracycline resistance gene tet(37), tigecycline resistance gene tetAB(60) and quaternary ammonium compounds (QACs) resistance gene galE [1, 23, 24].
In this study, we aimed to detect novel antimicrobial resistance genes from the human oral microbiome through a functional metagenomic approach. A human oral metagenomic library obtained from 50 healthy volunteers in Norway was constructed and used to screen against antimicrobials that are commonly in contact with oral bacteria. We have identified multiple resistance clones against triclosan, CTAB, sodium hypochlorite, chlorhexidine, and erythromycin.
Methods
Study participants and collection of saliva samples
Ethical approval was obtained from Regional Committees for Medical and Health Research Ethics (Project number 2018/1373/REK nord). Saliva samples were collected between October and November 2018 from 50 healthy volunteers visiting the University Dental Clinic at UiT The Arctic University of Norway who were invited to participate in the study. All participants gave their written consent to participate in the study. The following criteria were used for participation: no history of antibiotic use in the last three months prior to saliva sampling, no history of regular medication, nor chronic diseases. All volunteers gave their written consent to participate in the study. The volunteers were asked not to drink, eat or brush their teeth within an hour before the collection. A paraffin gum was used to stimulate saliva secretion during collection, and 2 mL of stimulated saliva was collected from each participant into a Saliva DNA Collection and Preservation Kit (Norgen Biotek Corp, Ontario, Canada). All samples were anonymized and stored at room temperature.
Extraction of oral metagenomic DNA and construction of the oral metagenomic library
Saliva metagenomic DNA was extracted from each sample by mixing 750 μl of each saliva in the preservation tube with 750 μl phosphate-buffered saline (PBS) buffer. This mixture was centrifuged for 10 min at 15,700×g. The supernatant was discarded and resuspended in 125 μl PBS and 25 μl MetaPolyzyme (Sigma-Aldrich, Norway). The samples were incubated at 35 °C for 4 h. The DNA samples were then extracted with QIAcube (Qiagen, Norway), following the protocol from QIAamp® DNA Mini QIAcube Kit.
For the construction of the oral metagenomic library, 10 μl of extracted DNA was aliquoted from each of the 50 samples, making up 500 μl. The pooled metagenomic DNA was partially digested for 2, 3, and 4 min at 37 °C with HindIII restriction enzyme to serve us large DNA fragments. The digested product was run on an agarose gel electrophoresis, and DNA fragments with a size of more than 1000 bp were extracted by using QIAgen gel extraction kit (Qiagen). The pSMART BAC HindIII vector (7.6 kb) was fully digested and dephosphorylated by using HindIII restriction enzyme and calf intestinal alkaline phosphatase (CIAP) enzyme (NEB, UK) at 37 °C for 60 min. Afterwards, the partially digested oral metagenome was ligated into a pre-digested pSMART BAC vector by using Anza T4 DNA Ligase Master Mix (ThermoFisher, Norway) and incubated for 16 h at 4 °C. The ligation product was desalted in an agarose cone. The desalted ligation product (2 µl) was then mixed with 20 μl BAC-Optimized Replicator (BacRep) Escherichia coli Electrocompetent cells (Lucigen, USA), and transferred to a pre-chilled 0.1 cm electroporation cuvette (Bio-Rad, Norway). The mixture was electroporated with the following settings: 1.8 kV, 25 μF, 200 Ω (MicroPulser Electroporator, Bio-Rad, Norway). A pre-warmed recovery medium (950 µl) (Lucigen, USA) was immediately added to the cells and incubated at 37 °C with shaking for 1 h, before plating 100 μl on Luria–Bertani (LB) Agar supplemented with 12.5 μg/ml chloramphenicol plate.
Determination of average insert size of the constructed oral metagenomic library
To determine the average insert size of the constructed oral metagenomics library, 10 random colonies from the LB Agar chloramphenicol control plate were subcultured into 5 ml LB broth containing 12.5 μg/ml chloramphenicol and incubated at 37 °C with shaking at 200 RPM for 18 h. The plasmids containing the insert from each clone were extracted by using QIAprep Spin Miniprep Kit (QIAgen, Norway), following the manufacturer’s protocol. After extraction, each plasmid was digested in 10 μl reaction, containing 1 µl CutSmart buffer (10x), 0.5 μl HindIII restriction enzyme, 1 μl plasmid, and 7.5 μl distilled water. Each reaction was digested at 37 °C for 30 min. To visualize the inserts, each digested product was run on agarose gel electrophoresis with 120 V for one hour. The average insert size of the constructed library was calculated based on the insert size of each sample, estimated from the gel.
Determination of minimum inhibitory concentration and screening of the oral metagenomic library
The minimum inhibitory concentration (MIC) of each antimicrobial was determined for E. coli BacRep containing pSMART BAC vector (with no insert), following the broth dilution method as described previously [38]. An overnight culture was set up by subculturing a single colony into 5 ml LB broth containing 12.5 μg/ml chloramphenicol and incubated for 18 h at 37 °C with shaking at 200 RPM. The overnight culture was diluted to the OD600 of 0.1. The MIC was determined in a 96-well microtiter plate by adding 10 μl of the diluted culture and 90 μl LB broth containing different concentrations of antimicrobial agents, shown in Table 1. The plates were incubated at 37 °C with shaking for 18–24 h, and the growth was determined by reading OD600 before and after incubation with a microplate spectrophotometer. This was repeated three times for each antimicrobial.
For the screening of the oral metagenomic library for resistance clones, 100 μl of the electroporated E. coli carrying pSMART with oral metagenome insert were spread on LB agar supplemented with 12.5 μg/ml chloramphenicol and antimicrobial with the MIC concentration determined in the previous step, then incubated overnight at 37 °C. All of the colonies grown on the screening plates of each antimicrobial were streaked onto a new antimicrobial containing plate, and also subcultured into 5 ml LB broth containing the antimicrobial to confirm the resistance.
Characterization of genes conferring antimicrobial resistance
The confirmed resistance clones were subcultured into 5 ml LB broth, containing chloramphenicol and antimicrobial of their resistance, and incubated at 37 °C for 18 h with shaking. The plasmids were extracted by using QIAprep Spin Miniprep Kit, digested with HindIII restriction enzyme, and visualized on an agarose gel to estimate the size of inserts.
All the inserts, except for the sodium hypochlorite-clone, were amplified by setting up PCR reactions with Platinum SuperFi Green PCR Master Mix (ThermoFisher Scientific, Norway), which can amplify up to 13 kb DNA, and SL1-SR4 primer pair (Lucigen, USA), which were the primers flanking the cloning site on the pSMART BAC vector. The 50-μl PCR reactions composed of 2 µl SL1 forward primer (10 μM), 2 µl SR4 reverse primer (10 μM), 25 µl 2 × Platinum SuperFi Green PCR Master Mix (Thermo Scientific, Norway), 20 µl molecular grade water, and 1 μl plasmid. The PCR cycle was programmed, as suggested by the manufacturer’s protocol. The PCR products were purified by using QIAquick PCR Purification Kit (Qiagen, Norway), then sent for Sanger sequencing from both ends with SL-1 and SR-4 primers at Genewiz, Germany. Additional primers were designed to extend the sequencing for samples that were not fully sequenced by the initial sequencing.
Sequencing data were aligned and manipulated by using BioEdit software version 7.2.0 (http://www.mbio.ncsu.edu/bioedit/bioedit.html). The contigs of each sample were assembled by using CAP3 contig assembly program [39]. The assembled sequences were compared with sequences in the nucleotide and protein databases by using BlastN and BlastX from the National Centre for Biotechnology Information (NCBI) [40]. The nucleotide sequences of all resistance clones were deposited in Genbank with the accession numbers MZ955857 to MZ955863.
Subcloning of putative genes conferring chlorhexidine and sodium hypochlorite resistance
The putative resistance genes were amplified from the plasmids extracted from chlorhexidine and sodium hypochlorite resistant clones by using primers listed in Additional file 1: Table S1. The 30-μl PCR reactions composed of 15 µl 2 × BioMix Red (Bioline, United Kingdom), 2 μl of each primer (10 μM), 1 μl extracted plasmid and 10 μl molecular grade water. The PCR products were purified by using QIAquick PCR Purification Kit (Qiagen, Norway). All purified products, except recA PCR amplicons, were digested with HindIII and ligated to a HindIII-predigest pSMART BAC vector by using Anza T4 DNA Ligase Master Mix. The HindIII-ligated products were electroporated into BacRep E. coli Electrocompetent cells and grew on LB agar containing 12.5 μg/ml chloramphenicol. For the recA PCR amplicons, it was digested with EcoRI and ligated to an EcoRI-predigested pUC19 vector instead as there was an internal HindIII restriction site in the recA gene. The pUC19-recA ligation product was introduced into Subcloning Efficiency™ DH5α Competent Cells (Thermo Scientific, Norway) by heat-shock transformation and grew on LB agar containing 100 μg/ml ampicillin. The listed of bacterial strains and plasmids from the subcloning of the putative resistance genes were shown in Additional file 2: Table S2.
Results
Screening of the constructed oral metagenomic library against antimicrobials used in dental practice
After the construction of the metagenomic library, the average insert size was calculated by determining the insert size from 10 random colonies, which showed the average insert size of the constructed library as 5500 bp (Additional file 3: Fig. S1).
The MICs of E. coli BacRep carrying an empty pSMART BAC vector towards different antimicrobials were determined and listed in Table 1. The oral metagenomic library was then screened against each antimicrobial based on these MICs in which 7 different resistant clones were identified, including Chlorhexidine-1 (Chx-1), Chlorhexidine-2 (Chx-2), Triclosan-1 (Tric-1), Triclosan-2 (Tric-2), Erythromycin-1 (Ery-1), Tetracycline-1 (Tet-1) and Sodium hypochlorite-1 (NaOCl-1). HindIII plasmid digestion was performed to estimate the insert size of each resistant clone (Fig. 1). Sequencing and bioinformatics analysis through BlastN and BlastX of each clone were performed and shown in Fig. 2 and Table 2.
Schematic representation of predicted ORFs found on insert DNA of each resistant clone. The open arrowed boxes represent ORFs, pointing in the probable direction of transcription. The known resistant genes and other genes are shown in green and blue, respectively. The dash boxes and arrow boxes represent the regions that are not found on the inserts, compared to the sequences in the database
Identification of genes conferring resistance in the identified resistant clones
Among 7 resistant clones identified from the screening, 4 clones were shown to carry previously known resistance genes. For Ery-1, it contained msr and mef(A) macrolide resistance genes, that encode for a macrolide efflux pump [41, 42]. The tetracycline-resistant clone Tet-1 carried tetAB(46), an ABC-transporter which was isolated previously from the human oral cavity [24, 43]. It has been proved to transport tetracyclines over the cell membrane in both Gram-positive and Gram-negative bacteria, but as a single-drug efflux pump [43]. Both Tric-1 and Tric-2 contained fabK (an isoform of fabI), encoding for an enzyme, enoyl-acyl carrier protein reductase II (ENR). ENR is involved in the fatty acid synthesis, and also the binding site for triclosan to inhibit the synthesis. Upregulation of ENR resulted in a lower inhibitory effect from triclosan, as shown previously [1].
For chlorhexidine resistant clones (Chx-1 and Chx-2), bioinformatics analysis showed that they each contain only one complete open reading frame with the size of 750 bp and 489 bp, respectively. Blast analysis of the ORF in Chx-1 did not show any match by blastN and blastP. A partial match was shown by blastX that it had a 236-bp region that showed 51% similarity to a part of type IV secretion system DNA-binding domain-containing protein from Kocuria indica. The complete ORF in Chx-2 was matched to accB gene, encoding acetyl-CoA carboxylase biotin carboxyl carrier protein. As there were also another 2 incomplete ORFs in Chx-2, the accB gene of Chx-2 was amplified and subcloned into pSMART BAC vector to confirm that it is the only gene responsible for the chlorhexidine resistance phenotype. The MIC of chlorhexidine against E. coli BacRep::pSMART-Chx-1, E. coli BacRep::pSMART-Chx-2 and E. coli BacRep::pSMART-accB were shown to increase two-fold (from 1.0 to 2.0 µg/ml), compared to the wild-type E. coli BacRep::pSMART.
Bioinformatic analysis showed that the insert DNA of NaOCl-1 clone contained multiple genes, (as shown in Fig. 2 and Table 2) none of them was reported as a gene conferring sodium hypochlorite resistance. Four putative genes, encoding methyltransferase, diaminopimelate epimerase (DapF), hemin ABC transporter and RecA, were amplified and subcloned to determine the gene that conferred sodium hypochlorite resistance. The subcloning results showed that only E. coli DH5α::pUC19-recA showed an increase in MIC against sodium hypochlorite from 0.040 to 0.050%, compared to the wild-type E. coli DH5α::pUC19.
The distribution of all detected resistance genes was determined by performing BlastN on the metagenomic sequencing data of each saliva oral metagenomic DNA. The results showed that 6 of 7 detected resistance genes could be found in all 50 subjects, and the last one (Chx-1) could be found in 4 out of 50 subjects (Additional file 4).
Discussion
Antimicrobial resistance is a major burden for the healthcare system worldwide. Several AMR genes have been discovered from the human oral microbiome previously. Some of them were found to be associated with mobile genetic elements, such as Tn916 family conjugative transposons [44, 45], that can facilitate that spreading to other oral bacteria, including pathogens. As the oral cavity is the entry point to both the respiratory and gastrointestinal tract, oral bacteria that contain these resistance genes can therefore easily wander through the body via the bloodstream or by swallowing, and thereby have the potential to transfer their resistance genes to other microbiomes [25]. Therefore, it is important to screen and identify resistance genes in the oral microbiome so that we can design effective strategies and guidelines for antimicrobial uses to limit the spreading of these genes.
In our study, we used functional metagenomics to screen the oral metagenome for resistance genes against the antimicrobials used in dental practices, where 7 resistance clones were identified. Four of them contained previously known AMR, which also could be found in the oral cavity, where one of them (fabI) was found by functional metagenomic screening [1, 43, 46, 47]. The rest did not contain known resistance genes: two of them showed resistance to chlorhexidine, and another clone had resistance against sodium hypochlorite. This is the first time that functional metagenomic screening identified resistance genes for both antimicrobials from the oral metagenome.
Chlorhexidine is a widely used broad-spectrum antimicrobial in dentistry (e.g. endodontology, periodontology, oral surgery) [48]. It is normally combined with gluconid or acetic acid to form water-soluble digluconate or diacetate salts [49]. The mode of action is dose-dependent; bacteriostatic at low concentrations, bactericidal in higher, both through binding to negatively charged membrane phospholipids, resulting in reduced membrane fluidity and osmoregulation [50, 51]. Therefore, in lower concentrations, it disrupts the membrane causing leakage of low-weight molecules, while, in higher concentrations, it causes cytolysis by forming precipitates and releasing intracellular components [49, 50].
For the Chx-2 clone, the gene responsible for resistance was accB, encoding for biotin carboxyl carrier protein (BCCP) which is a component of acetyl CoA carboxylase that catalyzes the first step in fatty acid and phospholipid biosynthesis [52]. BCCP was previously reported to be one of the proteins that were upregulated as a consequence of chlorhexidine exposure in a proteomic analysis of a resistant Pseudomonas aeruginosa [53]. In our study, we showed and confirmed that accB conferred chlorhexidine resistance by expressing a heterologous accB, recovered from the oral cavity, in an E. coli surrogate host. Overexpressing accB in E. coli could increase the rate of phospholipid biosynthesis, which is the main component in the bacterial cell membrane, allowing the bacteria to become less sensitive against chlorhexidine which targets phospholipids in the cell membrane.
Sodium hypochlorite is an irrigation agent that is widely used in endodontic procedures, such as root canal fillings, by dentists. It is antimicrobial mainly in wet environments as it ionizes to Na+ and OCl−. At pH levels between 4 and 7, its form is hypochlorous acid (HClO), while at pH > 9 it is OCl−—both of them reactive oxidizing agents [54]. Teeth with pulp necrosis have a pH value between 6–7.4 before treatment, which means that HClO is the most important form of sodium hypochlorite for the treatments [55]. However, it is the tissue-dissolving property that may be considered the most important one for the procedures, where peptide bonds are destroyed, followed by dissolving proteins that can be irrigated away from the root canal [56].
The antimicrobial effect of sodium hypochlorite has been shown to be a complex process. As it is an oxidative antimicrobial, it causes oxidative stress which damage both DNA and lipids [57, 58]. We found that recA, isolated from the oral metagenome in our study, could confer sodium hypochlorite resistance when in E. coli host. RecA plays an important role in homologous recombination and DNA repairs like SOS response that is activated by DNA damage from various environmental factors and antibiotics. Previously, it was demonstrated that mutations in recA and recB repair genes increased the sensitivity an E. coli strains to HClO [59, 60]. In our case, as the E. coli lab strains (DH5α and BacRep) were recA-deficient strains, introducing pSMART BAC and pUC19 plasmids containing recA would result in complementation of recA in these E. coli lab strains, which was shown to have higher MIC against NaOCl.
It is a common practice in functional genomic studies to use E. coli to test the metagenomic constructed libraries. In the current study, E. coli was utilised to express accB and recA genes. Although the two genes are housekeeping genes, it is advisable to assess the expression of these genes in oral bacteria given their heterogeneous nature.
The two newly recognized AMR genes were not associated with mobile genetic elements, which implies that they are not able to be transferred or spread by themselves. However, it has been shown that selective pressure from uses of antimicrobials could drive these housekeeping genes to be associated with mobile genetic elements like fabI gene found in IS1272 composite transposons [61], which give them the ability to be spread. Exposing bacteria to compounds like sodium hypochlorite could also lead to the spread of antibiotic resistance genes between bacteria, as it can induce an SOS-response in bacteria which promotes the spread of mobile genetic elements like integrative conjugative elements (SXT) that can facilitate an intercellular transposition [62].
Chlorhexidine is most used as a mouth rinse the days after a third mandibular molar surgery, to treat pericoronitis as well as it might be considered as an additional treatment for periodontitis patients with impaired access to adequate mechanical hygiene [63,64,65]. It is known that mouth rinses containing chlorhexidine reduce the amount of plaque in the oral cavity [21, 48], which is important as we know that the oral microbiota can play an important role in respiratory infections [66,67,68]. There is also a known relationship between the presence of oral bacteria, specifically viridans group streptococci, and infectious endocarditis (IE) [69,70,71]. A hot topic of discussion is the use of antibiotic prophylaxis in dental procedures that increase the risk of infectious endocarditis. In England, it has been observed that with more restrictive use of antibiotic prophylaxis, the incidence of IE has also increased [72]. Therefore, it is crucial for the health system to design a proper and well-balanced dental antimicrobial stewardship that can minimize the spread of AMR genes but still effectively prevent infection in dental treatments at the same time.
Conclusion
In conclusion, multiple clones conferring resistance to antimicrobials commonly used in dental practices were detected in the studied population, proved to contain known and novel AMR genes when screened by functional-based metagenomics. This emphasizes the importance of reducing all use of antimicrobials, as this will reduce the selective pressure that could drive the spread of AMR genes even from commensal oral bacteria to pathogens. There is a need for more studies in this field to increase our knowledge regarding the AMR crisis.
Availability of data and materials
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Abbreviations
- AMR:
-
Antimicrobial resistance
- CTAB:
-
Cetyltrimethylammonium bromide
- BCCP:
-
Biotin carboxyl carrier protein
- Chx:
-
Chlorhexidine
- Tric:
-
Triclosan
- Ery:
-
Erythromycin
- Tet:
-
Tetracycline
- NaOCl:
-
Sodium hypochlorite
- MIC:
-
Minimum inhibitory concentration
- BacRep:
-
BAC-optimized replicator
- HClO:
-
Hypochlorous acid
References
Tansirichaiya S, Reynolds LJ, Cristarella G, Wong LC, Rosendahl K, Roberts AP. Reduced susceptibility to antiseptics is conferred by heterologous housekeeping genes. Microb Drug Resist. 2018;24(2):105–12.
Paitan Y. Current trends in antimicrobial resistance of Escherichia coli. Curr Top Microbiol Immunol. 2018;416:181–211.
Harada K, Asai T. Role of antimicrobial selective pressure and secondary factors on antimicrobial resistance prevalence in Escherichia coli from food-producing animals in Japan. J Biomed Biotechnol. 2010;2010:180682.
Murray AK, Zhang L, Yin X, Zhang T, Buckling A, Snape J, et al. Novel Insights into selection for antibiotic resistance in complex microbial communities. mBio. 2018;9(4):e00969–18.
D’Costa VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C, et al. Antibiotic resistance is ancient. Nature. 2011;477(7365):457–61.
Warinner C, Rodrigues JFM, Vyas R, Trachsel C, Shved N, Grossmann J, et al. Pathogens and host immunity in the ancient human oral cavity. Nat Genet. 2014;46(4):336–44.
Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks ED, Johnston MD, et al. Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS ONE. 2012;7(4):e34953.
Huddleston JR. Horizontal gene transfer in the human gastrointestinal tract: potential spread of antibiotic resistance genes. Infect Drug Resist. 2014;7:167–76.
Bengtsson-Palme J, Kristiansson E, Larsson DGJ. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol Rev. 2018;42(1):fux053.
Lerminiaux NA, Cameron ADS. Horizontal transfer of antibiotic resistance genes in clinical environments. Can J Microbiol. 2019;65(1):34–44.
Crofts TS, Gasparrini AJ, Dantas G. Next-generation approaches to understand and combat the antibiotic resistome. Nat Rev Microbiol. 2017;15(7):422–34.
Tacconelli E, Sifakis F, Harbarth S, Schrijver R, van Mourik M, Voss A, et al. Surveillance for control of antimicrobial resistance. Lancet Infect Dis. 2018;18(3):e99–106.
McEwen SA, Collignon PJ. Antimicrobial resistance: a one health perspective. Microbiol Spectr. 2018;6(2):521–47.
Shaw L, Ribeiro ALR, Levine AP, Pontikos N, Balloux F, Segal AW, et al. The human salivary microbiome is shaped by shared environment rather than genetics: evidence from a large family of closely related individuals. mBio. 2017;8(5):e01237–17.
Baker JL, Bor B, Agnello M, Shi W, He X. Ecology of the oral microbiome: beyond bacteria. Trends Microbiol. 2017;25(5):362–74.
Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43(11):5721–32.
Verma D, Garg PK, Dubey AK. Insights into the human oral microbiome. Arch Microbiol. 2018;200(4):525–40.
Balachandran M, Cross KL, Podar M. Single-cell genomics and the oral microbiome. J Dent Res. 2020;99(6):613–20.
Krishnan K, Chen T, Paster BJ. A practical guide to the oral microbiome and its relation to health and disease. Oral Dis. 2017;23(3):276–86.
Solderer A, Kaufmann M, Hofer D, Wiedemeier D, Attin T, Schmidlin PR. Efficacy of chlorhexidine rinses after periodontal or implant surgery: a systematic review. Clin Oral Investig. 2019;23(1):21–32.
James P, Worthington HV, Parnell C, Harding M, Lamont T, Cheung A, et al. Chlorhexidine mouthrinse as an adjunctive treatment for gingival health. Cochrane Database Syst Rev. 2017;3:CD008676.
Goncalves LS, Rodrigues RC, Andrade Junior CV, Soares RG, Vettore MV. The effect of sodium hypochlorite and chlorhexidine as irrigant solutions for root canal disinfection: a systematic review of clinical trials. J Endod. 2016;42(4):527–32.
Diaz-Torres ML, McNab R, Spratt DA, Villedieu A, Hunt N, Wilson M, et al. Novel tetracycline resistance determinant from the oral metagenome. Antimicrob Agents Chemother. 2003;47(4):1430–2.
Reynolds LJ, Roberts AP, Anjum MF. Efflux in the oral metagenome: the discovery of a novel tetracycline and tigecycline ABC transporter. Front Microbiol. 2016;7:1923.
Diaz-Torres ML, Villedieu A, Hunt N, McNab R, Spratt DA, Allan E, et al. Determining the antibiotic resistance potential of the indigenous oral microbiota of humans using a metagenomic approach. FEMS Microbiol Lett. 2006;258(2):257–62.
Almeida VSM, Azevedo J, Leal HF, Queiroz ATL, da Silva Filho HP, Reis JN. Bacterial diversity and prevalence of antibiotic resistance genes in the oral microbiome. PLoS ONE. 2020;15(9):e0239664.
Moraes LC, So MV, Dal Pizzol TS, Ferreira MB, Montagner F. Distribution of genes related to antimicrobial resistance in different oral environments: a systematic review. J Endod. 2015;41(4):434–41.
Rocas IN, Siqueira JF Jr. Detection of antibiotic resistance genes in samples from acute and chronic endodontic infections and after treatment. Arch Oral Biol. 2013;58(9):1123–8.
Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner ACR, Yu W-H, et al. The human oral microbiome. J Bacteriol. 2010;192(19):5002–17.
Wade W, Thompson H, Rybalka A, Vartoukian S. Uncultured members of the oral microbiome. J Calif Dent Assoc. 2016;44(7):447–56.
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM. Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol. 1998;5(10):R245–9.
Boolchandani M, Patel S, Dantas G. Functional metagenomics to study antibiotic resistance. Methods Mol Biol. 2017;1520:307–29.
Tansirichaiya S, Reynolds LJ, Roberts AP. Functional metagenomic screening for antimicrobial resistance in the oral microbiome. Methods Mol Biol. 2021;2327:31–50.
Mullany P. Functional metagenomics for the investigation of antibiotic resistance. Virulence. 2014;5(3):443–7.
Ye SH, Siddle KJ, Park DJ, Sabeti PC. Benchmarking metagenomics tools for taxonomic classification. Cell. 2019;178(4):779–94.
Verma MK, Ahmed V, Gupta S, Kumar J, Pandey R, Mandhan V, et al. Functional metagenomics identifies novel genes ABCTPP, TMSRP1 and TLSRP1 among human gut enterotypes. Sci Rep. 2018;8(1):1397.
Marathe NP, Janzon A, Kotsakis SD, Flach CF, Razavi M, Berglund F, et al. Functional metagenomics reveals a novel carbapenem-hydrolyzing mobile beta-lactamase from Indian river sediments contaminated with antibiotic production waste. Environ Int. 2018;112:279–86.
Wiegand I, Hilpert K, Hancock REW. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc. 2008;3(2):163–75.
Huang X. CAP3: a DNA sequence assembly program. Genome Res. 1999;9(9):868–77.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10.
Chancey ST, Zahner D, Stephens DS. Acquired inducible antimicrobial resistance in Gram-positive bacteria. Future Microbiol. 2012;7(8):959–78.
Schroeder MR, Stephens DS. Macrolide resistance in Streptococcus pneumoniae. Front Cell Infect Microbiol. 2016;6:98.
Warburton PJ, Ciric L, Lerner A, Seville LA, Roberts AP, Mullany P, et al. TetAB46, a predicted heterodimeric ABC transporter conferring tetracycline resistance in Streptococcus australis isolated from the oral cavity. J Antimicrob Chemother. 2013;68(1):17–22.
Santoro F, Vianna ME, Roberts AP. Variation on a theme; an overview of the Tn916/Tn1545 family of mobile genetic elements in the oral and nasopharyngeal streptococci. Front Microbiol. 2014;5:535.
Lunde TM, Hjerde E, Al-Haroni M. Prevalence, diversity and transferability of the Tn916-Tn1545 family ICE in oral streptococci. J Oral Microbiol. 2021;13(1):1896874.
Villedieu A, Diaz-Torres ML, Roberts AP, Hunt N, McNab R, Spratt DA, et al. Genetic basis of erythromycin resistance in oral bacteria. Antimicrob Agents Chemother. 2004;48(6):2298–301.
Chaffanel F, Charron-Bourgoin F, Libante V, Leblond-Bourget N, Payot S. Resistance genes and genetic elements associated with antibiotic resistance in clinical and commensal isolates of Streptococcus salivarius. Appl Environ Microbiol. 2015;81(12):4155–63.
Loe H, Schiott CR. The effect of mouthrinses and topical application of chlorhexidine on the development of dental plaque and gingivitis in man. J Periodontal Res. 1970;5(2):79–83.
Lim KS, Kam PC. Chlorhexidine–pharmacology and clinical applications. Anaesth Intensive Care. 2008;36(4):502–12.
Karpinski TM, Szkaradkiewicz AK. Chlorhexidine–pharmaco-biological activity and application. Eur Rev Med Pharmacol Sci. 2015;19(7):1321–6.
Wand ME, Bock LJ, Bonney LC, Sutton JM. Mechanisms of increased resistance to chlorhexidine and cross-resistance to colistin following exposure of Klebsiella pneumoniae clinical isolates to chlorhexidine. Antimicrob Agents Chemother. 2016;61(1):e01162–16.
Karow M, Fayet O, Georgopoulos C. The lethal phenotype caused by null mutations in the Escherichia coli htrB gene is suppressed by mutations in the accBC operon, encoding two subunits of acetyl coenzyme A carboxylase. J Bacteriol. 1992;174(22):7407–18.
Hashemi MM, Holden BS, Coburn J, Taylor MF, Weber S, Hilton B, et al. Proteomic analysis of resistance of gram-negative bacteria to chlorhexidine and impacts on susceptibility to colistin, antimicrobial peptides, and ceragenins. Front Microbiol. 2019;10:210.
Abuhaimed TS, Abou Neel EA. Sodium hypochlorite irrigation and its effect on bond strength to dentin. Biomed Res Int. 2017;2017:1930360.
Tronstad L, Andreasen JO, Hasselgren G, Kristerson L, Riis I. pH changes in dental tissues after root canal filling with calcium hydroxide. J Endod. 1981;7(1):17–21.
Kristoffersen Ø, Fristad I. Natriumhypokloritt – anbefalinger og praktisk bruk. Nor Tannlegeforen Tid. 2007;117:656–60.
Miller RA, Britigan BE. Role of oxidants in microbial pathophysiology. Clin Microbiol Rev. 1997;10(1):1–18.
Small DA, Chang W, Toghrol F, Bentley WE. Toxicogenomic analysis of sodium hypochlorite antimicrobial mechanisms in Pseudomonas aeruginosa. Appl Microbiol Biotechnol. 2007;74(1):176–85.
Collao B, Morales EH, Gil F, Polanco R, Calderon IL, Saavedra CP. Differential expression of the transcription factors MarA, Rob, and SoxS of Salmonella typhimurium in response to sodium hypochlorite: down-regulation of rob by MarA and SoxS. Arch Microbiol. 2012;194(11):933–42.
Dukan S, Touati D. Hypochlorous acid stress in Escherichia coli: resistance, DNA damage, and comparison with hydrogen peroxide stress. J Bacteriol. 1996;178(21):6145–50.
Furi L, Haigh R, Al Jabri ZJ, Morrissey I, Ou HY, Leon-Sampedro R, et al. Dissemination of novel antimicrobial resistance mechanisms through the insertion sequence mediated spread of metabolic genes. Front Microbiol. 2016;7:1008.
Beaber JW, Hochhut B, Waldor MK. SOS response promotes horizontal dissemination of antibiotic resistance genes. Nature. 2004;427(6969):72–4.
Bouloux GF, Steed MB, Perciaccante VJ. Complications of third molar surgery. Oral Maxillofac Surg Clin N Am. 2007;19(1):117–28.
Caso A, Hung LK, Beirne OR. Prevention of alveolar osteitis with chlorhexidine: a meta-analytic review. Oral Surg Oral Med Oral Pathol Oral Radiol Endod. 2005;99(2):155–9.
Sanz M, Herrera D, Kebschull M, Chapple I, Jepsen S, Beglundh T, et al. Treatment of stage I–III periodontitis—the EFP S3 level clinical practice guideline. J Clin Periodontol. 2020;47(Suppl 22):4–60.
Perry SE, Huckabee ML, Tompkins G, Milne T. The association between oral bacteria, the cough reflex and pneumonia in patients with acute stroke and suspected dysphagia. J Oral Rehabil. 2020;47(3):386–94.
Scannapieco FA. Role of oral bacteria in respiratory infection. J Periodontol. 1999;70(7):793–802.
Nishizawa T, Niikura Y, Akasaka K, Watanabe M, Kurai D, Amano M, et al. Pilot study for risk assessment of aspiration pneumonia based on oral bacteria levels and serum biomarkers. BMC Infect Dis. 2019;19(1):761.
Sukumar S, Roberts AP, Martin FE, Adler CJ. Metagenomic insights into transferable antibiotic resistance in oral bacteria. J Dent Res. 2016;95(9):969–76.
Parahitiyawa NB, Jin LJ, Leung WK, Yam WC, Samaranayake LP. Microbiology of odontogenic bacteremia: beyond endocarditis. Clin Microbiol Rev. 2009;22(1):46–64.
DeSimone DC, Tleyjeh IM, Correa de Sa DD, Anavekar NS, Lahr BD, Sohail MR, et al. Incidence of infective endocarditis due to Viridans Group Streptococci before and after the 2007 American Heart Association’s Prevention Guidelines: an extended evaluation of the Olmsted County, Minnesota, population and nationwide inpatient sample. Mayo Clin Proc. 2015;90(7):874–81.
Dayer MJ, Jones S, Prendergast B, Baddour LM, Lockhart PB, Thornhill MH. Incidence of infective endocarditis in England, 2000–13: a secular trend, interrupted time-series analysis. Lancet. 2015;385(9974):1219–28.
Acknowledgements
We would like to thank the staff at the university dental clinics for facilitating the collection of saliva samples. The bioinformatics analysis were performed on resources provided by UNINETT Sigma2—the National Infrastructure for High Performance Computing and Data Storage in Norway (NN9724K). The publication charges for this article have been funded by a grant from the publication fund at UiT The Arctic University of Norway
Funding
The current study is supported by the Department of Clinical Dentistry, Faculty of Health Sciences, UiT The Arctic University of Norway.
Author information
Authors and Affiliations
Contributions
Collection of clinical samples and design of experiments were conceived by MAL and SUP. JOW and ENW collected the clinical samples and perform the laboratory work. SUP supervise the laboratory work and MAL did the overall supervision. Manuscript was drafted by JOW and critically reviewed by MAL and SUP. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The current study and method were carried out according to the current role and regulation and the ethical approval was obtained from the Regional Committees for Medical and Health Research Ethics in Norway (Project number 2018/1373/REK nord). All participants volunteered and signed an informed consent form to use their saliva samples in the study.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1
. Primers used in this study.
Additional file 2
. Bacterial strains and plasmids used in the subcloning of putative chlorhexidine and sodium hypochlorite resistance genes.
Additional file 3
. Estimation of the average insert size of the constructed human oral metagenomic library.
Additional file 4
. BlastN results of the metagenomic sequencing data of each saliva oral metagenomic DNA against the resistance genes detected in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wigand, J., Tansirichaiya, S., Winje, E. et al. Functional screening of a human saliva metagenomic DNA reveal novel resistance genes against sodium hypochlorite and chlorhexidine. BMC Oral Health 21, 632 (2021). https://doi.org/10.1186/s12903-021-02000-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12903-021-02000-5