Abstract
Background
Manipulations in Saccharomyces cerevisiae classically depend on use of auxotrophy selection markers. There are several disadvantages to this in a microbial cell factory setting: (1) auxotrophies must first be engineered in prototrophic strains, and many industrial strains are polyploid/aneuploid prototrophs (2) available strain auxotrophies must be paired with available repair plasmids (3) remaining auxotrophies must be repaired prior to development of industrial bioprocesses. Use of dominant antibiotic resistance markers can circumvent these problems. However, there are relatively few yeast antibiotic resistance marker vectors available; furthermore, available vectors contain only one expression cassette, and it is often desirable to introduce more than one gene at a time.
Results
To overcome these problems, eight new shuttle vectors have been developed. The plasmids are maintained in yeast under a 2 μm ori and in E. coli by a pUC ori. They contain two yeast expression cassettes driven by either (1) the constitutive TEF1 and PGK1 promoters, or (2) the constitutive TEF1 promoter and the inducible GAL10 or HXT7 promoters. Expression strength of these promoters over a typical production time frame in glucose/galactose medium was examined, and identified the TEF1 and HXT7 promoters as preferred promoters over long term fermentations. Selection is provided by either aphA1 (conferring resistance to G418 in yeast and kanamycin/neomycin in E. coli) or ble (conferring resistance to phleomycin in both yeast and E. coli). Selection conditions for these plasmids/antibiotics in defined media were examined, and selection considerations are reviewed. In particular, medium pH has a strong effect on both G418 and phleomycin selection.
Conclusions
These vectors allow manipulations in prototrophic yeast strains with expression of two gene cassettes per plasmid, and will be particularly useful for metabolic engineering applications. The vector set expands the (currently limited) selection of antibiotic marker plasmids available for use in yeast, and in addition makes available dual gene expression cassettes on individual plasmids using antibiotic selection. The resistance gene cassettes are flanked by loxP recognition sites to allow CreA-mediated marker removal and recycling, providing the potential for genomic integration of multiple genes. Guidelines for selection using G418 and phleomycin are provided.
Similar content being viewed by others
Background
Saccharomyces cerevisiae is a model organism and an industrial workhorse. It is readily transformed with plasmid expression vectors, and is therefore particularly amenable to functional analysis experiments and genetic engineering applications. Plasmid vectors may be integrative (yeast integrating plasmids, YIp), autonomously replicating high copy-number vectors (yeast episomal plasmids, YEp), or autonomously replicating low copy-number vectors (yeast centromeric plasmids, YCp) [1, 2]. Typically, selection for presence/genomic integration of the plasmid is performed by complementation/repair of a chromosomal mutation resulting in auxotrophy for an amino acid. Commonly-used markers include URA3, HIS3, LEU2, TRP1 and LYS2. A wide variety of yeast strains containing one or several mutations with low reversion rates conferring appropriate auxotrophies are available (e.g. ura3-52, his3-Δ1, leu2-Δ1, trp1-Δ1 and lys2-201[1]; see the Saccharomyces Genome Database, http://www.yeastgenome.org/). However, to use these strains one must have a plasmid bearing an appropriate auxotrophy gene, and vice-versa: the plasmid bearing a particular gene of interest must encode an auxotrophy repair gene for which the cognate auxotrophy is found in the strain of interest. This may not be the case, particularly in heavily engineered strains where more genes are being introduced, as auxotrophies may already be complemented/repaired. Furthermore, any remaining auxotrophies must be repaired in engineered strains for which industrial bioprocesses are being developed (e.g. [3]). In addition, wild-type and industrial strains are typically polyploid or aneuploid prototrophs [4]. Engineering in prototrophs requires use of dominant markers and/or time-consuming construction of auxotrophy mutants, which may be difficult or impossible in a complex genetic background.
An alternative to using auxotrophies is to use dominant resistance markers. Selection genes conferring resistance to various substances are available (reviewed in [2, 5]). Among these, resistance to antibiotics is most commonly used. The antibiotics G418 [6–10], hygromycin B [8–14], phleomycin [15], chloramphenicol [16], nourseothricin [8, 13, 14], bialaphos [13], zeocin [9], glufosinate [9] and aureobasidin A [17] can be used for selection of S. cerevisiae transformed with appropriate resistance genes, and shuttle vectors conferring resistance to several of these antibiotics (or equivalents) in E. coli have also been developed. However, the yeast community has historically used auxotrophy repair/complementation more widely than antibiotic selection because of the insensitivity of yeast to many antibiotics and the frequency of spontaneous resistant mutants [18]. Furthermore, the efficiency of transformation and transformant recovery is often lower using antibiotic marker selection. Nowadays, antibiotic resistance vectors that provide improved transformant selection are available (e.g. [19]), and greatly improved yeast transformation methods (e.g., [20]) as well as improved antibiotic selection methods (e.g. [21]) mean than efficiencies similar to those obtained using auxotrophy selection can be achieved.
Current antibiotic selection plasmids typically contain only one gene expression cassette. However, it is often desirable to introduce more than one gene at a time, particularly for metabolic engineering projects where multiple modifications and/or complex pathway reconstruction is required. Several dual gene expression cassette vectors have been developed [22–26], as have single gene expression vectors that can be used combinatorially for metabolic engineering applications [27]. However, all of these vectors rely on auxotrophy selection markers. Currently, to our knowledge there are no dual expression vector plasmids with antibiotic selection available. Here, we describe development of dual gene expression shuttle vectors for yeast transformation/engineering using dominant antibiotic marker genes. Antibiotic selection characteristics of S. cerevisiae strains bearing these plasmids were examined, and guidelines for antibiotic use in defined media are provided.
Results and discussion
Vector construction and promoter analysis
Partow et al. (2010) recently developed a pair of dual cassette expression vectors for use in glucose-containing media. They replaced the bi-directional galactose-inducible GAL1-GAL10 promoter region in pESC-Ura (Stratagene; now supplied by Agilent Technologies http://www.genomics.agilent.com) with a TEF1-PGK1 bi-directional promoter developed from the transcriptional elongation factor EF-1α (TEF1) and phosphoglycerate kinase (PGK1) promoters. These glycolysis gene promoters perform well on glucose-based media in S. cerevisiae CEN.PK-derived strains [22]. Two vectors (pSP-G1 and pSP-G2) were constructed, with the bi-directional promoter in both orientations relative to the multiple cloning site and terminator sequences (either ADH1 or CYC1 terminators from S. cerevisiae) [22].
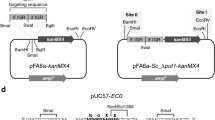
To develop dual gene expression shuttle vectors for yeast transformation/engineering with dominant antibiotic marker genes, we replaced the uracil auxotrophy selection marker (URA3) in pSP-G1 and pSP-G2 with either aphA1 or ble cassettes flanked by loxP recombination sites from pUG6 or pUG66 [28–30] respectively. The aphA1 gene confers resistance to kanamycin (in E. coli) and G418 (in yeast); the ble gene confers resistance to phleomycin in both E. coli and yeast. Gene replacement resulted in four different plasmids with all permutations of TEF1-PGK1 promoter orientation and resistance gene: pCEV-G1-Ph, pCEV-G1-Km, pCEV-G2-Ph and pCEV-G2-Km (Figure 1, Table 1).
Generic plasmid vector map for the pCEV dual gene cassette expression vectors. Each plasmid has two expression cassettes, each driven by a different promoter (P1 or P2), and a selection cassette with either aphA1 or ble genes. Promoter/selection combinations are shown in Figure 1. Each gene of interest can be optionally tagged using either a FLAG epitope tag (Expression Cassette 1) or a c-myc epitope tag (Expression Cassette 2). The selection cassette is controlled by the TEF promoter and terminator from Ashbya gossypii (P AgTEF1 and T AgTEF1 ). The terminator for Expression Cassette 1 is derived from the yeast alcohol dehydrogenase (AHD1) gene. The terminator for Expression Cassette 2 is derived from the yeast cytochrome C (CYC1) gene. The potential exists for integration of the expression cassettes onto the chromosome by amplification using primers with homologous arms; in this case, the selection marker cassette can be removed to recycle the marker via CRE-mediated recombination at the available loxP sites. A pUC origin of replication (Ori) is available for maintenance in E. coli and a 2 μm Ori for maintenance in S. cerevisiae. Selection in E. coli can be performed using either the aphA1/ble selection cassette, which is functional for selection in E. coli, or using the beta-lactamase (bla) ampicillin resistance gene. The dual promoter region is not to scale (size varies depending on which promoters are present) and the selection marker is also not to scale (can be either aphA1 or ble). Different vector components in these two regions, as well as the unique restriction enzyme sites found in each multiple cloning site (MCS), are shown in Table 2. Note that the MCS spans the epitope tag sites for each expression cassette; choice of restriction sites will determine whether or not the tag sequence is retained in the construct.
In contrast to Partow et al. (2010), who examined promoter strength in CEN.PK-derived strains, we observed unexpectedly poor expression strength when using the PGK1 expression cassette in our vectors with S288C-derived strains (data not shown). We therefore examined the relative strength of several promoters using a β-galactosidase reporter gene assay in an S288C-derived strain. As well as the TEF1 and PGK1 promoters, we included the GAL10 promoter, since the S288C-derived strain EPY210C [31] that we were using also has galactose-inducible modifications, and we were interested in using this promoter for future engineering experiments. In addition, we tested a high affinity hexose transporter (HXT7) promoter [22]. The HXT7 gene is up-regulated when glucose concentrations are low and repressed when glucose concentrations are high [22, 32–34], and therefore provides a potential alternative for driving gene expression in media with low/no glucose - including galactose or glucose/galactose medium used for GAL promoter-driven expression (e.g., [3, 35–37]). Available promoter:β-galactosidase yeast integrating plasmids were used for the TEF1 and PGK1 promoters [22], and similar plasmids were constructed for the HXT7 and GAL10 promoters (see Methods).
Reporter gene activity driven by the TEF1, PGK1, GAL10 and HXT7 promoters was examined over several days in cultures grown on rich medium containing 0.2% glucose + 1.8% galactose (Figure 2). This sugar combination is used to build up biomass prior to inducing expression of desired genes (e.g. [3, 35, 37]). In this medium, glucose is rapidly consumed within the first 6–8 hours of fermentation; growth then shifts to being galactose-based. This experiment confirmed that the PGK1 promoter drives relatively poor expression in S. cerevisiae S288C-derived strains growing on galactose. Over the short term (24 hr, by which time cultures are in stationary phase), the TEF1 promoter emerged as the strongest promoter, with the GAL10 and HXT7 promoters driving slightly lower, but similar levels of expression. By the third day, β-galactosidase activity was similar for the TEF1, GAL10 and HXT7 promoters. By day 7, the TEF1 and HXT7 promoters were still driving good levels of expression, while β-galactosidase activity driven by the GAL10 promoter was much weaker. Depending on the time scale of the fermentation and relative stability of the gene products, both the GAL10 and HXT7 promoters show utility for glucose/galactose fermentations using S288C-derived strains. We therefore replaced the weak PGK1 promoter with GAL10 or HXT7 promoters. This resulted in four new dual gene expression plasmids (Table 1, Figure 1). In plasmids containing the GAL10-controlled expression cassette (pCEV-G3-Km and pCEV-G3-Ph), the potential exists to generate a third expression cassette by inserting an MCS and a terminator sequence between the bidirectional GAL10-1 promoter region and the TEF1 promoter. This third expression cassette would be controlled by the GAL1 promoter. Genes already fused to a terminator can be placed under the control of the GAL1 promoter by insertion through the unique FseI restriction site in pCEV-G3-Ph, or the unique FseI, KroI, NaeI or NgoMIV restriction sites in pCEV-G3-Km.
β-Galactosidase activity driven by various promoters in the S. cerevisiae S288C-derived strain EPY210C growing on galactose. EPY210C [31] was used as a base for strain construction to test promoter expression strength using promoter:lacZ constructs on integrative plasmids. Plasmids pSF015 (HXT7 promoter), pSF016 (PGK1 promoter), and pSF019 (TEF1 promoter), bearing promoters amplified from S. cerevisiae CEN.PK, have been described previously [22]. We reconstructed a P GAL10 : lacZ fusion construct (pGAL10lac) by amplifying the relevant region from S. cerevisiae S288C genomic DNA (see Methods). The negative control was the promoterless lacZ plasmid pSF011 [22]. β-Galactosidase assays are described in the Methods. Bars are means of n = 3 biological replications; errors are standard deviations.
Selection characteristics on G418 and phleomycin
In the process of applying the expression vectors for yeast engineering projects, we learned a number of things about using G418 andas selective agents. In particular, we observed plasmid instability and poor selection on defined media. For all 2 μ plasmids, we observed spontaneous plasmid loss after 48 hr in non-selective medium (data not shown); we have previously observed this for 2 μ auxotrophy selection plasmids as well [31]. We have summarized our observations, along with a review of the relevant literature, in the sections below. In addition, we did a detailed study on the effects of pH on selection.
G418
The aphA1 gene from the E. coli transposon Tn903[38] encodes an aminoglycoside 3′-phosphotransferase activity which confers resistance against G418 (sold as Geneticin® by Invitrogen/Life Technologies) in S. cerevisiae and kanamycin in E. coli[4, 6, 18, 39]. It is also referred to as kan (for kanamycin resistance), neo (for neomycin resistance) or npt1 (for neomycin phosphotransferase). G418 and kanamycin are 2-deoxystreptamine aminoglycoside antibiotics; they act by inhibition of protein translation at the elongation step [40]. The level of resistance conferred depends on several factors, including expression strength and copy number. The kanMX module [39] contains aphA1 (kan) under the control of promoter and terminator sequences from the TEF gene of the filamentous fungus Ashbya gossypii. It confers resistance to G418 at 200 μg/mL in S. cerevisiae grown on YPD (YEPD) and kanamycin at 30 μg/mL in E. coli grown on LB. This cassette was subsequently flanked by loxP sites to allow facile marker gene removal during knock-out and tagging experiments via the CRE-loxP recombination system [28]. The kanMX loxP construct was used in the current work for plasmid assembly.
Medium composition affects the efficiency of G418 selection [18]. In particular, pH has a strong effect on aminoglycoside phosphotransferase activity [41]; therefore, medium pH may modulate selective pressure. Activity is higher at pH 7 than at decreased pH such is found in typical defined yeast media (often around 5.5; [42, 43]). While selection in complex YPD medium is effective at 200 μg/mL G418 in YPD, we encountered problems when using defined media. To examine how pH affects selective power under these conditions, we tested growth on defined medium with the pH adjusted to either 5 or 7 and supplemented with a range of G418 concentrations (Table 2). At both pH 5 and pH 7, selection was ineffective at 200 μg/mL G418. Effective selection could be achieved at 400 μg/mL G418 at both pH values. This indicates that the aminoglycoside transferase activity was in fact similar under both pH conditions, and that some other medium component/condition affects selectivity at concentrations below 400 μg/mL G418. In addition to pH, ammonium sulphate is known to interfere with G418 selection; it can be replaced with monosodium glutamate at 1 g/L to improve selection characteristics without modifying medium pH [44].
It should also be noted that G418 efficiency can decrease at high cell densities in mammalian cells [45]; this can presumably occur in yeast as well. In addition, commercial G418 preparations can vary in purity and potency, even in between batches [28]. In summary, the following considerations should be taken into account when using G418 as a selective agent:
-
1.
Medium pH should be considered, as aminoglycoside transferase activity decreases with decreasing pH.
-
2.
Medium composition affects selection conditions.
-
3.
Selection conditions may vary with strain.
-
4.
Loss of G418 resistance may occur over long incubation periods; this is most likely related to high culture density. Dosing in more antibiotic may improve selection over long cultivations to prevent plasmid loss.
-
5.
Plating density may also affect selection.
-
6.
Growth rates may decrease in the presence of G418.
-
7.
Adjusting the pH to neutral or increasing antibiotic concentrations may be required.
Phleomycin
The ble gene from the bacterial transposon Tn5 confers resistance to the metallo-glycopeptide class antibiotics bleomycin and phleomycin in both prokaryotes and eukaryotes [15, 46–48]. These antibiotics act by binding to DNA and causing strand breakage [49–51]. Efficient DNA binding and breakage requires metal ions (ferrous ions) and molecular oxygen as cofactors [50, 51]. Accordingly, S. cerevisiae is significantly more resistant to phleomycin under anaerobic conditions [15], and presence of EDTA decreases/removes toxicity [52]. Sensitivity to phleomycin depends on growth phase in S. cerevisiae: stationary phase cells are less sensitive than exponentially growing cells [52].
The Ble protein inactivates phleomycin family antibiotics by binding to them [53]. The ble gene was first developed as a selective marker for phleomycin resistance in S. cerevisiae by Gatignol et al. [15]. The aphA1 gene in the kanMX module was replaced by the ble gene to develop a second TEF1 promoter-driven dominant marker gene knock-out cassette [29]. This construct conferred resistance to 7.5 μg/mL phleomycin in S. cerevisiae (CEN.PK2-1C) growing on YPD. In early studies, incubation in non-selective medium prior to application of selection pressure was required to obtain transformed cells with reasonable efficiency [21], though Gueldener et al. [29] later found that pre-incubation did not enhance transformation efficiency. This may be due to genotypic differences between yeast strains (OL1 in the former study and CEN.PK-derived in the latter study), differences in expression characteristics (CYC1 promoter in the former vs. TEF1 in the latter) and/or differences in growth media. In our hands, we found that a three hour incubation in non-selective medium is optimal for efficient transformation in the S288C-derived strain EPY210C being selected on YPD (data not shown). Plating at high densities can also decrease transformation efficiency [21].
Most commercial preparations of phleomycin suggest using 10 μg/mL for selection of S. cerevisiae on YPD, as determined by Gatignol et al. [19]. However, we observed colonies of S288C-derived strains (without plasmid) on YPD plates containing phleomycin at 10 μg/mL after three days (data not shown). This suggests that spontaneous resistance can develop over that timeframe under those conditions. Increasing the concentration to 15 μg/mL inhibited growth significantly over three days; using 20 μg/mL completely inhibited development of spontaneous resistance over a seven day period (data not shown).
Phleomycin sensitivity decreases at lower pH values [54], such as commonly found in defined yeast media [42, 43]. We therefore examined selection efficiency in defined medium at pH 5 and 7, again using several different concentrations of phleomycin (Table 2). Selection was essentially ineffective at pH 5. Some selective pressure was observed at 60–80 µg/mL; selective pressure increased significantly at 100 µg/mL, but growth was not completely repressed in control strains even at 100 µg/mL. When the pH was adjusted to 7, selection was highly effective at concentrations at or above 40 µg/mL.
We also observed that the TEF1 promoter-driven ble module could confer resistance to phleomycin in E. coli (data not shown), as shown for other yeast ble constructs [15]. This allows dual antibiotic selection of transformed E. coli (using ampicillin at 100 μg/mL and phleomycin at 5 μg/mL).
In summary, the following considerations should be taken into account when using phleomycin as a selective agent:
-
1.
Phleomycin activity requires the presence of molecular oxygen and ferrous ions. Anaerobic fermentations may require significantly higher phleomyin concentrations.
-
2.
Plating at high densities can decrease transformation efficiency.
-
3.
Depending on strain/medium, efficient selection may require incubation on non-selective medium prior to application of selective pressure, and may also affect maintenance of selection.
-
4.
Sensitivity to phleomycin decreases below neutral pH. Phleomycin concentration may need to be increased significantly in media with lower pH values; ideally, medium pH should be adjusted to 7.
-
5.
Antibiotic titration/dosing may be required during prolonged cultivations to prevent development of spontaneous resistance and/or plasmid loss.
-
6.
Stationary phase cells are less sensitive to phleomycin than exponentially growing cells.
Conclusions
In summary, we have developed dominant marker plasmids containing two gene expression cassettes that provide selection in Saccharomyces cerevisiae via either phleomycin or G418 resistance. A caveat of using G418 and phleomycin for selection is the requirement for adjustment of medium pH to neutral and/or increasing antibiotic concentration to maintain selective pressure on defined media. Expression of genes in each plasmid can be controlled by a combination of the TEF1 promoter with either the PGK1, GAL10 or HXT7 promoters. Each promoter is followed by a multiple cloning site (MCS) and a transcription termination sequence. In addition, both expression cassettes encode epitope tags that can be used for C- or N-terminal tagging. The plasmids are shuttle plasmids with a 2 μm ori (yeast) and a pUC ori (E. coli). Selection in E. coli is conferred by a beta-lactamase (bla) ampicillin resistance gene as well as the aphA1 or ble genes. The TEF1, GAL10 and HXT7 promoters drive strong expression in galactose medium; TEF1 and HXT7 are preferred for high expression over prolonged fermentations. The HXT7 promoter might also be useful in other non-/low-glucose media (e.g. sucrose-based media). We have subsequently used these plasmids successfully for expression of isoprenoid production genes in yeast (data not shown). Genomic integration of the expression and selection cassettes could be readily achieved by amplification using primers with suitable homologous arms [55]. PCR-mediated integration of similarly large fragments has been demonstrated recently, and useful integration loci have been characterized [23, 27]. The presence of the loxP sites allows removal of selective marker genes by CreA-mediated recombination so they can be re-used for subsequent modifications [28, 55]. The aphA1 or ble genes can also be replaced by genes encoding alternative selection functions (e.g., see review in Introduction) to expand the available selection systems. The antibiotic marker vectors can also be used to pyramid further modifications into strains where available auxotrophy markers have been exhausted by insertion of multiple constructs. In addition, they may also be useful in other yeasts and filamentous fungi that are sensitive to phleomycin or G418; for example, the yeasts Schizosaccharomyces pombe[56] and Cryptococcus neoformans[57] and the filamentous fungi Aspergillus flavus[54], Neurospora crassa[58], Aspergillus nidulans[58] and Pleurotus ostreatus[59] are all sensitive to phleomycin. The plasmids are available through AddGene (http://www.addgene.org/) and Euroscarf (http://web.uni-frankfurt.de/fb15/mikro/euroscarf/). Detailed plasmid datasheets are available in Additional file 1 and annotated sequence files in Clone Manager format (Sci-Ed Software, http://www.scied.com) are available through the AddGene website (http://www.addgene.org/). Sequence data can also be downloaded from GenBank using the provided accession numbers (Table 1).
Methods
Strains and media
E. coli strain MG1655 (Coli Genetic Stock Centre CGSC#7740, New Haven CT, USA) or α-select (Bioline; Alexandria NSW, Australia) were used for plasmid manipulations; they were maintained on LB medium [60], and transformations were performed by chemical transformation according to the manufacturer’s instructions. S. cerevisiae S288C was kindly provided by the Australian Wine Research Organisation, and was originally sourced from the American Type Culture Collection (http://www.atcc.org/; ATCC Accession: 204508). The S288C-derived S. cerevisiae strain EPY210C [31] was used as a base for strain construction to test promoter expression strength (see below). S. cerevisiae strains were grown on YPD [43] for general growth and selection, and on YPD with the glucose replaced with 0.2% glucose and 1.8% galactose (YPDG) for promoter analysis experiments (see below). To examine the effect of pH on selective power in defined medium, yeast were grown in SD medium [43] with the pH adjusted to either 5 or 7 (using potassium hydroxide) and filter-sterilised. All media were supplemented with appropriate antibiotics at the concentrations stated. Antibiotics G418 sulfate and Phleomycin were sourced from Invivogen (San Diego, California; cat. # ant-gn-5 and ant-ph-5, respectively).
Plasmids and plasmid construction
The uracil auxotrophy selection marker in pSP-G1 and pSP-G2 [22] was replaced with either aphA1 or ble cassettes containing loxP recombination sites from pUG6 or pUG66 [28, 29] (sourced from Euroscarf), respectively. pUG6 and pUG66 were prepared for use as PCR templates by digesting with BglI. The digested plasmids were used as PCR templates with primers HA1-kanphleo and HA2-kanphleo (Table 3). For HA1-kanphleo, the 5′ sequence is homologous with sequence just downstream from the 2 μm ori pUG6/pUG66. The SalI site in pUG6 is destroyed, but the AccI site (underlined) remains for facile removal of the cassette. For HA2-kanphleo, the 5′ sequence is homologous with sequence at the end of the ADH1 terminator in pUG6/pUG66. The sequences in italics are the priming sites in pUG6/pUG66. Resulting PCR products have arms that are homologous to sequences upstream and downstream of the URA marker in pSP-G1 and pSP-G2; replacement of the URA3 gene with the kanMX or ble cassettes removes the F1 ori in pSP-G1/pSP-G2. PCR products were digested with DpnI. The vectors (pSP-G1/pSP-G2) were prepared by digestion with EcoRV and NcoI and then cleaned. Fragments were transformed into electro-competent E. coli MG1655 cells harbouring the λ Red recombinase plasmid pKD46 [61] for cloning by homologous recombination ('recombineering’; [62, 63]). Selection was performed on ampicillin (100 μg/mL) plus either kanamycin (30 μg/mL) or phleomycin (5 μg/mL). Clones were screened by colony PCR using primers 2muDown and ADH1-T F1 (Table 3). Plasmids isolated from positive clones were screened by restriction analysis and the insert fully sequenced using the same primers. Resulting plasmids were pCEV-G1-Ph, pCEV-G1-Km, pCEV-G2-Ph and pCEV-G2-Km.
Using pCEV-G2-Ph and pCEV-G2-Km as base plasmids, a second set of plasmids where the PGK1 promoter was replaced with the strong galactose-inducible GAL10 promoter or the HXT7 promoter (which is induced at low glucose concentrations) was also constructed. The complete GAL10-GAL1 promoter region was amplified from S. cerevisiae S288C using primers GAL10LacF and Gal10LacR (Table 3). The pSF011 promoterless target vector was opened using BamHI and XbaI restriction sites and the amplified fragment with homologous arms was cloned in by homologous recombination as described previously [64] using E. coli strain One Shot Omnimax 2 T1 (Life Technologies). The resulting plasmid was called pGAL10Lac. The GAL10 promoter was then PCR-amplified from pGAL10Lac using the primers GAL10P2B and GAL10P1A (Table 3). The HXT7 promoter was PCR amplified from pSF015 [22] using the primers HXT7P1B and HXT7P2A (Table 3). The PCR products and the vectors pCEV-G2-Km and pCEV-G2-Ph were digested with FseI and SpeI; Dpn I was also included in the PCR fragment digests to eliminate template DNA. The vectors were purified by gel extraction and the PCR products were purified through a column (QIAquick PCR purification kit, Qiagen). Ligation products were transformed into E. coli α-Select Silver Efficiency chemically competent cells (Bioline). Selection was performed on ampicillin (100 μg/mL). Colonies were screened by PCR using ADH1-T R1 paired with either SFB018 for pCEV-G3 constructs or SFB017 for pCEV-G4 constructs (Table 3). Plasmids were confirmed by restriction analysis and sequencing. The resulting plasmids were pCEV-G3-Km, pCEV-G3-Ph, pCEV-G4-Km and pCEV-G4-Ph.
The promoter:lacZ integrative plasmids pSF015 (HXT7 promoter), pSF016 (PGK1 promoter), and pSF019 (TEF1 promoter), as well as the promoterless lacZ control integrative plasmid pSF011, were kindly provided by Prof. Jens Nielsen (Chalmers University of Technology, Sweden). These constructs have been described previously [22]. The Partow et al. (2010) study also included GAL1 and GAL10 promoters isolated from S. cerevisiae CEN.PK; we reconstructed the P GAL10 :lacZ fusion construct by amplifying the relevant regions from S. cerevisiae S288C genomic DNA using GAL10LacF and GAL10LacR primers (Table 3). The pSF011 plasmid was opened using Bam HI and Xba I restriction sites and amplified fragments with homologous arms were cloned in by homologous recombination as described previously [64] using E. coli strain One Shot Omnimax 2 T1 (Life Technologies). The GAL promoter region was cloned as a complete region (P GAL1 + P GAL10 in divergent orientations) and inserted in the GAL10 orientation to produce pGAL10Lac.
Strain construction
S. cerevisiae transformation was performed using the lithium acetate method [20]. For YEp dominant marker plasmids, transformants were selected on YPD supplemented with 200 μg/mL G418 or 20 μg/mL phleomycin after a three hour pre-incubation period on non-selective medium. Integrative plasmids were linearized using NcoI prior to transformation. Selection was performed on uracil SD drop-out agar (Sigma-Aldrich; Sydney, Australia) supplemented with 2% glucose. For all transformations, biological replicates were generated as described previously [65]; briefly, three individual colonies were selected from the transformation plates and denoted as biological replicates for each new strain. Colony PCR was performed using primers CYC1-TR1 (GGGACCTAGACTTCAGGTTGTC) and Lac9434r (GAAGCCTGCGATGTCGGTTTC). Each biological replicate was purified by streaking out on uracil SD drop-out agar and inoculated into 5 mL SD uracil drop-out broth (Sigma-Aldrich; Sydney, Australia) supplemented with 2% glucose. Cultures were grown overnight at 30°C, 200 rpm. Glycerol stocks were prepared in 20% glycerol and stored at -80°C.
For antibiotic sensitivity testing, EPY210C was transformed with pCEV-G2-Ph and pCEV-G2-Km to generate EPY210C(pCEV-G2-Ph) and EPY201C(pCEV-G2-Km), respectively. Biological replicates and glycerol stocks were produced as described above. To confirm plasmid presence, plasmid was extracted from each strain and verified by PCR.
β-galactosidase assays
For β-galactosidase assays, glycerol stocks were streaked out on solid YPD and grown at 30°C. Single colonies (three per strain; technical replications) were inoculated into 5 mL YPD broths and cultured overnight at 30°C, 200 rpm. Overnight cultures were used to inoculate 30 mL YPDG to an OD660 of 0.2. Cultures were sampled at 24 hr and again at 3 days and 7 days. To extract protein, 5 mL culture was centrifuged (3000 × g, 4°C, 5 min) and the resulting pellet washed twice with ice-cold H2O. The cell pellet was resuspended in 1 mL extraction buffer (25 mM Tris–HCl, 1 mM DTT, 5 mM EDTA, pH 7.5) containing 20 μL of protease inhibitor cocktail (Sigma-Aldrich; Sydney, Australia) and the resulting suspension transferred to a pre-chilled 2 mL tube half filled with acid-washed glass beads (Ø = 0.5 mm). Cells were disrupted with a Mini Beadbeater-1 (Biospec Products, Bartlesville, OK, USA) using 4 x 20 s bead-beating bursts at 4800 oscillations per minute with 1 min cooling in ice-water between beatings. The disrupted cells were centrifuged (13 000 × g, 4°C, 5 min) and the supernatant collected. Protein concentration was determined by the method of Bradford [66] using a BioRad Protein Assay kit (BioRad, Gladesville, NSW, Australia) and bovine serum albumin as a standard. β-Galactosidase assays were performed using high-throughput microtitre plate method as described previously [67] with minor modifications, including use of a kinetic approach for the enzyme assay and standardisation to extract protein concentration. Briefly, 5 μL of supernatant extract was added in triplicate to microtitre plate wells containing 85 μL Z buffer [68]. The reaction was initiated by adding 10 μL 4 mg/mL O-nitrophenyl-β-d-galactoside (ONPG) prepared in phosphate buffer. The reaction was monitored at 37°C using a FLOUstar Omega plate-reading spectrophotometer (BMG Labtech, Mornington, VIC, Australia) for at least 30 min. The initial linear phase (normally within the first 8 min of the reaction) was used to calculate the rate. β-Galactosidase activity was expressed as ΔA450/min/mg protein. S. cerevisiae EPY210C transformed with linearised pSF011 was used as a negative control.
Antibiotic sensitivity testing
Antibiotic sensitivity was tested using solid SD medium with the pH adjusted to either 5 or 7, supplemented with G418 at 200, 400, 600 or 800 µg/mL, or with phleomycin at 20, 40, 60, 80 or 100 µg/mL. Wild type S288C was used as a negative control, EPY210C(pCEV-G2-Ph) and EPY201C(pCEV-G2-Km) were used as representative strains bearing phleomycin and G418 resistance plasmids, respectively. Strains were streaked out on SD medium with the pH adjusted to 7; for EPY210C(pCEV-G2-Ph), the medium was supplemented with 20 µg/mL phleomycin, and for EPY201C(pCEV-G2-Km) the medium was supplemented with 200 µg/mL G418. Plates were grown for 2 d at 30°C. Single colonies from all three strains were streaked out on the various SD media described above and incubated at 30°C for 3 days to test antibiotic selection characteristics.
References
Sherman F, et al: Yeast genetics. The encyclopedia of molecular biology and molecular medicine: volume 6. 1997, 302-325. Weinheim, Germany: VCH Publisher
Romanos MA, Scorer CA, Clare JJ: Foreign gene expression in yeast: a review. Yeast. 1992, 8: 423-488. 10.1002/yea.320080602
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM, Ho KA, Eachus RA, Ham TS, Kirby J, et al: Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature. 2006, 440: 940-943. 10.1038/nature04640
Hadfield C, Jordan BE, Mount RC, Pretorius GHJ, Burak E: G418-resistance as a dominant marker and reporter for gene expression in Saccharomyces cerevisiae. Curr Genet. 1990, 18: 303-313. 10.1007/BF00318211
Stansfield I, Stark MJR: Yeast gene analysis. 2007, San Diego, CA: Academic Press, 2
Jimenez A, Davies J: Expression of a transposable antibiotic-resistance element in Saccharomyces. Nature. 1980, 287: 869-871. 10.1038/287869a0
Bardazzi I, Casalone E: Construction of two new vectors for transformation of laboratory, natural and industrial Saccharomyces cerevisiae strains to trifluoroleucine and G418 resistance. Folia Microbiol (Praha). 2004, 49: 534-538. 10.1007/BF02931529
Taxis C, Knop M: System of centromeric, episomal, and integrative vectors based on drug resistance markers for saccharomyces cerevisiae. Biotechniques. 2006, 40: 73-78. 10.2144/000112040
Sadowski I, Lourenco P, Parent J: Dominant marker vectors for selecting yeast mating products. Yeast. 2008, 25: 595-599. 10.1002/yea.1604
Baruffini E, Serafini F, Lodi T: Construction and characterization of centromeric, episomal and GFP-containing vectors for saccharomyces cerevisiae prototrophic strains. J Biotechnol. 2009, 143: 247-254. 10.1016/j.jbiotec.2009.08.007
Gritz L, Davies J: Plasmid-encoded hygromycin B resistance: the sequence of hygromycin B phosphotransferase gene and its expression in Escherichia coli and Saccharomyces cerevisiae. Gene. 1983, 25: 179-188. 10.1016/0378-1119(83)90223-8
Kaster KR, Burgett SG, Ingolia TD: Hygromycin-B resistance as dominant selectable marker in yeast. Curr Genet. 1984, 8: 353-358. 10.1007/BF00419824
Goldstein AL, McCusker JH: Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast. 1999, 15: 1541-1553. 10.1002/(SICI)1097-0061(199910)15:14<1541::AID-YEA476>3.0.CO;2-K
Carter Z, Delneri D: New generation of loxP-mutated deletion cassettes for the genetic manipulation of yeast natural isolates. Yeast. 2010, 27: 765-775. 10.1002/yea.1774
Gatignol A, Baron M, Tiraby G: Phleomycin resistance encoded by the ble gene from transposon Tn5 as a dominant selectable marker in Saccharomyces cerevisiae. Mol Gen Genet. 1987, 207: 342-348. 10.1007/BF00331599
Hadfield C, Cashmore AM, Meacock PA: An efficient chloramphenicol-resistance marker for Saccharomyces cerevisiae and Escherichia coli. Gene. 1986, 45: 149-158. 10.1016/0378-1119(86)90249-0
Hashida-Okado T, Ogawa A, Kato I, Takesako K: Transformation system for prototrophic industrial yeasts using the AUR1 gene as a dominant selection marker. FEBS Lett. 1998, 425: 117-122. 10.1016/S0014-5793(98)00211-7
Webster TD, Dickson RC: Direct selection of Saccharomyces cerevisiae resistant to the antibiotic G418 following transformation with a DNA vector carrying the kanamycin-resistance gene of Tn903. Gene. 1983, 26: 243-252. 10.1016/0378-1119(83)90194-4
Gatignol A, Dassain M, Tiraby G: Cloning of Saccharomyces cerevisiae promoters using a probe vector based on phleomycin resistance. Gene. 1990, 91: 35-41. 10.1016/0378-1119(90)90159-O
Gietz RD, Woods RA: Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 2002, 350: 87-96.
Wenzel TJ, Migliazza A, Steensma HY, van den Berg JA: Efficient selection of phleomycin-resistant Saccharomyces cerevisiae transformants. Yeast. 1992, 8: 667-668. 10.1002/yea.320080810
Partow S, Siewers V, Bjørn S, Nielsen J, Maury J: Characterization of different promoters for designing a new expression vector in Saccharomyces cerevisiae. Yeast. 2010, 27: 955-964. 10.1002/yea.1806
Mikkelsen MD, Buron LD, Salomonsen B, Olsen CE, Hansen BG, Mortensen UH, Halkier BA: Microbial production of indolylglucosinolate through engineering of a multi-gene pathway in a versatile yeast expression platform. Metab Eng. 2012, 14: 104-111. 10.1016/j.ymben.2012.01.006
pESC vectors. http://www.genomics.agilent.com/article.jsp?pageId=596
Miller CA, Martinat MA, Hyman LE: Assessment of aryl hydrocarbon receptor complex interactions using pBEVY plasmids: expression vectors with bi-directional promoters for use in saccharomyces cerevisiae. Nucleic Acids Res. 1998, 26: 3577-3583. 10.1093/nar/26.15.3577
Li AM, Liu ZS, Li QX, Yu L, Wang DC, Deng XM: Construction and characterization of bidirectional expression vectors in saccharomyces cerevisiae. FEMS Yeast Res. 2008, 8: 6-9. 10.1111/j.1567-1364.2007.00335.x
Fang F, Salmon K, Shen MWY, Aeling KA, Ito E, Irwin B, Tran UPC, Hatfield GW, Da Silva NA, Sandmeyer S: A vector set for systematic metabolic engineering in Saccharomyces cerevisiae. Yeast. 2011, 28: 123-136. 10.1002/yea.1824
Güldener U, Heck S, Fielder T, Beinhauer J, Hegemann JH: A new efficient gene disruption cassette for repeated use in budding yeast. Nucleic Acids Res. 1996, 24: 2519-2524. 10.1093/nar/24.13.2519
Gueldener U, Heinisch J, Koehler GJ, Voss D, Hegemann JH: A second set of loxP marker cassettes for cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res. 2002, 30: e23- 10.1093/nar/30.6.e23
EUROpean saccharomyces cerevisiae ARchive for functional analysis. http://web.uni-frankfurt.de/fb15/mikro/euroscarf
Behrendorff JBYH, Vickers CE, Chrysanthopoulos P, Nielsen LK: 2, 2-Diphenyl-1-picrylhydrazyl as a screening tool for recombinant monoterpene biosynthesis. Microb Cell Fact. 2013, 12: 76. 10.1186/1475-2859-12-76
Diderich JA, Schepper M, van Hoek P, Luttik MA, van Dijken JP, Pronk JT, Klaassen P, Boelens HF, de Mattos MJ, van Dam K, Kruckeberg AL: Glucose uptake kinetics and transcription of HXT genes in chemostat cultures of Saccharomyces cerevisiae. J Biol Chem. 1999, 274: 15350-15359. 10.1074/jbc.274.22.15350
Sedlak M, Ho NWY: Characterization of the effectiveness of hexose transporters for transporting xylose during glucose and xylose co-fermentation by a recombinant Saccharomyces yeast. Yeast. 2004, 21: 671-684. 10.1002/yea.1060
Ye L, Berden JA, van Dam K, Kruckeberg AL: Expression and activity of the Hxt7 high-affinity hexose transporter of Saccharomyces cerevisiae. Yeast. 2001, 18: 1257-1267. 10.1002/yea.771
Westfall PJ, Pitera DJ, Lenihan JR, Eng D, Woolard FX, Regentin R, Horning T, Tsuruta H, Melis DJ, Owens A, et al: Production of amorphadiene in yeast, and its conversion to dihydroartemisinic acid, precursor to the antimalarial agent artemisinin. Proc Natl Acad Sci U S A. 2012, 109: E111-E118. 10.1073/pnas.1110740109
Ro D-K, Ouellet M, Paradise E, Burd H, Eng D, Paddon C, Newman J, Keasling J: Induction of multiple pleiotropic drug resistance genes in yeast engineered to produce an increased level of anti-malarial drug precursor, artemisinic acid. BMC Biotechnol. 2008, 8: 83- 10.1186/1472-6750-8-83
Lindahl AL, Olsson ME, Mercke P, Tollbom O, Schelin J, Brodelius M, Brodelius PE: Production of the artemisinin precursor amorpha-4, 11-diene by engineered Saccharomyces cerevisiae. Biotechnol Lett. 2006, 28: 571-580. 10.1007/s10529-006-0015-6
Oka A, Sugisaki H, Takanami M: Nucleotide sequence of the kanamycin resistance transposon Tn903. J Mol Biol. 1981, 147: 217-226. 10.1016/0022-2836(81)90438-1
Wach A, Brachat A, Pohlmann R, Philippsen P: New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast. 1994, 10: 1793-1808. 10.1002/yea.320101310
Magnet S, Blanchard JS: Molecular insights into aminoglycoside action and resistance. ChemInform. 2005, 105: 477-496. no-no
Lang-Hinrichs C, Berndorff D, Seefeldt C, Stahl U: G418 resistance in the yeast Saccharomyces cerevisiae: comparison of the neomycin resistance genes from Tn5 and Tn903. Appl Microbiol Biotechnol. 1989, 30: 388-394. 10.1007/BF00296629.
Verduyn C, Postma E, Scheffers WA, Van Dijken JP: Effect of benzoic acid on metabolic fluxes in yeasts: a continuous-culture study on the regulation of respiration and alcoholic fermentation. Yeast. 1992, 8: 501-517. 10.1002/yea.320080703
Sherman F: Getting started with yeast. Methods Enzymol. 2002, 350: 3-41.
Cheng TH, Chang CR, Joy P, Yablok S, Gartenberg MR: Controlling gene expression in yeast by inducible site-specific recombination. Nucleic Acids Res. 2000, 28: E108- 10.1093/nar/28.24.e108
G418 sulfate product insert. http://www.invivogen.com/PDF/G418_TDS.pdf
Genilloud O, Garrido MC, Moreno F: The transposon Tn5 carries a bleomycin-resistance determinant. Gene. 1984, 32: 225-233. 10.1016/0378-1119(84)90050-7
Collis CM, Hall RM: Identification of a Tn5 determinant conferring resistance to phleomycins, bleomycins, and tallysomycins. Plasmid. 1985, 14: 143-151. 10.1016/0147-619X(85)90074-5
Mazodier P, Cossart P, Giraud E, Gasser F: Completion of the nucleotide sequence of the central region of Tn5 confirms the presence of three resistance genes. Nucleic Acids Res. 1985, 13: 195-205. 10.1093/nar/13.1.195
Stern R, Rose JA, Friedman RM: Phleomycin-induced cleavage of deoxyribonucleic acid. Biochemistry. 1974, 13: 307-312. 10.1021/bi00699a012
Povirk LF, Hogan M, Dattagupta N, Buechner M: Copper(II).bleomycin, iron(III).bleomycin, and copper(II) phleomycin: comparative study of deoxyribonucleic acid binding. Biochemistry. 1981, 20: 665-671. 10.1021/bi00506a034
Sleigh MJ: The mechanism of DNA breakage by phleomycin in vitro. Nucleic Acids Res. 1976, 3: 891-901. 10.1093/nar/3.4.891
Moore CW: Control of in vivo (cellular) phleomycin sensitivity by nuclear genotype, growth phase, and metal ions. Cancer Res. 1982, 42: 929-933.
Gatignol A, Durand H, Tiraby G: Bleomycin resistance conferred by a drug-binding protein. FEBS Lett. 1988, 230: 171-175. 10.1016/0014-5793(88)80665-3
He ZM, Price MS, Obrian GR, Georgianna DR, Payne GA: Improved protocols for functional analysis in the pathogenic fungus Aspergillus flavus. BMC Microbiol. 2007, 7: 104- 10.1186/1471-2180-7-104
Da Silva NA, Srikrishnan S: Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Res. 2012, 12: 197-214. 10.1111/j.1567-1364.2011.00769.x
Belenguer P, Oustrin M-L, Tiraby G, Ducommun B: Effects of phleomycin-induced DNA damage on the fission yeast Schizosaccharomyces pombe cell cycle. Yeast. 1995, 11: 225-231. 10.1002/yea.320110305
Hua J, Meyer JD, Lodge JK: Development of positive selectable markers for the fungal pathogen Cryptococcus neoformans. Clin Diagn Lab Immunol. 2000, 7: 125-128.
Austin B, Hall RM, Tyler BM: Optimized vectors and selection for transformation of Neurospora crassa and Aspergillus nidulans to bleomycin and phleomycin resistance. Gene. 1990, 93: 157-162. 10.1016/0378-1119(90)90152-H
Kim B-G, Joh J-H, Yoo Y-B, Magae Y: Transformation of the edible basidiomycete, Pleurotus ostreatus to phleomycin resistance. Mycobiology. 2003, 31: 42-45. 10.4489/MYCO.2003.31.1.042. 10.4489/MYCO.2003.31.1.042
Sambrook J, Russell DW: Molecular cloning: a laboratory manual. 2001, Cold Spring Harbour, NY: Cold Spring Harbor Laboratory Press, 3
Datsenko KA, Wanner BL: One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A. 2000, 97: 6640-6645. 10.1073/pnas.120163297
Muyrers JPP, Zhang Y, Stewart AF: Techniques: recombinogenic engineering - new options for cloning and manipulating DNA. Trends Biochem Sci. 2001, 26: 325-331. 10.1016/S0968-0004(00)01757-6
Thomason L, Court DL, Bubunenko M, Costantino N, Wilson H, Datta S, Oppenheim A: Current protocols in molecular biology. Recombineering: genetic engineering in bacteria using homologous recombination. 2007, http://www.ncbi.nlm.nih.gov/pubmed/18265390
Oliner JD, Kinzler KW, Vogelstein B: In vivo cloning of PCR products in E. coli. Nucleic Acids Res. 1993, 21: 5192-5197. 10.1093/nar/21.22.5192
Bruschi M, Boyes S, Sugiarto H, Nielsen LK, Vickers CE: A transferrable sucrose utilization approach for non-sucrose-utilizing Escherichia coli strains. Biotech Adv. 2011, 30: 1001-1010.
Bradford MM: A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976, 72: 248-254. 10.1016/0003-2697(76)90527-3
Griffith KL, Wolf RE: Measuring beta-galactosidase activity in bacteria: cell growth, permeabilization, and enzyme assays in 96-well arrays. Biochem Biophys Res Commun. 2002, 290: 397-402. 10.1006/bbrc.2001.6152
Miller JH: Experiments in molecular genetics. 1972, Cold Spring Harbour Labratory: Cold Spring Harbour, NY
Acknowledgements
We thank Jens Nielsen and and Siavash Partow (Chalmers University of Technology, Sweden) for providing some of the plasmids used in this study. We also thank Dariella Nunez Bernal for useful discussions about culturing in the presence of G418. This research was supported by a Queensland State Government National and International Research Alliance Program (NIRAP) grant. CEV was supported by a Queensland State Government Smart Futures Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
CV conceived the research, designed the approach, managed the project and wrote the manuscript. SB carried out the molecular work for plasmid vector construction and the antibiotic testing. YZ carried out the molecular work for promoter analysis and the β-galactosidase assays. LN participated in experimental design and project management. All authors contributed to revising the manuscript. All authors read and approved the final manuscript.
Electronic supplementary material
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Vickers, C.E., Bydder, S.F., Zhou, Y. et al. Dual gene expression cassette vectors with antibiotic selection markers for engineering in Saccharomyces cerevisiae. Microb Cell Fact 12, 96 (2013). https://doi.org/10.1186/1475-2859-12-96
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1475-2859-12-96