Abstract
Proteasomes are highly conserved multienzyme complexes responsible for proteolytic degradation of the short-lived, regulatory, misfolded, and damaged proteins. They play an important role in the processes of brain plasticity, and decrease in their function is accompanied by the development of neurodegenerative pathology. Studies performed in different laboratories both on cultured mammalian and human cells and on preparations of the rat and rabbit brain cortex revealed a large number of proteasome-associated proteins. Since the identified proteins belong to certain metabolic pathways, multiple enrichment of the proteasome fraction with these proteins indicates their important role in proteasome functioning. Extrapolation of the experimental data, obtained on various biological objects, to the human brain suggests that the proteasome-associated proteins account for at least 28% of the human brain proteome. The proteasome interactome of the brain contains a large number of proteins involved in the assembly of these supramolecular complexes, regulation of their functioning, and intracellular localization, which could be changed under different conditions (for example, during oxidative stress) or in different phases of the cell cycle. In the context of molecular functions of the Gene Ontology (GO) Pathways, the proteins of the proteasome interactome mediate cross-talk between components of more than 30 metabolic pathways annotated in terms of GO. The main result of these interactions is binding of adenine and guanine nucleotides, crucial for realization of the nucleotide-dependent functions of the 26S and 20S proteasomes. Since the development of neurodegenerative pathology is often associated with regioselective decrease in the functional activity of proteasomes, a positive therapeutic effect would be obviously provided by the factors increasing proteasomal activity. In any case, pharmacological regulation of the brain proteasomes seems to be realized through the changes in composition and/or activity of the proteins associated with proteasomes (deubiquitinase, PKA, CaMKIIα, etc.).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Proteasomes are highly conserved multienzyme complexes, which are present in all prokaryotic and eukaryotic cells and cleave the short-lived, regulatory, misfolded, and damaged proteins [1-3]. The proteolytic (core) and regulatory particles of proteasomes, and the proteasome containing both particles, were named 20S, 19S, and 26S proteasomes in accordance with their sedimentation coefficients. Proteasomes play an important role in the processes of brain plasticity, and decrease in their functional activity is accompanied by the development of neurodegenerative pathology [2, 3].
In most cases, proteins subjected to proteasomal degradation first undergo ubiquitination, the ATP-dependent attachment of ubiquitin, an 8.5 kDa protein. The labeled (poly)ubiquitinated proteins are recognized by the 19S proteasome receptors. Then, deubiquitinases cleave the ubiquitin label, and the proteins subjected to proteolytic degradation enter the 20S proteasome [4, 5]. In addition to this protein delivery pathway, known as ATP- and ubiquitin-dependent proteasomal degradation [6], proteins can be degraded in proteasomes via the ATP- and ubiquitin-independent way [7-11]. In the latter case, one of the main structural prerequisites for such protein degradation is the presence of disordered regions, which initiate interaction with the 20S proteasome [12].
STRUCTURE AND FUNCTIONS OF PROTEASOMES
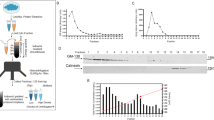
20S Proteasome (20S Core Particle). The 20S proteasome is a cylinder consisting of four heptameric rings, each of which is formed by seven α or seven β subunits encoded by fourteen different genes. The two outer rings of the cylinder, consisting of α subunits, function of a “gate” through which the client substrate proteins enter the inner catalytic region; they are also responsible for association of the 20S proteasome with the regulatory particles [13]. Two inner rings formed by β subunits exhibit several proteolytic activities: caspase-like activity (β1), trypsin-like activity (β2), and chymotrypsin-like activity (β5) [14, 15] (Fig. 1).
Structure of proteasomes (see explanations in the text). The image (map03050) has been adapted and taken from the KEGG (Kyoto Encyclopedia of Genes and Genomes; Kanehisa Laboratory) [15] open source with permission from the copyright holder.
19S Proteasome (19S Particle) and other Proteasome Regulators. 19S Particle. The regulatory 19S particle, also known as the proteasome activator 700 (PA700; Proteasome activator 700), consists of two subcomplexes: the lid and the base; it includes up to 20 different subunits with or without ATPase activity (Fig. 1). Subunits with ATPase activity are designated as Rpt subunits (Rpt, regulatory particle triphosphatases), and subunits lacking ATPase activity are designated as Rpn (Rpn, regulatory particle non-triphosphatases). In addition to 19S subunits (PA700), which can attach to the 20S core particle at one side or both sides simultaneously, there are other proteasome regulators. PA28 (or 11S particle) exists in two main forms: PA28αβ and PA28γ (or REGγ). PA28αβ is expressed in the cytoplasm; it consists of two subunits, α and β (with molecular mass of 28 kDa each), and is induced by γ-interferon. PA28αβ enhances the ability of 20S proteasome to cleave short peptides and oxidized substrates [15, 16]. The proteasome activator PA28γ is expressed in the nucleus; being attached to the 20S core particle, it acts as “a molecular sieve”, selecting proteins for degradation directly in the core part of the proteasome via the ATP- and ubiquitin-independent pathway. PA28γ is involved in regulation of such important cellular processes as cell growth and proliferation, apoptosis, DNA repair, immune response, and metabolism, thus maintaining cell homeostasis [17].
The 19S regulatory particle is responsible for recognition of the polyubiquitinated substrates, their unfolding, deubiquitination, and direction into the interior of the 20S core particle, where they are cleaved into oligopeptides. Active transport of substrates into the catalytic region is carried out using the energy of ATP hydrolysis. Six subunits of the base of the 19S regulatory particle (Rpt1-Rpt6) belong to ATPases of the AAA family (ATPases associated with various cellular activities). These subunits form a ring, as well as four subunits lacking ATPase activity (Rpn1, Rpn2, Rpn10, and Rpn13) [18]. The Rpn1, Rpn10, and Rpn13 subunits serve as ubiquitin receptors, recognizing client substrates labeled for elimination in the proteasome [19-22]. The subunits, ubiquitin receptors, differ in their affinity for different polyubiquitin chains [23, 24]. The lid is represented by nine different subunits (Rpn3, Rpn5-9, Rpn11, Rpn12, and Rpn15), which form a horseshoe-shaped structure [25-27]. The main function of the lid is substrate deubiquitination, which involves several deubiquitinases (one of them is the proteasome subunit Rpn11) [28-30]. The energy of ATP is necessary for stabilization of the complex of regulatory and catalytic core particles of the proteasome and, most importantly, for changes in the conformation of subunits that allow “gate” opening for the passage of protein substrates to the proteolytic cavity [31]. The energy of ATP hydrolysis is also used to unfold substrates during their movement into the proteolytic region [32, 33].
Other proteasome regulators. The PA200 proteasome activator is a phosphoprotein widely present in the cell as a proteasome-free pool. Under stressful conditions, PA200 is recruited by proteasomes. PA200 attachment to the 20S proteasome enhances its peptidase activity. PA200 can also interact with the 26S proteasome, forming a 19S-20S-PA200 hybrid proteasome. PA200 is involved in the key cell signaling pathways, it plays a role in DNA repair, providing genome stability. Expression of this factor sharply increases in the case of tumorigenic processes and, conversely, is suppressed in neurodegenerative diseases. Currently, this factor attracts much attention as a possible therapeutic target [34].
In higher vertebrates, cell stimulation with γ-interferon or other anti-inflammatory cytokines leads to immunoproteasome formation from the 15S preproteasome complex, which contains alternative catalytic subunits β1i, β2i, β5i, instead of β1, β2, and β5 subunits and has altered proteolytic activity and substrate specificity [35, 36] (Fig. 1). There is an increasing evidence that the role of immunoproteasomes is not limited by the immune response; immunoproteasomes play a certain role in oxidative stress, carcinogenesis, and neurodegenerative diseases, such as Parkinson’s, Alzheimer’s, Huntington’s, amyotrophic lateral sclerosis, autoimmune diseases (particularly, multiple sclerosis) [37, 38].
The Ecm29 regulator plays a key role in protecting cells from oxidative stress. Under oxidative stress there is a sharp increase in the number of 20S proteasomes in the cell, not only due to transcription regulation, but also due to dissociation of the 26S proteasome complexes. This is necessary for rapid elimination of the damaged proteins in an ubiquitin- and ATP-independent manner. The Ecm29 regulator accelerates dissociation of 26S proteasomes in response to oxidative stress, apparently by causing conformational changes and affecting protein–protein interactions between the 19S and 20S subcomplexes [39].
The proteasome regulator PI31 (Proteasomal Inhibitor of 31 kDa) was originally discovered as an inhibitor of peptide hydrolysis by the 20S proteasome in vitro. Later, it was found that in vivo it promoted proteasomal cleavage of the proteins. PI31 ribosylation promotes 26S proteasome assembly. Recently, it has been found that this factor also works as an adapter for proteasomal transport into neurons. Experiments, performed using mutant mice, have shown that knockout of this factor in the spinal motor neurons and in Purkinje cells caused axonopathy, neuronal degeneration, spinal and cerebellar neurological dysfunction. The authors suggest that the proteasome regulator PI31 could play a key role in protein homeostasis and synapse function and, accordingly, its dysfunction may lead to the development of neurodegenerative diseases in humans [40].
Proteasome assembly. Assembly of proteasomes is a complex regulated process, which has been well studied using yeast and human proteasomes by means of cryogenic electron microscopy. Assembly of the core particle in eukaryotes can be conditionally subdivided into three stages: formation of the α ring, formation of the β ring, dimerization of the semiproteasome, and maturation. These stages are mediated by five chaperones, known as Pba1-Pba4 (proteasome biogenesis associated 1-4) and Ump1 (underpinning maturation of proteasome 1) (in yeast) and, respectively, PAC1-PAC4 (proteasome assembly chaperone 1-4) and POMP (proteasome maturation protein) (in the case of human proteasomes) [41, 42].
Assembly of the 19S regulatory particle is also a multi-stage process; its two subcomplexes – the lid and the base – can be assembled separately. Five chaperones are involved in the base assembly (in yeast and humans, respectively): Nas2 (p27), Nas6 (p28), Hsm3 (S5b), Rpn14 (PAAF1) (proteasomal ATPase associated factor 1), and Adc17 (ATPase dedicated chaperone of 17 kDa). Two models for the assembly of the “base” of the regulatory subunit of the proteasome have been proposed. According to one of them, the “base” assembly does not depend on the core particle; another model suggests that the 20S particle serves as a “platform” for formation of the “base” of the 19S particle [41, 42].
The lid can be assembled in the absence of the base of the 19S particle and the 20S core particle. At the first stage, two intermediates are formed: one consists of Rpn5-6, Rpn8, Rpn9, and Rpn11 subunits, and the other consists of Rpn3, Rpn7, and Rpn15 subunits. After association of these intermediates, the Rpn12 subunit is attached. This is the trigger for conformational changes that allow the lid to attach to the “base” of the 19S particle. Association of the core and regulatory parts of the proteasome involves the Nas6 chaperone (p28) and the lid subunits Rpn5 and Rpn6 [41, 43].
Functional state of mitochondria can influence the proteasome assembly [44]. Defects in the respiratory complex I impair assembly of the 26S proteasome; they are reversible in the presence of pyruvate or aspartate [44].
Proteasome compartmentalization. In order to eliminate appropriate proteins at the right moment in the right place, proteasomes need to be dynamic not only in terms of their structure, but also in terms of their compartmentalization. Therefore, subcellular localization of proteasomes can be changed under changing conditions (for example, under oxidative stress) or in different phases of the cell cycle. Proteasomes are located in cytoplasm; some of them are associated with the cytoskeleton and membranes of endoplasmic reticulum. At the same time, many proteasomes are located in the nucleus. Moreover, experiments with mouse embryonic fibroblasts have shown that the newly synthesized proteasomes were located particularly in the nucleus, while the three-day-old proteasomes were mainly found in the cytoplasm, which indicates inflow of the newly synthesized proteasomes from the nucleus to the cytoplasm [45]. Interestingly, proteins involved in realization of nuclear functions (cyclins, inhibitors of cyclin-dependent kinases, transcription factors NF-κB, IκB, p53) were among the first identified physiological proteasome substrates [46-48].
Recently, the AKIRIN2 protein, an adapter for import of mature core particles into the nucleus, was discovered in mammals (in three different cell lines). AKIRIN2 directly binds to the fully assembled 20S proteasomes and promotes binding of importin 9, the factor required for transport to the nucleus. AKIRIN2 inhibits the 20S core particle and is cleaved when the 20S core particle enters the nucleus [49]. The mechanisms of proteasome import into the nucleus in yeast and mammals are similar in that the 20S core particle passes through the nuclear pore complexes either in an inhibited or immature state, probably to avoid degradation of the nuclear pore proteins rich in disordered sequences.
Using mammalian and yeast cells it has been shown that various stress factors (proteasome inhibition, oxidative stress, and others) cause proteasome accumulation in the nuclear or perinuclear loci (known as specific “membraneless organelles”). As soon as the stress factors cease to act, these “organelles” are disassembled [50-53].
If aberrant proteins are not eliminated in the nucleus in time due to proteasomal dysfunctions, they can accumulate in PML bodies (promyelocytic leukemia nuclear bodies), filling the “protein quality control compartments” in response to stress [54]. Proteasomes are recruited into the PML bodies for protein degradation; however, under unfavorable conditions, such as lack of ATP, the ubiquitin–proteasome system cannot function at full capacity. In this case, PML bodies with excess of proteasomes become toxic; this can lead to neurodegenerative diseases [55].
A new 20S proteasome complex, localized in the plasma membrane and exposed to the extracellular space, was found in neurons [56, 57]. The peptides formed as a result of the work of this complex can stimulate calcium signaling of the neurons.
Post-translational modifications of the proteasome. Among the post-translational modifications of proteasomes, phosphorylation is the most studied one. According to the PhosphoSitePlus database, more than 450 phosphosites have been found on each proteasome subunit of the human 26S proteasome [58]. Amino acid sequences surrounding proteasome phosphosites correspond to the recognition motifs of various protein kinases (MAPK, CDK, CaMK, GSK3, and some others) [58]. This suggests participation of various protein kinases in the regulation of proteasome function (and protein phosphatases, which reverse such regulation by dephosphorylation). Results of several studies suggest that phosphorylation of the 19S proteasome Rpn6 subunit by cAMP-dependent protein kinase (PKA) increases proteasome activity and increases degradation of toxic proteins. Activation of PKA in vivo by increasing the level of intracellular cAMP reduced accumulation of the phosphorylated tau protein and improved cognitive functions in mice with tauopathy [59]. Stimulation of 26S proteasome activity during phosphorylation by cGMP-dependent protein kinase also contributed to the increase in proteasomal degradation of proteins (including proteins involved in the development of neurodegenerative diseases) [60].
In addition to phosphorylation, other post-translational modifications have been reported. These include O-linked N-acetylglucosamination [61], ADP-ribosylation [62], acetylation, and myristylation [63, 64]. O-linked N-acetylglucosamination leads to inhibition of ATPase activity of the 26S complex and inhibits proteolytic activity of proteasomes. In mammals, the Rpt2 subunit of the 19S proteasome undergoes this type of modification (both in vitro and in vivo) [61]. ADP-ribosylation promotes 26S proteasome activity in both Drosophila and human cells. Tankyrase, the ADP-ribosyltransferase enzyme, as well as dp27 and dS5b chaperones involved in the assembly of the 19S proteasome, binds to the PI31 proteasome regulator. ADP-ribosylation of PI31 reduces its affinity for α-subunits of the 20S proteasome. This reduces the effect of PI31 on the 20S core particle. In addition, the PI31 modification increases binding and sequestration of dp27 and dS5b from the 19S regulatory particles, thereby promoting 26S assembly. Proteomic profiling of the mouse heart 26S proteasomes revealed N-terminal acetylation of five 19S proteasome subunits (Rpn1, Rpn5, Rpn6, Rpt3, and Rpt6) and five 20S proteasome subunits (α2, α5, α7, β3, and β4), as well as N-terminal myristylation of the Rpt2 subunits of 19S proteasomes [63]. Increase in acetylation of the 20S core particle subunits increased proteolytic activity of the mouse and human proteasomes [65, 66]. In yeast, the myristylated Rpt2 subunit directs the proteasome to control the quality of nuclear proteins. Mutations that block this modification lead to disruption of the intracellular localization of proteasomes [64].
Oxidative modification of sulfhydryl groups of the 20S proteasome (oxidation of cysteine residues Cys-SH to cysteine sulfonic acid Cys-SOH) results in subsequent S-glutathionylation (Cys-S-SG). This is accompanied by a partial loss of the chymotrypsin-like activity [67]. In vitro, glutaredoxin 2 exhibited deglutathionylase activity removing glutathione from the glutathionylated in vivo and in vitro 20S proteasomes. The other cytoplasmic redox proteins, thioredoxin 1 and thioredoxin 2, acted similarly [67].
PROTEASOME-ASSOCIATED PROTEINS
In addition to intrinsic proteasome proteins, the proteasome fractions isolated by various methods from various sources (from yeast to cells and tissues of higher vertebrates and humans) contain a significant amount of proteins associated with these particles [68-78].
In one of the first works devoted to the analysis of protein composition of the purified 19S, 20S, and 26S yeast proteasomes, all identified proteins were subdivided into several classes (according to terminology of the authors of the work) [70]:
(1) Proteasome subunits, as well as components of the ubiquitin–proteasome system (UPS) interacting with them (including ubiquitinases, deubiquitinases, etc.)
(2) Chaperone proteins involved in proteasome association/dissociation, separation of the densely packed ubiquitinated substrates during their preparation for proteolysis, and interaction with the unfolded (or misfolded) target proteins for subsequent proteolytic degradation.
(3) Proteins involved in regulation of transcription and translation, as well as in functioning of the cytoskeleton, RNA metabolism, cell division, signaling, and metabolism.
(4) Ribosome proteins and glycolytic enzymes. Considering the data that at least one third of all the newly synthesized proteins in mammalian cells undergo proteasomal degradation within a few minutes after translation [79], association of the protein synthesis machinery and UPS is important for the immediate elimination of the aberrant proteins. In addition, the substrate phosphorylation reactions involving glycolytic enzymes create potential opportunities for formation of the additional amounts of ATP that affect the nucleotide-sensitive interaction of proteins with proteasomes, as well as functioning of the subunits of the regulatory 19S particle that exhibit ATPase activity (Rpt 1-6).
Inconsistency of such classification of proteins associated with proteasomes is obvious, especially considering the last three classes. Multifunctional proteins, for example, glycolytic enzymes, in addition to their classical biochemical functions, can be also considered as chaperones [for example, glyceraldehyde-3-phosphate dehydrogenase (GAPDH)], and as proteins for degradation in proteasomes. The same is applicable to the regulators and components of the translational machine (ribosomal proteins). Participation of the proteins of these groups in specific metabolic pathways, including proteasomal degradation, is determined by their structural features at any given time. For example, the above-mentioned glycolytic enzyme, GAPDH, can perform chaperone functions, protecting the newly synthesized protein released from ribosomes against proteasome degradation [80]. However, when the cysteine residue (Cys247) is oxidized, GAPDH loses its ability to function as a chaperone [80], and the Hsp70 chaperone, binding to oxidized GAPDH, protects the cell from aggregation of this protein [81].
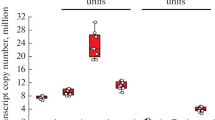
The studies performed using cultured mammalian and human cells [68, 70-75], preparations of the rat brain cortex [69] and rabbit brain and liver [76, 77] also revealed a large number of proteasome-associated proteins (table). Proteins that are not intrinsic UPS components belong to various functional groups, including: (i) components of the cytoskeleton and proteins involved in the transport of intracellular “cargo”; (ii) protective proteins; (iii) signal proteins and regulators of enzyme activity; (iv) regulators of gene expression, genome stability, and cell differentiation; (v) metabolic enzymes, including multifunctional proteins. Interestingly, abundance of a number of proteins associated with proteasomes was not lower and even exceeded abundance of the proteins, forming the proteasome structure [69, 76, 77]. The level of proteasome subunits that form the core part of proteasome varied in the range of 400-1000 units (arbitrary units of spectrum counting); for the subunits of the 19S regulatory particle, this parameter was in the range of 200-680 units [76]. For some proteins of the above functional groups, the abundance exceeded the ranges found for the proteasome subunits. For example: (i) cytoplasmic actin (P29751 Actin, cytoplasmic) – 1,777.6 units; (ii) heat shock protein (G1T9M9 Heat shock protein family A (Hsp70) member 8) – 509.2 units; (iii) calmodulin (P62160 Calmodulin) – 1386.3 units; (iv) heterogeneous nuclear ribonucleoprotein K (O19049 Heterogeneous nuclear ribonucleoprotein K) – 468.5 units; (v) GAPDH (P46406 glyceraldehyde-3-phosphate dehydrogenase) – 952.8 units. [76]. Taking into account the fact that the absolute amount of these proteins in the brain [82] differs from their abundance in the proteasome fraction, all this obviously indicates that the proteins isolated together with the proteasome fraction are not contaminants, but are components of the proteasome proteomes. This notion is also supported by multiple enrichment of the isolated brain proteasomes with proteins belonging to certain metabolic pathways (Fig. 2).
Functional groups of proteins associated with proteasomes
Proteasomes source | Method of isolation | Functional groups of proteins | Method of selective validation of the interaction with proteasome | References | ||||
|---|---|---|---|---|---|---|---|---|
I | II | III | IV | V | ||||
Saccharomyces cerevisiae | affinity purification based on anti-Flag M2* agarose | + | + | + | + | + | [70] | |
Saccharomyces cerevisiae | isotope tagging, in vivo cross-linking and tandem affinity purification SILAC**, quantitative analysis QTAX*** | + | + | + | + | + | reciprocal co-purification and immunoblotting | [72] |
Homo sapiens, 293HF-UbR/Rpn11-TB cells | affinity purification with the in vivo cross-linking XBAP**** | + | + | + | + | [74] | ||
Homo sapiens, K562 cells | affinity purification based on biotin-streptavidin complex and TEV protease***** | + | + | + | + | [75] | ||
Homo sapiens, extracellular 26S proteasomes of K562 | sucrose gradient centrifugation and ion-exchange chromatography | + | + | co-purification and immunoblotting | [73] | |||
Homo sapiens, 26S proteasomes of HEK293 expressing Rpn11-HTBH cell line | affinity purification MAP******-SILAC | + | + | + | + | [68] | ||
Rattus norvegicus, cortex cytosol and synaptosomes | affinity purification with the aid of glutathione sepharose, GST (glutathione S-transferase), and UBL (ubiquitin-like domain) tag | + | + | + | + | co-purification and immunoblotting | [69] | |
Rattus norvegicus, skeletal muscle | affinity purification with the aid of glutathione sepharose, GST (glutathione S-transferase), and UBL (ubiquitin-like domain) tag | + | + | + | + | + | [78] | |
Oryctolagus cuniculus, liver | high speed ultracentrifugation, ammonium sulfate fractionation | + | + | + | + | + | optical biosensor (surface plasmon resonance) | [77] |
Oryctolagus cuniculus, brain | high speed ultracentrifugation, ammonium sulfate fractionation | + | + | + | + | + | optical biosensor (surface plasmon resonance) | |
Enrichment of the proteasome fraction with certain groups of proteins identified in the brain [76, 77]. Data analysis by the Quick GO resource in the Explore Biology database showed distribution of the identified proteins among several metabolic pathways. In the context of extrapolation to the full-length human proteome, the highest enrichment was found in the case of proteins involved in the metabolism of glutamine and glutamate (P02747, 100-fold enrichment in the subproteome of proteins associated with proteasomes), glycolysis (P00024; 82-fold enrichment), and fructose metabolism (P02744; 37-fold enrichment), pyruvate metabolism (P02772; 34-fold enrichment), etc. Significant enrichment was also found for the metabolic pathways involved in the development of Parkinson’s disease (P00049; 12-fold enrichment) and Huntington’s disease (P00029; 8-fold enrichment). Metabolic pathway identifiers are presented according to the Quick GO resource nomenclature in terms of GO Slims (https://www.ebi.ac.uk/QuickGO) [83].
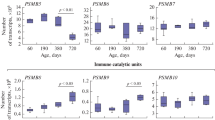
Fractionation of the brain 26S proteasome had a significant impact on the profile of proteins associated with the 20S core particle. The number of individual proteins identified in the fraction of 20S proteasomes of the rabbit brain almost doubled as compared to the fraction of brain 26S proteasomes [76, 77] mainly due to the metabolic enzymes, proteins involved in signal transduction and regulation of enzyme activity, protective proteins, and regulators of expression proteins genes, cell division, and differentiation [77]. During fractionation of the rabbit liver 26S proteasomes, the number of individual proteins in the fraction of 20S proteasomes remained virtually unchanged compared to the 26S fraction. It should be emphasized that comparison of the protein subproteomes of the 26S and 20S fractions of the rabbit brain and liver proteasomes revealed their high organ specificity. The pool of total proteins (n = 35) is mainly represented by metabolic and protective proteins, which account for more than 70% of the proteins. Interestingly, 10 of 35 proteasome-associated proteins common to all four fractions (fractions of brain and liver 26S and 20S proteasomes) belong to the so-called multifunctional proteins. These include GAPDH [84], α-enolase [85, 86], elongation factor 1-α 1 [87], aldolase [88, 89], glutathione peroxidase [90, 91], heat shock protein Hsp60 [92], lactate dehydrogenase [93], triose phosphate isomerase [94]. The significantly increased diversity of the repertoire of proteins associated with the 20S proteasome core particle of the brain after removal of the proteins of the 19S particle indicates that the protein components of the 19S particle play an important role in formation of the proteasome interactome and its regulation. In any case, profiles of the rat brain mitochondrial proteins bound to the Rpn10 subunit of the 19S particle changed significantly when the neurotoxin 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) and the neuroprotector isatin were administered to the animals [95]. Since the Rpn10 subunit of the 19S proteasome plays an important role in the recognition of substrates destined for proteolytic degradation in proteasomes [96, 97], this is consistent with the concept that the 19S proteasome subunits determine ordered delivery of the proteins undergoing proteolytic degradation in the 20S core particle. At the same time, it should be noted that, in contrast to the generally accepted viewpoint that the proteasome ubiquitin receptors, Rpn10 and Rpn13 subunits, are interchangeable in the context of proteasome functioning [25], profiles of the brain proteins bound to the Rpn10 and Rpn13 subunits in vitro differed significantly [98]. This is consistent with the results obtained by other authors on yeast [72]. According to their data, profiles of the proteins interacting with the subunits of the 19S proteasome Rpn1, Rpn10, Rpn11, and Rpt15 were not completely identical.
In the context of known data on the intracellular traffic of proteasomes and their translocation into various cell compartments depending on the functional state of the cell [42-45], existence of the large number of proteins interacting with proteasomes and forming the proteasome interactome is quite understandable.
Brief overview of the groups of proteins interacting with proteasomes. Being a widely distributed protein complex in the cells of the brain and peripheral tissues [64, 99], proteasomes are found in the nucleus and cytoplasm, where they are associated with various subcellular structures, including chromatin, cytoskeleton, nuclear envelope, plasma membrane, and cytosolic membrane. At the same time, their distribution in cells is uneven, and specific subcellular localization of proteasomes often depends on the cell type, growth status, and dynamically changing regulatory stimuli [64, 100]. In the context of molecular mechanisms of brain plasticity, localization of the UPS components serves as an important regulator of synaptic function, and neuronal proteasomes can interact with the intracellular membrane structures, including synaptic vesicles, Golgi apparatus vesicles, mitochondria, and lysosomes [101]. Local disruptions in proteasomal degradation are implicated in the development of many neurodegenerative diseases [102, 103].
(i) Cytoskeletal components and proteins involved in intracellular cargo transport and their role in proteasome localization in different cell compartments. One of the molecular motors, the dynein complex, plays a key role in proteasome mobility in axons [104, 105].
Redistribution of 26S proteasomes in neurons and other cell types is facilitated by interaction with the components of the cytoskeleton, which, in turn, undergoes remodeling with participation of proteasomes [106]. Neurons use the microtubule-dependent molecular motors to locate proteasomes at synapses.
In Drosophila neurons, the dynein light chain proteins (DYNLL1/2) serve as components of microtubule-dependent proteasome transport [105].
The conserved proteasome-binding protein PI31 plays an important adapter role in proteasome binding to the dynein light chain proteins (DYNLL1/2) [107, 108]. Phosphorylation by p38 MAPK increased PI31 binding to DYNLL1/2, stimulated formation of the proteasome–DYNLL1/2 complexes, and promoted directed proteasome movement in axons [108]. Inactivation of PI31 results in the impaired interaction of the dynein light chains with proteasomes and their transport into axons. This leads to the changes in the presynaptic zones and contributed to the development of defects in protein homeostasis on the periphery of neurons. In addition, PI31 tightly binds to the Ntc/FBXO7/PARK15 F-box protein [107, 108]; mutations in this protein are accompanied by proteasome dysfunction and cause development of the juvenile form of Parkinson’s disease [109]. Binding of another adapter protein, Ecm29, to myosins and kinesins, facilitates interaction of proteasomes with various cell compartments [110]. Knockdown of the heavy chain of motor protein kinesin 1 (KIF5B) leads to the impaired dendritic transport, impacts learning and memory processes [111] and anterograde movement of proteasomes to axons [101]. Factors promoting axon growth stimulate retrograde transport of proteasomes from the growing axon terminals, which is regulated by phosphorylation of the proteasome adapter protein Ecm29, interacting with dynein [104].
(ii) Protective proteins. In addition to participation in the assembly of proteasomes, cellular chaperones are involved in proteolytic degradation of proteins by proteasomes. Members of the Hsp70 family are directly involved in the processes of protein degradation by delivering client protein substrates to proteasome. Involvement of Hsc70/Hsp70 in protein degradation by the 26S proteasome is mediated by cochaperone CHIP (carboxyl terminus of Hsc70 interacting protein). It acts as a ubiquitin ligase, and the BAG1 (BCL2-associated athanogene) protein coordinates binding of the Hsp70-substrate complex to the 26S proteasome [112-114]. In the case of proteolytic elimination of the oxidized proteins by proteasomes, cells use proteolytic capabilities of the 20S proteasome, by dissociating the 26S proteasome. An important role in this process belongs to the heat shock protein 70 (Hsp70) that promotes increase in the number of free 20S proteasomes and prevents accumulation of the oxidized proteins in the cells under oxidative stress [115]. During formation of the cellular response to oxidative stress, Hsp70 can interact with both oxidized proteins and 20S proteasome, which, unlike the 26S proteasome, is able to recognize and cleave unfolded proteins in an ATP- and ubiquitin-independent manner [116]. The 20S proteasome recognizes its substrates by their unstructured hydrophobic regions exposed to the surface as a result of unfolding of the protein molecule [7-11]. At the same time, the 20S proteasome is much more resistant to oxidative stress than the 26S proteasome [117, 118]. Resistance of the 20S proteasome to oxidative damage is mediated by Hsp90 [119]. Hsp90 also binds to the oxidized calmodulin [120] (calmodulin abundance in brain proteasomes exceeds the level of a number of proteasome subunits [76]) and promotes degradation of this regulatory protein by the 20S proteasome.
(iii) Proteins involved in signal transduction and regulation of enzyme activity. Phosphorylation plays an important role in the regulation of proteasomes and their subcellular localization. Retrograde transport of proteasomes depends on the stage of neuron development and increases with the growth of axons [104]. This process is stimulated by cAMP and brain-derived neurotrophic factor (BDNF) without affecting the anterograde direction. The regulatory mechanism includes BDNF/cAMP-dependent activation of PKA and phosphorylation of the adapter protein Ecm29, which increases interaction of proteasomes with dynein. The α-subunit of calcium calmodulin-dependent protein kinase II (CaMKIIα) [58], which is associated with proteasomes in the brain [121], is considered as the main regulator of synapses. Translocation of this enzyme to synapse promotes accumulation of proteasomes in the spines and their postsynaptic redistribution. CaMKIIα autophosphorylation increases proteasome binding and mobilization of the latter into spines. The effect of CaMKIIα on proteasomes is realized via non-catalytic and catalytic mechanisms. In the first case, the activated (autophosphorylated) CaMKIIα binds more efficiently to proteasomes, facilitating their mobilization to the spines. In the second case, CaMKIIα stimulates proteasome activity by phosphorylation of the serine residue (Ser120) of the Rpt6 subunit. However, CaMKIIα translocation rather than its kinase activity is important for degradation of polyubiquitinated spine proteins [121]. According to other authors, blockade of this phosphorylation in the mutant protein with the S120A amino acid substitution or inhibition of CaMKIIα reduces synaptic activity and spine growth [122, 123]. Fear conditioning was accompanied by the increased phosphorylation of the Ser120 Rpt6 subunit of the proteasome regulatory particle and the proteasome activity in the amygdala of Long Evans rats [124]. Administration of a specific inhibitor of CaMKII, myr-AIP (myristoylated autocamtide-2 related inhibitory peptide), led to the significant decrease in the learning-induced increase in Rpt6 Ser120 phosphorylation and proteasome activity, without affecting the levels of protein polyubiquitination. The specific PKA inhibitor did not exhibit such effect. These and other data suggest that CaMKII is involved in memory formation by regulating Rpt6 phosphorylation and proteasome function [124, 125].
The polo-like protein kinase 1 (polo-like kinase 1, an enzyme of the serine-threonine kinase family involved in regulation of the cell cycle, cell responses to DNA damage, etc.) activates the 20S proteasome by phosphorylation of α-subunits [126]. However, the role of this kinase, which some authors consider as proteasome protein kinase [58], in brain plasticity remains unclear. Inhibition of this enzyme is known to block cell cycle progression in some gliomas [127].
Calmodulin, an activator of the calcium calmodulin-dependent protein kinase II, has been detected during proteomic profiling of the 26S and 20S fractions of the brain proteasomes [76, 77]. Taking into consideration the available data on dissociation of calmodulin from CaMKII [128], presence of this protein in proteasomes may indicate its potential role as a regulator of CaMKII and, possibly, other calmodulin-dependent proteasome-associated enzymes. It is also possible that this protein binds to proteasomes for proteolytic degradation. The latter assumption is supported by the fact that the Ca2+-free calmodulin undergoes degradation by either 26S or 20S proteasomes that does not require ubiquitination [120, 129].
(iv) Regulators of gene expression, genome stability, and cell differentiation. A significant number of proteasomes are located in the nucleus, where they play a key role in cell cycle regulation, transcription, chromatin remodeling, epigenetic control, RNA splicing, DNA damage repair, and quality control of nuclear proteins [64, 130]. In this context, regulators of gene expression, genome stability, and differentiation found in the proteasome fraction could be considered as potential substrates subject to proteolytic degradation. The eEF1A elongation factor, which plays an important role in the long-term synaptic plasticity [131], binds to proteasomes [76, 77, 132]. The level of this protein is reduced in hippocampus of the patients with Alzheimer’s disease [133]. On the other hand, there are reports that the eEF1A elongation factor binds aberrant proteins released from ribosomes and delivers them to proteasomes for subsequent degradation [132]. eEF1A interacts predominantly with the Rpt1 subunit of the regulatory 19S particle, as well as with ubiquitinated proteins [132]. The strongest interaction was observed with ATP depletion; upon deletion of the gene encoding the Rpt1 subunit, the eEF1A binding to proteasome decreased, but did not completely disappear [132]. This indicates that eEF1A could also bind to other proteasome components. The latter seems to explain the fact that eEF1A has been found in the brain both in the 26S proteasome fraction and in the 20S proteasome fraction [76, 77].
(v) Metabolic enzymes. Metabolic enzymes, associated with proteasomes and found in proteasomes isolated from various biological objects [69, 70, 72, 73, 75-78], are involved in almost all types of metabolism. At the same time, only glycolytic enzymes found in proteasomes (Fig. 3) and supplying them with ATP energy in reactions of substrate phosphorylation, apparently, could be considered as functionally significant. In any case, it is difficult to “offer a job in their specialty” for the subunits of the mitochondrial ATP synthase complex [73, 78] or transport ATPases [76, 77] in proteasomes. It is likely that some of these enzymes (for example, the previously mentioned GAPDH), which do not undergo degradation in proteasomes, could perform some noncanonical functions, acting, for example, as chaperones or 20S proteasome inhibitor proteins [134]. On the other hand, the lactate dehydrogenase subunits [76, 77], which are a part of the lactate oxidase complex found in the rat brain neurons [135], could serve as one of the links between the components of the proteasome interactome and mitochondria.
Enrichment of the proteasome fraction with glycolytic enzymes identified in the rabbit brain [76, 77]. Red color indicates glycolytic reactions, involving the detected proteins. Blocks filled with green show identified proteins, which are designated by numbers in the enzyme classification, as well as by KEGG and UniProtKB identifiers: Phosphoglucomutase-1 (EC 5.4.2.2; KEGG identifier is K01835, UniProtKB identifier is P36871), ATP-dependent 6-phosphofructokinase (EC 2.7.1.11; KEGG-identifier K00850, UniProtKB identifier – P308237), Fructose-bisphosphate aldolase A (EC 4.1.2.13, KEGG-identifier K01623, UniProtKB identifier – P04075), Triosephosphate isomerase (EC 5.3.1.1; KEGG-identifier K01803, UniProtKB ID – P60174), Glyceraldehyde-3-phosphate dehydrogenase (EC 1.2.1.12; KEGG-ID K00134, UniProtKB ID – P04406), Phosphoglycerate kinase 2 (EC 2.7.2.3; KEGG-ID K00927, UniProtKB ID – P07205), Alpha-enolase (EC 4.2.1.11; KEGG-ID K01689, UniProtKB ID – P06733), Pyruvate kinase PKM (EC 2.7.1.40; KEGG-ID K00873, UniProtKB ID – P14618), L-lactate dehydrogenase (EC 1.1.1.27; ID KEGG – K00016, ID UniProtKB – P00338). The image has been adapted and taken from the KEGG (Kyoto Encyclopedia of Genes and Genomes; Kanehisa Laboratory) open resource with permission from the copyright holder. The ID of the original KEGG glycolysis map is map00010 [15].
CONCLUSIONS
The proteasome interactome contains a large number of proteins that are involved in the assembly of these supramolecular complexes, regulation of their functioning, and intracellular localization. Extrapolation of the data, obtained using various biological objects, to the human brain shows that the proteasome-associated proteins account for at least 28% of the human brain proteome (Fig. 4).
Functional classes (Protein Classes) of the proteasome-associated proteins identified in the rabbit brain [76, 77] and extrapolated to the human proteome. Proteins associated with proteasome fall into several functional classes according to the analysis of the GO Slims resource (https://www.ebi.ac.uk/QuickGO). Blue color indicates contribution of the proteins in the identified proteasome-associated subproteome, green color indicates contribution of an identical functional class in the full-length human proteome. Functional classes of the proteins (in parentheses) are listed in the Quick GO Explore Biology resource nomenclature.
In the context of molecular functions of Gene Ontology (GO) Pathways, proteins of the proteasome interactome provide interaction between the components of more than 30 metabolic pathways annotated in terms of GO (Fig. 5) [136]. The main result of these interactions is binding of adenine and guanine nucleotides, required for the nucleotide-dependent functions of 26S and 20S proteasomes [31, 137, 138] in the cells of the central nervous system and peripheral tissues.
Molecular functions of brain proteins [76, 77] associated with proteasomes. The tree structure of distribution and relationship of functional classes of proteasome-associated proteins was reconstructed using the PANTHER™ Protein Class resource (version 17.0, updated on 2022-02-22; http://pantherdb.org/) [136]. Analysis of functional classes was performed with a False discovery rate (FDR) < 0.001 correction against the full-length human proteome. Blocks of classes, to which the identified proteasome-associated proteins belong, are highlighted in yellow; blocks of functional classes corresponding to or associated with the functional activities of proteins associated with proteasomes are marked in white. Molecular function identifiers are given in the Gene Ontology nomenclature. Color indicators in blocks reflect direction of the influence (activity) of neighboring or related blocks.
Involvement of proteasomes in pathogenesis of various diseases of the central nervous system and peripheral tissues makes these supramolecular complexes an attractive target for targeted pharmacological regulation. Although there is a clear interest in the field of development and use of proteasome inhibitors for treatment of various types of cancer, pharmacological regulation of functional activity of the brain proteasomes has also received some attention. Since the development of neurodegenerative pathology is frequently associated with regioselective decrease in the functional activity of proteasomes, a positive therapeutic effect could be obviously provided by the factors that increase activity of proteasomes. The possibility of pharmacological inhibition of deubiquitinase activity is currently considered as one of the main approaches [109]. Blockade of these enzymes facilitates entry of the ubiquitinated substrates into proteasomes for subsequent proteolytic degradation. Another approach, suggesting posttranslational modification of subunits of the proteasome complex, is associated with a number of components of the proteasome subproteome (PKA, CaMKIIα, etc.) [139]. In any case, pharmacological regulation of the brain proteasomes seems to be realized through changes in the composition and/or activity of the proteasome-associated proteins. The results of studies conducted on model objects in vitro and in vivo [139] inspire certain optimism in terms of feasibility of this approach.
Abbreviations
- CaMKII:
-
calcium calmodulin-dependent protein kinase II
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
- GO:
-
Gene Ontology
- PKA:
-
cAMP-dependent protein kinase
- UPS:
-
ubiquitin–proteasome system
References
Tanaka, K. (2009) The proteasome: overview of structure and functions, Proc. Jpn. Acad. Ser. B Phys. Biol. Sci., 85, 12-36, https://doi.org/10.2183/pjab.85.12.
Schmidt, M., and Finley, D. (2014) Regulation of proteasome activity in health and disease, Biochim. Biophys. Acta, 1843, 13-25, https://doi.org/10.1016/j.bbamcr.2013.08.012.
Goldberg, A. L. (2003) Protein degradation and protection against misfolded or damaged proteins, Nature, 426, 895-899, https://doi.org/10.1038/nature02263.
Hershko, A., and Ciechanover, A. (1992) The ubiquitin system for protein degradation, Annu. Rev. Biochem., 61, 761-807, https://doi.org/10.1146/annurev.bi.61.070192.003553.
Driscoll, J., and Goldberg, A. L. (1990) The proteasome (multicatalytic protease) is a component of the 1500-kDa proteolytic complex which degrades ubiquitin-conjugated proteins, J. Biol. Chem., 265, 4789-4792, https://doi.org/10.1016/S0021-9258(19)34041-4.
Schwartz, A. L., and Ciechanover, A. (2009) Targeting proteins for destruction by the ubiquitin system: implications for human pathobiology, Annu. Rev. Pharmacol. Toxicol., 49, 73-96, https://doi.org/10.1146/annurev.pharmtox.051208.165340.
Ben-Nissan, G., and Sharon, M. (2014) Regulating the 20S proteasome ubiquitin-independent degradation pathway, Biomolecules, 4, 862-884, https://doi.org/10.3390/biom4030862.
Orlowski, M., and Wilk, S. (2003) Ubiquitin-independent proteolytic functions of the proteasome, Arch. Biochem. Biophys., 415, 1-5, https://doi.org/10.1016/s0003-9861(03)00197-8.
Erales, J., and Coffino, P. (2014) Ubiquitin-independent proteasomal degradation, Biochim. Biophys. Acta, 1843, 216-221, https://doi.org/10.1016/j.bbamcr.2013.05.008.
Sánchez-Lanzas, R., and Castaño, J. G. (2014) Proteins directly interacting with mammalian 20S proteasomal subunits and ubiquitin-independent proteasomal degradation, Biomolecules, 4, 1140-1154, https://doi.org/10.3390/biom4041140.
Buneeva, O. A. and Medvedev, A. E. (2018) Ubiquitin-independent protein degradation in proteasomes, Biomed. Khim., 64, 134-148, https://doi.org/10.18097/PBMC20186402134.
Uversky, V. N., Oldfield, C. J., and Dunker, A. K. (2005) Showing your ID: intrinsic disorder as an ID for recognition, regulation and cell signaling, J. Mol. Recognit., 18, 343-384, https://doi.org/10.1002/jmr.747.
Sharon, M., Witt, S., Felderer, K., Rockel, B., Baumeister, W., and Robinson, C. V. (2006) 20S proteasomes have the potential to keep substrates in store for continual degradation, J. Biol. Chem., 281, 9569-9575, https://doi.org/10.1074/jbc.M511951200.
Bard, J. A. M., Goodall, E. A., Greene, E. R., Jonsson, E., Dong, K. C., and Martin, A. (2018) Structure and function of the 26S proteasome, Annu. Rev. Biochem., 87, 697-724, https://doi.org/10.1146/annurev-biochem-062917-011931.
Kanehisa, M., Sato, Y., and Kawashima, M. (2022) KEGG mapping tools for uncovering hidden features in biological data, Protein Sci., 31, 47-53, https://doi.org/10.1002/pro.4172.
Pickering, A. M., and Davies, K. J. A. (2012) Differential roles of proteasome and immunoproteasome regulators Pa28 alpha beta, Pa28 gamma and Pa200 in the degradation of oxidized proteins, Arch. Biochem. Biophys., 523, 181-190, https://doi.org/10.1016/j.abb.2012.04.018.
Funderburk, K. E., Kang, J., and Li, H. J. (2022) Regulation of life & death by REGγ, Cells, 11, 2281, https://doi.org/10.3390/cells11152281.
Díaz-Villanueva, J. F., Díaz-Molina, R., and García-González, V. (2015) Protein folding and mechanisms of proteostasis, Int. J. Mol. Sci., 16, 17193-17230, https://doi.org/10.3390/ijms160817193.
Fu, H., Sadis, S., Rubin, D. M., Glickman, M., van Nocker, S., Finley, D., and Vierstra, R. D. (1998) Multiubiquitin chain binding and protein degradation are mediated by distinct domains within the 26 S proteasome subunit Mcb1, J. Biol. Chem., 273, 1970-1981, https://doi.org/10.1074/jbc.273.4.1970.
Schreiner, P., Chen, X., Husnjak, K., Randles, L., Zhang, N., Elsasser, S., Finley, D., Dikic, I., Walters, K. J., and Groll, M. (2008) Ubiquitin docking at the proteasome through a novel pleckstrin- homology domain interaction, Nature, 453, 548-552, https://doi.org/10.1038/nature06924.
Husnjak, K., Elsasser, S., Zhang, N., Chen, X., Randles, L., Shi, Y., Hofmann, K., Walters, K. J., Finley, D., and Dikic, I. (2008) Proteasome subunit Rpn13 is a novel ubiquitin receptor, Nature, 453, 481-488, https://doi.org/10.1038/nature06926.
Shi, Y., Chen, X., Elsasser, S., Stocks, B. B., Tian, G., Lee, B. H., Shi, Y., Zhang, N., de Poot, S. A. H., Tuebing, F., Sun, S., Vannoy, J., Tarasov, S. G., Engen, J. R., Finley, D., and Walters, K. J. (2016) Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome, Science, 351, 6275, https://doi.org/10.1126/science.aad9421.
Martinez-Fonts, K., Davis, C., Tomita, T., Elsasser, S., Nager, A. R., Shi, Y., Finley, D., and Matouschek, A. (2020) The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates, Nat. Commun., 11, 477, https://doi.org/10.1038/s41467-019-13906-8.
Liu, Z., Dong, X., Yi, H. W., Yang, J., Gong, Z., Wang, Y., Liu, K., Zhang, W. P., and Tang, C. (2019) Structural basis for the recognition of K48-linked Ub chain by proteasomal receptor Rpn13, Cell. Discov., 5, 19, https://doi.org/10.1038/s41421-019-0089-7.
Kish-Trier, E., and Hill, C. P. (2013) Structural biology of the proteasome, Annu. Rev. Biophys., 42, 29-49, https://doi.org/10.1146/annurev-biophys-083012-130417.
Da Fonseca, P. C., and Morris, E. P., (2008) Structure of the human 26S proteasome: subunit radial displacements open the gate into the proteolytic core, J. Biol. Chem., 283, 23305-23314, https://doi.org/10.1074/jbc.M802716200.
Sahu, I., and Glickman, M. H. (2021) Proteasome in action: substrate degradation by the 26S proteasome, Biochem. Soc. Trans., 49, 629-644, https://doi.org/10.1042/BST20200382.
Leggett, D. S., Hanna, J., Borodovsky, A., Crosas, B., Schmidt, M., Baker, R. T., Walz, T., Ploegh, H., and Finley, D. (2002) Multiple associated proteins regulate proteasome structure and function, Mol. Cell, 10, 495-507, https://doi.org/10.1016/s1097-2765(02)00638-x.
Lander, G. C., Estrin, E., Matyskiela, M. E., Bashore, C., Nogales, E., and Martin, A. (2012) Complete subunit architecture of the proteasome regulatory particle, Nature, 482, 186-191, https://doi.org/10.1038/nature10774.
Guterman, A., and Glickman, M. H. (2004) Complementary roles for Rpn11 and Ubp6 in deubiquitination and proteolysis by the proteasome, J. Biol. Chem., 279, 1729-1738, https://doi.org/10.1074/jbc.M307050200.
Smith, D. M., Chang, S. C., Park, S., Finley, D., Cheng, Y., and Goldberg, A. L. (2007) Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s alpha ring opens the gate for substrate entry, Mol. Cell., 27, 731-744, https://doi.org/10.1016/j.molcel.2007.06.033.
Gallastegui, N., and Groll, M. (2010) The 26S proteasome: assembly and function of a destructive machine, Trends Biochem. Sci., 35, 634-642, https://doi.org/10.1016/j.tibs.2010.05.005.
Kleijnen, M. F., Roelofs, J., Park, S., Hathaway, N. A., Glickman, M., King, R. W., and Finley, D. (2007) Stability of the proteasome can be regulated allosterically through engagement of its proteolytic active sites, Nat. Struct. Mol. Biol., 14, 1180-1188, https://doi.org/10.1038/nsmb1335.
Yazgili, A. S., Ebstein, F., and Meiners, S. (2022) The proteasome activator PA200/PSME4: an emerging new player in health and disease, Biomolecules, 12, 1150, https://doi.org/10.3390/biom12081150.
Griffin, T. A., Nandi, D., Cruz, M., Fehling, H. J., Kaer, L. V., Monaco, J. J., and Colbert, R. A. (1998) Immunoproteasome assembly: cooperative incorporation of interferon gamma (IFN-gamma)-inducible subunits, J. Exp. Med., 187, 97-104, https://doi.org/10.1084/jem.187.1.97.
Kloetzel, P. M., and Ossendorp, F. (2004) Proteasome and peptidase function in MHC-class-I-mediated antigen presentation, Curr. Opin. Immunol., 16, 76-81, https://doi.org/10.1016/j.coi.2003.11.004.
Johnston-Carey, H. K., Pomatto, L. C., and Davies, K. J. (2015) The immunoproteasome in oxidative stress, aging, and disease, Crit. Rev. Biochem. Mol. Biol., 51, 268-281, https://doi.org/10.3109/10409238.2016.1172554.
Ferrington, D. A., and Gregerson, D. S. (2012) Immunoproteasomes: structure, function, and antigen presentation, Prog. Mol. Biol. Transl. Sci., 109, 75-112, https://doi.org/10.1016/B978-0-12-397863-9.00003-1.
Wang, X., Chemmama, I. E., Yu, C., Huszagh, A., Xu, Y., Viner, R., Block, S. A., Cimermancic, P., Rychnovsky, S. D., Ye, Y., Sali, A., and Huang, L. (2017) The proteasome-interacting Ecm29 protein disassembles the 26S proteasome in response to oxidative stress, J. Biol. Chem., 292, 16310-16320, https://doi.org/10.1074/jbc.M117.803619.
Minis, A., Rodriguez, J. A., Levin, A., Liu, K., Govek, E. E., Hatten, M. E., and Steller, H. (2019) The proteasome regulator PI31 is required for protein homeostasis, synapse maintenance, and neuronal survival in mice, Proc. Natl. Acad. Sci. USA, 116, 24639-24650, https://doi.org/10.1073/pnas.1911921116.
Budenholzer, L., Cheng, C. L., Li, Y., and Hochstrasser, M. (2017) Proteasome structure and assembly, J. Mol. Biol., 429, 3500-3524, https://doi.org/10.1016/j.jmb.2017.05.027.
Livneh, I., Cohen-Kaplan, V., Cohen-Rosenzweig, C., Avni, N., and Ciechanover, A. (2016) The life cycle of the 26S proteasome: from birth, through regulation and function, and onto its death, Cell Res., 26, 869-885, https://doi.org/10.1038/cr.2016.86.
Rousseau, A., and Bertolotti, A. (2018) Regulation of proteasome assembly and activity in health and disease, Nat. Rev. Mol. Cell Biol., 19, 697-712, https://doi.org/10.1038/s41580-018-0040-z.
Meul, T., Berschneider, K., Schmitt, S., Mayr, C. H., Mattner, L. F., Schiller, H. B., Yazgili, A. S., Wang, X., Lukas, C., Schlesser, C., Prehn, C., Adamski, J., Graf, E., Schwarzmayr, T., Perocchi, F., Kukat, A., Trifunovic, A., Kremer, L., Prokisch, H., Popper, B., von Toerne, C., Hauck, S. M., Zischka, H., and Meiners, S. (2020) Mitochondrial regulation of the 26S proteasome, Cell Rep., 32, 108059, https://doi.org/10.1016/j.celrep.2020.108059.
Enenkel, C., Kang, R. W., Wilfling, F., and Ernst, O. P. (2022) Intracellular localization of the proteasome in response to stress conditions, J. Biol. Chem., 298, 102083, https://doi.org/10.1016/j.jbc.2022.102083.
Herter, J. R., and Fuchs, S. Y. (2002) Recognition of substrate and Skp1 by the Homologue of Slimb (HOS) ubiquitin ligase receptor D role of the F-box, Med. Sci. Monit., 8, BR283-BR288.
Dawson, S., Hastings, R., Takayanagi, K., Reynolds, S., Low, P., Billett, M., and Mayer, R. J. (1997) The 26S-proteasome: regulation and substrate recognition, Mol. Biol. Rep., 24, 39-44, https://doi.org/10.1023/a:1006800522814.
Devine, T., and Dai, M. S. (2013) Targeting the ubiquitin-mediated proteasome degradation of p53 for cancer therapy, Curr. Pharm. Des., 19, 3248-3262, https://doi.org/10.2174/1381612811319180009.
de Almeida, M., Hinterndorfer, M., Brunner, H., Grishkovskaya, I., Singh, K., Schleiffer, A., Jude, J., Deswal, S., Kalis, R., Vunjak, M., Lendl, T., Imre, R., Roitinger, E., Neumann, T., Kandolf, S., Schutzbier, M., Mechtler, K., Versteeg, G. A., Haselbach, D., and Zuber, J. (2021) AKIRIN2 controls the nuclear import of proteasomes in vertebrates, Nature, 599, 491-496, https://doi.org/10.1038/s41586-021-04035-8.
Uriarte, M., Sen Nkwe, N., Tremblay, R., Ahmed, O., Messmer, C., Mashtalir, N., et al. (2021) Starvation-induced proteasome assemblies in the nucleus link amino acid supply to apoptosis, Nat. Commun., 12, 6984, https://doi.org/10.1038/s41467-021-27306-4.
Yasuda, S., Tsuchiya, H., Kaiho, A., Guo, Q., Ikeuchi, K., Endo, A., et al. (2020) Stress- and ubiquitylation-dependent phase separation of the proteasome, Nature, 578, 296-300, https://doi.org/10.1038/s41586-020-1982-9.
Kaganovich, D., Kopito, R., and Frydman, J. (2008) Misfolded proteins partition between two distinct quality control compartments, Nature, 454, 1088-1095, https://doi.org/10.1038/nature07195.
Gu, Z. C., Wu, E., Sailer, C., Jando, J., Styles, E., Eisenkolb, I., et al. (2017) Ubiquitin orchestrates proteasome dynamics between proliferation and quiescence in yeast, Mol. Biol. Cell, 28, 2479-2491, https://doi.org/10.1091/mbc.E17-03-0162.
Fabunmi, R. P., Wigley, W. C., Thomas, P. J., and DeMartino, G. N. (2001) Interferon gamma regulates accumulation of the proteasome activator PA28 and immunoproteasomes at nuclear PML bodies, J. Cell Sci., 114 (Pt 1), 29-36, https://doi.org/10.1242/jcs.114.1.29.
Mediani, L., Guillen-Boixet, J., Vinet, J., Franzmann, T. M., Bigi, I., Mateju, D., et al. (2019) Defective ribosomal products challenge nuclear function by impairing nuclear condensate dynamics and immobilizing ubiquitin, EMBO J., 38, 101341, https://doi.org/10.15252/embj.2018101341.
Ramachandran, K. V., and Margolis, S. S. (2017) A mammalian nervous-system-specific plasma membrane proteasome complex that modulates neuronal function, Nat. Struct. Mol. Biol., 24, 419-430, https://doi.org/10.1038/nsmb.3389.
Türker, F., Cook, E. K., and Margolis, S. S. (2021) The proteasome and its role in the nervous system, Cell. Chem. Biol., 28, 903-917, https://doi.org/10.1016/j.chembiol.2021.04.003.
Guo, X., Huang, X., and Chen, M. J. (2017) Reversible phosphorylation of the 26S proteasome, Protein Cell, 8, 255-272, https://doi.org/10.1007/s13238-017-0382-x.
Myeku, N., Clelland, C. L., Emrani, S., Kukushkin, N. V., Yu, W. H., Goldberg, A. L., et al. (2016) Tau-driven 26S proteasome impairment and cognitive dysfunction can be prevented early in disease by activating cAMP-PKA signaling, Nat. Med., 22, 46-53, https://doi.org/10.1038/nm.4011.
VerPlank, J. J. S., Tyrkalska, S. D., Fleming, A., Rubinsztein, D. C., and Goldberg, A. L. (2020) CGMP via PKG activates 26S proteasomes and enhances degradation of proteins, including ones that cause neurodegenerative diseases, Proc. Natl. Acad. Sci. USA, 117, 14220-14230, https://doi.org/10.1073/pnas.2003277117.
Zhang, F., Su, K., Yang, X., Bowe, D. B., Paterson, A. J., and Kudlow, J. E. (2003) O-GlcNAc modification is an endogenous inhibitor of the proteasome, Cell, 115, 715-725, https://doi.org/10.1016/S0092-8674(03)00974-7.
Cho-Park, P. F., and Steller, H. (2013) Proteasome regulation by ADP-ribosylation, Cell, 153, 614-627, https://doi.org/10.1016/j.cell.2013.03.040.
Gomes, A. V., Zong, C., Edmondson, R. D., Li, X., Stefani, E., Zhang, J., Jones, R. C., Thyparambil, S., Wang, G. W., Qiao, X., Bardag-Gorce, F., and Ping, P. (2006) Mapping the murine cardiac 26S proteasome complexes, Circ. Res., 99, 362-371, https://doi.org/10.1161/01.RES.0000237386.98506.f7.
Guo, X. (2022) Localized proteasomal degradation: from the nucleus to cell periphery, Biomolecules, 12, 229, https://doi.org/10.3390/biom12020229.
Wang, D., Fang, C., Zong, N. C., Liem, D. A., Cadeiras, M., Scruggs, S. B., Yu, H., Kim, A. K., Yang, P., Deng, M., Lu, H., and Ping, P. (2013) Regulation of acetylation restores proteolytic function of diseased myocardium in mouse and human, Mol. Cell. Proteomics, 12, 3793-3802, https://doi.org/10.1074/mcp.M113.028332.
Bi, M., Du, X., Jiao, Q., Chen, X., and Jiang, H. (2021) Expanding the role of proteasome homeostasis in Parkinson’s disease: beyond protein breakdown, Cell Death Dis., 12, 154, https://doi.org/10.1038/s41419-021-03441-0.
Silva, G. M., Netto, L. E., Discola, K. F., Piassa-Filho, G. M., Pimenta, D. C., Bárcena, J. A., and Demasi, M. (2008) Role of glutaredoxin 2 and cytosolic thioredoxins in cysteinyl-based redox modification of the 20S proteasome, FEBS J., 275, 2942-2955, https://doi.org/10.1111/j.1742-4658.2008.06441.
Wang, X., Chen, C. F., Baker, P. R., Chen, P. L., Kaiser, P., and Huang, L. (2007) Mass spectrometric characterization of the affinity-purified human 26S proteasome complex, Biochemistry, 46, 3553-3565, https://doi.org/10.1021/bi061994u.
Tai, H. C., Besche, H., Goldberg, A. L., and Schuman, E. M. (2010) Characterization of the brain 26S proteasome and its interacting proteins, Front. Mol. Neurosci., 3, 12, https://doi.org/10.3389/fnmol.2010.00012.
Verma, R., Chen, S., Feldman, R., Schieltz, D., Yates, J., Dohmen, J., and Deshaies, R. J. (2000) Proteasomal proteomics: identification of nucleotide-sensitive proteasome-interacting proteins by mass spectrometric analysis of affinity-purified proteasomes, Mol. Biol. Cell, 11, 3425-3439, https://doi.org/10.1091/mbc.11.10.3425.
Guerrero, C., Tagwerker, C., Kaiser, P., and Huang, L. (2006) An integrated mass spectrometry-based proteomic approach: quantitative analysis of tandem affinity-purified in vivo cross-linked protein complexes (QTAX) to decipher the 26 S proteasome-interacting network, Mol. Cell. Proteomics, 5, 366-378, https://doi.org/10.1074/mcp.M500303-MCP200.
Guerrero, C., Milenkovic, T., Przulj, N., Kaiser, P., and Huang, L. (2008) Characterization of the proteasome interaction network using a QTAX-based tag-team strategy and protein interaction network analysis, Proc. Natl. Acad. Sci. USA, 105, 13333-13338, https://doi.org/10.1073/pnas.0801870105.
Zaikova, Y. Y., Kulichkova, V. A., Ermolaeva, Y. B., Bottrill, A., Barlev, N. A., and Tsimoha, A. S. (2013) Characterization of extracellular proteasomes and its interacting proteins by iTRAQ mass spectrometry, Cell Tissue Biol., 7, 253-265, https://doi.org/10.1134/S1990519X13030139.
Yu, C., Yang, Y., Wang, X., Guan, S., Fang, L., Liu, F., Walters, K. J., Kaiser, P., and Huang, L. (2016) Characterization of dynamic UbR-proteasome subcomplexes by in vivo cross-linking (X) assisted bimolecular tandem affinity purification (XBAP) and label-free quantitation, Mol. Cell. Proteomics, 15, 2279-2292, https://doi.org/10.1074/mcp.M116.058271.
Artamonova, T. O., Khodorkovskiĭ, M. A., and Tsimokha, A. S. (2014) Mass spectrometric analysis of proteasomes affinity purified from the human myelogenous leukemia cells K562, Bioorg. Khim., 40, 720-734, https://doi.org/10.1134/s1068162014060041.
Buneeva, O., Kopylov, A., Kaloshina, S., Zgoda, V., and Medvedev, A. (2021) 20S and 26S proteasome-binding proteins of the rabbit brain: a proteomic dataset, Data Brief, 38, 107276, https://doi.org/10.1016/j.dib.2021.107276.
Buneeva, O. A., Kopylov, A. T., Zgoda, V. G., Gnedenko, O. V., Kaloshina, S. A., Medvedeva, M. V., Ivanov, A. S., and Medvedev, A. E. (2022) Comparative analysis of proteins associated with 26S and 20S proteasomes isolated from rabbit brain and liver [in Russian], Biomed. Khim., 68, 18-31, https://doi.org/10.18097/PBMC20226801018.
Besche, H. C., Haas, W., Gygi, S. P., and Goldberg, A. L. (2009) Isolation of mammalian 26S proteasomes and p97/VCP complexes using the ubiquitin-like domain from HHR23B reveals novel proteasome-associated proteins, Biochemistry, 48, 2538-2549, https://doi.org/10.1021/bi802198q.
Schubert, U., Antón, L. C., Gibbs, J., Norbury, C. C., Yewdell, J. W., and Bennink, J. R. (2000) Rapid degradation of a large fraction of newly synthesized proteins by proteasomes, Nature, 404, 770-774, https://doi.org/10.1038/35008096.
Jia, J., Arif, A., Willard, B., Smith, J. D., Stuehr, D. J., Hazen, S. L., and Fox, P. L. (2012) Protection of extraribosomal RPL13a by GAPDH and dysregulation by S-nitrosylation, Mol. Cell, 47, 656-663, https://doi.org/10.1016/j.molcel.2012.06.006.
Lazarev, V. F., Nikotina, A. D., Mikhaylova, E. R., Nudler, E., Polonik, S. G., Guzhova, I. V., and Margulis, B. A. (2016) Hsp70 chaperone rescues C6 rat glioblastoma cells from oxidative stress by sequestration of aggregating GAPDH, Biochem. Biophys. Res. Commun., 470, 766-771, https://doi.org/10.1016/j.bbrc.2015.12.076.
Ishihama, Y., Oda, Y., Tabata, T., Sato, T., Nagasu, T., Rappsilber, J., and Mann, M. (2005) Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein, Mol. Cell. Proteomics, 4, 1265-1272, https://doi.org/10.1074/mcp.M500061-MCP200.
Binns, D., Dimmer, E., Huntley, R., Barrell, D., O’Donovan, C., and Apweiler, R. (2009) QuickGO: a web-based tool for gene ontology searching, Bioinformatics, 25, 3045-3046, https://doi.org/10.1093/bioinformatics/btp536.
Kohr, M. J., Murphy, E., and Steenbergen, C. (2014) Glyceraldehyde-3-phosphate dehydrogenase acts as a mitochondrial trans-S-nitrosylase in the heart, PLoS One, 9, 111448, https://doi.org/10.1371/journal.pone.0111448.
Qiao, G., Wu, A., Chen, X., Tian, Y., and Lin, X. (2021) Enolase 1, a moonlighting protein, as a potential target for cancer treatment, Int. J. Biol. Sci., 17, 3981-3992, https://doi.org/10.7150/ijbs.63556.
Didiasova, M., Schaefer, L., and Wygrecka, M. (2019) When place matters: shuttling of enolase-1 across cellular compartments, Front. Cell Dev. Biol., 7, 61, https://doi.org/10.3389/fcell.2019.00061.
Ejiri, S. (2002) Moonlighting functions of polypeptide elongation factor 1: from actin bundling to zinc finger protein R1-associated nuclear localization, Biosci. Biotechnol. Biochem., 66, 1-21, https://doi.org/10.1271/bbb.66.1.
Pirovich, D. B., Da’dara, A. A., and Skelly, P. J. (2021) Multifunctional fructose 1,6-bisphosphate aldolase as a therapeutic target, Front. Mol. Biosci., 8, 719678, https://doi.org/10.3389/fmolb.2021.719678.
Gizak, A., Wiśniewski, J., Heron, P., Mamczur, P., Sygusch, J., and Rakus, D. (2019) Targeting a moonlighting function of aldolase induces apoptosis in cancer cells, Cell Death Dis., 10, 712, https://doi.org/10.1038/s41419-019-1968-4.
Conrad, M., Schneider, M., Seiler, A., and Bornkamm, G. W. (2007) Physiological role of phospholipid hydroperoxide glutathione peroxidase in mammals, Biol Chem., 388, 1019-1025, https://doi.org/10.1515/BC.2007.130.
Haraguchi, C. M., Mabuchi, T., Hirata, S., Shoda, T., Yamada, A. T., Hoshi, K., and Yokota, S. (2003) Spatiotemporal changes of levels of a moonlighting protein, phospholipid hydroperoxide glutathione peroxidase, in subcellular compartments during spermatogenesis in the rat testis, Biol. Reprod., 69, 885-895, https://doi.org/10.1095/biolreprod.102.013524.
Jeffery, C. J. (2018) Protein moonlighting: what is it, and why is it important?, Philos. Trans. R. Soc. Lond. B Biol Sci., 373, 20160523, https://doi.org/10.1098/rstb.2016.0523.
Brighenti, E., Carnicelli, D., Brigotti, M., and Fiume, L. (2017) The inhibition of lactate dehydrogenase A hinders the transcription of histone 2B gene independently from the block of aerobic glycolysis, Biochem. Biophys. Res. Commun., 485, 742-745, https://doi.org/10.1016/j.bbrc.2017.02.119.
Auer, J., Camoin, L., Courtot, A. M., Hotellier, F., and De Almeida, M. (2004) Evidence that P36, a human sperm acrosomal antigen involved in the fertilization process is triosephosphate isomerase, Mol. Reprod. Dev., 68, 515-523, https://doi.org/10.1002/mrd.20107.
Medvedev, A. E., Buneeva, O. A., Kopylov, A. T., Tikhonova, O. V., Medvedeva, M. V., Nerobkova, L. N., Kapitsa, I. G., and Zgoda, V. G. (2017) Brain mitochondrial subproteome of Rpn10-binding proteins and its changes induced by the neurotoxin MPTP and the neuroprotector isatin, Biochemistry (Moscow), 82, 330-339, https://doi.org/10.1134/S0006297917030117.
Deveraux, Q., Ustrell, V., Pickart, C., and Rechsteiner, M. (1994) A 26S protease subunit that binds ubiquitin conjugates, J. Biol. Chem., 269, 7059-7061.
Hamazaki, J., Sasaki, K., Kawahara, H., Hisanaga, S., Tanaka, K., and Murata, S. (2007) Rpn10-mediated degradation of ubiquitinated proteins is essential for mouse development, Mol. Cell. Biol., 19, 6629-6638, https://doi.org/10.1128/MCB.00509-07.
Buneeva, O. A., Kopylov, A. T., and Medvedev, A. E. (2020) Qualitative difference of mitochondrial subproteoms of brain RPN10- and RPN13-binding proteins, Biomed. Khim., 66, 138-144, https://doi.org/10.18097/PBMC20206602138.
Asano, S., Fukuda, Y., Beck, F., Aufderheide, A., Förster, F., Danev, R., and Baumeister, W. (2015) Proteasomes. A molecular census of 26S proteasomes in intact neurons, Science, 347, 439-442, https://doi.org/10.1126/science.1261197.
Wójcik, C., and DeMartino, G. N. (2003) Intracellular localization of proteasomes, Int. J. Biochem. Cell Biol., 35, 579-589, https://doi.org/10.1016/s1357-2725(02)00380-1.
Otero, M. G., Alloatti, M., Cromberg, L. E., Almenar-Queralt, A., Encalada, S. E., Pozo Devoto, V. M., Bruno, L., Goldstein, L. S., and Falzone, T. L. (2014) Fast axonal transport of the proteasome complex depends on membrane interaction and molecular motor function, J. Cell Sci., 127, 1537-1549, https://doi.org/10.1242/jcs.140780.
Oddo, S. (2008) The ubiquitin-proteasome system in Alzheimer’s disease, J. Cell. Mol. Med., 12, 363-373, https://doi.org/10.1111/j.1582-4934.2008.00276.x.
Yi, J. J., and Ehlers, M. D. (2007) Emerging roles for ubiquitin and protein degradation in neuronal function, Pharmacol. Rev., 59, 14-39, https://doi.org/10.1124/pr.59.1.4.
Hsu, M. T., Guo, C. L., Liou, A. Y., Chang, T. Y., Ng, M. C., Florea, B. I., Overkleeft, H. S., Wu, Y. L., Liao, J. C., and Cheng, P. L. (2015) Stage-dependent axon transport of proteasomes contributes to axon development, Dev. Cell, 35, 418-431, https://doi.org/10.1016/j.devcel.2015.10.018.
Kreko-Pierce, T., and Eaton, B. A. (2017) The Drosophila LC8 homolog cut up specifies the axonal transport of proteasomes, J. Cell Sci., 130, 3388-3398, https://doi.org/10.1242/jcs.207027.
Ibañez-Vega, J., Del Valle Batalla, F., Saez, J. J., Soza, A., and Yuseff, M. I. (2019) Proteasome dependent actin remodeling facilitates antigen extraction at the immune synapse of B cells, Front. Immunol., 10, 225, https://doi.org/10.3389/fimmu.2019.00225.
Bader, M., Benjamin, S., Wapinski, O. L., Smith, D. M., Goldberg, A. L., and Steller, H. (2011) A conserved F box regulatory complex controls proteasome activity in Drosophila, Cell, 145, 371-382, https://doi.org/10.1016/j.cell.2011.03.021.
Liu, K., Jones, S., Minis, A., Rodriguez, J., Molina, M., and Steller, H. (2019) PI31 is an adaptor protein for proteasome transport in axons and required for synaptic development, Dev. Cell, 50, 509-524, https://doi.org/10.1016/j.devcel.2019.06.009.
Buneeva, O., and Medvedev, A. (2022) Atypical ubiquitination and Parkinson’s disease, Int. J. Mol. Sci., 23, 3705, https://doi.org/10.3390/ijms23073705.
Gorbea, C., Pratt, G., Ustrell, V., Bell, R., Sahasrabudhe, S., Hughes, R. E., and Rechsteiner, M. (2010) A protein interaction network for Ecm29 links the 26 S proteasome to molecular motors and endosomal components, J. Biol. Chem., 285, 31616-31633, https://doi.org/10.1074/jbc.M110.154120.
Zhao, J., Fok, A. H. K., Fan, R., Kwan, P.Y., Chan, H. L., Lo, L. H., Chan, Y. S., Yung, W. H., Huang, J., Lai, C. S. W., and Lai, K. O. (2020) Specific depletion of the motor protein KIF5B leads to deficits in dendritic transport, synaptic plasticity and memory, eLife, 9, e53456, https://doi.org/10.7554/eLife.53456.
Kundrat, L., and Regan, L. (2010) Balance between folding and degradation for Hsp90-dependent client proteins: a key role forCHIP, Biochemistry, 49, 7428-7438, https://doi.org/10.1021/bi100386w.
Luders, J., Demand, J., and Hohfeld, J. (2000) The ubiquitin-related BAG-1 provides a link between the molecular chaperones Hsc70/Hsp70 and the proteasome, J. Biol. Chem, 275, 4613-4617, https://doi.org/10.1074/jbc.275.7.4613.
Murata, S., Minami, Y., Minami, M., Chiba, T., and Tanaka, K. (2001) CHIP is a chaperone-dependent E3 ligase that ubiquitylates unfolded protein, EMBO Rep., 2, 1133-1138, https://doi.org/10.1093/embo-reports/kve246.
Reeg, S., Jung, T., Castro, J. P., Davies, K. J. A., Henze, A., and Grune, T. (2016) The molecular chaperone Hsp70 promotes the proteolytic removal of oxidatively damaged proteins by the proteasome, Free Radic. Biol. Med., 99, 153-166, https://doi.org/10.1016/j.freeradbiomed.2016.08.002.
Davies, K. J. (2001) Degradation of oxidized proteins by the 20S proteasome, Biochimie, 83, 301-310, https://doi.org/10.1016/s0300-9084(01)01250-0.
Reinhecke, T., Sitte, N., Ullrich, O., Kuckelkorn, U., Davies, K. J., and Grune, T. (1998) Comparative resistance of the 20S and 26S proteasome to oxidative stress, Biochem. J., 335, 637-642, https://doi.org/10.1042/bj3350637.
Reinheckel, T., Ullrich, O., Sitte, N., and Grune, T. (2000) Differential impairment of 20S and 26S proteasome activities in human hematopoietic K562 cells during oxidative stress, Arch. Biochem. Biophys., 377, 65-68, https://doi.org/10.1006/abbi.2000.1717.
Conconi, M., Petropoulos, I., Emod, I., Turlin, E., Biville, F., and Friguet, B. (1998) Protection from oxidative inactivation of the 20S proteasome by heat-shock protein 90, Biochem. J., 333, 407-415, https://doi.org/10.1042/bj3330407.
Whittier, J. E., Xiong, Y., Rechsteiner, M. C., and Squier, T. C. (2004) Hsp90 enhances degradation of oxidized calmodulin by the 20 S proteasome, J. Biol. Chem., 279, 46135-46142, https://doi.org/10.1074/jbc.M406048200.
Bingol, B., Wang, C. F., Arnott, D., Cheng, D., Peng, J., and Sheng, M. (2010) Autophosphorylated CaMKIIα acts as a scaffold to recruit proteasomes to dendritic spines, Cell, 140, 567-578, https://doi.org/10.1016/j.cell.2010.01.024.
Djakovic, S. N., Marquez-Lona, E. M., Jakawich, S. K., Wright, R., Chu, C., Sutton, M. A., and Patrick, G. N. (2012) Phosphorylation of Rpt6 regulates synaptic strength in hippocampal neurons, J. Neurosci., 32, 5126-5131, https://doi.org/10.1523/JNEUROSCI.4427-11.2012.
Hamilton, A. M., Oh, W. C., Vega-Ramirez, H., Stein, I. S., Hell, J. W., Patrick, G. N., and Zito, K. (2012) Activity-dependent growth of new dendritic spines is regulated by the proteasome, Neuron, 74, 1023-1030, https://doi.org/10.1016/j.neuron.2012.04.031.
Jarome, T. J., Kwapis, J. L., Ruenzel, W. L., and Helmstetter, F. J. (2013) CaMKII, but not protein kinase A, regulates Rpt6 phosphorylation and proteasome activity during the formation of long-term memories, Front. Behav. Neurosci., 7, 115, https://doi.org/10.3389/fnbeh.2013.00115.
Jarome, T. J., Ferrara, N. C., Kwapis, J. L., and Helmstetter, F. J. (2016) CaMKII regulates proteasome phosphorylation and activity and promotes memory destabilization following retrieval, Neurobiol. Learn. Mem., 128, 103-109, https://doi.org/10.1016/j.nlm.2016.01.001.
Feng, Y., Longo, D. L., and Ferris, D. K. (2001) Polo-like kinase interacts with proteasomes and regulates their activity, Cell Growth Differ., 12, 29-37.
Metselaar, D. S., du Chatinier, A., Meel, M. H., Ter Huizen, G., Waranecki, P., Goulding, J. R., Bugiani, M., Koster, J., Kaspers, G. J. L., and Hulleman, E. (2022) AURKA and PLK1 inhibition selectively and synergistically block cell cycle progression in diffuse midline glioma, iScience, 25, 104398, https://doi.org/10.1016/j.isci.2022.104398.
Colbran, R. J., and Brown, A. M. (2004) Calcium/calmodulin-dependent protein kinase II and synaptic plasticity, Curr. Opin. Neurobiol., 14, 318-327, https://doi.org/10.1016/j.conb.2004.05.008.
Tarcsa, E., Szymanska, G., Lecker, S., O'Connor, C. M., and Goldberg, A. L. (2000) Ca2+-free calmodulin and calmodulin damaged by in vitro aging are selectively degraded by 26 S proteasomes without ubiquitination, J. Biol. Chem., 275, 20295-20301, https://doi.org/10.1074/jbc.M001555200.
Kito, Y., Matsumoto, M., Hatano, A., Takami, T., Oshikawa, K., Matsumoto, A., and Nakayama, K. I. (2020) Cell cycle-dependent localization of the proteasome to chromatin, Sci. Rep., 10, 5801, https://doi.org/10.1038/s41598-020-62697-2.
Beckelman, B. C., Day, S., Zhou, X., Donohue, M., Gouras, G. K., Klann, E., Keene, C. D., and Ma, T. (2016) Dysregulation of elongation factor 1A expression is correlated with synaptic plasticity impairments in Alzheimer’s disease, J. Alzheimers Dis., 54, 669-678, https://doi.org/10.3233/JAD-160036.
Chuang, S. M., Chen, L., Lambertson, D., Anand, M., Kinzy, T. G., and Madura, K. (2005) Proteasome-mediated degradation of cotranslationally damaged proteins involves translation elongation factor 1A, Mol. Cell. Biol., 25, 403-413, https://doi.org/10.1128/MCB.25.1.403-413.2005.
Beckelman, B. C., Zhou, X., Keene, C. D., and Ma, T. (2016) Impaired eukaryotic elongation factor 1A expression in Alzheimer’s disease, Neurodegener. Dis., 16, 39-43, https://doi.org/10.1159/000438925.
Olshina, M. A., Arkind, G., Deshmukh, F. K., Fainer, I., Taranavsky, M., Hayat, D., Ben-Dor, S., Ben-Nissan, G., and Sharon, M. (2020) Regulation of the 20S proteasome by a novel family of inhibitory proteins, Antioxid. Redox Signal., 32, 636-655, https://doi.org/10.1089/ars.2019.7816.
Hashimoto, T., Hussien, R., Cho, H. S., Kaufer, D., and Brooks, G. A. (2008) Evidence for the mitochondrial lactate oxidation complex in rat neurons: demonstration of an essential component of brain lactate shuttles, PLoS One, 3, e2915, https://doi.org/10.1371/journal.pone.0002915.
Thomas, P.D., Ebert, D., Muruganujan, A., Mushayahama, T., Albou, L.P., and Mi, H. (2022) PANTHER: Making genome-scale phylogenetics accessible to all, Protein Sci., 31, 8-22, https://doi.org/10.1002/pro.4218.
Rabl, J., Smith, D. M., Yu, Y., Chang, S. C., Goldberg, A. L., and Cheng, Y. (2008) Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases, Mol. Cell, 30, 360-368, https://doi.org/10.1016/j.molcel.2008.03.004.
Smith, D. M., Fraga, H., Reis, C., Kafri, G., and Goldberg, A. L. (2011) ATP binds to proteasomal ATPases in pairs with distinct functional effects, implying an ordered reaction cycle, Cell, 144, 526-538, https://doi.org/10.1016/j.cell.2011.02.005.
Myeku, N., and Duff, K. E. (2018) Targeting the 26S proteasome to protect against proteotoxic diseases, Trends Mol. Med., 24, 18-29, https://doi.org/10.1016/j.molmed.2017.11.006.
Funding
The work was carried out within the framework of the Program of Fundamental Scientific Research in the Russian Federation for a long-term period (2021-2030) (no. 122030100170-5).
Author information
Authors and Affiliations
Contributions
O. A. Buneeva – analysis of own and literature data, writing sections of the review. A. T. Kopylov – bioinformatic analysis of proteomic data arrays, preparation of illustrations, participation in writing sections of the review on proteasome-associated proteins. A. E. Medvedev – concept of the review, writing individual sections of the review, scientific editing.
Corresponding author
Ethics declarations
The authors declare no conflict of interests. This work did not involve the use of humans and animals as objects of research. The results of the original scientific research summarized in the review were carried out with the approval of the corresponding ethics committees indicated in each cited article.
Rights and permissions
Open access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Buneeva, O.A., Kopylov, A.T. & Medvedev, A.E. Proteasome Interactome and Its Role in the Mechanisms of Brain Plasticity. Biochemistry Moscow 88, 319–336 (2023). https://doi.org/10.1134/S0006297923030033
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297923030033