Abstract—Proteasomes are key components of the ubiquitin–proteasome system. Various forms of proteasomes are known. During aging, disturbances in the functioning of proteasomes have been revealed, as well as increased expression of their particular forms. Considering these data, we studied the expression of genes encoding the constitutive and immune subunits of proteasomes in cerebral cortex samples from C57BL/6 mice at the ages of 60, 190, 380, and 720 days. In addition, the contents of constitutive and immune proteasome subunits, chymotrypsin-like and caspase-like activities of proteasome pools, as well as the activity of the β5i immune subunit were studied in tissue homogenates. The chymotrypsin-like activity and the activity of the β5i subunit of different forms of proteasomes separated by electrophoresis in native gel were characterized. Compared with samples from young animals, in the cerebral cortex of animals at an age of 720 days the following changes in the expression patterns of proteasome genes were revealed: a decreased expression of the PSMB5 gene encoding constitutive proteasome subunit β5; increased expression of genes encoding immune proteasome subunits β5i and β1i. In tissue homogenates of aged mice, an increase in the content of immune subunits β1i and β2i was shown. In samples from old animals, chymotrypsin-like activity was decreased and a tendency to a decrease in caspase-like activity of proteasomes as well as the β5i subunit activity was revealed. Analysis of the activity of native complexes in tissues obtained from old animals revealed decreased chymotrypsin-like activity of 26S and 20S proteasomes containing the β5i subunit. Based on the obtained data, it can be assumed that changes in the pool of nonconstitutive proteasomes reflect aging-associated adaptive processes in the mouse brain.
Similar content being viewed by others
INTRODUCTION
The ubiquitin–proteasome system maintains cell and tissue homeostasis via degradation of most intracellular proteins [1]. Central elements of the ubiquitin-proteasome system are 20S proteasomes. These are multisubunit protein complexes that carry out proteolysis. The functional activity and substrate specificity of proteasomes depend on the set of proteolytic subunits within the complex. Proteasomes can contain only constitutive subunits (constitutive proteasomes), only immune, or immune and constitutive catalytic subunits simultaneously (here, all proteasomes containing immune subunits are termed “nonconstitutive proteasomes”). The ratio of constitutive and nonconstitutive proteasomes is different in different organs. Thus, in the cerebral cortex, most of the proteasome pool is represented by constitutive proteasomes, while immune organs, in contrast, are enriched with nonconstitutive proteasomes. The proteasome pool is dynamic: under stress, accumulation of protein aggregates, or foreign proteins in cells, the content of nonconstitutive proteasomes increases and the activity profile of the complexes changes [2]. The forms of proteasomes, in addition, differ in the presence/absence of activators, which provide access of substrates to the proteolytic chamber, and affect the activity of proteasomes. Thus, 20S proteasomes carrying the 19S activator are known as 26S proteasomes. Such complexes specifically recognize and degrade proteins labeled with ubiquitin. Proteasomes, including nonconstitutive ones, can also interact with other activators, for example, with 11S, which apparently affects their activity with respect to certain substrates [3]. In addition, the activity of proteasomes can be regulated by their interaction with other proteins (except the activators) and post-translational modifications [3].
Aging is associated with the accumulation of potentially toxic protein aggregates in cells [4]. In this regard, the functional changes that occur in the proteasome pool with age are of great interest and are being actively studied. Previous studies of a proteasome pool in different organs, including the central nervous system performed on aged F344BN and Wistar rats, revealed an increase in the expression of proteasomes with immune catalytic subunits [5–7]. Giannini et al. [8] reported that in the cerebral cortex of aging Sprague Dawley rats, the expression of immune proteasome subunits was increased and the efficiency of fluorogenic substrate hydrolysis by 20S and 26S proteasomes was decreased. At the same time, the degradation of polyubiquitinated model substrates by 26S proteasomes of old animals was slightly increased compared to those isolated from young animals. It should be noted that there are few works on the study of aging-associated changes in the functional activity and composition of the proteasome pool of the mouse cerebral cortex.
We have studied the expression of proteasome genes, their subunit composition, and the activity of individual forms (including those containing immune subunits) in the cerebral cortex of C57BL/6 mice at different stages of natural aging.
EXPERIMENTAL
Animals. Male С57BL/6 animals were used in the experiments. Mice were kept in the SPF (Specific Pathogen Free) vivarium of the Institute of Physiologically Active Substances of the Russian Academy of Sciences under conditions of artificially regulated daylight hours at a temperature of 22–26°C and free access to food and water. Manipulations with mice were carried out in full accordance with the Rules of laboratory practice in the Russian Federation dated April 1, 2016 No. 199n.
Obtaining mouse brain samples. Cerebral cortex samples fromC57BL/6 mice at an age of 60, 190, and 380 days (n = 6 for each age) and 720 days (n = 4) were used. All brain tissues were obtained surgically following terminal anesthesia of the animal and labeled as described previously [9]. The cerebral cortex samples were placed in 1.5-mL test tubes, frozen in liquid nitrogen, and then stored at –80°C.
RNA isolation and cDNA preparation. Isolation of total RNA from animal tissue samples was performed using the GeneJET RNA Purification kit (Thermo Fisher Scientific, United States) according to the manufacturer’s recommendations. The RNA concentration was determined spectrophotometrically using a NanoDrop instrument (Thermo Fisher Scientific). To remove DNA, RNA samples were treated with DNase using a RapidOut DNA Removal kit (Thermo Fisher Scientific). To obtain cDNA, 1.5 µg of total RNA was used. The reverse transcription reaction was performed with Maxima H Minus reverse transcriptase (Thermo Fisher Scientific) and oligo(dT)20 primer.
Estimation of expression levels of proteasome genes. The quantity of transcripts of proteasome subunit genes: PSMB5, PSMB6, PSMB7, PSMB8, PSMB9, and PSMB10 and of the gene encoding β-actin (Actb), was evaluated by real-time PCR using a Luminaris Color HiGreen qPCR Master Mix kit (Thermo Fisher Scientific). Comparative analysis of the absolute levels of expression of proteasome genes was performed using a previously developed system for quantitative real-time PCR [10].
Tissue lysis and immunoblotting. Animal cerebral cortex tissues were homogenized in buffer: 50 mM Tris-HCl (pH 7.5), 100 mM NaCl, 5 mM MgCl2, 1 mM EDTA, 1 mM dithiothreitol, 10% glycerol, 10 mM Na2S2O5, 2 mM ATP with 50 µl of buffer per 10 mg of tissue. The resulting homogenates were clarified by centrifugation at 13 000 g in an Eppendorf 5418R centrifuge (Eppendorf, Germany) for 30 min. The supernatant was collected and the concentration of total protein was determined by the Lowry method. Proteins were separated by electrophoresis in 12% SDS-PAGE and transferred onto a nitrocellulose membrane (Bio-Rad, United States). The transfer efficiency was evaluated by staining the membranes with a 0.1% Ponceau Rouge solution (Sigma-Aldrich, United States).
To identify proteasome subunits, membranes were incubated with primary antibodies (Table 1) for 2 h, then washed with phosphate buffer saline (PBS) containing 0.1% Tween 20 (Thermo Fisher Scientific) and incubated with the corresponding secondary antibodies (Table 1) conjugated with horseradish peroxidase (HRP). For additional confirmation of presence of the target protein and semiquantitative assessment of its content in the samples, commercial preparations of constitutive and immune 20S proteasomes (Boston Biochem) were applied to the gel. To normalize the signal, the membranes were incubated with primary antibodies to β-actin and corresponding secondary HRP-labeled antibodies (Table 1). Membranes were stained with anti-β-actin antibodies after washing the membranes from previous antibodies, first with a buffer: PBS, 2% SDS, 100 mM β-mercaptoethanol, and then additionally with PBS for 1 h. The target proteins were detected using the ECL Prime kit (GE Healthcare, United Kingdom) and the obtained images were analyzed using the ImageJ software (https://imagej.net/software/fiji/).
Determination of proteasome activity. Chymotrypsin-like, caspase-like activities of the proteasomes, as well as the specific activity of the β5i subunit in clarified homogenates of mouse cerebral cortex samples were assessed as described previously [9]. To determine the chymotrypsin-like activity of proteasomes, the fluorogenic substrate Suc-LLVY-AMC (Sigma-Aldrich) was used; the caspase-like activity was determined using Z-LLE-AMC (Sigma-Aldrich) substrate. The specific activity of the β5i subunit was determined by hydrolysis of the fluorogenic substrate Ac-ANW-AMC (Boston Biochem). The measurements were carried out on a VersaFluor Fluorometer (Bio-Rad) in at least three repetitions for each sample.
Determination of proteasome activity in native PAAG. Proteasome activity in nondenaturing PAAG was determined as described previously [9]. Clarified homogenates of animal cerebral cortex samples (~20 µg of total protein) were applied to a gradient 4‒20% PAAG. Electrophoresis was carried out for 36 h at 4°C (12 h at 80 V, 12 h at 140 V, 12 h at 240 V). After that the gel was placed for 30 min in a solution containing a fluorogenic substrate (Suc-LLVY-AMC to assess chymotrypsin-like activity or Ac-ANW-AMC to assess the activity of the β5i subunit). The gel was analyzed under ultraviolet light and photographed, then the images were analyzed using the ImageJ software. To confirm equal protein load between samples, the gel was stained with Coomassi G-250 (Serva, Germany).
Statistical analysis of the results. To assess the significance of the observed changes after qPCR, one-way analysis of the variance (ANOVA) was used using Tukey’s test as a correction for multiple comparisons. Statistical analysis of the immunoblotting results and measurement of proteasome activity was performed by comparing each point with the data in the control group using the Mann–Whitney test and the GraphPad Prism V8.4.3 software (GraphPad Software, United States). The differences were considered statistically significant at p < 0.05.
RESEARCH RESULTS
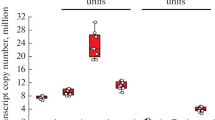
To study the changes in the proteasome pool of the cerebral cortex that accompany aging in mice, we used samples from C57BL/6 animals of four age groups: 60, 190, 380, and 720 days. Using the qPCR system that we developed earlier [10], we assessed the expression of the PSMB5‒7 genes, encoding constitutive catalytic subunits, and the PSMB8‒10 genes that code for immune catalytic proteasome subunits. It was shown that the expression of all the studied genes in samples of 60, 190, and 380-day-old mice did not differ significantly (Fig. 1) and corresponded to the data we obtained earlier [9]. However, in samples from mice at the age of 720 days compared with samples from 380 days old animals, significant changes in the levels of transcripts of immune and constitutive proteasome subunits were revealed. Thus, an almost two-fold increase in the expression of the PSMB8 gene encoding the immune subunit β5i, and the PSMB9 gene (immune subunit β1i), as well as a decrease (up to 2.5 folds) in expression of the PSMB5 gene encoding the constitutive subunit β5 was observed (Fig. 1).
The dynamics of expression levels of proteasome subunit genes in cerebral cortex samples from C57BL/6 mice of different ages (n = 6 in groups of animals at an age of 60, 190 and 380 days, n = 4, at the age of 720 days). The median of the values in the sample is indicated. Statistical significance of differences was assessed using ANOVA (Tukey’s test).
The obtained data indicate that during aging in the cells of the cerebral cortex of C57BL/6 mice the pool of proteasomes changes. A redistribution of proteasome forms in favor of an increase in nonconstitutive ones occurs. To elucidate specific age-related changes in proteasome patterns, we studied the contents of constitutive catalytic subunits β1, β2, and β5 and immune catalytic subunits β1i, β2i, and β5i in clarified homogenates of cerebral cortex samples from C57BL/6 mice of different ages by immunoblotting (Fig. 2).
The content of proteasome subunits β1, β2, β5, β1i, and β2i in clarified homogenates of the cerebral cortex samples from C57BL/6 mice of different ages. (a) Immunoblotting of homogenates with antibodies to β1, β2, β5, β1i, and β2i subunits. In each experiment cortical samples from three mice of the same age were used. Serial dilutions of commercial mouse constitutive (m20S) and immune (m20Si) proteasomes (Boston Biochem, UK) were used as standard samples. (b) Analysis of the obtained results using the ImageJ software. Data are presented as the mean ± standard deviation (SD). Statistical significance of differences was assessed using the Mann–Whitney test; *p < 0.05.
As can be seen from the results presented in Fig. 2, with age, the content of constitutive proteasome subunits β1 and β2 was stable. In homogenates of the cerebral cortex from mice at an age of 380 and 720 days, a significant increase in the content of immune subunits β1i and β2i was revealed. It should be noted that the antibodies used in the study practically did not recognize the β5i subunit, both in animal tissue samples and in commercial preparations of immune proteasomes (data not provided). Despite the obtained data on a decrease in the level of expression of the gene encoding the constitutive β5 subunit (Fig. 1), a increased amount of this subunit was revealed in tissue slightly homogenates of old mice (Fig. 2).
Based on the data on the increase in the content of proteasome immune subunits, one can assume an increase in the total chymotrypsin-like activity, a decrease in the caspase-like activity, and an increase in the activity of immune subunits in tissue samples of old animals. However, a decrease in chymotrypsin-like activity, as well as a trend towards a decrease in caspase-like activity and the activity of the β5i subunit was revealed in cerebral cortex samples from 720 days old mice (Fig. 3).
Chymotrypsin-like, caspase-like activity of proteasomes, as well as the activity of the β5i subunit in cerebral cortex samples from C57BL/6 mice of different ages. The fluorogenic substrates Suc-LLVY-AMC, Z-LLE-AMC, and Ac-ANW-AMC, were used to determine the chymotrypsin-like (ChLA) and caspase-like (CLA) activities of the proteasomes, as well as the activity of the β5i (β5i) subunit, respectively. The proteasome activity in the group of young animals (60 days) was taken as 100%. Results are presented as the mean ± SD. The statistical significance of the differences was assessed using the Mann–Whitney test; *p < 0.05.
The interaction of proteasomes with activators changes the molecular weight of proteasomes and allows separation of various forms of proteasomes by electrophoresis in native conditions. Electrophoretic analysis of animal tissue samples under nondenaturing conditions revealed free 20S proteasomes and 20S proteasomes with attached activators: 26S proteasomes with one or two 19S activators and, probably, 20S proteasomes with one or two 11S activators (Fig. 4) [9, 11, 12]. Fluorogenic substrates were used to determine their chymotrypsin-like activity, as well as the activity of the built-in β5i subunits (Fig. 4).
The distribution of chymotrypsin-like activity and activity of the β5i subunit between the 20S and 26S proteasomes in cerebral cortex lysates of C57BL/6 mice of different ages (n = 4 for each group). (a) The proteasome activity in the gel regions corresponding to the mobility of 20S and 26S proteasomes in native PAAG (see the Experimental section) was determined using the corresponding fluorogenic substrates. (b) The chymotrypsin-like activity and the activity of the β5i subunit were assessed via analysis of the optical density of the signals in the (a) using the ImageJ software. Results are presented as the mean ± SD.
As a result, it was shown that in the cerebral cortex of C57BL/6 animals, there is a tendency to a decrease in chymotrypsin-like activity both in the localization regions of 26S and 20S proteasomes with age. It should be noted that the total chymotrypsin-like activity of proteasomes in the 26S region was higher than in the 20S region. The activity of the β5i subunit was found in both areas of proteasome localization; at the same time, by the 720th day, it was decreased both in the 26S and in the 20S regions, despite the activation of the expression of the gene encoding the β5i subunit.
DISCUSSION
Aging is associated with the accumulation of oxidized and damaged proteins in cells and tissues, which, in turn, can provoke the development of inflammation, stress, and, as a result, various diseases [13–15]. At the same time, the accumulation of oxidized and damaged proteins is considered as a marker of disturbances in the functioning of intracellular proteolysis systems. We studied the proteasome pool of the cerebral cortex from healthy C57BL/6 mice of four age groups: 60, 190, 380, and 720 days, and analyzed the dynamics of changes in the studied parameters throughout almost the entire life of the animals. The data we obtained on a decrease in the expression of genes of constitutive subunits and an increase in the expression of genes of immune proteasome subunits are consistent with the results obtained using other animal models [4‒8, 16]. In the lysates of the cerebral cortex of mice at the age of 720 days, compared to samples from 60-day-old animals, an increase (on average by 2–3 folds) in the amount of immune subunits β1i and β2i was revealed (Fig. 2). Thus, changes occur in the pool of intracellular proteasomes that lead to a redistribution in favor of an increase in the fraction of proteasomes with immune subunits.
It is known that stress and the formation of protein aggregates within the cell lead to changes in the pool of proteasomes, in particular, to an increase in the expression of nonconstitutive complexes, which are characterized by an increased efficiency of degradation of oxidized and damaged proteins [14, 17–20]. Thus, proteasome induction may reflect the activation of compensatory mechanisms aimed at maintaining proteostasis. Besides, it is known that aging is accompanied by the development of neuroinflammation [21], which can also stimulate the assembly of proteasomes containing immune subunits, although the efficiency of immunoproteasome induction by interferon-γ decreases with age [22]. We have shown that despite an increase in the contents of nonconstitutive proteasomes during aging, the total chymotrypsin-like activity exhibited by the β5, β5i, and β1i subunits and the activity of the β5i subunit individually decreases both in the 26S and 20S proteasomes (Fig. 4). The obtained data suggests that, on the one hand, aging stimulates the synthesis of nonconstitutive proteasomes, and, on the other hand, their deactivation, probably due to post-translational modifications and/or interaction with protein aggregates, as well as endogenous inhibitors, such as PI31, what is observed in various age-related diseases [19, 23, 24]. Thus, despite an increase in the expression of immune subunits, the activity of nonconstitutive proteasomes, at least when measured using fluorogenic substrates, does not increase with age.
In general, it can be assumed that aging is accompanied by the activation of compensatory mechanisms. These include the ubiquitin-proteasome system and increased expression of non-constitutive proteasomes. However, a decrease in their activity does not allow to effectively neutralize the accumulation of oxidized and damaged proteins in cells. In this regard, it is important to mention that the analysis of tissue samples from centenarians and long-lived animals, such as naked mole rats and giant mollusks, revealed an increased activity of proteasomes [4, 25, 26]. In addition, long-lived primates have an increased level of gene expression of immune proteasome subunits and high proteasome activity [27]. Based on these data, one can assume a prognostic significance of the proteasome activity, as well as the use of strategies aimed at modulating the activity of various forms of proteasomes [28–31] to prolong life and improve its quality.
REFERENCES
Ciechanover A., Kwon Y.T. 2015. Degradation of misfolded proteins in neurodegenerative diseases: Therapeutic targets and strategies. Exp. Mol. Med. 47, e147.
Ferrington D.A., Gregerson D.S. 2012. Immunoproteasomes: Structure, function, and antigen presentation. Prog. Mol. Biol. Transl. Sci. 109, 75–112.
Morozov A.V., Karpov V.L. 2018. Biological consequences of structural and functional proteasome diversity. Heliyon. 4, e00894.
Chondrogianni N., Petropoulos I., Franceschi C., Friguet B., Gonos E.S. 2000. Fibroblast cultures from healthy centenarians have an active proteasome. Exp. Gerontol. 35, 721–728.
Husom A.D., Peters E.A., Kolling E.A., Fugere N.A., Thompson L.V., Ferrington D.A. 2004. Altered proteasome function and subunit composition in aged muscle. Arch. Biochem. Biophys. 421, 67–76.
Ferrington D.A., Husom A.D., Thompson L.V. 2005. Altered proteasome structure, function, and oxidation in aged muscle. FASEB J. 19, 644–646.
Gavilán M.P., Castaño A., Torres M., Portavella M., Caballero C., Jiménez S., García-Martínez A., Parrado J., Vitorica J., Ruano D. 2009. Age-related increase in the immunoproteasome content in rat hippocampus: Molecular and functional aspects. J. Neurochemistry. 108, 260–272.
Giannini C., Kloß A., Gohlke S., Mishto M., Nicholson T.P., Sheppard P.W., Kloetzel P.M., Dahlmann B. 2013. Poly-Ub-substrate-degradative activity of 26S proteasome is not impaired in the aging rat brain. PLoS One. 7, e64042.
Morozov A.V., Burov A.V., Funikov S.Yu., Teterina E.V., Astakhova T.M., Erokhov P. A., Ustyugov A.A., Karpov V.L. 2023. Changes in the activity and content of individual forms of proteasomes in samples of the cerebral cortex during pathology development in 5xFAD mice.Mol. Biol. (Moscow). 57 (5), 885–896.
Funikov S.Y., Spasskaya D.S., Burov A.V., Teterina E.V., Ustyugov A.A., Karpov V.L., Morozov A.V. 2021. Structures of the mouse central nervous system contain different quantities of proteasome gene transcripts. Mol. Biol. (Moscow). 55 (1), 47–55. https://doi.org/10.1134/S0026893320060047
Erokhov P.A., Lyupina Y.V., Radchenko A.S., Kolacheva A.A., Nikishina Y.O., Sharova N.P. 2017. Detection of active proteasome structures in brain extracts: Proteasome features of August rat brain with violations in monoamine metabolism. Oncotarget. 8, 70941–70957.
Morozov A.V., Kulikova A.A., Astakhova T.M., Mitkevich V.A., Burnysheva K.M., Adzhubei A.A., Erokhov P.A., Evgen’ev M.B., Sharova N.P., Karpov V.L., Makarov A.A. 2016. Amyloid-β increases activity of proteasomes capped with 19S and 11S regulators. J. Alzheimers Dis. 54, 763–776.
Chondrogianni N., Stratford F.L., Trougakos I.P., Friguet B., Rivett A.J., Gonos E.S. 2003. Central role of the proteasome in senescence and survival of human fibroblasts: Induction of a senescence-like phenotype upon its inhibition and resistance to stress upon its activation. J. Biol. Chem. 278, 28026–28037.
López-Otín C., Blasco M.A., Partridge L., Serrano M., Kroemer G. 2013. The hallmarks of aging. Cell. 153, 1194–1217.
Saez I., Vilchez D. 2014. The mechanistic links between proteasome activity, aging and agerelated diseases. Curr. Genomics. 15, 38–51.
Ly D.H., Lockhart D.J., Lerner R.A., Schultz P.G. 2000. Mitotic misregulation and human aging. Science. 287, 2486–2492.
Pickering A.M., Koop A.L., Teoh C.Y., Ermak G., Grune T., Davies K.J. 2010. The immunoproteasome, the 20S proteasome and the PA28αβ proteasome regulator are oxidative-stress-adaptive proteolytic complexes. Biochem. J. 432, 585–595.
Pickering A.M., Davies K.J.A. 2012. Degradation of damaged proteins: The main function of the 20S proteasome. Prog. Mol. Biol. Transl. Sci. 109, 227–248.
Grune T., Jung T., Merker K., Davies K.J. 2004. Decreased proteolysis caused by protein aggregates, inclusion bodies, plaques, lipofuscin, ceroid, and “aggresomes” during oxidative stress, aging, and disease. Int. J. Biochem. Cell Biol. 36, 2519–2530.
Abi Habib J., De Plaen E., Stroobant V., Zivkovic D., Bousquet M.P., Guillaume B., Wahni K., Messens J., Busse A., Vigneron N., Van den Eynde B.J. 2020. Efficiency of the four proteasome subtypes to degrade ubiquitinated or oxidized proteins. Sci. Rep. 10, 15765.
Sparkman N.L., Johnson R.W. 2008. Neuroinflammation associated with aging sensitizes the brain to the effects of infection or stress. Neuroimmunomodulation. 15, 323–330.
Stratford F.L., Chondrogianni N., Trougakos I.P., Gonos E.S., Rivett A.J. 2006. Proteasome response to interferon-γ is altered in senescent human fibroblasts. FEBS Lett. 580, 3989–3994.
Powell S.R., Wang P., Divald A., Teichberg S., Haridas V., McCloskey T.W., Davies K.J., Katzeff H. 2005. Aggregates of oxidized proteins (lipofuscin) induce apoptosis through proteasome inhibition and dysregulation of proapoptotic proteins. Free Radic. Biol. Med. 38, 1093–1101.
Johnston-Carey H.K., Pomatto L.C.D., Davies K.J.A. 2015. The immunoproteasome in oxidative stress, aging, and disease. Crit. Rev. Biochem. Mol. Biol. 51, 268–281.
Pérez V.I., Buffenstein R., Masamsetti V., Leonard S., Salmon A.B., Mele J., Andziak B., Yang T., Edrey Y., Friguet B., Ward W., Richardson A., Chaudhuri A. 2009. Protein stability and resistance to oxidative stress are determinants of longevity in the longest-living rodent, the naked mole-rat. Proc. Natl. Acad. Sci. U. S. A. 106, 3059–3064.
Ungvari Z., Csiszar A., Sosnowska D., Philipp E.E., Campbell C.M., McQuary P.R., Chow T.T., Coelho M., Didier E.S., Gelino S., Holmbeck M.A., Kim I., Levy E., Sonntag W.E., Whitby P.W., Austad S.N., Ridgway I. 2013. Testing predictions of the oxidative stress hypothesis of aging using a novel invertebrate model of longevity: the giant clam (Tridacna derasa). J. Gerontol. A Biol. Sci. Med. Sci. 68, 359–367.
Pickering A.M., Lehr M., Miller R.A. 2015. Lifespan of mice and primates correlates with immunoproteasome expression. J. Clin. Invest. 125, 2059–2068.
Chondrogianni N., Voutetakis K., Kapetanou M., Delitsikou V., Papaevgeniou N., Sakellari M., Lefaki M., Filippopoulou K., Gonos E.S. 2015. Proteasome activation: an innovative promising approach for delaying aging and retarding age-related diseases. Ageing Res. Rev. 23, 37–55.
Chondrogianni N., Voutetakis K., Kapetanou M., Delitsikou V., Papaevgeniou N., Sakellari M., Lefaki M., Filippopoulou K., Gonos E.S. 2005. Overexpression of proteasome β5 assembled subunit increases the amount of proteasome and confers ameliorated response to oxidative stress and higher survival rates. J. Biol. Chem. 280, 11840–11850.
Tonoki A., Kuranaga E., Tomioka T., Hamazaki J., Murata S., Tanaka K., Miura M. 2009. Genetic evidence linking age-dependent attenuation of the 26S proteasome with the aging process. Mol. Cell Biol. 29, 1095–1106.
Vilchez D., Morantte I., Liu Z., Douglas P.M., Merkwirth C., Rodrigues A.P., Manning G., Dillin A. 2012. RPN-6 determines C. elegans longevity under proteotoxic stress conditions Nature. 489, 263–268.
Funding
The work was supported by the Russian Science Foundation grant No. 18-74-10095.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Statement on the welfare of animals. Experiments with animals were carried out in accordance with the Rules of laboratory practice in the Russian Federation dated April 1, 2016 no. 199n.
Conflict of interest. The authors declare that they have no conflicts of interest.
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burov, A.V., Funikov, S.Y., Astakhova, T.M. et al. Dynamic Changes in the Activities and Contents of Particular Proteasome Forms in the Cerebral Cortex of C57BL/6 Mice during Aging. Mol Biol 57, 897–904 (2023). https://doi.org/10.1134/S0026893323050035
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893323050035