Summary
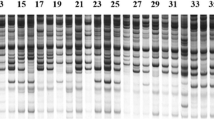
The importance of neem (Azadirachta indica A. Juss.) as a medicinal tree species has been acknowledged worldwide. Superior trees with desired traits such as high azadirachtin content have been identified and micropropagated. Somaclonal variants that may arise in vitro, however, pose limitations to large-scale micropropagation. It is, therefore, imperative to establish genetic uniformity of such plantlets by ensuring strict quality checks at various stages of in vitro culture. This is the first study that evaluates the applicability of amplified fragment length polymorphism (AFLP) markers in establishing clonal fidelity of tissue culture(TC)-raised neem plants. Seven AFLP primer combinations generated a total of 334 amplified fragments across the mother plant, TC progenies, and other neem accessions that were included as controls. Two hundred and thirty-nine amplified fragments were monomorphic across the mother tree and its TC progenies. No extra band was detected in the TC plantlets that was absent in the mother tree, indicating that the TC plantlets regenerated through nodal explants are indeed true-to-type. Ninety-five AFLP fragments were detected in the controls, which allowed their discrimination from the elite mother tree and its TC progenies. Similarity matrix based on Jaccard's coefficient revealed that the pair-wise value between the mother tree and its TC plantlets was ‘1’, indicating perfect similarity. Phenetic dendrogram based on UPGMA (unweighted pair group method of arithmetic averages) analysis further confirmed the true-to-type nature of TC progenies, since a tie was observed between the mother tree and its TC plantlets. On the contrary, the control neem accessions were distinct from the mother and its TC progenies. AFLP markers proved to be an ideal tool for routine analysis and certification of genetic fidelity of micropropagated plants prior to commercialization, especially in tree species because of their long generation time.
Similar content being viewed by others
References
Andrade, L. B.; Echeverrigaray, S.; Fracaro, F.; Pauletti, G. F.; Rota, L. The effect of growth regulators on shoot propagation and rooting of common lavender (Lavandula vera DC). Plant Cell Tiss. Organ Cult. 8:235–242; 1999.
Beck, S. L.; Dunlop, R.; Staden, J. van Micropropagation of Acacia mearnsii from ex-vitro material. Plant Growth Regul. 26:143–148; 1998.
Breyne, P.; Boerjan, W.; Gerats, T.; Van-Montagu, M.; Van-Gysel, A. Applications of AFLPTM in plant breeding, molecular biology and Genetics. Belg. J. Bot. 129:107–117; 1997.
Cecchini, E.; Natali, L.; Cavallini, A.; Durante, M. DNA variations in regenerated plants of pea (Pisum sativum L.). Theor. Appl. Genet. 84:874–879; 1992.
Cuenca, S.; Amo-Marco, J. B. In-vitro propagation of Centaurea spachii from inflorescence stems. Plant Growth Regul. 26:143–148; 2000.
D'Amato, F. Cytogenetics of plant cell and tissue cultures and their regenerants. Crit. Rev. Plant Sci. 3:73–112; 1985.
Das, S.; Rajagopal, J.; Bhatia, S.; Srivastava, P. S.; Lakshmikumaran, M. Assessment of genetic variation within Brassica campestris cultivars using AFLP and RAPD markers. J. Biosci. 24:433–440; 1999.
Doyle, J. J.; Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12:13–15; 1990.
Durante, M.; Geri, C.; Grisvard, J.; Guille, E.; Parenti, R.; Buiatti, M. Variation in DNA complexity in Nicotiana glauca tissue cultures. I. Pith tissue de-differentiation in vitro. Protoplasma 114:114–118; 1983.
Earle, E. D.; Demarly, Y. Variability in plants regenerated from tissue culture. New York: Praeger, 1982.
Eeswara, J. P.; Stuchbury, T.; Allan, E. J.; Mordue, A. J. A standard procedure for micropropagation of the neem tree (Azadirachta indica A. Juss.). Plant Cell Rep. 17:215–219; 1998.
Ganeshkumar, M.; Jayakumar, R.; Raghupathy, A.; Rajasekharan, B. Liquid chromatographic determination and monitoring of azadirachtin in neem ecotypes. Pestology 18:11–14; 1994.
Goto, S.; Thakur, R. C.; Ishii, K. Determination of genetic stability in longterm micropropagated shoots of Pinus thunbergii Parl. using RAPD markers. Plant Cell Rep. 18:193–197; 1998.
Hashmi, G.; Huettel, R.; Meyer, R.; Krusberg, L.; Hammerschlag, F. RAPD analysis of somaclonal variants derived from embryo callus cultures of peach. Plant Cell Rep. 16:624–627; 1997.
Hirochika, H. Activation of tobacco retrotransposons during tissue culture. EMBO J. 12:2521–2528; 1993.
Hirochika, H.; Sugimoto, K.; Otsuki, Y.; Tsugawa, H.; Kanda, M. Retrotransposons of rice involved in mutations induced by tissue culture. Proc. Natl Acad. Sci. USA 93:7783–7788; 1996.
Jaccard, P. Nouvelles researches sur la distribution florale. Bull. Soc. Vaud. Sci. Nat. 44:223–270; 1908.
Kathiravan, K.; Ignacimuthu, S. Micropropagation of Canavalia virosa (Roxb.) Wight and Arn.: a medicinal plant. Phytomorphology 49:61–66; 1999.
Kidwell, K. K.; Osborn, T. C. Variation among alfalfa somaclones in copy number of repeated DNA sequences. Genome 36:906–912; 1993.
Larkin, P. J.; Scowcroft, W. R. Somaclonal variations—a novel source of variability from cell cultures for plant improvement. Theor. Appl. Genet. 60:197–214; 1981.
Leroy, X. J.; Leon, K.; Charles, G.; Branchard, M. Cauliflower somatic embryogenesis and analysis of regenerant stability by ISSRs. Plant Cell Rep. 19:1102–1107; 2000.
Lisowska, K.; Wysokinska, H. In-vitro propagation of Catalpa ovata G. Don. Plant Cell Tiss. Organ Cult. 60:171–176; 2000.
Matsumoto, S.; Fukui, H. Identification of rose cultivars and clonal plants by random amplified polymorphic DNA. Sci. Hortic. (Amst) 67:49–54; 1996.
Munthali, M. T.; Newbury, H. J.; Ford-Lloyd, B. V. The detection of somaclonal variants of beet using RAPD. Plant Cell Rep. 15:474–478; 1996.
Murthy, B. N. S.; Saxena, P. K. Somatic embryogenesis and plant regeneration of neem (Azadirachta indica A. Juss.). Plant Cell Rep. 17:469–475; 1998.
Nelke, M.; Nowak, J.; Wright, J. M.; McClean, N. L. DNA fingerprinting of red clover (Trifolium pratense L.) with Jeffreys probes: detection of somaclonal variation and other applications. Plant Cell Rep. 13:72–78; 1993.
Nobre, J.; Santos, C.; Romano, A. Micropropagation of the Mediterranean species Viburnum tinus. Plant Cell Tiss. Organ Cult. 60:75–78; 2000.
Pal, M. Clonal approaches for yield improvement in neem: strategies and protocols for selective use of genetic diversity. Indian Forester 121:1033–1039; 1995.
Peschke, V. M.; Phillips, R. L. Genetic implications of somaclonal variation in plants. Adv. Genet. 30:41–75; 1992.
Phillips, R. L.; Kaeppler, S. M.; Olhoft, P. Genetic stability of plant tissue cultures: breakdown of normal controls. Proc. Natl Acad. Sci. USA 91:5222–5226; 1994.
Rangaswamy, S.; Paramar, B. S. Azadirachtin—a content of seeds of neem ecotypes in relation to agroecological regions of India. Pestic. Res. J. 7:140–148; 1995.
Rani, V.; Parida, A.; Raina, S. N. Random amplified polymorphic DNA (RAPD) markers for genetic analysis in micropropagated plants of Populus deltoides Marsh. Plant Cell Rep. 14:459–462; 1995.
Rani, V.; Raina, S. N. Genetic fidelity of organized meristem-derived micropropagated plants: a critical appraisal. In Vitro Cell Dev. Biol. Plant 36:319–330; 2000.
Rohlf, F. J., NTSYS-pc. Numerical taxonomy and multivariate analysis system. Version 2.0. New York: Exeter Publications; 1998.
Rostiana, O.; Niwa, M.; Marubashi, W. Efficiency of inter-simple sequence repeat PCR for detecting somaclonal variation among leaf culture regenerated plants of horseradish. Breed. Sci. 49:245–250; 1999.
Salvi, N. D.; George, L.; Eapen, S. Plant regeneration from leaf base callus of turmeric and random amplified polymorphic DNA analysis of regenerated plants. Plant Cell Tiss. Organ Cult. 66:113–119; 2001.
Schmutterer, H. The neem tree Azadirachta indica A. Juss. and other Meliacious plants: source of unique natural products for integrated pest management, medicine, industry and other purposes. Weinheim, Germany: VCH Press, 1995:696.
Shenoy, V. B.; Vasil, I. K. Biochemical and molecular analysis of plants derived from embryogenic tissue cultures of napier grass (Pennisetum purpureum K. Schum). Theor. Appl. Genet. 83: 947–955; 1992.
Singh, A.; Negi, M. S.; Rajagopal, J.; Bhatia, S.; Tomar, U. K.; Srivastava, P. S.; Lakshmikumaran, M. Assessment of genetic diversity in Azadirachta indica using AFLP markers. Theor. Appl. Genet. 99:272–279; 1999.
Sneath, P. H. A.; Sokal, R. R. Numerical taxonomy. San Francisco: Freeman Press; 1973.
Varshney, A.; Lakshmikumaran, M.; Srivastava, P. S.; Dhawan, V. Establishment of genetic fidelity of in-vitro raised Lilium bulblets through RAPD markers. In Vitro Cell Dev. Biol. Plant 36:383–391; 2000.
Vendrame, W. A.; Kochert, G.; Wetzstein, H. Y. AFLP analysis of variation in pecan somatic embryos. Plant Cell Rep. 18:853–857; 1999.
Venkateswarlu, B.; Katyal, J. C.; Choudhari, J.; Mukhopadhyaya, K. Azadirachtin content in the neem seed samples collected from different dryland regions. Neem Newsletter 14:7–11; 1997.
Venkateswarlu, B.; Katyal, J. C.; Choudhari, J.; Mukhopadhyaya, K. Micropropagation of plus neem (Azadirachta indica A. Juss) and evaluation of field transferred plants. Indian Forester 124:537–543; 1998.
Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; van de Lee, T.; Hornes, M.; Frijters, A.; Plot, J.; Peleman, J.; Kuiper, M.; Zabeau, M. AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res. 23: 4407–4414; 1995.
Zabeau, M.; Vos, P. Selective restriction fragment amplification: a general method for DNA fingerprinting. European patent application of 92402629 (publ. no. 0534858A1); 1993.
Zhu, J.; Gale, M. D.; Quarrie, S.; Jackson, M. T.; Bryan, G. J. AFLP markers for study of rice biodiversity. Theor. Appl. Genet. 96:602–611; 1998.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, A., Negi, M.S., Moses, V.K. et al. Molecular analysis of micropropagated neem plants using aflp markers for ascertaining clonal fidelity. In Vitro Cell Dev Biol -Plant 38, 519–524 (2002). https://doi.org/10.1079/IVP2002341
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1079/IVP2002341