Abstract
Slavum lentiscoides and Chaetogeoica ovagalla are two aphid species from the subtribe Fordina of Fordini within the subfamily Eriosomatinae, and they produce galls on their primary host plants Pistacia. We assembled chromosome-level genomes of these two species using Nanopore long-read sequencing and Hi-C technology. A 332 Mb genome assembly of S. lentiscoides with a scaffold N50 of 19.77 Mb, including 11,747 genes, and a 289 Mb genome assembly of C. ovagalla with a scaffold N50 of 11.85 Mb, containing 14,492 genes, were obtained. The Benchmarking Universal Single-Copy Orthologs (BUSCO) benchmark of the two genome assemblies reached 93.7% (91.9% single-copy) and 97.0% (95.3% single-copy), respectively. The high-quality genome assemblies in our study provide valuable resources for future genomic research of galling aphids.
Similar content being viewed by others
Background & Summary
Some insects can induce abnormal growth and development of host plants and form highly specialized structures termed galls. Galls benefit insect inducers and their offspring by providing abundant nutrition and protection against natural enemies and adverse abiotic factors1. Gall formation at the early stage is generally induced by insect stimuli, including feeding and oviposition1. The molecular mechanisms of gall induction and development have been found to be inseparable from insect effectors2,3 and phytohormones4,5,6. Aphids (Hemiptera: Aphidoidea) are an important group of plant-sapping insects that comprise over 5,000 species, 10–20% of which can induce galls on their primary host plants7,8. Galling aphid species mainly belong to Adelgidae, Phylloxeridae, and several subfamilies of Aphididae, including Eriosomatinae, Hormaphidinae, Tamaliinae, Thelaxinae, and Aphidinae9,10. To date, reference genomes are available for more than 40 aphid species. Among them, only seven true gall-forming species have been sequenced and assembled11,12,13,14,15,16. The lack of genomic resources for galling aphids has greatly hindered our understanding of the genetic basis for adaptive evolution of gall induction in aphids.
The tribe Fordini of subfamily Eriosomatinae is a typical lineage of true gall-inducing aphids. Species from one subtribe, Melaphidina, induce galls on Rhus (Anacardiaceae). The Rhus galls usually contain high concentrations of tannins and have economic and medicinal values17. The genome of one representative species, Schlechtendalia chinensis, has been reported recently15. Aphids within the other subtribe, Fordina, induce galls on Pistacia (Anacardiaceae), which is an economically significant genus of plants. Pistacia vera (cultivated pistachio) produces edible nuts that are of great commercial importance. Seeds of other wild Pistacia species also possess economic value and can be utilized for local consumption, oil extraction, soap production, and most importantly, as rootstock sources for pistachio trees18. Extracts from Pistacia galls exhibit anti-inflammatory and antioxidant activities19,20. However, no genomic data are available for this galling group.
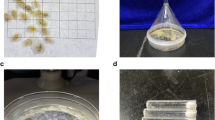
Producing more high-quality genome assemblies encompassing different lineages is foundational to the genomic study of galling aphids. In this study, we generated chromosome-level genome assemblies of two galling species from Fordina, Slavum lentiscoides Mordvilko and Chaetogeoica ovagalla (Zhang). Both species induce closed bag-like galls on the main veins of Pistacia spp. leaves8 (Fig. 1a,b). A total of 332.26 Mb and 289.72 Mb assembled sequences were anchored to 17 and 27 pseudo-chromosomes for S. lentiscoides and C. ovagalla, with 11,747 and 14,492 predicted protein-coding genes, respectively.
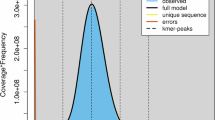
Aphid galls and genome features of Slavum lentiscoides and Chaetogeoica ovagalla. Galls and aphids living in galls of S. lentiscoides (a) and C. ovagalla (b). K-mer distribution plots for genomes of S. lentiscoides (c) and C. ovagalla (d). Hi-C interaction heatmaps for genomes of S. lentiscoides (e) and C. ovagalla (f).
Methods
Sample collection
S. lentiscoides samples within galls were collected on a pistachio tree (P. vera) from Andijan, Uzbekistan (40.719°N, 72.436°E) in September 2019. The galls of C. ovagalla were obtained on Pistacia chinensis from Qingdao, China (36.193°N, 120.573°E) in July 2018. The aphids from fresh galls were rapidly frozen and stored in liquid nitrogen at the National Animal Collection Resource Center, Institute of Zoology, Chinese Academy of Sciences, Beijing, China.
Genome sequencing
Total genomic DNA was extracted from over 0.2 grams of aphids within a single gall using a standard CTAB method. For Oxford Nanopore sequencing, the concentration and quality of DNA were checked using 1% agarose gel electrophoresis, NanoDrop spectrophotometry (Thermo Fisher Scientific, Lafayette, USA), and Qubit fluorometry (Invitrogen, Darmstadt, Germany). Large-sized segment libraries were selected (≥30 kb) with the BluePippinTM System and processed by the ONT Template Prep Kit (SQK-LSK109, Oxford Nanopore Technologies [ONT], Oxford, UK) protocol. DNA fragments were end-repaired and 3′-adenylated using the NEB Next FFPE DNA Repair Mix kit (New England Biolabs [NEB], Ipswich, USA). The Nanopore sequencing adapters were ligated using the NEBNext Quick Ligation Module (E6056) (NEB). The final library was sequenced on R9 flow cells using a PromethION DNA sequencer (ONT) with an ONT sequencing reagent kit (EXP-FLP001.PRO.6). The raw signal data were called, and the FAST5 files were converted into FASTQ files using MinKNOW software v2.0 (ONT). Short reads (<2kb) and reads with low-quality bases and adapter sequences were removed. Shotgun sequencing reads were used to estimate genome size and to correct the genome assembly. Paired-end libraries were prepared following the instructions for sequencing with the NovaSeq 6000 Reagent Kit on a NovaSeq 6000 platform, with an insert size of 350 bp. For Hi-C sequencing, nuclear DNA from tissue cells was cross-linked and enzymatically digested with DpnII. The DNA fragments with interaction relationships were captured with streptavidin beads and prepared for sequencing. Hi-C libraries with 300–700 bp insert sizes were constructed on the PE150 Illumina platform. Total RNA was extracted from the same colony for RNA extraction and sequencing. RNA-seq libraries were constructed using the NEBNext Ultra RNA Library Prep Kit for Illumina (NEB) and were then sequenced on the NovaSeq 6000 platform with a 150-bp paired-end output.
We finally generated for the first time chromosome-level genome assemblies of S. lentiscoides and C. ovagalla using a combination of Nanopore long reads (44.14 Gb and 55.02 Gb, respectively), Illumina short reads (58.13 Gb and 36.40 Gb, respectively), and Hi-C sequencing data (40.35 Gb and 45.54 Gb, respectively).
Karyotype analysis
We dissected young embryos of aphids on slides. The embryos were kept in 0.7% sodium citrate for 30 min and then fixed in Carnoy’s fixative for 15 min. After fixation, a coverslip was placed on the slide and vertically pressed down to disperse the cell mass as much as possible. The treated slide was frozen at −80 °C for 10 min, followed by air drying. The air-dried slide was stained in 5% Giemsa solution for 15 min and then washed with a tiny stream of distilled water. The slide was observed and subjected to photomicrography using a Leica DM6B photomicroscope (camera magnification 100 × 10). The result showed that S. lentiscoides has a diploid chromosome number of 2n = 34 (Fig. S1), while the karyotype of C. ovagalla was not obtained due to sample limitations.
Genome assembly
Before genome assembly, we extracted Illumina paired sequencing reads with approximately 50× coverage to estimate genome size and heterozygosity. The 17 k-mer frequency spectra obtained with Jellyfish v1.1.1021 and GenomeScope22 suggested that the heterozygosity of the S. lentiscoides and C. ovagalla genomes was 0.99% and 0.70%, the estimated genome sizes were 293.69 Mb and 276.95 Mb, and the repeat sequence content was 41.0% and 40.5%, respectively (Table 1, Fig. 1c,d).
Different assembly strategies were used to assemble the genomes of these two aphid species. For S. lentiscoides, the Nanopore long reads were initially cleaned using Canu v1.523 for sequence error correction and were then assembled from the error-corrected reads with the parameters “useGrid = false, genomeSize = 300 m, minReadLength = 2000, minOverlapLength = 500, corOutCoverage = 135, corMinCoverage = 2”. We used the Redundans24 pipeline to remove bubble contigs from the draft assembly. For C. ovagalla, the long reads were analyzed using Nextdenovo v2.125 for correction with the parameter “-seed_cutoff = 30k”, and then the corrected reads were assembled by SMARTdenovo (https://github.com/ruanjue/smartdenovo) with the parameters “-k 21 -J 5000 -t 20”. We found that the Canu assembly pipeline run obviously more slowly than the latter methods. These two draft aphid genome assemblies were subjected to two rounds of error correction using RACON26 with Nanopore reads, followed by three rounds of additional polishing using Pilon27 with Illumina sequencing data. We further removed a few contigs with microbial contamination by BLAST searches against the NCBI nt database, and no contigs related to the mitochondrial genome were found. After assembly and correction, we produced draft genome assemblies of S. lentiscoides and C. ovagalla, comprising a total of 336.27 Mb and 293.08 Mb of sequences, with 313 and 128 contigs, in which the contig N50 lengths were 6.79 Mb and 5.38 Mb, respectively (Table 1).
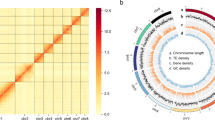
The Hi-C paired-end reads were mapped to the Nanopore assembly using BWA aln v0.7.10-r78928. The unique read pairs around the DpnII site were determined. Then, we applied LACHESIS software29 to cluster, order, and orient contigs from the draft assembly. Invalid read pairs were filtered with HiC-Pro30 using default settings. The predicted chromosomal genome was divided into bins of equal length (500 kb), and the number of valid Hi-C read pairs between bins was then used to represent the interaction signals between bins. Finally, we constructed a heatmap based on these interaction signals by R. For C. ovagalla, considering the variability in chromosome numbers observed in Aphididae31, we initially established a range of 10 to 30 for the cluster number. After careful consideration and analysis, we determined that a cluster number of 27 was the most appropriate choice. As a result, 40.35 Gb and 45.54 Gb of Hi-C clean reads were generated to build chromosome-level assemblies. After clustering, ordering, and orientation of the contigs, we obtained two final assemblies with chromosome-anchored sizes of 332.26 Mb and 289.72 Mb and scaffold N50 lengths of 19.77 Mb and 11.85 Mb for S. lentiscoides and C. ovagalla, respectively (Table 1). Therein, 128 and 86 contigs could be anchored to 17 and 27 linkage groups, with 98.81% and 98.85% anchoring rates, respectively (Table 1, Fig. 1e,f). The chromosome circus plots were generated using Circos v0.69-932 (Fig. 2).
Genome annotation
Before gene prediction, a de novo repeat library was built by using LTR_FINDER33 and RepeatScout34, and the library was classified by PASTEClassifier35. Transposable element sequences in the genome were predicted with RepeatMasker36 using the de novo library and Repbase library37. Consequently, we identified 87.71 Mb and 61.25 Mb of repeat sequences, accounting for 25.94% and 20.90% of the genome assemblies of S. lentiscoides and C. ovagalla, respectively (Table 1).
The protein-coding genes were predicted by integrating the evidence generated via the de novo, homology-based, and RNA-seq-based methods. For ab initio prediction, we used Augustus v2.438 to generate the de novo predictions with the default parameters. For the homology-based analysis, GeMoMa v1.3.139 was used to query the protein sequences against a database of four species (Myzus persicae, Aphis gossypii, Rhopalosiphum maidis, and Acyrthosiphon pisum). For RNA-seq annotation, RNA-seq reads were aligned to the genome assembly using HISAT2 v2.0.440 and StringTie v1.2.341 with the default parameters. TransDecoder v2.042, GeneMarkS-T v5.143, and PASA v2.0.244 were used to generate transcript predictions. Finally, we integrated three types of evidence for gene prediction by EVidenceModeler (EVM) v 1.1.145. By integrating ab initio-based, homology-based, and transcriptome-based evidence, we predicted 11,747 genes in the S. lentiscoides genome, and 14,492 genes in the genome of C. ovagalla. The predicted genes were then searched for homology-based functions by BLAST searches against the NCBI non-redundant protein sequences (nr), Eukaryotic Orthologous Groups (KOG), TrEMBL, Kyoto Encyclopedia of Genes and Genomes (KEGG), and the Gene Ontology (GO) databases with BLASTP v2.2.3146 (-evalue 1e-5). The GO terms were assigned using blast2go v5.047. A total of 99.22% (11,656 genes) and 89.73% (13,004 genes) of the predicted genes of these two genomes were supported by functional annotation from the above five protein databases. Among these genes of S. lentiscoides and C. ovagalla, 98.11% and 89.15% showed homology to proteins in NCBI nr, 98.89% and 67.84% in TrEMBL, 67.34% and 57.00% in KOG, 48.76% and 43.27% in KEGG, and 38.11% and 43.17% in GO, respectively (Table 2).
Three types of noncoding RNAs (ncRNAs) were identified. The microRNA (miRNA) and ribosomal RNA (rRNA) genes were predicted by BLASTN searches against the Rfam database48. The transfer RNA (tRNA) genes were identified using tRNAscan-SE49. Finally, we identified 27 and 31 miRNAs, 66 and 68 rRNAs, and 200 and 172 tRNAs within the genomes of S. lentiscoides and C. ovagalla, respectively (Table 1).
Data Records
The genome raw data, RNA sequencing data, and genome assemblies are available at the National Center for Biotechnology Information (NCBI) under the BioProject accession numbers PRJNA765394 and PRJNA832539. The Illumina WGS data was archived with the accession numbers SRR1604696350 and SRR2399932551. The Nanopore WGS data was deposited with the accession numbers SRR1604696452 and SRR2399932653. The RNA-Seq data was archived with the accession numbers SRR1604696154 and SRR2399932355. The Hi-C data was archived under the accession numbers SRR1604696256 and SRR2399932457. The genome assemblies have been deposited at GenBank under the accession numbers GCA_032441835.158 and GCA_032441825.159. The genome annotation files have been deposited at the Figshare60.
Technical Validation
The quality and completeness of draft assemblies were assessed by three methods as follows. The Illumina reads were mapped to the draft assembly with BWA-MEM v0.7.10-r78961, and the mapping ratio was calculated with SAMTOOLS v1.362. In addition, the Core Eukaryotic Genes Mapping Approach (CEGMA v2.563) and BUSCO v3.1.064 were used to assess the completeness of the genome assembly.
The S. lentiscoides and C. ovagalla genomes contained 96.77% and 97.58% of highly conserved Core Eukaryotic Genes (CEGs), respectively. They showed a high representation of conserved complete Benchmarking Universal Single-Copy Orthologs (BUSCOs) (S. lentiscoides: 93.70% complete BUSCOs of insecta_odb10; C. ovagalla: 97.00% complete BUSCOs of insecta_odb10), mapped with 96.78% and 97.19% of Illumina short reads, respectively, which indicated that these genome assemblies were of high quality and near-complete.
Code availability
This paper does not report original code. If no detailed parameters were mentioned for the software, default parameters were used according to the software introduction.
References
Stone, G. N. & Schönrogge, K. The adaptive significance of insect gall morphology. Trends Ecol. Evol. 18, 512–522 (2003).
Thorpe, P., Cock, P. J. & Bos, J. Comparative transcriptomics and proteomics of three different aphid species identifies core and diverse effector sets. BMC Genom. 17, 1–18 (2016).
Cambier, S. et al. Gall wasp transcriptomes unravel potential effectors involved in molecular dialogues with oak and rose. Front. Physiol. 10, 926 (2019).
Yamaguchi, H. et al. Phytohormones and willow gall induction by a gall-inducing sawfly. New Phytol. 196, 586–595 (2012).
Tooker, J. F. & Helms, A. M. Phytohormone dynamics associated with gall insects, and their potential role in the evolution of the gall-inducing habit. J. Chem. Ecol. 40, 742–753 (2014).
Hirano, T. et al. Reprogramming of the developmental program of Rhus javanica during initial stage of gall induction by Schlechtendalia chinensis. Front. Plant Sci. 11, 471 (2020).
Chakrabarti, S. Diversity and biosystematics of gall-inducing aphids (Hemiptera: Aphididae) and their galls in the Himalaya. Orient. Insects 41, 35–54 (2007).
Blackman, R. L. & Eastop, V. F. Aphids on the world’s plants: an online identification and information guide. http://www.aphidsonworldsplants.info/ (2024).
Wool, D. Galling aphids: specialization, biological complexity, and variation. Annu. Rev. Entomol. 49, 175–192 (2004).
Chen, J. & Qiao, G. X. Galling aphids (Hemiptera: Aphidoidea) in China: diversity and host specificity. Psyche 2012, 621934 (2012).
Dial, D. T. et al. Whole-genome sequence of the Cooley spruce gall adelgid, Adelges cooleyi (Hemiptera: Sternorrhyncha: Adelgidae). G3-Genes Genomes Genet. 14, jkad224 (2023).
Rispe, C. et al. The genome sequence of the grape phylloxera provides insights into the evolution, adaptation, and invasion routes of an iconic pest. BMC Biol. 18, 1–25 (2020).
Stern, D. L. & Han, C. Gene structure-based homology search identifies highly divergent putative effector gene family. Genome Biol. Evol. 14, evac069 (2022).
Smith, T. E. et al. Elucidation of host and symbiont contributions to peptidoglycan metabolism based on comparative genomics of eight aphid subfamilies and their Buchnera. PLoS Genet. 18, e1010195 (2022).
Wei, H. Y. et al. Chromosome-level genome assembly for the horned-gall aphid provides insights into interactions between gall-making insect and its host plant. Ecol. Evol. 12, e8815 (2022).
Korgaonkar, A. et al. A novel family of secreted insect proteins linked to plant gall development. Curr. Biol. 31, 1836–1849 (2021).
Zhang, C. X., Tang, X. D. & Cheng, J. A. The utilization and industrialization of insect resources in China. Entomol. Res. 38, S38–S47 (2008).
Kafkas, S., Kafkas, E. & Perl-Treves, R. Morphological diversity and a germplasm survey of three wild Pistacia species in Turkey. Genet. Resour. Crop Ev. 49, 261–270 (2002).
Ahmed, Z. B. et al. Study of the antioxidant activity of Pistacia atlantica Desf. gall extracts and evaluation of the responsible compounds. Biochem. Syst. Ecol. 100, 104358 (2022).
Giner-Larza, E. M. et al. Anti-inflammatory triterpenes from Pistacia terebinthus galls. Planta Medica 68, 311–315 (2002).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Pryszcz, L. P. & Gabaldón, T. Redundans: an assembly pipeline for highly heterozygous genomes. Nucleic Acids Res. 44, e113 (2016).
Hu, J. et al. An efficient error correction and accurate assembly tool for noisy long reads. Preprint at https://doi.org/10.1101/2023.03.09.531669 (2023).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 1–11 (2015).
Gavrilov-Zimin, I. A., Stekolshchikov, A. V. & Gautam, D. C. General trends of chromosomal evolution in Aphidococca (Insecta, Homoptera, Aphidinea+Coccinea). Comp. Cytogenet. 9, 335–422 (2015).
Krzywinski, M. et al. Circos: An information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358 (2005).
Hoede, C. et al. PASTEC: an automatic transposable element classification tool. PLoS One 9, e91929 (2014).
Chen, N. Using Repeat Masker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 5, 4.10.11–14.10.14 (2004).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile. DNA 6, 11 (2015).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Keilwagen, J. et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Res. 44, e89 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. & Papanicolaou, A. TransDecoder (find coding regions within transcripts). Google Scholar (2016).
Tang, S., Lomsadze, A. & Borodovsky, M. Identification of protein coding regions in RNA transcripts. Nucleic Acids Res. 43, e78 (2015).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res. 33, D121–D124 (2005).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR16046963 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR23999325 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR16046964 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR23999326 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR16046961 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR23999323 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR16046962 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR23999324 (2023).
Institute of Zoology, Chinese Academy of Sciences. GenBank https://identifiers.org/insdc.gca:GCA_032441835.1 (2023).
Institute of Zoology, Chinese Academy of Sciences. GenBank https://identifiers.org/insdc.gca:GCA_032441825.1 (2023).
Xu, S. The genome annotation files of galling aphids Slavum lentiscoides and Chaetogeoica ovagalla. Figshare. https://doi.org/10.6084/m9.figshare.25602348.v1 (2024).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://doi.org/10.48550/arXiv.1303.3997 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Acknowledgements
All authors thank Baocheng Guo for his suggestion on data mining. The work was supported by the National Natural Science Foundation of China (No. 32030014), the Youth Innovation Promotion Association of Chinese Academy of Sciences (No. 2020087), and the Key Collaborative Research Program of the Alliance of International Science Organizations (Grant No. ANSO-CR-KP-2020-04).
Author information
Authors and Affiliations
Contributions
J.C., G.Q. and L.J. designed the research. G.Q. and L.J. identified voucher specimens. S.X. conducted karyotype experiments and all data analyses. Z.Z. and M.Z. assisted in genome assembly and annotation. J.C. guided all data analyses. S.X. and J.C. wrote the manuscript and all authors contributed to revisions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, S., Jiang, L., Zou, Z. et al. Two chromosome-level genome assemblies of galling aphids Slavum lentiscoides and Chaetogeoica ovagalla. Sci Data 11, 803 (2024). https://doi.org/10.1038/s41597-024-03653-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03653-x
- Springer Nature Limited