Abstract
Balanced immunity is pivotal for health and homeostasis. CD4+ helper T (Th) cells are central to the balance between immune tolerance and immune rejection. Th cells adopt distinct functions to maintain tolerance and clear pathogens. Dysregulation of Th cell function often leads to maladies, including autoimmunity, inflammatory disease, cancer, and infection. Regulatory T (Treg) and Th17 cells are critical Th cell types involved in immune tolerance, homeostasis, pathogenicity, and pathogen clearance. It is therefore critical to understand how Treg and Th17 cells are regulated in health and disease. Cytokines are instrumental in directing Treg and Th17 cell function. The evolutionarily conserved TGF-β (transforming growth factor-β) cytokine superfamily is of particular interest because it is central to the biology of both Treg cells that are predominantly immunosuppressive and Th17 cells that can be proinflammatory, pathogenic, and immune regulatory. How TGF-β superfamily members and their intricate signaling pathways regulate Treg and Th17 cell function is a question that has been intensely investigated for two decades. Here, we introduce the fundamental biology of TGF-β superfamily signaling, Treg cells, and Th17 cells and discuss in detail how the TGF-β superfamily contributes to Treg and Th17 cell biology through complex yet ordered and cooperative signaling networks.
Similar content being viewed by others
Introduction
Our immune system, consisting of the innate and adaptive arms, has evolved to achieve two principal goals to maintain health. One is to recognize and tolerate entities that are deemed innocuous, or “self”. The other is to recognize and reject entities that are deemed nocuous or “nonself”. T cells are fundamental to achieve these principal goals because of T cells’ ability to recognize, with high specificity, nearly infinite numbers of antigens derived from an entity, regardless of “self” or “nonself”. For proper immunity, T cells must also properly distinguish “self” and “nonself”. Thymic negative selection eliminates T cells reacting strongly to “self”. Innate immunity enables a productive T-cell response against “nonself”. In addition, cytokines play crucial roles in balancing tolerance and immunity following the productive T-cell response. TGF-β has been recognized as a chief cytokine of “Yin-Yang” function, as it is critical for both tolerance and immunity. TGF-β was initially regarded as an immune regulatory cytokine because it suppresses proinflammatory cytotoxic and Th cells and promotes immunosuppressive Treg cells. TGF-β was later found to induce Th17 cells of both immune regulatory and pathogenic functions. It is now clear that TGF-β regulates T-cell-mediated tolerance and immunity through both Treg and Th17 cells in a context-dependent manner. Emerging evidence suggests a broad function for the TGF-β superfamily in Treg and Th17 cell biology. TGF-β superfamily signaling pathways crosstalk and interact with a myriad of other factors and pathways through multilayered mechanisms to translate complex environmental cues into defined and precise responses. Such a property of the TGF-β superfamily enables intricate control of Treg and Th17 cell function in a context-dependent manner, highlighting TGF-β’s fundamental role in immune balance. This article aims to familiarize readers with the current understanding of the TGF-β superfamily signaling and the biology of Treg and Th17 cells and to further discuss the involvement of the TGF-β superfamily in Treg and Th17 cell biology.
TGF-β superfamily signaling is regulated at multiple molecular levels
The TGF-β superfamily consists of over 35 members, including TGF-βs, Activins, BMPs (bone morphogenetic proteins), Nodal, and GDFs (growth differentiation factors). These secreted proteins play pleiotropic roles in controlling the development, homeostasis, proliferation, differentiation, and functions of diverse cell types in health and disease [1,2,3,4,5,6,7]. In the immune system, TGF-β is the best studied. Activin [8,9,10,11] and BMP [12,13,14,15,16,17] also contribute to immune regulation. TGF-β has three homologs: TGF-β1, TGF-β2, and TGF-β3. While the biochemical properties of TGF-β1, TGF-β2, and TGF-β3 are similar, their expression pattern and function are tissue- and cell-type specific. Germline deletion of TGF-β2 and TGF-β3 leads to embryo lethality [18, 19]. In contrast, while TGF-β1 deletion perturbs endothelial differentiation and yolk sac hematopoiesis to a certain extent [20], TGF-β1 knockout mice can be born but succumb to severe multifocal inflammation shortly after birth [21,22,23]. Therefore, the principal function of TGF-β1 is to control hematopoiesis and immune function, which agrees with its preferential expression in immune cells when compared to other isoforms [24]. TGF-β1 is produced by and regulates the function of both innate immune cells, including macrophages [25,26,27] and dendritic cells (DCs) [28,29,30], and adaptive immune cells, including T and B cells [31, 32]. To control the immune responses, TGF-β1 mainly targets T cells because T-cell-specific deletion of TGF-β receptors results in a systemic, multifocal, and lethal inflammatory disease resembling the phenotypes of TGF-β1-deficient mice [33,34,35].
TGF-β activation via complex posttranslational mechanisms
While the transcriptional and translational regulation of the Tgfb gene is poorly understood, the generation of biologically active TGF-β at the posttranslational level has been studied extensively [36, 37]. TGF-β is synthesized as a precursor molecule composed of a signal peptide, a pro-domain (latency-associated-peptide, LAP), and the mature polypeptide (TGF-β) [36, 37]. After signal peptide removal, latent LAP-TGF-β dimerizes and forms a large latent complex by binding covalently to LTBP (large-latent-TGF-β-binding-protein) through disulfide bonds to be deposited in the extracellular matrix [36]. Latent LAP-TGF-β can also disulfide-link with membrane-bound GARP (glycoprotein-A repetitions predominant protein), a protein expressed on the surface of Treg cells and platelets to position TGF-β at the cell membrane [38]. Therefore, TGF-β secretion and function tend to be localized. Unlike TGF-β, Activins and some BMPs are not secreted as a latent complex. Mature TGF-β only becomes active to induce signal transduction after being freed from LAP by proteolytic cleavage via various integrins and proteases and extracellular matrix proteins in cell-type- and context-dependent manners [39] (Fig. 1).
TGF-β activation and signaling. Inactive LAP-TGF-β is produced and associates with LTBP in the extracellular matrix or with GARP on the cell membrane. Active TGF-β is released from LAP-TGF-β by proteases, integrins and low pH. Activated TGF-β binds to its receptor to activate r-Smad proteins. Activated r-Smad proteins interact with co-Smad4 to translocate into the nucleus to control target gene expression by interacting with various transcription factors (TFs) and coregulators. i-Smad and SKI/SnoN proteins negatively regulate TGFβR and Smad function. E3-ubiquitin ligases, including Smurf and Arkadia, target protein degradation to modulate TGF-β signaling. TGF-β binding to its receptor also activates MAP kinase pathways, including Ras-ERK, PI3K-AKT-mTOR, and TAK1, to program cellular responses independent of Smad pathways
TGF-β signaling via specific receptors and Smad proteins
Active TGF-β binds to its specific receptor on target cells to signal and program cellular function (Fig. 1). The signaling mechanisms of the TGF-β superfamily are conserved in both immune and nonimmune cells [40,41,42]. TGF-β receptor (TGFβR) is a transmembrane heterotetrametric complex consisting of two copies of ligand-specific receptor I and receptor II. Upon TGF-β binding, TGFβRII phosphorylates TGFβRI (also known as ALK (activin receptor-like kinase)-5) and activates ALK-5’s serine/threonine domain. Activated ALK-5 phosphorylates and activates receptor-associated (r)-Smad (suppressor of mothers against decapentaplegic) proteins, including r-Smad2 and r-Smad3, and promotes their disassociation from SARA (Smad anchor for receptor) [43, 44]. Activated r-Smad proteins can then translocate to the nucleus with or without associating with co-Smad4, a common Smad protein that can bind to r-Smad proteins activated by TGF-βs, Activins, and BMPs [42]. Smad-containing complexes, which often include other transcription factors and epigenetic regulators, bind to target loci in the genome to regulate gene expression positively or negatively. In addition to co-Smad4, r-Smad2 and r-Smad3 can also bind to a nuclear protein called TIF1γ (transcriptional intermediary factor 1γ), also known as TRIM33 (tripartite motif-containing 33), to control target gene expression [45]. Like TGF-β, Activins and BMPs bind to and activate their specific receptors and r-Smads to control target gene expression. The major type I receptor for Activin is ALK-4, which activates r-Smad2 and r-Smad3. The major BMP type I receptors are ALK-1, ALK-2, ALK-3, and ALK-6, which activate r-Smad1, r-Smad5, and r-Smad8. In addition to binding to ALK-5, TGF-β can bind to ALK1 or ALK2 to stimulate epithelial cell proliferation and migration [46] and to activate r-Smad1 and r-Smad5 during epithelial-to-mesenchymal transition [47]. Therefore, TGF-β can crosstalk with other members of the same family, especially Activins and BMPs, in a context-dependent manner, which can be of importance for immune regulation [8,9,10,11,12,13,14,15,16,17]. However, not all Smad proteins promote TGF-β signaling; inhibitory (i) Smad6 (i-Smad6) and i-Smad7 dampen TGF-β signaling by associating with type I receptors to prevent r-Smad activation or to disrupt the association between r-Smad and co-Smad proteins [7, 42, 48].
As the major transducers of the TGF-β signaling pathway, Smad proteins have two highly conserved MH (mad homology) domains. N-terminal MH1 and C-terminal MH2 mediate nuclear localization and protein‒protein interactions, respectively. Therefore, the MH1 and MH2 domains are important for Smads to bind to DNA and other proteins. The binding of Smads to various genetic loci and proteins enables TGF-β signaling pathways to crosstalk with a plethora of other signaling pathways to control diverse cellular functions [42, 48]. r-Smad3 and the co-Smad4 complex interact with c-Jun and c-Fos at the AP1 binding site to establish crosstalk between the pathways of TGF-β and JNK (c-Jun N-terminal kinase), a MAPK (mitogen activated protein kinase) [49]. r-Smads and co-Smad4 cooperate with LEF-1 (lymphoid enhancer-binding factor 1) and β-catenin in Wnt signaling [50]. In addition, TGF-β signaling regulates the transcription of HES-1 (hairy and enhancer of split-1) of the Notch pathway through the direct binding of r-Smad3 to NICD (notch intracellular domain) [51]. TGF-β signaling also converges with Hedgehog signaling through the co-Smad4 and GLI (glioma-associated oncogene) interaction, involving r-Smad2 and histone acetyltransferase PCAF (p300/CREB-binding protein-associated factor), to activate target genes [52]. Since the MAPK, Wnt, Notch, and Hedgehog pathways are important for T-cell functions, it is plausible that TGF-β exerts broad effects on T cells by corroborating these pathways, which warrants further study. Smad function can be regulated through posttranslational mechanisms [53,54,55]. For example, the MH2 domain of r-Smad can be dephosphorylated by the protein phosphatase PPM1A (protein phosphatase, Mg2+/Mn2+ dependent, 1 A)/PP2Cα (protein phosphatase-2Cα) [56] and can be phosphorylated by casein kinase Iγ2 [57] to promote r-Smad degradation. R-Smad’s linker region can be phosphorylated by p38 MAPK and ROCK (Rho-associated coiled coil kinase) [58], JNK [59], and casein kinase Iϵ [60] to positively or negatively regulate TGF-β responses. In addition, mono- and polyubiquitylation, sumoylation, acetylation, methylation, and polyadenosine diphosphate (ADP)-ribosylation of Smads regulate Smad function [53]. Notably, r-Smads, co-Smad4, i-Smads, and the activated Smad complex can be ubiquitinated by various E3 ubiquitin ligases. The best-known E3 ubiquitin ligases that regulate TGF-β signaling pathways belong to the HECT (homologous to E6AP C-terminus) family and Smurf (Smad ubiquitin regulatory factors) family, including Smurf1 and Smurf2. These E3 ubiquitin ligases target TGF-β-activated r-Smad2 and r-Smad3 and BMP-activated r-Smad1 and r-Smad5 [61, 62]. r-Smad3 is also targeted by other ubiquitin E3 ligases, including SCF (Skp1-Cullin-F-box) proteins and U-box CHIP (carboxyl terminus of Hsc70-interacting protein) [63,64,65]. Nuclear RNF (ring finger protein) 111, also known as Arkadia, is an E3 ligase that mediates the degradation of i-Smad7, SnoN, and SKI, which can lead to enhanced TGF-β signaling [66, 67]. In contrast to polyubiquitination-induced, proteosome-mediated degradation, monoubiquitylation leads to different functional outcomes of Smad proteins. The monoubiquitylation of r-Smad2 by Itch (Itchy E3 ubiquitin protein ligase) promotes the stable interaction between r-Smad2 and TGFβRI. However, the monoubiquitylation of r-Smad3 interferes with Smad3’s ability to bind to target loci [68, 69].
TGF-β signaling pathways also regulate gene transcription epigenetically by interacting with chromatin remodelers, histone modifiers, DNA modifiers, chromatin readers, and long noncoding RNAs [70]. r-Smad2 interacts with SMARCA4 (SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily A, member 4; also known as BRG1) to epigenetically modify chromatin structures for the expression of a majority of the TGF-β-targeted genes, except i-Smad7 and SnoN, to achieve a tightly controlled feedback loop of TGF-β signaling [71]. HAT (histone acetyltransferase) and HDAC (histone deacetylase) are importantly involved in TGF-β signaling [72]. r-Smad1, r-Smad2, r-Smad3, r-Smad5, and co-Smad4 can all recruit HAT or HDAC through the MH1 domain to control target gene expression in a context-dependent manner [73,74,75]. In addition to modifying histone acetylation, Smad proteins regulate histone methylation. In response to Nodal stimulation, r-Smad2 and r-Smad3 recruit histone demethylase JMJD3 (Jumonji domain-containing protein 3) to the promoter region to reduce inhibitory histone markers of H3K27me3 [76].
TGF-β signaling through Smad-independent pathways
In addition to Smad-dependent pathways, TGF-β receptors can signal through Smad-independent pathways, including ERK (extracellular signal-regulated kinase), JNK, p38 MAPK, PI3K (phosphatidylinositol-3 kinase), AKT, and Rho family GTPases [48, 77] (Fig. 1). TGF-β-activated ERK signaling requires ShcA. Activated TGFβRI recruits and phosphorylates ShcA on tyrosine and serine residues. Phosphorylated ShcA assembles the ShcA-Grb2-SOS complex and activates Ras for MEK1/2 and ERK signaling [78]. Activated TGFβR recruits TAK1 (TGF-β activated kinase 1) to further activate the p38 MAPK and JNK pathways. TGF-β stimulates the PI3K-AKT-mTOR pathway through an indirect association of activated TGFβRI and TGFβRII with p85, a regulatory unit of PI3K [79]. TGF-β can also induce the ubiquitination of p85 through TRAF6, resulting in AKT-mTOR activation [80], which is important for cell proliferation, metabolism, and migration [81, 82].
TGF-β signaling has evolved to carry out broad functions through diverse mechanisms by integrating various environmental stimuli in a cell-type- and context-dependent manner. Since TGF-β and its signal are central to a myriad of cellular functions that are vital for development, health, and disease, TGF-β and its signal must be carefully regulated. Such regulation is achieved by imposing controls at every step of the signaling process to ensure proper outcomes in response to the unique combination of stimuli in a specific niche. In the following sections, we will discuss the roles of TGF-β signaling in regulating Treg and Th17 cells. We will first introduce the important aspects of the biology of Treg and Th17 cells and then discuss in detail how TGF-β signaling regulates their functions.
TREG Cells are central to immune tolerance and homeostasis
Approximately a century ago, the concept that mechanisms must exist to prevent autoimmunity and to uphold tolerance was proposed by Paul Ehrlich, albeit in an absolute sense denying the possibility of autoimmunity. The very existence of autoimmunity was not accepted until after the 1960s [83]. With the realization that self-tolerance is not given and can be broken to result in autoimmune diseases, much effort has been devoted to understanding how tolerance is established and maintained. Decades of investigation have informed us that multiple mechanisms contribute to tolerance, including clonal selection, anergy induction, and active immune suppression through cells and cytokines [84,85,86,87]. In the 1970s, the existence of a cell type with active immune suppression function was proposed [88,89,90]. It was not until the 1980s that Sakaguchi et al. identified Lyt-1high,2,3low T cells enriched for immune-suppressive activity [91, 92]. In 1995, a seminal work revealed that CD4+CD25+ cells are markers for a T-cell population that is highly enriched for immune-suppressive function [93], allowing for extensive characterization of these cells.
The genetic underpinning of suppressor T cells was revealed in the early 2000s. Monogenetic loss-of-function mutation of an X-linked gene called Foxp3 (forkhead box p3) was found to be accountable for multifocal lymphoproliferation autoimmunity syndrome in scurfy mice and IPEX (immunodysregulation, polyendocrinopathy, and enteropathy, X-linked) human patients [94,95,96,97]. It was later found that the Foxp3 gene is predominantly, if not exclusively, expressed in CD4+CD25+ T cells [98] and that Foxp3 is sufficient and required for the generation and function of CD4+CD25+ suppressor T cells [99, 100]. CD4+CD25+ T cells with Foxp3 expression and immunosuppressive activity are now defined as regulatory T (Treg) cells. Of note, while Foxp3 expression alone can mark mouse Treg cells with good certainty, it is often unreliable for human Treg cells, as Foxp3 can be upregulated, often transiently, in activated human CD4+ T cells with no suppressive function [101,102,103]. In this review, we will focus on discussing mouse Treg cells.
Treg cell classification, generation, and maintenance
Treg cells can develop in the thymus. Thymus-derived Treg cells are known as tTreg cells. In addition, Treg cells can be generated extrathymically at peripheral sites. Periphery-generated Treg cells are known as pTreg cells. Moreover, Treg cells can be induced in cell culture when CD4+ T cells are activated in the presence of TGF-β. TGF-β-induced Treg cells in culture are known as iTreg cells [104, 105]. The expression of Foxp3 is induced in tTreg cells during thymic development in response to moderately strong TCR stimulation in conjunction with interleukin-2 (IL-2) produced by activated self-reactive thymocytes [106,107,108,109]. tTreg cells stably express Foxp3 and other Treg cell markers by extensive epigenetic modification at relevant genetic loci for systemic self-tolerance [110]. pTreg cells can be generated from Foxp3– CD4+ T cells that are exposed to factors, including TGF-β and IL-2, in peripheral tissues [111, 112]. These pTreg cells accumulate mostly at barrier sites (such as the intestines) where they respond to innocuous antigens derived from food or commensal microbes, metabolites, and environmental factors [113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129]. Non-Treg cells can be induced to become iTreg cells in culture by TGF-β [130], retinoic acid [122], and IL-2 [131] following TCR stimulation. While iTreg cells can express high levels of Foxp3 and are immunosuppressive, they generally lack the epigenetic modifications of tTreg cells to stabilize Treg cell phenotypes [132, 133]. In addition to the populations of Foxp3+ Treg cells defined by the above taxonomy [105], Treg cells can be further or alternatively categorized into naive, activated, effector, and memory Treg cells based on the surface expression of CD62L, CD44, CD127, CD69 and Klrg [134]. Tissue-resident Treg cells have distinct genetic programs from tTreg cells and bear unique markers, including CD103 and chemokine receptors, to regulate immune responses in a tissue-specific manner [134, 135].
Stable Foxp3 expression is important for Treg cell maintenance, identity, and function [136,137,138,139] (Fig. 2). Cytokines are central to Treg cell homeostasis. γc-dependent cytokines are critical for the development and homeostasis of tTregs as well as pTreg and iTreg cells. TGF-β is especially important for inducing pTreg and iTreg cells [109, 140]. In addition, PI3K-mTOR-AKT, an important signaling axis regulated by TCR, costimulation, and cytokines, including TGF-β, plays a pivotal role in the development, function, and maintenance of Treg cells by regulating immune metabolism [141, 142] and the expression of genes critical for Foxp3 expression, including FOXOs [143,144,145]. Mechanisms also exist to specifically control stable Foxp3 expression independent of its induction. Three CNSs (conserved noncoding DNA sequences), namely, CNS1-3, in Foxp3 loci are critical for Foxp3 expression [146]. While CNS1 and CNS3 are important for TGF-β- and TCR/CD28-induced Foxp3 expression, respectively, CNS2 is specifically required to stabilize Foxp3 expression in the progeny of dividing Treg cells. Foxp3, Runx1-Cbfβ, and GATA3 bind to CNS2 to promote stable Foxp3 expression [146,147,148,149]. In addition, the posttranslational modifications of Foxp3 proteins, including phosphorylation, O-GlcNAcylation, acetylation, ubiquitylation, and methylation, are important for controlling the functions of Foxp3 and therefore Treg cells [150, 151]. It is thus predicted that perturbations in the abovementioned mechanisms through genetic, pharmacological, and environmental means alter Treg cell homeostasis and function, with or without affecting the stability of Foxp3 expression [152,153,154,155,156,157,158,159,160].
TGF-β signaling promotes Treg cell function for tolerance and homeostasis. TGFβR and Smad proteins promote Foxp3 expression and Treg cell generation by activating CNS1 of the Foxp3 locus. The promoter, CNS2, and CNS3 regions are also important for the induction and stability of Foxp3 expression by integrating a myriad of upstream signaling pathways, including TCR, CD28, CD25, AHR and various transcription factors. The Arkadia/SKI/SnoN pathway is important for TGF-β-induced Foxp3 expression. Foxp3-expressing Treg cells suppress the function of T cells, macrophages, and DCs through TGF-β, IL-10, granzymes, CTLA-4, and CD25. Treg cells promote tissue repair through CCR6-mediated chemotaxis and amphiregulin- and CCN3-mediated tissue regeneration to maintain tolerance and homeostasis in health and disease
The mechanisms of Treg cell function
Treg cells control adaptive and innate immune responses through broad mechanisms in both antigen-specific and antigen-nonspecific manners [161] (Fig. 2). Treg cells carry out their functions through cytokine modulation, cytolysis, metabolic disruption, and modulation of DC maturation and function [140, 162]. Treg cells can directly suppress T-cell function by acting as a sink for IL-2, which is critical for the survival and proliferation of activated effector T cells [163, 164], as CD25 (IL2Ra) is highly expressed by Treg cells. Treg cells, especially activated Treg cells, produce large amounts of TGF-β and IL-10 to suppress both adaptive and innate immunity [84]. High levels of CTLA-4 (cytotoxic T-lymphocyte associated protein-4) on the surface of Treg cells can downregulate costimulatory molecules on antigen-presenting cells (APCs) and induce tolerogenic DCs [165, 166] to reduce the ability of APCs to activate non-Treg cells [101, 167]. Treg cells produce elevated levels of granzymes that are suggested to target APCs for disruption [168]. Treg cells are also found to form more stable interactions with APCs than non-Treg cells [169, 170] to prevent non-Treg cells from interacting with APCs. Of note, Foxp3 reprograms Treg cell metabolism to adapt to low-glucose, high-lactate environments [171]. This allows Treg cells to utilize different energy resources than other activated immune cells for better fitness in a specific niche, especially under inflammatory conditions [172, 173]. Treg cells migrate to and reside in a specific niche through chemotaxis by sensing chemokine gradients and expressing relevant chemokine receptors and integrins [174,175,176]. Interestingly, the molecular program of Treg cells seems to be quite flexible and heterogeneous to allow Treg cells to tailor their function to suppress specific responses. Treg cells can express T-bet, a Th1 cell differentiation factor [177], with an enhanced ability to suppress Th1 responses [178]. Treg cells can express IRF4 (interferon regulatory factor 4), a Th2 differentiation factor [179], with an enhanced ability to suppress Th2 responses [180]. Treg cells can express RORγt, a Th17 differentiation factor [181], with an enhanced ability to suppress Th17 responses [182]. The mechanisms for these observations are not entirely clear. Two mutually nonexclusive possibilities are that (1) existing tTreg cells and/or residential Treg cells activated under Th1-, Th2-, and Th17-skewing conditions adopt respective molecular programs upon activation, as the epigenetic program of Th differentiation of Treg cells is “poised and flexible” but not “fixed” to allow such adaptation [157,158,159,160, 183, 184]. Adapted Treg cells, likely antigen specific, become better fitted in a microenvironment with enhanced suppressive function toward ongoing responses. (2) In a specific microenvironment with ongoing inflammation, a fraction of antigen-specific non-Treg cells are converted into pTreg cells by acquiring Foxp3 expression and suppressive function while maintaining their Th program acquired before conversion. These converted pTreg cells will likely have enhanced function toward ongoing responses. In fact, studies have shown that a fraction of CD4+ T cells, or pTreg cell precursors, in the intestines express RORγt before acquiring Foxp3 and Treg cell properties [185, 186].
While the cardinal function of Treg cells is to suppress immune responses, Treg cells are also important for promoting tissue repair through both indirect and direct mechanisms. The indirect mechanism can be attributed to the suppressive function of Treg cells [187]. Inflammatory cells, including neutrophils, macrophages, and activated conventional T cells, infiltrate into the site of inflammation and release tissue-damaging cytokines to adversely affect tissue repair. By suppressing the effector function of inflammatory cells, Treg cells promote tissue repair indirectly [187, 188]. More importantly, Treg cells can promote tissue repair through direct, immune-suppression-independent mechanisms. During inflammation, activated Treg cells migrate to inflammatory tissues, a process that requires CCR6, a chemokine receptor [189,190,191]. Tissue-infiltrating Treg cells produce amphiregulin, the epidermal growth factor receptor (EGFR) ligand, to promote tissue repair in acutely injured skeletal muscle and in acutely influenza virus-infected lungs [192,193,194]. In addition, CCN3, a growth regulatory protein implicated in tissue regeneration, is produced by Treg cells to promote oligodendrocyte differentiation and myelination to facilitate CNS tissue regeneration and to ameliorate neuro-immune pathologies, including EAE and MS [195, 196]. The myriad of Treg cell functions in immune suppression and tissue repair enable the broad involvement of Treg cells in health and disease.
Maintaining immune homeostasis
Immune homeostasis is the state in which the immune system maintains a balance between immune activation and immune tolerance. As one of the pillars of tolerance, proper Treg cell function is central to immune homeostasis [161, 197, 198]. Dysregulation of Treg cell function caused by genetic or environmental factors invariantly disrupts immune homeostasis and tolerance and often causes autoimmunity [199]. Therefore, enhancing Treg cell function can benefit disease treatment. For instance, low-dose IL-2 has been used to treat T1D by specifically promoting Treg cell function [200, 201]. More recently, adoptive Treg cell therapy has gained much attention for the treatment of autoimmune diseases in an antigen-specific manner [202,203,204]. In graft-versus-host or host-versus-graft diseases that can be viewed as a form of autoimmunity introduced through transplantation, enhancing Treg cell function benefits transplant tolerance [205, 206]. During allergic diseases resulting from tolerance breakdown mostly due to environmental factors, promoting Treg cell function helps to broadly restrain hyperactive T cells, eosinophils, mast cells, basophils, antibody isotype switching, inflammatory DCs, and inflammatory cell migration to tissues [207].
In addition to preventing and restraining autoimmune inflammation, Treg cells are important for establishing and restoring homeostasis during host–microorganism interactions. During the host‒pathogen interaction and pathogen clearance response, the activation of innate and adaptive immunity by pathogens causes inflammatory and immune pathology in hosts. Excessive immune activation often causes pathologies that can be debilitating or lethal. Proper Treg cell function is important to restore immune homeostasis during and after pathogen clearance. Treg cell-mediated suppression is necessary to temper inflammatory responses during infection and regain immune homeostasis after infection and pathogen clearance [208,209,210]. In addition, Treg cells establish tolerance to obligate microbiota [211]. Treg cells can achieve this by actively suppressing both innate and adaptive immunity through antigen-dependent and antigen-independent mechanisms [162] and by promoting tissue repair [187].
In addition to autoimmune and infection-induced perturbation of immune homeostasis, the aging process is often associated with a systemic inflammatory syndrome, also known as inflammaging. Inflammaging is a naturally occurring inflammation that progresses with age and contributes to immune senescence, immunological aging, and age-associated morbidities and mortalities, including infection, cancer, and autoimmunity in the elderly [212,213,214]. While many factors can contribute to inflammaging, reduced Treg cell function has recently been associated with inflammaging [215]. Aged Treg cells are more severely senesced and less proliferative than aged non-Treg cells. As a result, aged Treg cells are not optimal in restraining non-Treg cell function. Treg and non-Treg cell functions are off-balanced during aging, which may contribute to inflammaging [215]. Therefore, bolstering Treg cell function will help to establish, maintain, and restore immune and tissue homeostasis under normal and pathological conditions.
Establishing tumor tolerance
Tumors develop when the immune system fails to eradicate tumor cells. One reason for such a failure is due to the immune-suppressive TME (tumor microenvironment) established during coevolution of the tumor cells and the host. Treg cells are enriched in many tumor types and contribute to the TME and tumor tolerance [216,217,218,219,220,221]. Treg cells can be induced by self- and tumor-antigens in the presence of TGF-β, which is often produced by transformed cells, and can be clonally expanded in the TME [222, 223], aided by the Treg cells’ unique ability to metabolically adapt to low-glucose and high-lactate environments in tumors [171]. In addition to inducing Treg cells, tumors attract Treg cells by secreting chemokines, including CCL1, CCL5, CCL22, and CCL28, as well as by inducing chemokine receptor expression on Treg cells [220]. Strategies to target Treg cell generation, recruitment, and function promise to benefit cancer immunotherapy [224,225,226].
Treg cells impose critical and broad functions for tolerance, immune homeostasis, and tissue repair in health and disease. Much research has been focused on understanding how their generation and function are controlled. TGF-β emerges as a central regulator of Treg cell biology. In the following section, we will discuss in detail how TGF-β signaling controls various aspects of Treg cell biology in health and disease.
TGF-β controls TREG Cell generation, homeostasis, and function through cell-type and context-dependent mechanisms
The varying roles of TGF-β in Treg cell generation and homeostasis under different contexts
The interest in TGF-β in Treg cell function stems from early studies demonstrating that TGF-β1 deletion led to a multifocal lethal inflammatory disease early in life in mice in the 1990s [21,22,23]. The ensuing studies revealed that TGF-β is an immune regulatory cytokine that inhibits activation-induced T-cell proliferation and more potently suppresses Th cell differentiation and effector function [227]. The relationship between TGF-β and Treg cells became clear when a seminal study revealed that TGF-β promotes CD4+CD25- conventional T cells to differentiate into CD4+CD25+ cells upon activation in culture [130]. TGF-β-induced CD4+CD25+ cells are anergic and do not produce Th1 or Th2 cytokines, yet these cells express Foxp3 and produce TGF-β. Importantly, TGF-β-induced CD4+CD25+ cells are immunosuppressive and inhibit antigen-driven CD4+ T-cell expansion during lung inflammation [130]. Therefore, under culture conditions, TGF-β is sufficient to induce the generation of Treg cells that phenotypically resemble Treg cells identified in vivo. A later study using a Foxp3-mRFP reporter mouse strain found that such an induction occurs by promoting de novo Foxp3 expression in activated naive CD4+ T cells [132]. One mechanism by which TGF-β promotes Foxp3 expression is through r-Smad3 and NFAT (nuclear factor of activated T cells) binding to a Foxp3 enhancer region in T cells [228]. This enhancer region was later found to be the CNS1 region [146]. Other mechanisms can be through downregulating inhibitory i-Smad7 at the transcriptional level through Foxp3, which results in boosted TGF-β signaling through enhancing the r-Smad3 and co-Smad4 response to promote Foxp3 expression [229], suggesting that TGF-β promotes iTreg cell generation through a self-enhancing feedforward mechanism. While naive T cells are readily converted into Treg cells by TGF-β stimulation after activation in culture, TGF-β is incapable of converting predifferentiated Th cells into iTregs in culture [230], which could be due to the altered molecular context in differentiated Th cells and reduced TGFβRI expression in activated T cells [231].
Because TGF-β signaling can promote Foxp3 expression and Treg cell function both in culture and in vivo, whether and how TGF-β pathways are required for Treg cell function in vivo are important questions to be addressed. Studies were conducted in which components of TGF-β signaling pathways in T cells were deleted under various conditions. In one study, T-cell-specific knockout of TGFβRII led to reduced Treg cells in the periphery but not in the thymus [35]. In a mixed bone marrow chimeric mouse model, where both wild-type and TGFβRII-knockout T cells coexist, TGFβRII was later found to be required for maintaining Treg cells in the periphery in a cell-intrinsic manner [232]. Detailed analysis of T-cell-specific TGFβRI knockout mice revealed that TGFβR is critical for tTreg cell generation in the thymus [33]. Such a TGF-β-mediated effect can occur through a direct mechanism because TGF-β is abundant in the thymus due to thymocyte apoptosis during thymic selection [233]. The defective tTreg cell generation due to TGFβRI deletion can be compensated for, however, by the enhanced IL-2 response of TGFβRI-deficient Treg cells under inflammatory conditions in these mice [33]. Another mechanism by which TGFβR is required for tTreg cell generation may be indirect, through increased death of TGFβR-deficient tTreg cells under inflammatory conditions [234]. IFN-γ, an inflammatory cytokine suppressed by TGF-β, impairs the homeostasis of TGFβR-deficient Treg cells. IFN-γ deletion largely restored the TGFβRII-deficient Treg cell population in the periphery in a BDC2.5 NOD mouse model [235]. Therefore, under inflammatory conditions, TGF-β promotes Treg cell generation and maintenance in the thymus and periphery through both direct and indirect mechanisms.
The aforementioned findings also provide further understanding of how TGF-β may be involved in Treg cell generation and homeostasis in the absence of inflammation. Available evidence suggests that TGFβR is dispensable for the maintenance and function of existing Treg cells under homeostatic conditions. Unlike deleting floxed Tgfbr2 alleles in developing thymocytes using a Cd4-Cre transgene, deleting floxed Tgfbr2 alleles in mature T cells in the periphery using a distal-Lck-Cre transgene does not lead to autoimmunity or lymphoproliferation under steady state [236]. These findings indicate that lymphopenia-driven T-cell proliferation and inflammation are restrained by TGF-β signaling, which is important for Treg cell survival. In agreement, Treg cell-specific deletion of TGFβRII did not lead to systemic inflammation and did not apparently perturb tTreg cell populations [233, 237]. It is therefore plausible that TGF-β is critical for tTreg cell generation and maintenance in a context-dependent manner, depending on the inflammatory status of the niche. Further studies are warranted to understand why TGFβR is important for the generation and maintenance of tTreg cells, especially under inflammatory conditions.
The finding that TGFβR is required for the generation and maintenance of Treg cells in a context-dependent manner prompts the question of what sources of TGF-β are critical for Treg cell homeostasis in vivo. TGF-β is broadly produced by many cell types, including immune cells and nonimmune cells, in a localized manner. Of interest, T cells, especially Treg cells, produce TGF-β1 in a membrane-bound form [238, 239]. Deletion of TGF-β1 specifically in T cells or in Treg cells does not lead to early-onset autoimmunity, unlike in T-cell-specific TGFβR knockout mice [239, 240], although T-cell-specific TGF-β1 knockout mice develop immunopathology later in life [240]. Nonetheless, Treg cell homeostasis is not obviously perturbed in these mice. In fact, the tTreg population is slightly increased, suggesting that endogenously generated TGF-β1 by tTreg cells restrains tTreg cell homeostatic proliferation but is dispensable for their generation or maintenance under noninflammatory, homeostatic conditions. A study, however, challenges these findings by showing that the deletion of TGF-β1 in Treg cells led to impaired Treg cell homeostasis and autoimmune syndrome in mice [241]. Further examination revealed that the findings in this study were due to cryptic genetic manipulation, which led to undue Treg cell death [242]. Thus, the notion that TGF-β1 is dispensable for Treg cell homeostasis stands. Nonetheless, in TGF-β1−/− mice, where systemic inflammation occurs, TGF-β1 was shown to be important for Treg cells to be maintained in the periphery and to maintain the stability of Foxp3 expression [243]. Therefore, it becomes obvious that TGF-β controls Treg cell generation and homeostasis in a context- and microenvironment-dependent manner; while TGF-β is largely dispensable for Treg cell homeostasis under noninflammatory conditions, inflammation makes Treg cells sensitive to the loss of TGF-β and its signal for their generation, maintenance, and stability. The mechanisms underlying such a dichotomous role of TGF-β in Treg cells need further investigation to understand how Treg cells can be maintained under inflammatory conditions when needed most.
Multilayered mechanisms of TGF-β-promoted Treg cell generation and homeostasis
While TGF-β promotes both tTreg and iTreg cell generation in a context-dependent manner, TGF-β has differential effects on the genetic programs of tTreg and iTreg cells. Some of the tTreg genes, including Il2ra, Socs2, Tnfrsf18, and Ctla4, are not enhanced by TGF-β [244]. iTreg cells rely on continuous TGF-β signaling to maintain Foxp3 expression, without which they have a brief life and reduced Foxp3 stability when compared with Treg cells isolated in vivo [245]. While r-Smad2 and r-Smad3 are required for the generation of iTreg cells induced by TGF-β [246, 247], tTreg cell generation and homeostasis are unperturbed when r-Smad2 and r-Smad3 are deleted [246,247,248,249]. In addition to r-Smad2 and r-Smad3, the Smad-independent, TAK1-dependent TGFβR signaling pathway contributes to tTreg cell homeostasis [248]. Similarly, T-cell-specific knockout of co-Smad4 does not obviously affect tTreg cell generation even when TGFβRII is absent. However, iTreg cell generation is severely reduced when co-Smad4 is deleted [232, 250]. T-cell-specific knockout of Arkadia, an E3 ligase that mediates the degradation of i-Smad7, SKI, and SnoN, which are inhibitory to TGF-β signaling, led to reduced generation of iTreg cells in culture and decreased generation of intestinal RORγt+Foxp3+ pTreg cells but not the generation of tTreg cells [251]. One of the mechanisms underlying these findings could be through co-Smad4, as co-Smad4 is required for iTreg cell generation [232, 250], and the function of co-Smad4 is suppressed by i-Smad7 [252], SKI, and SnoN [67, 253]. The notion that TGF-β functions differently in tTreg, pTreg and iTreg cells is further supported by the findings that CNS1 is critical for iTreg and pTreg cell generation but can be compensated for by CNS3 in tTreg cells [146]. It was also reported that mice lacking CNS1 generated tTreg cells normally but generated fewer pTreg cells in the intestines [254]. These findings suggest that the generation of tTreg cells and pTreg and iTreg cells utilize distinct mechanisms for Foxp3 expression: TGF-β signaling is important for pTreg and iTreg cells but is much less important for tTreg cell generation under noninflammatory, homeostatic conditions. While TGF-β1 is the predominant cytokine that can promote iTreg cell generation, Activin A, another TGF-β superfamily member, can also promote iTreg cell differentiation in culture by synergizing with TGF-β1 but not on its own, indicating an interaction between TGF-β and Activin A in controlling Treg cell function. Activin A does so by promoting Smad and p38 MAPK signaling initiated by TGF-β [255, 256]. How important Activin A is in the biology of tTreg and pTreg cells in vivo remains to be addressed. In addition, whether other members of the TGF-β superfamily are also involved in Treg cell generation needs to be elucidated since their signaling can impact co-Smad4 function, which is critically involved in iTreg cell generation.
Involvement of TGF-β in Treg cell function
The role for TGF-β in regulating Treg cell function has been under debate. The first connection between TGF-β and Treg cell suppressive function was from a study using a T-cell transfer-induced colitis mouse model. In this study, the suppressive function of Treg cells was found to be abrogated by anti-TGF-β1 antibodies [257]. In addition, it was found that Treg cells mediate their suppressive function through surface-bound TGF-β1 in culture [258]. Since then, various TGF-β- and TGFβR-deficient (TGF-β1−/−, TGFβRI−/−, and TGFβRII−/−) mouse models have been generated to investigate the relationship between Treg cell function and TGF-β. TGF-β1 was found to be indispensable for Treg cell-mediated function [240]. TGF-β1−/− Τreg cells failed to suppress the colitis induced by cotransferred naive wild-type CD4+ T cells, which displayed enhanced Th1 cell differentiation in recipient mice [240]. However, it was later found that Treg cell-specific knockout of TGF-β1 did not lead to systemic inflammation and had a negligible effect on immune homeostasis and EAE development [239]. These findings suggest that while Treg cell-produced TGF-β is dispensable for immune homeostasis under steady-state conditions, Treg cell-produced TGF-β may become important under certain inflammatory conditions. Of interest, a study where TGFβRI is specifically deleted in Treg cells revealed that TGF-β signaling is important for certain functions of Treg cells [237]: TGFβRI deletion in Treg cells leads to reduced T-bet but increased RORγt expression and showed an increased ability to suppress Th1 cells but reduced control of Th17 cells in aged mouse lungs and GI tracts and during EAE in young mice. In addition, TGFβRI is critical for Treg cell recruitment and retention in the gastrointestinal tract. TGFβRI deletion in Treg cells leads to reduced expression of the tissue residential protein CD103 and colon chemotaxis protein GPR15 and increased expression of GPR174, a LysoPS (lysophosphatidylserine) receptor that negatively regulates Treg cell homeostasis [259], limiting Treg cell accumulation in the intestines [237]. This study reveals a tissue-specific role of TGF-β signaling in controlling Treg cell function, emphasizing how TGF-β may control Treg cells in a microenvironment-dependent manner. While TGF-β promotes the expression of amphiregulin in fibroblasts [260], whether and how TGF-β signaling is indeed involved in Treg cell-mediated tissue repair remains to be addressed.
Collectively, available evidence suggests that TGF-β and related signals are not always essential for Treg cell generation, homeostasis, or function under all circumstances, as initially thought. Nonetheless, TGF-β and related signals are indeed required for iTreg cell differentiation in culture and pTreg cell generation in vivo, and such a requirement becomes inflammation-dependent for tTreg cells. Therefore, TGF-β and related signals are critical for Treg cells in a cell-type- and context-dependent manner (Fig. 2). Much study is needed to fully understand how TGF-β signaling controls the generation and function of various Treg cell subsets through genetic and epigenetic mechanisms at the genomic and proteomic levels with or without crosstalk with other factors that sense complex environmental cues.
Th17 cells are broadly involved in immune pathogenicity and regulation
In the 1980s, Mosmann and Coffman’s work established a paradigm that, in response to different cytokines, naive CD4+ T cells may differentiate de novo into functionally distinct effector Th cells, namely, IFN-γ-producing Th1 cells and IL-4-producing Th2 cells, to regulate different immune responses [261]. A later study found that the p40 subunit of IL-12, a Th1 cell differentiating factor, not only pairs with the p35 subunit to form IL-12 but also pairs with the p19 subunit to form IL-23 [262]. Therefore, observations made by targeting p40 may not be entirely attributed to the function of IL-12 and thus Th1 cells. Indeed, it was later found that IL-23 but not IL-12 is crucial for the induction of autoimmune EAE (experimental autoimmune encephalomyelitis) [263]. Subsequent studies revealed that IL-23, but not IL-12, promoted an IL-17-producing T-cell population, now called Th17 cells, in a p19-dependent manner [264,265,266]. Th17 cells are distinct from Th1 cells.
Th17 cell classification, generation, and maintenance
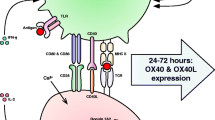
Th17 cells have diverse, sometimes opposite, functions. Th17 cells promote inflammation and autoimmune diseases, clear pathogens, and maintain barrier function of the mucosa [267, 268]. Therefore, Th17 cells can be both pathogenic and nonpathogenic. The dichotomous function of Th17 cells appears to be dictated by the microenvironment in which they reside. Nonpathogenic Th17 cells accumulate in the intestines under homeostasis to maintain mucosal integrity. Pathogenic Th17 cells can accumulate in the skin, nervous system, and skeleton during immune pathologies, including psoriasis, MS (multiple sclerosis), RA (rheumatoid arthritis), and AS (ankylosing spondylitis) [269]. In addition, intestinal nonpathogenic Th17 cells have been shown to be stem cell-like precursors for pathogenic Th17 cells in the spinal cord that cause EAE [270]. Therefore, the generation and maintenance of stem cell-like, nonpathogenic Th17 cells in vivo appears to depend on the microenvironment in the intestines, without which Th17 cells differentiate into pathogenic Th17 cells by default. The functional dichotomy of Th17 cells can also be observed in Th17 cells differentiated in culture. The TGF-β1 + IL-6 cytokine combination induces IL-10-producing Th17 cells with less pathogenic function [271]. IL-1β + IL-6 + IL-23, TGF-β + IL-6 + IL-23, and Activin-A + IL-6 cytokine combinations induce Th17 cells of potent pathogenic function with much less IL-10 production [269, 272, 273]. In agreement, nonpathogenic Th17 cells and pathogenic Th17 cells have different gene expression patterns. Pathogenic Th17 cells express proinflammatory genes, including Tbx21, Csf2, Il18r1, Il22, Il23r, Il33, and Cxcl3. In contrast, nonpathogenic Th17 cells express high levels of immunomodulatory genes, including Il4, Cd5l, Il9, Il10, Ccl20, Ahr, and Maf [121, 269, 274, 275] (Fig. 3 A). Interestingly, IL-23 can convert nonpathogenic Th17 cells into pathogenic Th17 cells [274]. Therefore, the first-identified Th17-driven cytokine, IL-23, is not only important in promoting Th17 cell proliferation and survival [276] but also in endowing Th17 cells with pathogenic function. Single-cell RNA-seq analysis of Th17 cells isolated from EAE-diseased mice revealed that Il10 and Cd5l are nonpathogenic Th17 cell markers that are coexpressed with proinflammatory genes, including Gpr65, Toso, and Plzp [277, 278]. It therefore appears that nonpathogenic Th17 cells have “dual intentions”, agreeing with the observation that these cells have the properties of stem cells and precursors [270]. In addition, pathogenic and nonpathogenic Th17 cells can be discriminated in EAE by GM-CSF+IFN-γ+CXCR6+IL-17+ and TCF1+SLAMF6+IL-17+, respectively [270].
TGF-β superfamily signaling controls the biology of Th17 cells with broad functions in health and disease. A Upon culturing in the presence of different cytokine combinations, activated CD4+ T cells can be differentiated into Th17 (pTh17) cells of high pathogenicity and Th17 (cTh17) cells of low pathogenicity. cTh17 and pTh17 cells bear different molecular signatures. B By integrating various signaling pathways, including TCR, interleukins, and AHR, the TGF-β superfamily members TGF-β and Activin A control RORγt expression and function as well as Il17 expression through Smad and interacting proteins during Th17 cell differentiation. Differentiated Th17 cells regulate immunity, autoimmunity, cancer, and homeostasis through the secretion of cytokines
Cytokines are critical to control, both positively and negatively, Th17 cell differentiation (Fig. 3B). IFN-γ and IL-4 are dispensable for Th17 cell differentiation. Instead, they inhibit the development of Th17 cells [265, 266]. The Th1 cell-polarizing cytokine IL-12 rapidly and irreversibly shuts down the Il17a/f locus in already-differentiated Th17 cells [275]. TGF-β can promote Foxp3 expression and thus Treg cell generation to suppress Th17 cell generation [250, 279, 280]. Nonetheless, TGF-β promotes Th17 cells in the absence of Foxp3 in vivo [186]. IL-6 and IL-21 promote Th17 cell differentiation through STAT3 [268, 281, 282]. IL-21’s function appears complex and context-dependent because when IL-21 signaling was blocked, the generation and function of Th17 cells were unaltered [283,284,285] during EAE but were reduced in gut inflammation [286]. Although IL-23 is not the differentiation factor for Th17 cells, productive and sustained Th17 cell responses only develop in the presence of IL-23, as revealed by studies with Il23p19−/− and Il23r−/− mice [263]. After the initial induction of Th17 cells, the availability of IL-23 becomes the limiting factor that determines whether Th17 cells, especially pathogenic Th17 cells, are sustained during the inflammatory response [287].
Combined signaling of TGF-β and IL-6/STAT3 and IL-21/STAT3 drives Th17 cell differentiation by promoting the expression of the transcription factors RORγt and RORα [181, 288,289,290,291,292,293], which in turn controls Th17 cell differentiation through shared and distinct mechanisms [294]. RORα and RORγt bind to the RORE (ROR response element) in the Il17a/f gene loci [291, 295] to directly promote IL-17 expression. STAT3 also binds directly to and transactivates the Il17 and Il21 promoters [296, 297]. Therefore, STAT3 and RORγt cooperate for Th17 cell generation. Some loci targeted by STAT3 in Th17 cells are also the targets of STAT5, a signal transducer of IL-2 that inhibits Th17 cell differentiation [298]. On these genetic loci, STAT3 promotes permissive histone modifications, yet STAT5 promotes repressive histone modifications.
In addition to RORs, other transcription factors are also important for controlling Th17 cell differentiation. IRF4, which is associated with the differentiation of the Th1 and Th2 cell subsets [299,300,301], is required for the differentiation of Th17 cells through RORγt-dependent and RORγt-independent mechanisms [302]. ETS1, the prototype member of the Ets family of transcription factors, inhibits Th17 cell differentiation by interacting with IL-2/STAT5 signaling because ETS1 deletion leads to increased Th17 cell differentiation with reduced IL-2 production and STAT5 signaling [303]. BATF (basic leucine zipper ATF-like transcription factor) and IRF4 form a complex to increase chromatin accessibility [304]. STAT3 then starts a transcriptional program that is eventually turned on by RORγt for Th17 cell differentiation [304]. Fosl2 limits the plasticity of Th17 cells [304]. c-Maf is importantly involved in Th17 cell generation in a context-dependent manner. c-Maf was found to function in a negative feedback loop to limit Th17 cell differentiation, where it is induced by both STAT3 and IRF4 and represses BATF [304, 305]. c-Maf is also critical for the maintenance, expansion, and function of differentiated Th17 cells by promoting the production of IL-21 [306] and RORγt [307].
Molecules other than cytokines also regulate Th17 cell differentiation. The host defense peptide cathelicidin promotes Th17 cell generation by enhancing AHR (aryl hydrocarbon receptor) and RORγt expression in a TGF-β1-dependent manner [308]. Although AHR activated by 2,3,7,8-tetrachlorodibenzo-p-dioxin induces Treg cells, AHR activated by 6-formylindolo[3,2-b]carbazole promotes Th17 cells to contribute to EAE [129, 244, 309]. Oxygen sensing appears important for Th17 cell differentiation because HIF-1 (hypoxia-inducible factor 1), a key metabolic sensor of oxygen, promotes Th17 cell differentiation through the direct transactivation of Rorc [310, 311]. Microbial and metabolic products, including retinoic acid, short-chain fatty acids, and bile acids, can also regulate Th17 cell differentiation [120,121,122,123,124,125,126,127,128, 147, 312]. Acetyl-CoA carboxylase, a key enzyme of de novo fatty acid synthesis, influences the Th17 and Treg cell balance through the glycolytic and lipogenic pathways [313]. CD5L, a signature marker of nonpathogenic Th17 cells, regulates lipidome saturation to restrain Th17 cell pathogenicity [278]. The active form of vitamin D (1,25-dihydroxyvitamin D3), the main ligand for the vitamin D receptor, has been found to “severely impair” the production of IL-17 and IL-17F by Th17 cells [314]. How cytokines and noncytokines deploy shared and distinct mechanisms to regulate Th17 cell differentiation is an important, albeit complex, question to be fully elucidated.
All-encompassing Th17 cell function in health and disease
Contributions to autoimmune and inflammatory disease
Pathogenic Th17 cells contribute broadly to autoimmune and inflammatory diseases [268]. Th17 cells are important in causing the immune pathologies of diseases, including psoriasis [315], RA [316], MS [317], IBD (inflammatory bowel disease) [318], asthma [319, 320], Graves’ disease [321], transplant rejection [322, 323], and allergy [324]. T cells in human psoriatic skin lesions predominantly show a Th17 cell phenotype with high CCR6 expression [325]. This is in line with the observation that CCL20/CCR6 signaling is important for the chemoattraction of inflammatory cells to inflammatory tissues, including the skin. In RA patients, the expression of TNF, IL-1, and IL-17 is predictive of joint destruction [316]. Th17 cell-produced RANKL promotes RA by inducing osteoclastogenesis [326,327,328], which in turn promotes cartilage and bone destruction and resorption independently of TNF and IL-1 [329, 330]. In MS, IL-17 and IL-6 are among the most highly produced cytokines [317, 331]. In opticospinal forms of MS, IL-17 and CXCL8 (IL-8), a target of IL-17 and a strong neutrophil chemoattractant, are elevated and positively associated with spinal lesions [332]. Th17 cells effectively transmigrate across the blood‒brain barrier (BBB) and infiltrate into the CNS parenchyma through IL-17- and IL-22-mediated disruption of BBB tight junctions [333].
Promotion of pathogen clearance
Th17 cells play important roles in the pathogen clearance response, especially when type-1 and type-2 immunity are imbalanced [334]. Th17 cells traffic to the sites of infection due to high CCR6 expression. Diverse pathogens, including viruses, bacteria, and fungi-like microbes, can induce strong Th17 responses [335,336,337,338,339,340,341,342,343,344,345,346,347,348,349,350,351,352]. In humans, Th17 cells can recruit B cells through CXCL13 and promote antibody production by B cells for pathogen clearance [353,354,355,356].
Maintenance of tissue and immune homeostasis
In addition to being pathogenic for tissue damage, Th17 cells also promote tissue repair and homeostasis. Such an effect of Th17 cells can be mediated by IL-22, a member of the IL-10 family of cytokines [357], by promoting the regeneration of epithelial tissues [358]. During inflammation, Th17 cells migrate to the sites of inflammation via CCR6 [359, 360] and upregulate IL-22 [361] to limit liver damage in concanavalin A-induced hepatitis [362] and intestinal damage in T-cell-induced IBD [363]. Of note, IL-22 has also been found to be a pathogenic cytokine in psoriasis [364, 365] due to the IL-22-promoted excessive regeneration of epithelial cells. Thus, IL-22-producing Th17 cells may function in a cell-type- and context-dependent manner for tissue repair and homeostasis. Th17 cells, especially nonpathogenic Th17 cells, are important for establishing tolerance to commensal microbiota for homeostasis by coordinating with Treg cell functions [303]. In addition, the tissue repair function of Th17 cells will help maintain the barrier integrity of the mucosa to prevent microbes from translocating and the barriers from being breached [366,367,368,369,370,371,372]. Thus, Th17 cells in the mucosa are important for homeostasis by both preventing invasion of microbiota and promoting epithelial barrier integrity.
Participation in tumor immunity
Th17 cells and their cytokines are also involved in tumor development and cancer [373,374,375,376,377]. Th17 cells can differentiate into Th1-like Th17 cells (secreting both IFN-γ and IL-17) with stem cell-like properties to reject tumors [378, 379]. Compared with other Th cells, Th17 cells are long-lived with increased self-renewal ability. Such properties of Th17 cells seem to be due to their unique metabolic programming. Th17 cells utilize mitochondrial oxidative phosphorylation, which protects Th17 cells from apoptosis while enhancing their persistence in the periphery and TME [380]. Of note, Th17 cells have also been found to promote tumor formation induced by colonic inflammation in mice [381]. Therefore, the functions of Th17 cells in tumor development are complex and remain to be further characterized [382].
Th17 cells have both pathogenic and nonpathogenic properties. This allows Th17 cells to play broad and diverse roles in health and disease (Fig. 3B). The molecular mechanisms underlying the generation and function of Th17 cells have been under intensive investigation since their discovery. We now know that the TGF-β superfamily plays essential and discrete roles in controlling the biology of both pathogenic and nonpathogenic Th17 cells. In the following section, we will discuss in detail how TGF-β signaling pathways control Th17 cell function.
TGF-β superfamily members deploy shared and unique mechanisms to control Th17 cell differentiation and function
Importance of TGF-β in Th17 cell generation
During early studies defining Th17 cells as a new subset of Th cells that are proinflammatory and pathogenic and cause autoimmune neuroinflammation, TGF-β was not implicated in Th17 cell biology. Surprisingly, a seminal study demonstrated that TGF-β (a well-known immunosuppressive cytokine), in combination with IL-6, potently promoted IL-17 production and therefore Th17 cell differentiation under culture conditions [383,384,385]. This study demonstrated that TGF-β can promote inflammation by inducing IL-17 production [384]. The reconciliation of the seemingly contradictory roles of TGF-β in promoting both iTreg and Th17 cell differentiation in culture came from a later study showing that TGF-β balances Th17 and iTreg cell differentiation in a dose- and cytokine-milieu-dependent manner. In the lamina propria and during T-cell activation in the presence of TGF-β, Foxp3 and RORγt are coexpressed in CD4+ T cells and continuously counterbalance each other [279]. At low concentrations, TGF-β synergizes with IL-6 and IL-21 to favor Th17 differentiation by promoting IL-23R expression. However, high concentrations of TGF-β repress the expression of Th17 signatures, including IL-23R and IL-22, but promote the expression of Foxp3, which inhibits the activity of RORγt to favor iTreg differentiation [279]. In addition, Foxp3 antagonizes Th17 cell differentiation by inhibiting RORγt and RORα [250] and by interacting with Runx1 to suppress the Il17 locus [280]. In agreement with the notion that TGF-β stimulation is compatible with Th17 cell differentiation, it was also found that TGF-β induces and maintains the expression of IL-6Rα, whose signaling suppresses Foxp3 expression and activates STAT3 to induce RORγt expression and Th17 cell differentiation [268, 384]. These findings suggest that while high concentrations of TGF-β promote Foxp3 expression to restrict the Th17 cell program, TGF-β nonetheless permits Th17 cell differentiation when IL-6 and IL21 are present. IL-6 and IL21 promote Th17 cell differentiation by enhancing the Th17 program and antagonizing the Treg cell program [386].
Diverse functions of TGF-β signaling pathways in Th17 cell generation
Much effort has been devoted to understanding the molecular mechanisms through which TGF-β promotes Th17 cell differentiation. One study found that, in a dose-dependent manner, TGF-β suppresses the IL-6- and IL-21-promoted expression of SOCS3 (suppressor of cytokine signaling 3), a negative feedback inhibitor of STAT3. Interfering with TGFβR function with a dominant-negative form of TGFβII or the pharmacological TGFβRI inhibitor SB505124 leads to increased IL-6-induced SOCS3 expression and reduced STAT3 activation and thus fewer Th17 cells [387].
While TGF-β signaling is important for Th17 cell differentiation, the roles of the highly homologous r-Smad2 and r-Smad3 in Th17 cell differentiation seem distinct. r-Smad2 was found to positively regulate IL-17 expression by interacting with RORγt without affecting Rorc expression and to be required for Th17 cell differentiation [247]. T-cell-specific r-Smad2 knockout mice had reduced EAE with decreased Th17 cells [247]. In agreement, another study showed that Smad2 is important for Th17 cell generation, partly by regulating IL-6R expression [388]. Although closely related to r-Smad2, r-Smad3 was found to suppress Th17 cell differentiation by interfering with RORγt transcriptional activity [246]. r-Smad3 deletion led to increased RORγt expression and increased Th17 cell differentiation both in culture and in vivo, suggesting that r-Smad3 suppresses Th17 cell differentiation by interfering with the expression and function of RORγt [246]. Further study revealed additional mechanisms underlying the different roles of r-Smad2 and r-Smad3 in controlling Th17 cell differentiation. While r-Smad2 can be phosphorylated by ERK at the linker region and can associate with STAT3 and p300 as a coactivator to promote RORγt function, unphosphorylated r-Smad3 interacts with STAT3 and PIAS3 (protein inhibitor of activated STAT3) to repress Rorc and IL17a gene expression [389]. In addition, r-Smad2 is associated with TRIM33, a factor required for Th17 cell differentiation. TRIM33-deficient T cells developed less severe EAE [390]. TRIM33 associates with both Il17a and Il10 loci in the presence of r-Smad2 to promote Il17 expression and to suppress Il10 expression, therefore facilitating pathogenic Th17 cell generation [390]. The Th17 cell-promoting function of r-Smad2 appears to dominate r-Smad3’s suppressive function because knocking out both r-Smad2 and r-Smad3 results in less Th17 cell differentiation without affecting RORγt expression [249]. These observations were, however, disputed by a study showing that neither r-Smad2 nor r-Smad3 alone is required for Th17 cell differentiation in culture and in EAE [391]. In addition, the combination of r-Smad2 deletion and r-Smad3 inhibition by a pharmacological inhibitor did not significantly affect Th17 cell differentiation [391]. Instead, Th17 cell differentiation mostly depends on the JNK and p38 pathways [391]. The reason for the discrepancies is unclear but could be due to the different mouse strains and experimental approaches used in these studies. Nonetheless, these findings suggest that the roles for r-Smad proteins in Th17 cell differentiation could be nuanced and complex. Multiple mechanisms are used by Smad proteins to control RORγt and IL-17 expression at both the protein and gene levels. The crosstalk between the TGF-β/r-Smad and MAPK signaling pathways is important for Treg and Th17 cell differentiation. TCR stimulation-induced MEKK2/MEKK3 and ERK activation leads to the phosphorylation of the linker regions of r-Smad2 and r-Smad3. Such phosphorylation suppressed the transactivation function of r-Smad2 and r-Smad3 in response to TGF-β stimulation [392]. Deletion of both MEKK2 and MEKK3 leads to an enhancement of TGF-β-promoted Treg and Th17 cell differentiation in vitro and in vivo [392]. Interestingly, MEKK2 and MEKK3 double knockout mice developed more severe EAE than wild-type mice, suggesting that MEKK2 and MEKK3 are preferentially required to dampen the TGF-β-controlled Th17 program [392]. The function of co-Smad4 in T cells was initially perplexing. In contrast to what was predicted for a protein that is central to TGFβR signaling, T-cell-specific depletion of co-Smad4 did not yield autoimmune symptoms or apparent T-cell activation as in T-cell-specific TGFβR knockout mice [250, 393]. Instead, T-cell-specific co-Smad4 deletion led to spontaneous development of cancer with increased Th17 cell differentiation [250, 393], suggesting that Smad4 has functions beyond promoting TGF-β signaling in T cells. A later study found that co-Smad4 depletion rescued lethal autoimmune disease in T-cell-specific TGFβRΙΙ-deficient mice [232], suggesting that co-Smad4 counterbalances TGFβR signaling. Indeed, closer examination revealed that co-Smad4-deficient T cells readily differentiated into Th17 cells in the presence of IL-6 and IL-21 in culture even when TGF-β signaling was abrogated, although TGF-β + IL-6-promoted Th17 cell differentiation was not obviously affected [250, 253, 394, 395]. In addition, during EAE development, although TGFβRII-deficient T cells fail to differentiate into Th17 cells, simultaneous knockout of co-Smad4 fully restores Th17 cell differentiation [253]. These findings suggest that co-Smad4 restrains Th17 cell differentiation in the absence of TGF-β stimulation. Without TGF-β, SKI is critical for the co-Smad4-mediated effects by associating with co-Smad4 on the Rorc locus but not the Il17 locus and recruiting HDAC to the Rorc locus to restrain Rorc expression. Because SKI is very sensitive to TGF-β-induced protein degradation [396,397,398], low concentrations of TGF-β trigger SKI degradation and thus alleviate SKI/co-Smad4-complex-mediated suppression of Rorc expression [253]. Therefore, an important mechanism through which TGF-β promotes Th17 cell differentiation is to disrupt the SKI/co-Smad4 suppressive complex to allow Rorc expression and Th17 cell differentiation. Such a function of co-Smad4 appears to be context dependent. While co-Smad4 is indeed required to suppress Th17 cell differentiation under normal conditions, co-Smad4 is also required to promote pathogenic Th17 cell differentiation under febrile temperature [395]. High temperature promotes the heat shock response and the sumoylation of co-Smad4 and its nuclear translocation. Interestingly, febrile temperature induced co-Smad4 binding to the Il17 loci [395]. These findings suggest that Smad proteins control Th17 cell differentiation in a context-dependent manner by integrating various environmental cues to positively and negatively regulate Th17 cell differentiation. Protein‒protein interactions have emerged as an important way to control Smad function. It would be of interest to further investigate how the interactomes of r-Smad2, r-Smad3, and co-Smad4 are dynamically regulated during Th17 cell differentiation under various conditions to identify critical factors for Th17 cell differentiation.
Distinct roles for TGF-β superfamily members in regulating pathogenic and nonpathogenic functions of Th17 cells
Much effort has been devoted to understanding how TGF-β1 + IL-6 induces Th17 cells with low pathogenic function. Recently, we have seen increasing interest in understanding how pathogenic Th17 cell function is controlled due to its critical role in causing pathology [273, 274, 399,400,401,402,403,404,405]. TGF-β1 + IL-6-induced Th17 cells upregulate TGF-β3 in response to IL-23 to promote the pathogenic program, suggesting that TGF-β3 signaling can endow and enhance the pathogenic program of differentiating and differentiated Th17 cells [274]. However, whether TGF-β signaling is indeed involved in the de novo generation of pathogenic Th17 cells remains uncertain. In fact, a study suggests that TGF-β signaling is dispensable for the generation of pathogenic Th17 cells, particularly those differentiated by IL-6 + IL-1β + IL-23 [272]. Nonetheless, SKI degradation, a particularly sensitive readout for TGF-β signaling to allow Th17 cell differentiation [253], occurred under IL-6 + IL-1β + IL-23-polarizing conditions. Such SKI degradation was found to not be due to TGF-β signaling but rather to Activin A, a TGF-β superfamily member that is highly induced by IL-6 + IL-1β + IL-23. Activin A + IL-6 was shown to be sufficient to drive the differentiation of pathogenic Th17 cells that resemble IL-6 + IL-1β + IL-23-induced pathogenic Th17 cells. In addition, Activin A and its specific receptor I (ALK4) are critical for the generation of pathogenic Th17 cells to induce EAE. Furthermore, while TGF-β/ALK-5 signaling potently suppresses ERK activation, which is important for the pathogenic program of Th17 cells, Activin A/ALK4 signaling does not after ERK activation [406]. Therefore, different TGF-β superfamily members contribute to the generation and reinforcement of nonpathogenic and pathogenic Th17 cells through distinct and shared mechanisms. SKI suppresses the differentiation of both nonpathogenic and pathogenic Th17 cells. Ectopic SKI expression inhibits the generation of Th17 cells, especially pathogenic Th17 cells, both in culture and in vivo during EAE development [407]. SKI controls Th17 cell differentiation in a dose-dependent manner. Moderate stabilization of SKI in Arkadia-deficient T cells did not lead to substantial inhibition of Th17 cell differentiation [251], as certain levels of SKI expression can be tolerated for Th17 cell differentiation [406]. It is also possible that Arkadia knockout results in more complex effects, including the stabilization of the SnoN and Smad proteins and other uncharacterized targets that may affect co-Smad4 function, rather than only stabilizing the SKI protein [251].
Localized TGF-β production for Th17 cell generation and function
While both TGF-β and Activin A can be produced by various cell types, T-cell-generated TGF-β and Activin A play nonredundant roles in Th17 cell function. Both TGF-β1 and Activin A produced by Th17 effector T cells are important for Th17 cells [239, 406]. In agreement, TGF-β1 is required for Th17 cell stability and for maintaining the nonpathogenic program. A study using IL-17-producing cell-specific TGFβ1 knockout and fate-mapping systems (Tgfb1fl/flIl17aCreR26YFP) revealed that autocrine TGFβ1 in Th17 cells maintains their stability by repressing the expression of IL-12Rβ2 and IL-27Rα [408]. TGF-β1-deficient Th17 cells tend to produce IFN-γ and become more pathogenic, exacerbating tissue inflammation in an adoptive EAE transfer model [408].
The aforementioned studies highlight the important functions of different TGF-β superfamily members and their signaling in controlling Th17 cell generation and function. However, whether Th17 cells can possibly be generated independent of TGF-β signaling remains incompletely answered. In mice doubly deficient in STAT6 and T-bet, Th17 cells are readily induced by IL-6 when TGFβRII signaling is attenuated by the expression of a dominant negative form of TGFβRII [409]. The purported mechanism was that failed Th1 and Th2 cell differentiation of STAT4/T-bet-double-knockout CD4+ T cells led to Th17 cell differentiation by default in the presence of IL-6. This posited mechanism is questionable because blocking Th1 and Th2 cell differentiation by other means does not result in Th17 cell differentiation by IL-6 stimulation alone. It would be interesting to assess whether Activin A-related signaling contributes to these observations. The abovementioned studies also highlight that Th17 cells are functionally heterogeneous with varying cytokine production profiles depending on the microenvironment in which they reside. Such high heterogeneity of Th17 cells could be due to different differentiation trajectories or functional plasticity, allowing dynamic adaptation to the changing environment. While TGF-β superfamily cytokine signaling is clearly instrumental in the generation and function of Th17 cells (Fig. 3B), the molecular underpinnings (especially the crosstalk with the pathways that sense environmental cues, including oxygen, metabolites, and chemicals) of TGF-β-controlled Th17 cell generation, maintenance, and function remain poorly defined and warrant further investigation.
The above discussion suggests that although Treg and Th17 cells are seemingly distinct cell types, their differentiation programs are related and share common pathways, especially the TGF-β pathway. In fact, conversion between Treg and Th17 cells can occur. An earlier study found that IL-6 abrogates Treg cell suppressive activity [386]. IL-6 was later shown to convert established Treg cells into Th17-like cells [250, 410]. The downregulation of Foxp3, which leads to the instability and plasticity of established Treg cells, is important to allow Treg-to-Th17 cell conversion because high levels of Foxp3 suppress RORγt to restrain Th17 cell differentiation [250, 279]. The attenuation and downregulation of Foxp3 expression are often associated with Th17 cell conversion of established Treg cells during immune pathologies [411], including type I diabetes, systemic autoimmunity, autoimmune arthritis, and neuroinflammation [149, 152, 412, 413]. Of note, while high levels of Foxp3 suppress Th17 cell differentiation [279], Foxp3 and IL-17 are not mutually exclusive, as cells expressing both can be found in vitro and in vivo [186, 250, 410]. How is TGF-β involved in Treg-Th17 conversion? Current evidence suggests that the strength of TGF-β signaling is important. High doses of TGF-β promote high levels of Foxp3 to favor Treg cell differentiation over Th17 cell differentiation even in the presence of IL-6 [253, 279]. In addition, while Activin-A promotes Th17 cell differentiation to promote inflammation [269], it does not promote Treg cell differentiation on its own [414], suggesting a potential role for Activin-A favoring Th17 cells over Treg cells. Therefore, it is reasonable to believe that strong TGF-β signaling will favor not only Treg cell generation but also their stability for immune homeostasis and suppression. Weak TGF-β and/or other TGF-β superfamily member signaling will permit Th17 cell differentiation and/or Treg-to-Th17 transdifferentiation under inflammatory conditions. What signaling molecules sense and interpret the strong vs. weak TGF-β signaling for Treg cell stability and Treg-Th17 cell transdifferentiation and how other TGF-β superfamily members are involved in Treg cell stability and Treg-Th17 cell transdifferentiation warrant further investigation to help understand the etiology and to develop treatments for immune diseases.
Concluding remarks
The central canon for TGF-β’s function in Treg and Th17 cells is to balance the immune response in a context- and microenvironment-dependent manner through complex signaling pathways and molecular mechanisms. From an operational perspective, TGF-β signaling in Treg and Th17 cells must be intricate, sometimes nuanced, to support its ability to integrate and respond to a plethora of environmental cues through dose- and context-dependent mechanisms. This ability of TGF-β signaling allows T cells to correctly interpret cellular and molecular contexts and mount defined, precise responses in a malleable way to adapt to the ever-changing microenvironment. TGF-β accomplishes these daunting tasks by wiring and rewiring cell-intrinsic pathways involving different cellular factors in varying combinations. When studying TGF-β signaling and response, cellular and molecular contexts matter. Future research efforts should focus on revealing how TGF-β signaling components control T-cell responses in cell-type- and niche-specific contexts that are meaningful for immunity and diseases.
To fully appreciate the intricate, context-dependent roles of TGF-β signaling in T-cell function and to understand some of the seemingly conflicting observations, it is important to consider the following: First, the source of TGF-βs can be diverse and redundant, as TGF-βs with similar biochemical properties can be produced by broad cell types in secreted or membrane-bound forms. Therefore, the cell types involved and their physical position relative to Treg and Th17 cells in a specific niche should be considered. Second, to signal, mature TGF-β needs to be freed from LAP, a process that involves complex regulation. Activating TGF-β involves various mechanisms, including acidification, proteases, plasmin, matrix metalloproteases, thrombospondin-1, and integrins [415, 416]. Conditions, such as inflammation, that mobilize these mechanisms will impact the availability of active TGF-β and influence the reliance and responsiveness of T cells to TGF-β to achieve balanced responses. Third, under complex conditions in the microenvironment in vivo, other TGF-β superfamily members may also be involved in TGF-β signaling in Treg and Th17 cells in a context-dependent manner. TGF-β superfamily members can cooperate through shared or unique components and pathways to mitigate or exacerbate some of the effects observed under various settings. Finally, due to broad crosstalk between TGF-β signaling pathways and other signaling pathways, transcription factors, and epigenetic regulators, the “molecular contexts” in a cell will likely substantially affect the signaling and functional outputs of TGF-β stimulation. Therefore, it is important to comprehensively understand these “molecular contexts” using multiomics approaches, ideally at the single-cell level, to fully appreciate how intricately TGF-β signaling functions in a context-dependent fashion.
References
Gordon KJ, Blobe GC. Role of transforming growth factor-β superfamily signaling pathways in human disease. Biochimica et Biophysica Acta (BBA) - Mol Basis Dis. 2008;1782:197–228. https://doi.org/10.1016/j.bbadis.2008.01.006
Poniatowski ŁA, Wojdasiewicz P, Gasik R, Szukiewicz D. Transforming growth factor Beta family: insight into the role of growth factors in regulation of fracture healing biology and potential clinical applications. Mediators Inflamm. 2015;2015:137823. https://doi.org/10.1155/2015/137823
Zhang Y, Alexander PB, Wang XF. TGF-β Family signaling in the control of cell proliferation and survival. Cold Spring Harb Perspect Biol. 9, https://doi.org/10.1101/cshperspect.a022145 (2017).
Wu MY, Hill CS. TGF-β superfamily signaling in embryonic development and homeostasis. Dev Cell. 2009;16:329–43. https://doi.org/10.1016/j.devcel.2009.02.012
Xu X, Zheng L, Yuan Q, Zhen G, Crane JL, Zhou X, et al. Transforming growth factor-β in stem cells and tissue homeostasis. Bone Res. 2018;6:2. https://doi.org/10.1038/s41413-017-0005-4
Sanjabi S, Oh SA, Li MO. Regulation of the immune response by TGF-β: from conception to autoimmunity and infection. Cold Spring Harb Perspect Biol. 9, https://doi.org/10.1101/cshperspect.a022236. (2017).
Batlle E, Massagué J. Transforming growth factor-β signaling in immunity and cancer. Immunity. 2019;50:924–40.
Morianos I, Papadopoulou G, Semitekolou M, Xanthou G. Activin-A in the regulation of immunity in health and disease. J Autoimmun. 2019;104:102314. https://doi.org/10.1016/j.jaut.2019.102314
Robson NC, Phillips DJ, McAlpine T, Shin A, Svobodova S, Toy T, et al. Activin-A: a novel dendritic cell–derived cytokine that potently attenuates CD40 ligand–specific cytokine and chemokine production. Blood. 2008;111:2733–43. https://doi.org/10.1182/blood-2007-03-080994
Jones KL, et al. Activin A is a critical component of the inflammatory response, and its binding protein, follistatin, reduces mortality in endotoxemia. Proc Natl Acad Sci. 2007;104:16239–44. https://doi.org/10.1073/pnas.0705971104
Ogawa K, Funaba M, Chen Y, Tsujimoto M. Activin A functions as a Th2 cytokine in the promotion of the alternative activation of macrophages1. J Immunol. 2006;177:6787–94. https://doi.org/10.4049/jimmunol.177.10.6787
Cortez MA, Masrorpour F, Ivan C, Zhang J, Younes AI, Lu Y, et al. Bone morphogenetic protein 7 promotes resistance to immunotherapy. Nat Commun. 2020;11:4840. https://doi.org/10.1038/s41467-020-18617-z
Zhong S, Li H, Wang YS, Wang Y, Ji G, Li HY, et al. Bmp8a is a novel player in regulation of antiviral immunity. bioRxiv, 2020.2008.2009.243519, https://doi.org/10.1101/2020.08.09.243519 (2020).
Miyazono K, Kamiya Y, Morikawa M. Bone morphogenetic protein receptors and signal transduction. J Biochem. 2009;147:35–51. https://doi.org/10.1093/jb/mvp148
Kersten C, Sivertsen EA, Hystad ME, Forfang L, Smeland EB, Myklebust JH. BMP-6 inhibits growth of mature human B cells; induction of Smad phosphorylation and upregulation of Id1. BMC Immunol. 2005;6:9. https://doi.org/10.1186/1471-2172-6-9
Martínez VG, Hernández-López C, Valencia J, Hidalgo L, Entrena A, Zapata AG, et al. The canonical BMP signaling pathway is involved in human monocyte-derived dendritic cell maturation. Immunol Cell Biol. 2011;89:610–8. https://doi.org/10.1038/icb.2010.135
Yoshioka Y, Ono M, Osaki M, Konishi I, Sakaguchi S. Differential effects of inhibition of bone morphogenic protein (BMP) signalling on T-cell activation and differentiation. Eur J Immunol. 2012;42:749–59. https://doi.org/10.1002/eji.201141702
Sanford LP, Ormsby I, Gittenberger-de Groot AC, Sariola H, Friedman R, Boivin GP, et al. TGFbeta2 knockout mice have multiple developmental defects that are non-overlapping with other TGFbeta knockout phenotypes. Development. 1997;124:2659–70.
Proetzel G, Pawlowski SA, Wiles MV, Yin M, Boivin GP, Howles PN, et al. Transforming growth factor–β3 is required for secondary palate fusion. Nat Genet. 1995;11:409–14.
Dickson MC, Martin JS, Cousins FM, Kulkarni AB, Karlsson S, Akhurst RJ. Defective haematopoiesis and vasculogenesis in transforming growth factor-beta 1 knock out mice. Development. 1995;121:1845–54. https://doi.org/10.1242/dev.121.6.1845
Shull MM, Ormsby I, Kier AB, Pawlowski S, Diebold RJ, Yin M, et al. Targeted disruption of the mouse transforming growth factor-beta 1 gene results in multifocal inflammatory disease. Nature. 1992;359:693–9. https://doi.org/10.1038/359693a0
Kulkarni AB, Huh CG, Becker D, Geiser A, Lyght M, Flanders KC, et al. Transforming growth factor beta 1 null mutation in mice causes excessive inflammatory response and early death. Proc Natl Acad Sci. 1993;90:770–4.
Christ M, McCartney-Francis NL, Kulkarni AB, Ward JM, Mizel DE, Mackall CL, et al. Immune dysregulation in TGF-beta 1-deficient mice. J Immunol. 1994;153:1936–46. https://doi.org/10.4049/jimmunol.153.5.1936
Li MO, Wan YY, Sanjabi S, Robertson AK, Flavell RA. Transforming growth factor-beta regulation of immune responses. Annu Rev Immunol. 2006;24:99–146. https://doi.org/10.1146/annurev.immunol.24.021605.090737
Ueshima E, Fujimori M, Kodama H, Felsen D, Chen J, Durack JC, et al. Macrophage-secreted TGF-β(1) contributes to fibroblast activation and ureteral stricture after ablation injury. Am J Physiol Ren Physiol. 2019;317:F52–f64. https://doi.org/10.1152/ajprenal.00260.2018
Wynn TA, Barron L. Macrophages: master regulators of inflammation and fibrosis. Semin Liver Dis. 2010;30:245–57. https://doi.org/10.1055/s-0030-1255354
Gong D, Shi W, Yi SJ, Chen H, Groffen J, Heisterkamp N. TGFβ signaling plays a critical role in promoting alternative macrophage activation. BMC Immunol. 2012;13:31. https://doi.org/10.1186/1471-2172-13-31
Speck S, Lim J, Shelake S, Matka M, Stoddard J, Farr A, et al. TGF-β signaling initiated in dendritic cells instructs suppressive effects on Th17 differentiation at the site of neuroinflammation. PLOS ONE. 2014;9:e102390. https://doi.org/10.1371/journal.pone.0102390
Lukas D, et al. TGF-β inhibitor Smad7 regulates dendritic cell-induced autoimmunity. Proc Natl Acad Sci. 2017;114:E1480–E1489. https://doi.org/10.1073/pnas.1615065114
Bobr A, Igyarto BZ, Haley KM, Li MO, Flavell RA, Kaplan DH. Autocrine/paracrine TGF-β1 inhibits Langerhans cell migration. Proc Natl Acad Sci USA. 2012;109:10492–7. https://doi.org/10.1073/pnas.1119178109
Cazac BB, Roes J. TGF-β receptor controls B cell responsiveness and induction of IgA in vivo. Immunity. 2000;13:443–51. https://doi.org/10.1016/S1074-7613(00)00044-3
Bjarnadóttir K, Benkhoucha M, Merkler D, Weber MS, Payne NL, Bernard C, et al. B cell-derived transforming growth factor-β1 expression limits the induction phase of autoimmune neuroinflammation. Sci Rep. 2016;6:34594. https://doi.org/10.1038/srep34594
Liu Y, Zhang P, Li J, Kulkarni AB, Perruche S, Chen W. A critical function for TGF-β signaling in the development of natural CD4 + CD25+Foxp3+ regulatory T cells. Nat Immunol. 2008;9:632–40. https://doi.org/10.1038/ni.1607
Marie JC, Liggitt D, Rudensky AY. Cellular mechanisms of fatal early-onset autoimmunity in mice with the T cell-specific targeting of transforming growth factor-β receptor. Immunity. 2006;25:441–54. https://doi.org/10.1016/j.immuni.2006.07.012
Li MO, Sanjabi S, Flavell RA. Transforming growth factor-β controls development, homeostasis, and tolerance of T cells by regulatory T cell-dependent and-independent mechanisms. Immunity. 2006;25:455–71.
Miyazono K, Olofsson A, Colosetti P, Heldin C-H. A role of the latent TGF‐beta 1‐binding protein in the assembly and secretion of TGF‐beta 1. EMBO J. 1991;10:1091–101.
Derynck R, Budi EH. Specificity, versatility, and control of TGF-beta family signaling. Sci Signal. 2019;12:eaav5183. https://doi.org/10.1126/scisignal.aav5183
Wang R, Zhu J, Dong X, Shi M, Lu C, Springer TA. GARP regulates the bioavailability and activation of TGFβ. Mol Biol Cell. 2012;23:1129–39. https://doi.org/10.1091/mbc.e11-12-1018
Robertson IB, Rifkin DB. Regulation of the bioavailability of TGF-beta and TGF-beta-related proteins. Cold Spring Harb Perspect Biol. 8, https://doi.org/10.1101/cshperspect.a021907 (2016).
Schmierer B, Hill CS. TGFβ–SMAD signal transduction: molecular specificity and functional flexibility. Nat Rev Mol Cell Biol. 2007;8:970–82. https://doi.org/10.1038/nrm2297
Wrighton KH, Lin X, Feng X-H. Phospho-control of TGF-β superfamily signaling. Cell Res. 2009;19:8–20. https://doi.org/10.1038/cr.2008.327
Derynck R, Zhang YE. Smad-dependent and Smad-independent pathways in TGF-β family signalling. Nature. 2003;425:577–84. https://doi.org/10.1038/nature02006
Wu G, Chen YG, Ozdamar B, Gyuricza CA, Chong PA, Wrana JL, et al. Structural basis of Smad2 recognition by the Smad anchor for receptor activation. Science. 2000;287:92–97. https://doi.org/10.1126/science.287.5450.92
Tang WB, Ling GH, Sun L, Liu FY. Smad anchor for receptor activation (SARA) in TGF-beta signaling. Front Biosci (Elite Ed). 2010;2:857–60. https://doi.org/10.2741/e147
He W, Dorn DC, Erdjument-Bromage H, Tempst P, Moore MA, Massagué J. Hematopoiesis controlled by distinct TIF1γ and Smad4 branches of the TGFβ pathway. Cell. 2006;125:929–41. https://doi.org/10.1016/j.cell.2006.03.045
Goumans MJ, Valdimarsdottir G, Itoh S, Rosendahl A, Sideras P, ten Dijke P. Balancing the activation state of the endothelium via two distinct TGF-beta type I receptors. EMBO J. 2002;21:1743–53. https://doi.org/10.1093/emboj/21.7.1743
Ramachandran A, Vizán P, Das D, Chakravarty P, Vogt J, Rogers KW, et al. TGF-beta uses a novel mode of receptor activation to phosphorylate SMAD1/5 and induce epithelial-to-mesenchymal transition. Elife. 7, https://doi.org/10.7554/eLife.31756 (2018).
Attisano L, Wrana JL. Signal transduction by the TGF-beta superfamily. Science. 2002;296:1646–7. https://doi.org/10.1126/science.1071809
Zhang Y, Feng XH, Derynck R. Smad3 and Smad4 cooperate with c-Jun/c-Fos to mediate TGF-beta-induced transcription. Nature. 1998;394:909–13. https://doi.org/10.1038/29814
Nishita M, Hashimoto MK, Ogata S, Laurent MN, Ueno N, Shibuya H, et al. Interaction between Wnt and TGF-β signalling pathways during formation of Spemann’s organizer. Nature. 2000;403:781–5. https://doi.org/10.1038/35001602
Blokzijl A, Dahlqvist C, Reissmann E, Falk A, Moliner A, Lendahl U, et al. Cross-talk between the Notch and TGF-β signaling pathways mediated by interaction of the Notch intracellular domain with Smad3. J Cell Biol. 2003;163:723–8. https://doi.org/10.1083/jcb.200305112
Nye MD, Almada LL, Fernandez-Barrena MG, Marks DL, Elsawa SF, Vrabel A, et al. The transcription factor GLI1 interacts with SMAD proteins to modulate transforming growth factor β-induced gene expression in a p300/CREB-binding protein-associated factor (PCAF)-dependent manner *. J Biol Chem. 2014;289:15495–506. https://doi.org/10.1074/jbc.M113.545194
Xu P, Lin X, Feng X-H. Posttranslational regulation of Smads. Cold Spring Harb Perspect Biol. 2016;8:a022087.
Wrighton KH, Feng X-H. To (TGF)β or not to (TGF)β: Fine-tuning of Smad signaling via post-translational modifications. Cell Signal. 2008;20:1579–91. https://doi.org/10.1016/j.cellsig.2008.02.003
Mori S, Matsuzaki K, Yoshida K, Furukawa F, Tahashi Y, Yamagata H, et al. TGF-β and HGF transmit the signals through JNK-dependent Smad2/3 phosphorylation at the linker regions. Oncogene. 2004;23:7416–29. https://doi.org/10.1038/sj.onc.1207981
Lin X, Duan X, Liang YY, Su Y, Wrighton KH, Long J, et al. PPM1A functions as a Smad phosphatase to terminate TGFbeta signaling. Cell. 2006;125:915–28. https://doi.org/10.1016/j.cell.2006.03.044
Guo X, Waddell DS, Wang W, Wang Z, Liberati NT, Yong S, et al. Ligand-dependent ubiquitination of Smad3 is regulated by casein kinase 1 gamma 2, an inhibitor of TGF-beta signaling. Oncogene. 2008;27:7235–47. https://doi.org/10.1038/onc.2008.337
Kamaraju AK, Roberts AB. Role of Rho/ROCK and p38 MAP kinase pathways in transforming growth factor-beta-mediated Smad-dependent growth inhibition of human breast carcinoma cells in vivo. J Biol Chem. 2005;280:1024–36. https://doi.org/10.1074/jbc.M403960200
Mori S, Matsuzaki K, Yoshida K, Furukawa F, Tahashi Y, Yamagata H, et al. TGF-beta and HGF transmit the signals through JNK-dependent Smad2/3 phosphorylation at the linker regions. Oncogene. 2004;23:7416–29. https://doi.org/10.1038/sj.onc.1207981
Waddell DS, Liberati NT, Guo X, Frederick JP, Wang XF. Casein kinase Iepsilon plays a functional role in the transforming growth factor-beta signaling pathway. J Biol Chem. 2004;279:29236–46. https://doi.org/10.1074/jbc.M400880200
Zhu H, Kavsak P, Abdollah S, Wrana JL, Thomsen GH. A SMAD ubiquitin ligase targets the BMP pathway and affects embryonic pattern formation. Nature. 1999;400:687–93. https://doi.org/10.1038/23293
Zhang Y, Chang C, Gehling DJ, Hemmati-Brivanlou A, Derynck R. Regulation of Smad degradation and activity by Smurf2, an E3 ubiquitin ligase. Proc Natl Acad Sci. 2001;98:974–9. https://doi.org/10.1073/pnas.98.3.974
Fukuchi M, Imamura T, Chiba T, Ebisawa T, Kawabata M, Tanaka K, et al. Ligand-dependent degradation of Smad3 by a ubiquitin ligase complex of ROC1 and associated proteins. Mol Biol Cell. 2001;12:1431–43. https://doi.org/10.1091/mbc.12.5.1431
Ray D, et al. Transforming growth factor β facilitates β-TrCP-mediated degradation of Cdc25A in a Smad3-dependent manner. Mol Cell Biol. 2005;25:3338–47. https://doi.org/10.1128/MCB.25.8.3338-3347.2005
Xin H, Xu X, Li L, Ning H, Rong Y, Shang Y, et al. CHIP controls the sensitivity of transforming growth factor-β signaling by modulating the basal level of Smad3 through ubiquitin-mediated degradation. J Biol Chem. 2005;280:20842–50.
Nagano Y, Mavrakis KJ, Lee KL, Fujii T, Koinuma D, Sase H, et al. Arkadia induces degradation of SnoN and c-Ski to enhance transforming growth factor-β signaling. J Biol Chem. 2007;282:20492–501.
Levy L, Howell M, Das D, Harkin S, Episkopou V, Hill CS. Arkadia activates Smad3/Smad4-dependent transcription by triggering signal-induced SnoN degradation. Mol Cell Biol. 2007;27:6068–83.
Bai Y, Yang C, Hu K, Elly C, Liu Y-C. Itch E3 ligase-mediated regulation of TGF-β signaling by modulating Smad2 phosphorylation. Mol Cell. 2004;15:825–31. https://doi.org/10.1016/j.molcel.2004.07.021
Huang Y-T, Cheng AC, Tang HC, Huang GC, Cai L, Lin TH, et al. USP7 facilitates SMAD3 autoregulation to repress cancer progression in p53-deficient lung cancer. Cell Death Dis. 2021;12:880. https://doi.org/10.1038/s41419-021-04176-8
Bai J, Xi Q. Crosstalk between TGF-β signaling and epigenome. Acta Biochimica et Biophysica Sin. 2017;50:60–67. https://doi.org/10.1093/abbs/gmx122
Xi Q, He W, Zhang XH-F, Le H-V, Massagué J. Genome-wide impact of the BRG1 SWI/SNF chromatin remodeler on the transforming growth factor β transcriptional program. J Biol Chem. 2008;283:1146–55.
Gaarenstroom T, Hill CS. TGF-β signaling to chromatin: How Smads regulate transcription during self-renewal and differentiation. Semin Cell Dev Biol. 2014;32:107–18. https://doi.org/10.1016/j.semcdb.2014.01.009
Shioda T, Lechleider RJ, Dunwoodie SL, Li H, Yahata T, de Caestecker MP, et al. Transcriptional activating activity of Smad4: roles of SMAD hetero-oligomerization and enhancement by an associating transactivator. Proc Natl Acad Sci. 1998;95:9785–90.
Kim DW, Lassar AB. Smad-dependent recruitment of a histone deacetylase/Sin3A complex modulates the bone morphogenetic protein-dependent transcriptional repressor activity of Nkx3.2. Mol Cell Biol. 2003;23:8704–17. https://doi.org/10.1128/mcb.23.23.8704-8717.2003
Tang X, Li G, Su F, Cai Y, Shi L, Meng Y, et al. HDAC8 cooperates with SMAD3/4 complex to suppress SIRT7 and promote cell survival and migration. Nucleic Acids Res. 2020;48:2912–23. https://doi.org/10.1093/nar/gkaa039
Dahle Ø, Kumar A, Kuehn MR. Nodal signaling recruits the histone demethylase Jmjd3 to counteract polycomb-mediated repression at target genes. Sci Signal. 2010;3:ra48–ra48.
Zhang YE. Non-Smad signaling pathways of the TGF-β family. Cold Spring Harb Perspect Biol. 9, https://doi.org/10.1101/cshperspect.a022129 (2017).
Lee MK, Pardoux C, Hall MC, Lee PS, Warburton D, Qing J, et al. TGF-beta activates Erk MAP kinase signalling through direct phosphorylation of ShcA. EMBO J. 2007;26:3957–67. https://doi.org/10.1038/sj.emboj.7601818
Yi JY, Shin I, Arteaga CL. Type I transforming growth factor beta; receptor binds to and activates phosphatidylinositol 3-kinase. J Biol Chem. 2005;280:10870–6. https://doi.org/10.1074/jbc.M413223200
Hamidi A, et al. TGF-beta promotes PI3K-AKT signaling and prostate cancer cell migration through the TRAF6-mediated ubiquitylation of p85. Sci Signal. 2017;10:eaal4186. https://doi.org/10.1126/scisignal.aal4186
Fruman DA, Chiu H, Hopkins BD, Bagrodia S, Cantley LC, Abraham RT. The PI3K pathway in human disease. Cell. 2017;170:605–35. https://doi.org/10.1016/j.cell.2017.07.029
Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease. Cell. 2017;168:960–76.
Silverstein AM. Autoimmunity versus horror autotoxicus: the struggle for recognition. Nat Immunol. 2001;2:279–81.
Taylor A, Verhagen J, Blaser K, Akdis M, Akdis CA. Mechanisms of immune suppression by interleukin-10 and transforming growth factor-beta: the role of T regulatory cells. Immunology. 2006;117:433–42. https://doi.org/10.1111/j.1365-2567.2006.02321.x
Schietinger A, Greenberg PD. Tolerance and exhaustion: defining mechanisms of T cell dysfunction. Trends Immunol. 2014;35:51–60. https://doi.org/10.1016/j.it.2013.10.001
Xing Y, Hogquist KA. T-cell tolerance: central and peripheral. Cold Spring Harb Perspect Biol. 4, https://doi.org/10.1101/cshperspect.a006957 (2012).
Klinker MW, Lundy SK. Multiple mechanisms of immune suppression by B lymphocytes. Mol Med. 2012;18:123–37. https://doi.org/10.2119/molmed.2011.00333
Fudenberg HH. Genetically determined immune deficiency as the predisposing cause of “auto-immunity” and lymphoid neoplasia. J Immunol. 1971;107:937–937.
Cunningham AJ. Active suppressor mechanism maintaining tolerance to some self components. Nature. 1975;254:143–4. https://doi.org/10.1038/254143a0
Allison AC, Denman AM, Barnes RD. Cooperating and controlling functions of thymus-derived lymphocytes in relation to autoimmunity. Lancet. 1971;2:135–40. https://doi.org/10.1016/s0140-6736(71)92306-3
Sakaguchi S, Fukuma K, Kuribayashi K, Masuda T. Organ-specific autoimmune diseases induced in mice by elimination of T cell subset. I. Evidence for the active participation of T cells in natural self-tolerance; deficit of a T cell subset as a possible cause of autoimmune disease. J Exp Med. 1985;161:72–87. https://doi.org/10.1084/jem.161.1.72
Sakaguchi S, Takahashi T, Nishizuka Y. Study on cellular events in post-thymectomy autoimmune oophoritis in mice. II. Requirement of Lyt-1 cells in normal female mice for the prevention of oophoritis. J Exp Med. 1982;156:1577–86. https://doi.org/10.1084/jem.156.6.1577
Sakaguchi S, Sakaguchi N, Asano M, Itoh M, Toda M. Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J Immunol. 1995;155:1151–64.
Brunkow ME, Jeffery EW, Hjerrild KA, Paeper B, Clark LB, Yasayko SA, et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat Genet. 2001;27:68–73. https://doi.org/10.1038/83784
Owen CJ, Jennings CE, Imrie H, Lachaux A, Bridges NA, Cheetham TD, et al. Mutational analysis of the FOXP3 gene and evidence for genetic heterogeneity in the immunodysregulation, polyendocrinopathy, enteropathy syndrome. J Clin Endocrinol Metab. 2003;88:6034–9. https://doi.org/10.1210/jc.2003-031080
Kobayashi I, Shiari R, Yamada M, Kawamura N, Okano M, Yara A, et al. Novel mutations of FOXP3 in two Japanese patients with immune dysregulation, polyendocrinopathy, enteropathy, X linked syndrome (IPEX). J Med Genet. 2001;38:874–6. https://doi.org/10.1136/jmg.38.12.874
Bennett CL, Christie J, Ramsdell F, Brunkow ME, Ferguson PJ, Whitesell L, et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat Genet. 2001;27:20–21. https://doi.org/10.1038/83713
Khattri R, Cox T, Yasayko SA, Ramsdell F. An essential role for Scurfin in CD4 + CD25 + T regulatory cells. Nat Immunol. 2003;4:337–42. https://doi.org/10.1038/ni909
Hori S, Nomura T, Sakaguchi S. Control of regulatory T cell development by the transcription factor Foxp3. Science. 2003;299:1057–61. https://doi.org/10.1126/science.10794901079490. [pii]
Fontenot JD, Gavin MA, Rudensky AY. Foxp3 programs the development and function of CD4 + CD25+ regulatory T cells. Nat Immunol. 2003;4:330–6.
Corthay A. How do regulatory T cells work? Scand J Immunol. 2009;70:326–36. https://doi.org/10.1111/j.1365-3083.2009.02308.x
Akimova T, Hancock WW. How little is known about the role of human FOXP3+ Tregs in tumors. Expert Opin Ther Targets. 2018;22:655–8. https://doi.org/10.1080/14728222.2018.1499728
Ziegler SF. FOXP3: of mice and men. Annu Rev Immunol. 2006;24:209–26. https://doi.org/10.1146/annurev.immunol.24.021605.090547
Abbas AK, Benoist C, Bluestone JA, Campbell DJ, Ghosh S, Hori S, et al. Regulatory T cells: recommendations to simplify the nomenclature. Nat Immunol. 2013;14:307–8. https://doi.org/10.1038/ni.2554
Shevach EM, Thornton AM. tTregs, pTregs, and iTregs: similarities and differences. Immunol Rev. 2014;259:88–102. https://doi.org/10.1111/imr.12160
Burchill MA, Yang J, Vang KB, Moon JJ, Chu HH, Lio CW, et al. Linked T cell receptor and cytokine signaling govern the development of the regulatory T cell repertoire. Immunity. 2008;28:112–21. https://doi.org/10.1016/j.immuni.2007.11.022
Hemmers S, Schizas M, Azizi E, Dikiy S, Zhong Y, Feng Y, et al. IL-2 production by self-reactive CD4 thymocytes scales regulatory T cell generation in the thymus. J Exp Med. 2019;216:2466–78. https://doi.org/10.1084/jem.20190993
Lio CW, Hsieh CS. A two-step process for thymic regulatory T cell development. Immunity. 2008;28:100–11. https://doi.org/10.1016/j.immuni.2007.11.021
Josefowicz SZ, Lu LF, Rudensky AY. Regulatory T cells: mechanisms of differentiation and function. Annu Rev Immunol. 2012;30:531–64. https://doi.org/10.1146/annurev.immunol.25.022106.141623
Morikawa H, Sakaguchi S. Genetic and epigenetic basis of Treg cell development and function: from a FoxP3-centered view to an epigenome-defined view of natural Treg cells. Immunol Rev. 2014;259:192–205. https://doi.org/10.1111/imr.12174
Joetham A, Schedel M, O'Connor BP, Kim S, Takeda K, Abbott J, et al. Inducible and naturally occurring regulatory T cells enhance lung allergic responses through divergent transcriptional pathways. J Allergy Clin Immunol. 2017;139:1331–42. https://doi.org/10.1016/j.jaci.2016.06.051
Josefowicz SZ, Niec RE, Kim HY, Treuting P, Chinen T, Zheng Y, et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature. 2012;482:395–9. https://doi.org/10.1038/nature10772
Curotto de Lafaille MA, Lafaille JJ. Natural and adaptive foxp3+ regulatory T cells: more of the same or a division of labor? Immunity. 2009;30:626–35. https://doi.org/10.1016/j.immuni.2009.05.002
Apostolou I, von Boehmer H. In vivo instruction of suppressor commitment in naive T cells. J Exp Med. 2004;199:1401–8. https://doi.org/10.1084/jem.20040249
Cobbold SP, Castejon R, Adams E, Zelenika D, Graca L, Humm S, et al. Induction of foxP3+ regulatory T cells in the periphery of T cell receptor transgenic mice tolerized to transplants. J Immunol. 2004;172:6003–10. https://doi.org/10.4049/jimmunol.172.10.6003
Kretschmer K, Apostolou I, Hawiger D, Khazaie K, Nussenzweig MC, von Boehmer H. Inducing and expanding regulatory T cell populations by foreign antigen. Nat Immunol. 2005;6:1219–27. https://doi.org/10.1038/ni1265
Mucida D, Kutchukhidze N, Erazo A, Russo M, Lafaille JJ, Curotto de Lafaille MA. Oral tolerance in the absence of naturally occurring Tregs. J Clin Invest. 2005;115:1923–33. https://doi.org/10.1172/JCI24487
Chai JN, Peng Y, Rengarajan S, Solomon BD, Ai TL, Shen Z, et al. Helicobacter species are potent drivers of colonic T cell responses in homeostasis and inflammation. Sci Immunol. 2, https://doi.org/10.1126/sciimmunol.aal5068 (2017).
Xu M, Pokrovskii M, Ding Y, Yi R, Au C, Harrison OJ, et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature. 2018;554:373–7. https://doi.org/10.1038/nature25500
Benson MJ, Pino-Lagos K, Rosemblatt M, Noelle RJ. All-trans retinoic acid mediates enhanced T reg cell growth, differentiation, and gut homing in the face of high levels of co-stimulation. J Exp Med. 2007;204:1765–74. https://doi.org/10.1084/jem.20070719
Coombes JL, Siddiqui KR, Arancibia-Cárcamo CV, Hall J, Sun CM, Belkaid Y, et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism. J Exp Med. 2007;204:1757–64. https://doi.org/10.1084/jem.20070590
Mucida D, Park Y, Kim G, Turovskaya O, Scott I, Kronenberg M, et al. Reciprocal TH17 and regulatory T cell differentiation mediated by retinoic acid. Science. 2007;317:256–60. https://doi.org/10.1126/science.1145697
Schambach F, Schupp M, Lazar MA, Reiner SL. Activation of retinoic acid receptor-alpha favours regulatory T cell induction at the expense of IL-17-secreting T helper cell differentiation. Eur J Immunol. 2007;37:2396–9. https://doi.org/10.1002/eji.200737621
Sun CM, Hall JA, Blank RB, Bouladoux N, Oukka M, Mora JR, et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J Exp Med. 2007;204:1775–85. https://doi.org/10.1084/jem.20070602
Smith PM, Howitt MR, Panikov N, Michaud M, Gallini CA, Bohlooly-Y M, et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science. 2013;341:569–73. https://doi.org/10.1126/science.1241165
Furusawa Y, Obata Y, Fukuda S, Endo TA, Nakato G, Takahashi D, et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature. 2013;504:446–50. https://doi.org/10.1038/nature12721
Hang S, Paik D, Yao L, Kim E, Trinath J, Lu J, et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature. 2019;576:143–8. https://doi.org/10.1038/s41586-019-1785-z
Song X, Sun X, Oh SF, Wu M, Zhang Y, Zheng W, et al. Microbial bile acid metabolites modulate gut RORgamma(+) regulatory T cell homeostasis. Nature. 2020;577:410–5. https://doi.org/10.1038/s41586-019-1865-0
Quintana FJ, Basso AS, Iglesias AH, Korn T, Farez MF, Bettelli E, et al. Control of T(reg) and T(H)17 cell differentiation by the aryl hydrocarbon receptor. Nature. 2008;453:65–71. https://doi.org/10.1038/nature06880
Chen W, Jin W, Hardegen N, Lei KJ, Li L, Marinos N, et al. Conversion of peripheral CD4 + CD25- naive T cells to CD4 + CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J Exp Med. 2003;198:1875–86. https://doi.org/10.1084/jem.20030152
Zheng SG, Wang J, Wang P, Gray JD, Horwitz DA. IL-2 is essential for TGF-beta to convert naive CD4 + CD25- cells to CD25+Foxp3+ regulatory T cells and for expansion of these cells. J Immunol. 2007;178:2018–27. https://doi.org/10.4049/jimmunol.178.4.2018
Wan YY, Flavell RA. Identifying Foxp3-expressing suppressor T cells with a bicistronic reporter. Proc Natl Acad Sci. 2005;102:5126–31. https://doi.org/10.1073/pnas.0501701102
Floess S, Freyer J, Siewert C, Baron U, Olek S, Polansky J, et al. Epigenetic control of the foxp3 locus in regulatory T cells. PLoS Biol. 2007;5:e38. https://doi.org/10.1371/journal.pbio.0050038
Yuan X, Cheng G, Malek TR. The importance of regulatory T-cell heterogeneity in maintaining self-tolerance. Immunol Rev. 2014;259:103–14. https://doi.org/10.1111/imr.12163
Feuerer M, Hill JA, Mathis D, Benoist C. Foxp3+ regulatory T cells: differentiation, specification, subphenotypes. Nat Immunol. 2009;10:689–95.
Kim JM, Rasmussen JP, Rudensky AY. Regulatory T cells prevent catastrophic autoimmunity throughout the lifespan of mice. Nat Immunol. 2007;8:191–7. https://doi.org/10.1038/ni1428
Wan YY, Flavell RA. Regulatory T-cell functions are subverted and converted owing to attenuated Foxp3 expression. Nature. 2007;445:766–70. https://doi.org/10.1038/nature05479
Williams LM, Rudensky AY. Maintenance of the Foxp3-dependent developmental program in mature regulatory T cells requires continued expression of Foxp3. Nat Immunol. 2007;8:277–84. https://doi.org/10.1038/ni1437
Gavin MA, Rasmussen JP, Fontenot JD, Vasta V, Manganiello VC, Beavo JA, et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature. 2007;445:771–5.
Wan YY. Regulatory T cells: immune suppression and beyond. Cell Mol Immunol. 2010;7:204–10. https://doi.org/10.1038/cmi.2010.20
Lo YC, Lee CF, Powell JD. Insight into the role of mTOR and metabolism in T cells reveals new potential approaches to preventing graft rejection. Curr Opin Organ Transpl. 2014;19:363–71. https://doi.org/10.1097/MOT.0000000000000098
Chapman NM, Chi H. mTOR signaling, Tregs and immune modulation. Immunotherapy. 2014;6:1295–311. https://doi.org/10.2217/imt.14.84
Kerdiles YM, Stone EL, Beisner DR, McGargill MA, Ch'en IL, Stockmann C, et al. Foxo transcription factors control regulatory T cell development and function. Immunity. 2010;33:890–904. https://doi.org/10.1016/j.immuni.2010.12.002
Merkenschlager M, von Boehmer H. PI3 kinase signalling blocks Foxp3 expression by sequestering Foxo factors. J Exp Med. 2010;207:1347–50. https://doi.org/10.1084/jem.20101156
Ouyang W, Beckett O, Ma Q, Paik JH, DePinho RA, Li MO. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat Immunol. 2010;11:618–27. https://doi.org/10.1038/ni.1884
Zheng Y, Josefowicz S, Chaudhry A, Peng XP, Forbush K, Rudensky AY. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature. 2010;463:808–12. https://doi.org/10.1038/nature08750
Basu R, Hatton RD, Weaver CT. The Th17 family: flexibility follows function. Immunol Rev. 2013;252:89–103. https://doi.org/10.1111/imr.12035
Kitoh A, Ono M, Naoe Y, Ohkura N, Yamaguchi T, Yaguchi H, et al. Indispensable role of the Runx1-Cbfbeta transcription complex for in vivo-suppressive function of FoxP3+ regulatory T cells. Immunity. 2009;31:609–20. https://doi.org/10.1016/j.immuni.2009.09.003
Wang Y, Su MA, Wan YY. An essential role of the transcription factor GATA-3 for the function of regulatory T cells. Immunity. 2011;35:337–48. https://doi.org/10.1016/j.immuni.2011.08.012
Deng G, Song X, Fujimoto S, Piccirillo CA, Nagai Y, Greene MI. Foxp3 post-translational modifications and Treg suppressive activity. Front Immunol. 2019;10:2486. https://doi.org/10.3389/fimmu.2019.02486
Li Z, Li D, Tsun A, Li B. FOXP3+ regulatory T cells and their functional regulation. Cell Mol Immunol. 2015;12:558–65. https://doi.org/10.1038/cmi.2015.10
Chou WC, Guo Z, Guo H, Chen L, Zhang G, Liang K, et al. AIM2 in regulatory T cells restrains autoimmune diseases. Nature. 2021;591:300–5. https://doi.org/10.1038/s41586-021-03231-w
Shrestha S, Yang K, Guy C, Vogel P, Neale G, Chi H. Treg cells require the phosphatase PTEN to restrain TH1 and TFH cell responses. Nat Immunol. 2015;16:178–87. https://doi.org/10.1038/ni.3076
Huynh A, DuPage M, Priyadharshini B, Sage PT, Quiros J, Borges CM, et al. Control of PI(3) kinase in Treg cells maintains homeostasis and lineage stability. Nat Immunol. 2015;16:188–96. https://doi.org/10.1038/ni.3077
Luo CT, Liao W, Dadi S, Toure A, Li MO. Graded Foxo1 activity in Treg cells differentiates tumour immunity from spontaneous autoimmunity. Nature. 2016;529:532–6. https://doi.org/10.1038/nature16486
Wei J, Long L, Yang K, Guy C, Shrestha S, Chen Z, et al. Autophagy enforces functional integrity of regulatory T cells by coupling environmental cues and metabolic homeostasis. Nat Immunol. 2016;17:277–85. https://doi.org/10.1038/ni.3365
Miyao T, Floess S, Setoguchi R, Luche H, Fehling HJ, Waldmann H, et al. Plasticity of Foxp3(+) T cells reflects promiscuous Foxp3 expression in conventional T cells but not reprogramming of regulatory T cells. Immunity. 2012;36:262–75. https://doi.org/10.1016/j.immuni.2011.12.012
Hori S. Lineage stability and phenotypic plasticity of Foxp3(+) regulatory T cells. Immunol Rev. 2014;259:159–72. https://doi.org/10.1111/imr.12175
Zhang Z, Zhou X. Foxp3 instability helps ttregs distinguish self and non-self. Front Immunol. 2019;10:2226. https://doi.org/10.3389/fimmu.2019.02226
Murai M, Krause P, Cheroutre H, Kronenberg M. Regulatory T-cell stability and plasticity in mucosal and systemic immune systems. Mucosal Immunol. 2010;3:443–9. https://doi.org/10.1038/mi.2010.27
Lu L, Barbi J, Pan F. The regulation of immune tolerance by FOXP3. Nat Rev Immunol. 2017;17:703–17. https://doi.org/10.1038/nri.2017.75
Vignali DA, Collison LW, Workman CJ. How regulatory T cells work. Nat Rev Immunol. 2008;8:523–32. https://doi.org/10.1038/nri2343
Spolski R, Li P, Leonard WJ. Biology and regulation of IL-2: from molecular mechanisms to human therapy. Nat Rev Immunol. 2018;18:648–59. https://doi.org/10.1038/s41577-018-0046-y
Ross SH, Cantrell DA. Signaling and function of interleukin-2 in T lymphocytes. Annu Rev Immunol. 2018;36:411–33. https://doi.org/10.1146/annurev-immunol-042617-053352
Oderup C, Cederbom L, Makowska A, Cilio CM, Ivars F. Cytotoxic T lymphocyte antigen-4-dependent down-modulation of costimulatory molecules on dendritic cells in CD4 + CD25+ regulatory T-cell-mediated suppression. Immunology. 2006;118:240–9. https://doi.org/10.1111/j.1365-2567.2006.02362.x
Cederbom L, Hall H, Ivars F. CD4 + CD25+ regulatory T cells down-regulate co-stimulatory molecules on antigen-presenting cells. Eur J Immunol. 2000;30:1538–43. https://doi.org/10.1002/1521-4141(200006)30:6<1538::AID-IMMU1538>3.0.CO;2-X
Sojka DK, Huang YH, Fowell DJ. Mechanisms of regulatory T-cell suppression - a diverse arsenal for a moving target. Immunology. 2008;124:13–22. https://doi.org/10.1111/j.1365-2567.2008.02813.x
Cao X, Cai SF, Fehniger TA, Song J, Collins LI, Piwnica-Worms DR, et al. Granzyme B and perforin are important for regulatory T cell-mediated suppression of tumor clearance. Immunity. 2007;27:635–46. https://doi.org/10.1016/j.immuni.2007.08.014
Jang JE, Hajdu CH, Liot C, Miller G, Dustin ML, Bar-Sagi D. Crosstalk between regulatory T cells and tumor-associated dendritic cells negates anti-tumor immunity in pancreatic cancer. Cell Rep. 2017;20:558–71. https://doi.org/10.1016/j.celrep.2017.06.062
Onishi Y, Fehervari Z, Yamaguchi T, Sakaguchi S. Foxp3+ natural regulatory T cells preferentially form aggregates on dendritic cells in vitro and actively inhibit their maturation. Proc Natl Acad Sci USA. 2008;105:10113–8. https://doi.org/10.1073/pnas.0711106105
Angelin A, Gil-de-Gómez L, Dahiya S, Jiao J, Guo L, Levine MH, et al. Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab. 2017;25:1282–193:e1287. https://doi.org/10.1016/j.cmet.2016.12.018
Li X, Yang Y, Zhang B, Lin X, Fu X, An Y, et al. Lactate metabolism in human health and disease. Signal Transduct Target Ther. 2022;7:305. https://doi.org/10.1038/s41392-022-01151-3
Pucino V, Certo M, Bulusu V, Cucchi D, Goldmann K, Pontarini E, et al. Lactate buildup at the site of chronic inflammation promotes disease by inducing CD4(+) T cell metab rewiring. Cell Metab. 2019;30:1055–1074:e1058. https://doi.org/10.1016/j.cmet.2019.10.004
Bromley SK, Mempel TR, Luster AD. Orchestrating the orchestrators: chemokines in control of T cell traffic. Nat Immunol. 2008;9:970–80. https://doi.org/10.1038/ni.f.213
Iellem A, Mariani M, Lang R, Recalde H, Panina-Bordignon P, Sinigaglia F, et al. Unique chemotactic response profile and specific expression of chemokine receptors CCR4 and CCR8 by CD4(+)CD25(+) regulatory T cells. J Exp Med. 2001;194:847–53. https://doi.org/10.1084/jem.194.6.847
Chen D, Bromberg JS. T regulatory cells and migration. Am J Transpl. 2006;6:1518–23. https://doi.org/10.1111/j.1600-6143.2006.01372.x
Szabo SJ, Kim ST, Costa GL, Zhang X, Fathman CG, Glimcher LH. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell. 2000;100:655–69. https://doi.org/10.1016/s0092-8674(00)80702-3
Koch MA, Thomas KR, Perdue NR, Smigiel KS, Srivastava S, Campbell DJ. T-bet(+) Treg cells undergo abortive Th1 cell differentiation due to impaired expression of IL-12 receptor beta2. Immunity. 2012;37:501–10. https://doi.org/10.1016/j.immuni.2012.05.031
Honma K, Kimura D, Tominaga N, Miyakoda M, Matsuyama T, Yui K. Interferon regulatory factor 4 differentially regulates the production of Th2 cytokines in naive vs. effector/memory CD4+T cells. Proc Natl Acad Sci USA. 2008;105:15890–5. https://doi.org/10.1073/pnas.0803171105
Zheng Y, Chaudhry A, Kas A, deRoos P, Kim JM, Chu TT, et al. Regulatory T-cell suppressor program co-opts transcription factor IRF4 to control T(H)2 responses. Nature. 2009;458:351–6. https://doi.org/10.1038/nature07674
Ivanov II, McKenzie BS, Zhou L, Tadokoro CE, Lepelley A, Lafaille JJ, et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17 + T helper. cells Cell. 2006;126:1121–33. https://doi.org/10.1016/j.cell.2006.07.035
Yang BH, Hagemann S, Mamareli P, Lauer U, Hoffmann U, Beckstette M, et al. Foxp3(+) T cells expressing RORgammat represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol. 2016;9:444–57. https://doi.org/10.1038/mi.2015.74
Wei G, et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4 + T cells. Immunity. 2009;30:155–67. https://doi.org/10.1016/j.immuni.2008.12.009.
Ohkura N, Sakaguchi S. Transcriptional and epigenetic basis of Treg cell development and function: its genetic anomalies or variations in autoimmune diseases. Cell Res. 2020;30:465–74. https://doi.org/10.1038/s41422-020-0324-7
Ohnmacht C. ToLerance To The Intestinal Microbiota Mediated by ROR(gammat)(+) Cells. Trends Immunol. 2016;37:477–86. https://doi.org/10.1016/j.it.2016.05.002
van der Veeken J, Campbell C, Pritykin Y, Schizas M, Verter J, Hu W, et al. Genetic tracing reveals transcription factor Foxp3-dependent and Foxp3-independent functionality of peripherally induced Treg cells. Immunity. 2022;55:1173–1184:e1177. https://doi.org/10.1016/j.immuni.2022.05.010
Li J, Tan J, Martino MM, Lui KO. Regulatory T-cells: potential regulator of tissue repair and regeneration. Front Immunol. 2018;9:585. https://doi.org/10.3389/fimmu.2018.00585
Burzyn D, Benoist C, Mathis D. Regulatory T cells in nonlymphoid tissues. Nat Immunol. 2013;14:1007–13. https://doi.org/10.1038/ni.2683
Yamazaki T, Yang XO, Chung Y, Fukunaga A, Nurieva R, Pappu B, et al. CCR6 regulates the migration of inflammatory and regulatory T cells. J Immunol. 2008;181:8391–401. https://doi.org/10.4049/jimmunol.181.12.8391
Turner JE, Paust HJ, Steinmetz OM, Peters A, Riedel JH, Erhardt A, et al. CCR6 recruits regulatory T cells and Th17 cells to the kidney in glomerulonephritis. J Am Soc Nephrol. 2010;21:974–85. https://doi.org/10.1681/ASN.2009070741
Cook KW, Letley DP, Ingram RJ, Staples E, Skjoldmose H, Atherton JC, et al. CCL20/CCR6-mediated migration of regulatory T cells to the Helicobacter pylori-infected human gastric mucosa. Gut. 2014;63:1550–9. https://doi.org/10.1136/gutjnl-2013-306253
Burzyn D, Kuswanto W, Kolodin D, Shadrach JL, Cerletti M, Jang Y, et al. A special population of regulatory T cells potentiates muscle repair. Cell. 2013;155:1282–95. https://doi.org/10.1016/j.cell.2013.10.054
Castiglioni A, Corna G, Rigamonti E, Basso V, Vezzoli M, Monno A, et al. FOXP3 + T cells recruited to sites of sterile skeletal muscle injury regulate the fate of satellite cells and guide effective tissue regeneration. PLoS One. 2015;10:e0128094. https://doi.org/10.1371/journal.pone.0128094
Arpaia N, Green JA, Moltedo B, Arvey A, Hemmers S, Yuan S, et al. A distinct function of regulatory T cells in tissue protection. Cell. 2015;162:1078–89. https://doi.org/10.1016/j.cell.2015.08.021
Dombrowski Y, O'Hagan T, Dittmer M, Penalva R, Mayoral SR, Bankhead P, et al. Regulatory T cells promote myelin regeneration in the central nervous system. Nat Neurosci. 2017;20:674–80. https://doi.org/10.1038/nn.4528
de la Vega Gallardo N, Dittmer M, Dombrowski Y, Fitzgerald DC. Regenerating CNS myelin: Emerging roles of regulatory T cells and CCN proteins. Neurochem Int. 2019;130:104349. https://doi.org/10.1016/j.neuint.2018.11.024
Sakaguchi S, Ono M, Setoguchi R, Yagi H, Hori S, Fehervari Z, et al. Foxp3 + CD25 + CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol Rev. 2006;212:8–27. https://doi.org/10.1111/j.0105-2896.2006.00427.x
Sakaguchi S, Yamaguchi T, Nomura T, Ono M. Regulatory T cells and immune tolerance. Cell. 2008;133:775–87. https://doi.org/10.1016/j.cell.2008.05.009
Alroqi FJ, Chatila TA. T regulatory cell biology in health and disease. Curr Allergy Asthma Rep. 2016;16:27. https://doi.org/10.1007/s11882-016-0606-9
Hartemann A, Bensimon G, Payan CA, Jacqueminet S, Bourron O, Nicolas N, et al. Low-dose interleukin 2 in patients with type 1 diabetes: a phase 1/2 randomised, double-blind, placebo-controlled trial. Lancet Diabetes Endocrinol. 2013;1:295–305. https://doi.org/10.1016/S2213-8587(13)70113-X
Zhou P. Emerging mechanisms and applications of low-dose IL-2 therapy in autoimmunity. Cytokine Growth Factor Rev. 2022;67:80–88. https://doi.org/10.1016/j.cytogfr.2022.06.003
Kasagi S, Zhang P, Che L, Abbatiello B, Maruyama T, Nakatsukasa H, et al. In vivo-generated antigen-specific regulatory T cells treat autoimmunity without compromising antibacterial immune response. Sci Transl Med. 2014;6:241ra278. https://doi.org/10.1126/scitranslmed.3008895
Selck C, Dominguez-Villar M. Antigen-specific regulatory T cell therapy in autoimmune diseases and transplantation. Front Immunol. 2021;12:661875. https://doi.org/10.3389/fimmu.2021.661875
Sarmay G. Biologia Futura: Emerging antigen-specific therapies for autoimmune diseases. Biol Futur. 2021;72:15–24. https://doi.org/10.1007/s42977-021-00074-4
Gorantla VS, Schneeberger S, Brandacher G, Sucher R, Zhang D, Lee WP, et al. T regulatory cells and transplantation tolerance. Transpl Rev (Orlando). 2010;24:147–59. https://doi.org/10.1016/j.trre.2010.04.002
Pilat N, Wiletel M, Weijler AM, Steiner R, Mahr B, Warren J, et al. Treg-mediated prolonged survival of skin allografts without immunosuppression. Proc Natl Acad Sci USA. 2019;116:13508–16. https://doi.org/10.1073/pnas.1903165116
Palomares O, Yaman G, Azkur AK, Akkoc T, Akdis M, Akdis CA. Role of Treg in immune regulation of allergic diseases. Eur J Immunol. 2010;40:1232–40. https://doi.org/10.1002/eji.200940045
Taylor MD, van der Werf N, Harris A, Graham AL, Bain O, Allen JE, et al. Early recruitment of natural CD4+ Foxp3+ Treg cells by infective larvae determines the outcome of filarial infection. Eur J Immunol. 2009;39:192–206. https://doi.org/10.1002/eji.200838727
Oldenhove G, Bouladoux N, Wohlfert EA, Hall JA, Chou D, Dos Santos L, et al. Decrease of Foxp3+ Treg cell number and acquisition of effector cell phenotype during lethal infection. Immunity. 2009;31:772–86. https://doi.org/10.1016/j.immuni.2009.10.001
Kulkarni OP, Lichtnekert J, Anders HJ, Mulay SR. the immune system in tissue environments regaining homeostasis after injury: Is “Inflammation” always inflammation? Mediators Inflamm. 2016;2016:2856213. https://doi.org/10.1155/2016/2856213
Belkaid Y, Tarbell K. Regulatory T cells in the control of host-microorganism interactions (*). Annu Rev Immunol. 2009;27:551–89. https://doi.org/10.1146/annurev.immunol.021908.132723.
Fulop T, Larbi A, Dupuis G, Le Page A, Frost EH, Cohen AA, et al. Immunosenescence and inflamm-aging as two sides of the same coin: friends or foes? Front Immunol. 2017;8:1960. https://doi.org/10.3389/fimmu.2017.01960
Kennedy BK, Berger SL, Brunet A, Campisi J, Cuervo AM, Epel ES, et al. Geroscience: linking aging to chronic disease. Cell. 2014;159:709–13. https://doi.org/10.1016/j.cell.2014.10.039
Furman D, Campisi J, Verdin E, Carrera-Bastos P, Targ S, Franceschi C, et al. Chronic inflammation in the etiology of disease across the life span. Nat Med. 2019;25:1822–32. https://doi.org/10.1038/s41591-019-0675-0
Guo Z, Wang G, Wu B, Chou WC, Cheng L, Zhou C, et al. DCAF1 regulates Treg senescence via the ROS axis during immunological aging. J Clin Invest. 2020;130:5893–908. https://doi.org/10.1172/JCI136466
Ha TY. The role of regulatory T cells in cancer. Immune Netw. 2009;9:209–35. https://doi.org/10.4110/in.2009.9.6.209
Halvorsen EC, Mahmoud SM, Bennewith KL. Emerging roles of regulatory T cells in tumour progression and metastasis. Cancer Metastasis Rev. 2014;33:1025–41. https://doi.org/10.1007/s10555-014-9529-x
Fridman WH, Pages F, Sautes-Fridman C, Galon J. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12:298–306. https://doi.org/10.1038/nrc3245
Yamaguchi T, Sakaguchi S. Regulatory T cells in immune surveillance and treatment of cancer. Semin Cancer Biol. 2006;16:115–23. https://doi.org/10.1016/j.semcancer.2005.11.005
Ohue Y, Nishikawa H. Regulatory T (Treg) cells in cancer: Can Treg cells be a new therapeutic target? Cancer Sci. 2019;110:2080–9. https://doi.org/10.1111/cas.14069
Facciabene A, Motz GT, Coukos G. T-regulatory cells: key players in tumor immune escape and angiogenesis. Cancer Res. 2012;72:2162–71. https://doi.org/10.1158/0008-5472.CAN-11-3687
Nishikawa H, Kato T, Tanida K, Hiasa A, Tawara I, Ikeda H, et al. CD4 + CD25 + T cells responding to serologically defined autoantigens suppress antitumor immune responses. Proc Natl Acad Sci USA. 2003;100:10902–6. https://doi.org/10.1073/pnas.1834479100
Chen Y, Kuchroo VK, Inobe J, Hafler DA, Weiner HL. Regulatory T cell clones induced by oral tolerance: suppression of autoimmune encephalomyelitis. Science. 1994;265:1237–40. https://doi.org/10.1126/science.7520605
Dwarakanath BS, Farooque A, Gupta S. Targeting regulatory T cells for improving cancer therapy: Challenges and prospects. Cancer Rep. (Hoboken). 2018;1:e21105. https://doi.org/10.1002/cnr2.1105
Cinier J, Hubert M, Besson L, Di Roio A, Rodriguez C, Lombardi V, et al. Recruitment and expansion of tregs cells in the tumor environment-how to target them? Cancers (Basel). 13, https://doi.org/10.3390/cancers13081850 (2021).
Chen BJ, Zhao JW, Zhang DH, Zheng AH, Wu GQ. Immunotherapy of cancer by targeting regulatory T cells. Int Immunopharmacol. 2022;104:108469. https://doi.org/10.1016/j.intimp.2021.108469
Gorelik L, Flavell RA. Transforming growth factor-β in T-cell biology. Nat Rev Immunol. 2002;2:46–53. https://doi.org/10.1038/nri704
Tone Y, Furuuchi K, Kojima Y, Tykocinski ML, Greene MI, Tone M. Smad3 and NFAT cooperate to induce Foxp3 expression through its enhancer. Nat Immunol. 2008;9:194–202. https://doi.org/10.1038/ni1549
Fantini MC, Becker C, Monteleone G, Pallone F, Galle PR, Neurath MF. Cutting Edge: TGF-β induces a regulatory phenotype in CD4 + CD25 − T cells through Foxp3 induction and down-regulation of Smad7. J Immunol. 2004;172:5149–53. https://doi.org/10.4049/jimmunol.172.9.5149
Hadaschik EN, Enk AH. TGF-β1-induced regulatory T cells. Hum Immunol. 2015;76:561–4. https://doi.org/10.1016/j.humimm.2015.06.015
Tu E, Chia C, Chen W, Zhang D, Park SA, Jin W, et al. T cell receptor-regulated TGF-beta type I receptor expression determines T cell quiescence and activation. Immunity. 2018;48:745–759:e746. https://doi.org/10.1016/j.immuni.2018.03.025
Gu A-D, Zhang S, Wang Y, Xiong H, Curtis TA, Wan YY. A critical role for transcription factor Smad4 in T cell function that is independent of transforming growth factor β receptor signaling. Immunity. 2015;42:68–79. https://doi.org/10.1016/j.immuni.2014.12.019
Konkel JE, Jin W, Abbatiello B, Grainger JR, Chen W. Thymocyte apoptosis drives the intrathymic generation of regulatory T cells. Proc Natl Acad Sci. 2014;111:E465–E473. https://doi.org/10.1073/pnas.1320319111
Ouyang W, Beckett O, Ma Q, Li MO. Transforming growth factor-β signaling curbs thymic negative selection promoting regulatory T cell development. Immunity. 2010;32:642–53. https://doi.org/10.1016/j.immuni.2010.04.012
Ishigame H, Zenewicz LA, Sanjabi S, Licona-Limón P, Nakayama M, Leonard WJ, et al. Excessive Th1 responses due to the absence of TGF-beta signaling cause autoimmune diabetes and dysregulated Treg cell homeostasis. Proc Natl Acad Sci USA. 2013;110:6961–6. https://doi.org/10.1073/pnas.1304498110
Zhang N, Bevan MJ. TGF-beta signaling to T cells inhibits autoimmunity during lymphopenia-driven proliferation. Nat Immunol. 2012;13:667–73. https://doi.org/10.1038/ni.2319
Konkel JE, Zhang D, Zanvit P, Chia C, Zangarle-Murray T, Jin W, et al. Transforming growth factor-β signaling in regulatory T cells controls T Helper-17 cells and tissue-specific immune responses. Immunity. 2017;46:660–74. https://doi.org/10.1016/j.immuni.2017.03.015
Green EA, Gorelik L, McGregor CM, Tran EH, Flavell RA. CD4 + CD25 + T regulatory cells control anti-islet CD8 + T cells through TGF-beta-TGF-beta receptor interactions in type 1 diabetes. Proc Natl Acad Sci USA. 2003;100:10878–83. https://doi.org/10.1073/pnas.1834400100
Gutcher I, Donkor MK, Ma Q, Rudensky AY, Flavell RA, Li MO. Autocrine transforming growth factor-beta1 promotes in vivo Th17 cell differentiation. Immunity. 2011;34:396–408. https://doi.org/10.1016/j.immuni.2011.03.005
Li MO, Wan YY, Flavell RA. T cell-produced transforming growth factor-beta1 controls T cell tolerance and regulates Th1- and Th17-cell differentiation. Immunity. 2007;26:579–91. https://doi.org/10.1016/j.immuni.2007.03.014
Turner JA, Stephen-Victor E, Wang S, Rivas MN, Abdel-Gadir A, Harb H, et al. Regulatory T cell-derived TGF-beta1 controls multiple checkpoints governing allergy and auto. Immun Immun. 2020;53:1202–1214:e1206. https://doi.org/10.1016/j.immuni.2020.10.002
Velegraki M, Salem M, Ansa-Addo EA, Wu BX, Li Z. Autocrine transforming growth factor beta1 in regulatory T cell biology-gone but not missed. Immunity. 2021;54:395–6. https://doi.org/10.1016/j.immuni.2021.02.007
Marie JC, Letterio JJ, Gavin M, Rudensky AY. TGF-beta1 maintains suppressor function and Foxp3 expression in CD4 + CD25+ regulatory T cells. J Exp Med. 2005;201:1061–7. https://doi.org/10.1084/jem.20042276
Hill JA, Feuerer M, Tash K, Haxhinasto S, Perez J, Melamed R, et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity. 2007;27:786–800. https://doi.org/10.1016/j.immuni.2007.09.010
Selvaraj RK, Geiger TL. A kinetic and dynamic analysis of Foxp3 induced in T cells by TGF-β. J Immunol. 2007;178:7667–77.
Martinez GJ, Zhang Z, Chung Y, Reynolds JM, Lin X, Jetten AM, et al. Smad3 differentially regulates the induction of regulatory and inflammatory T cell differentiation *. J Biol Chem. 2009;284:35283–6. https://doi.org/10.1074/jbc.C109.078238
Martinez GJ, Zhang Z, Reynolds JM, Tanaka S, Chung Y, Liu T, et al. Smad2 positively regulates the generation of Th17 cells. J Biol Chem. 2010;285:29039–43. https://doi.org/10.1074/jbc.C110.155820
Gu A-D, Wang Y, Lin L, Zhang SS, Wan YY. Requirements of transcription factor Smad-dependent and -independent TGF-β signaling to control discrete T-cell functions. Proc Natl Acad Sci. 2012;109:905–10. https://doi.org/10.1073/pnas.1108352109
Takimoto T, Wakabayashi Y, Sekiya T, Inoue N, Morita R, Ichiyama K, et al. Smad2 and Smad3 are redundantly essential for the TGF-beta-mediated regulation of regulatory T plasticity and Th1 development. J Immunol. 2010;185:842–55. https://doi.org/10.4049/jimmunol.0904100
Yang XO, Nurieva R, Martinez GJ, Kang HS, Chung Y, Pappu BP, et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity. 2008;29:44–56. https://doi.org/10.1016/j.immuni.2008.05.007
Xu H, Wu L, Nguyen HH, Mesa KR, Raghavan V, Episkopou V, et al. Arkadia-SKI/SnoN signaling differentially regulates TGF-β–induced iTreg and Th17 cell differentiation. J Exp Med 218, https://doi.org/10.1084/jem.20210777 (2021).
Nakao A, Okumura K, Ogawa H. Smad7: a new key player in TGF-beta-associated disease. Trends Mol Med. 2002;8:361–3. https://doi.org/10.1016/s1471-4914(02)02376-6
Zhang S, Takaku M, Zou L, Gu AD, Chou WC, Zhang G, et al. Reversing SKI-SMAD4-mediated suppression is essential for TH17 cell differentiation. Nature. 2017;551:105–9. https://doi.org/10.1038/nature24283
Schlenner SM, Weigmann B, Ruan Q, Chen Y, von Boehmer H. Smad3 binding to the foxp3 enhancer is dispensable for the development of regulatory T cells with the exception of the gut. J Exp Med. 2012;209:1529–35. https://doi.org/10.1084/jem.20112646
Chen W, ten Dijke P. Immunoregulation by members of the TGFβ superfamily. Nat Rev Immunol. 2016;16:723–40. https://doi.org/10.1038/nri.2016.112
Huber S, Stahl FR, Schrader J, Lüth S, Presser K, Carambia A, et al. Activin a promotes the TGF-β-induced conversion of CD4 + CD25 − T cells into Foxp3+ induced regulatory T cells. J Immunol. 2009;182:4633–40.
Read S, Malmström V, Powrie F. Cytotoxic T lymphocyte–associated antigen 4 plays an essential role in the function of CD25 + CD4+ regulatory cells that control intestinal inflammation. J Exp Med. 2000;192:295–302.
Nakamura K, Kitani A, Strober W. Cell contact–dependent immunosuppression by Cd4 + Cd25+regulatory T cells is mediated by cell surface–bound transforming growth factor β. J Exp Med. 2001;194:629–44. https://doi.org/10.1084/jem.194.5.629
Barnes MJ, Li CM, Xu Y, An J, Huang Y, Cyster JG. The lysophosphatidylserine receptor GPR174 constrains regulatory T cell development and function. J Exp Med. 2015;212:1011–20. https://doi.org/10.1084/jem.20141827
Zhou Y, Lee JY, Lee CM, Cho WK, Kang MJ, Koff JL, et al. Amphiregulin, an epidermal growth factor receptor ligand, plays an essential role in the pathogenesis of transforming growth factor-beta-induced pulmonary fibrosis. J Biol Chem. 2012;287:41991–42000. https://doi.org/10.1074/jbc.M112.356824
Mosmann TR, Coffman RL. TH1 and TH2 cells: different patterns of lymphokine secretion lead to different functional properties. Annu Rev Immunol. 1989;7:145–73. https://doi.org/10.1146/annurev.iy.07.040189.001045
Oppmann B, Lesley R, Blom B, Timans JC, Xu Y, Hunte B, et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity. 2000;13:715–25. https://doi.org/10.1016/s1074-7613(00)00070-4
Cua DJ, Sherlock J, Chen Y, Murphy CA, Joyce B, Seymour B, et al. Interleukin-23 rather than interleukin-12 is the critical cytokine for autoimmune inflammation of the brain. Nature. 2003;421:744–8. https://doi.org/10.1038/nature01355
Langrish CL, Chen Y, Blumenschein WM, Mattson J, Basham B, Sedgwick JD, et al. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. J Exp Med. 2005;201:233–40. https://doi.org/10.1084/jem.20041257
Harrington LE, Hatton RD, Mangan PR, Turner H, Murphy TL, Murphy KM, et al. Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat Immunol. 2005;6:1123–32. https://doi.org/10.1038/ni1254
Park H, Li Z, Yang XO, Chang SH, Nurieva R, Wang YH, et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat Immunol. 2005;6:1133–41. https://doi.org/10.1038/ni1261
Dong C. TH17 cells in development: an updated view of their molecular identity and genetic programming. Nat Rev Immunol. 2008;8:337–48. https://doi.org/10.1038/nri2295
Korn T, Bettelli E, Oukka M, Kuchroo VK. IL-17 and Th17 cells. Annu Rev Immunol. 2009;27:485–517. https://doi.org/10.1146/annurev.immunol.021908.132710
Wu B, Wan Y. Molecular control of pathogenic Th17 cells in autoimmune diseases. Int Immunopharmacol. 2020;80:106187 https://doi.org/10.1016/j.intimp.2020.106187
Schnell A, Huang L, Singer M, Singaraju A, Barilla RM, Regan B, et al. Stem-like intestinal Th17 cells give rise to pathogenic effector T cells during autoimmunity. Cell. 2021;184:6281–698:e6223. https://doi.org/10.1016/j.cell.2021.11.018
McGeachy MJ, Bak-Jensen KS, Chen Y, Tato CM, Blumenschein W, McClanahan T, et al. TGF-beta and IL-6 drive the production of IL-17 and IL-10 by T cells and restrain T(H)-17 cell-mediated pathology. Nat Immunol. 2007;8:1390–97. https://doi.org/10.1038/ni1539
Ghoreschi K, Laurence A, Yang XP, Tato CM, McGeachy MJ, Konkel JE, et al. Generation of pathogenic T(H)17 cells in the absence of TGF-beta signalling. Nature. 2010;467:967–71. https://doi.org/10.1038/nature09447
Wu B, Zhang S, Guo Z, Bi Y, Zhou M, Li P, et al. The TGF-beta superfamily cytokine Activin-A is induced during autoimmune neuroinflammation and drives pathogenic Th17 cell differentiation. Immunity. 2021;54:308–323:e306. https://doi.org/10.1016/j.immuni.2020.12.010
Lee Y, Awasthi A, Yosef N, Quintana FJ, Xiao S, Peters A, et al. Induction and molecular signature of pathogenic TH17 cells. Nat Immunol. 2012;13:991–9. https://doi.org/10.1038/ni.2416
Lee YK, Mukasa R, Hatton RD, Weaver CT. Developmental plasticity of Th17 and Treg cells. Curr Opin Immunol. 2009;21:274–80. https://doi.org/10.1016/j.coi.2009.05.021
Stritesky GL, Yeh N, Kaplan MH. IL-23 promotes maintenance but not commitment to the Th17 lineage. J Immunol. 2008;181:5948–55. https://doi.org/10.4049/jimmunol.181.9.5948
Gaublomme JT, Yosef N, Lee Y, Gertner RS, Yang LV, Wu C, et al. Single-cell genomics unveils critical regulators of Th17 cell pathogenicity. Cell. 2015;163:1400–12. https://doi.org/10.1016/j.cell.2015.11.009
Wang C, Yosef N, Gaublomme J, Wu C, Lee Y, Clish CB, et al. CD5L/AIM regulates lipid biosynthesis and restrains Th17 cell pathogenicity. Cell. 2015;163:1413–27. https://doi.org/10.1016/j.cell.2015.10.068
Zhou L, Lopes JE, Chong MM, Ivanov II, Min R, Victora GD, et al. TGF-β-induced Foxp3 inhibits TH17 cell differentiation by antagonizing RORγt function. Nature. 2008;453:236–40. https://doi.org/10.1038/nature06878
Zhang F, Meng G, Strober W. Interactions among the transcription factors Runx1, RORgammat and Foxp3 regulate the differentiation of interleukin 17-producing T cells. Nat Immunol. 2008;9:1297–306. https://doi.org/10.1038/ni.1663
Akira S. Roles of STAT3 defined by tissue-specific gene targeting. Oncogene. 2000;19:2607–11. https://doi.org/10.1038/sj.onc.1203478
Korn T, Bettelli E, Gao W, Awasthi A, Jäger A, Strom TB, et al. IL-21 initiates an alternative pathway to induce proinflammatory TH17 cells. Nature. 2007;448:484–7.
Coquet JM, Chakravarti S, Smyth MJ, Godfrey DI. Cutting edge: IL-21 is not essential for Th17 differentiation or experimental autoimmune encephalomyelitis. J Immunol. 2008;180:7097–101. https://doi.org/10.4049/jimmunol.180.11.7097
Sonderegger I, Kisielow J, Meier R, King C, Kopf M. IL-21 and IL-21R are not required for development of Th17 cells and autoimmunity in vivo. Eur J Immunol. 2008;38:1833–8. https://doi.org/10.1002/eji.200838511
Holmdahl R. IL-21 and autoimmune disease–hypothesis and reality? Eur J Immunol. 2008;38:1800–2. https://doi.org/10.1002/eji.200838529
Fina D, Sarra M, Fantini MC, Rizzo A, Caruso R, Caprioli F, et al. Regulation of gut inflammation and th17 cell response by interleukin-21. Gastroenterology. 2008;134:1038–48. https://doi.org/10.1053/j.gastro.2008.01.041
Veldhoen M, Hocking RJ, Flavell RA, Stockinger B. Signals mediated by transforming growth factor-beta initiate autoimmune encephalomyelitis, but chronic inflammation is needed to sustain disease. Nat Immunol. 2006;7:1151–6. https://doi.org/10.1038/ni1391
Chi X, Jin W, Zhao X, Xie T, Shao J, Bai X, et al. RORgammat expression in mature TH17 cells safeguards their lineage specification by inhibiting conversion to TH2 cells. Sci Adv. 2022;8:eabn7774. https://doi.org/10.1126/sciadv.abn7774
Nurieva R, Yang XO, Martinez G, Zhang Y, Panopoulos AD, Ma L, et al. Essential autocrine regulation by IL-21 in the generation of inflammatory T cells. Nature. 2007;448:480–3. https://doi.org/10.1038/nature05969
Zhou L, Ivanov II, Spolski R, Min R, Shenderov K, Egawa T, et al. IL-6 programs T(H)-17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nat Immunol. 2007;8:967–74. https://doi.org/10.1038/ni1488
Yang XO, Pappu BP, Nurieva R, Akimzhanov A, Kang HS, Chung Y, et al. T helper 17 lineage differentiation is programmed by orphan nuclear receptors ROR alpha and ROR gamma. Immunity. 2008;28:29–39. https://doi.org/10.1016/j.immuni.2007.11.016
Harris TJ, Grosso JF, Yen HR, Xin H, Kortylewski M, Albesiano E, et al. Cutting edge: An in vivo requirement for STAT3 signaling in TH17 development and TH17-dependent autoimmunity. J Immunol. 2007;179:4313–7. https://doi.org/10.4049/jimmunol.179.7.4313
Yang XO, Panopoulos AD, Nurieva R, Chang SH, Wang D, Watowich SS, et al. STAT3 regulates cytokine-mediated generation of inflammatory helper T cells. J Biol Chem. 2007;282:9358–63. https://doi.org/10.1074/jbc.C600321200
Hall JA, Pokrovskii M, Kroehling L, Kim BR, Kim SY, Wu L, et al. Transcription factor RORalpha enforces stability of the Th17 cell effector program by binding to a Rorc cis-regulatory element. Immunity. 2022;55:2027–2043:e2029. https://doi.org/10.1016/j.immuni.2022.09.013
Akimzhanov AM, Yang XO, Dong C. Chromatin remodeling of interleukin-17 (IL-17)-IL-17F cytokine gene locus during inflammatory helper T cell differentiation. J Biol Chem. 2007;282:5969–72. https://doi.org/10.1074/jbc.C600322200
Wei L, Laurence A, Elias KM, O’Shea JJ. IL-21 is produced by Th17 cells and drives IL-17 production in a STAT3-dependent manner. J Biol Chem. 2007;282:34605–10. https://doi.org/10.1074/jbc.M705100200
Chen Z, Laurence A, Kanno Y, Pacher-Zavisin M, Zhu BM, Tato C, et al. Selective regulatory function of Socs3 in the formation of IL-17-secreting T cells. Proc Natl Acad Sci USA. 2006;103:8137–42. https://doi.org/10.1073/pnas.0600666103
Laurence A, Tato CM, Davidson TS, Kanno Y, Chen Z, Yao Z, et al. Interleukin-2 signaling via STAT5 constrains T helper 17 cell generation. Immunity. 2007;26:371–81. https://doi.org/10.1016/j.immuni.2007.02.009
Lohoff M, Mittrücker HW, Prechtl S, Bischof S, Sommer F, Kock S, et al. Dysregulated T helper cell differentiation in the absence of interferon regulatory factor 4. Proc Natl Acad Sci USA. 2002;99:11808–12. https://doi.org/10.1073/pnas.182425099
Rengarajan J, Mowen KA, McBride KD, Smith ED, Singh H, Glimcher LH. Interferon regulatory factor 4 (IRF4) interacts with NFATc2 to modulate interleukin 4 gene expression. J Exp Med. 2002;195:1003–12. https://doi.org/10.1084/jem.20011128
Hu CM, Jang SY, Fanzo JC, Pernis AB. Modulation of T cell cytokine production by interferon regulatory factor-4. J Biol Chem. 2002;277:49238–46. https://doi.org/10.1074/jbc.M205895200
Brüstle A, Heink S, Huber M, Rosenplänter C, Stadelmann C, Yu P, et al. The development of inflammatory T(H)-17 cells requires interferon-regulatory factor 4. Nat Immunol. 2007;8:958–66. https://doi.org/10.1038/ni1500
Moisan J, Grenningloh R, Bettelli E, Oukka M, Ho IC. Ets-1 is a negative regulator of Th17 differentiation. J Exp Med. 2007;204:2825–35. https://doi.org/10.1084/jem.20070994
Ciofani M, Madar A, Galan C, Sellars M, Mace K, Pauli F, et al. A validated regulatory network for Th17 cell specification. Cell. 2012;151:289–303. https://doi.org/10.1016/j.cell.2012.09.016
Rutz S, Noubade R, Eidenschenk C, Ota N, Zeng W, Zheng Y, et al. Transcription factor c-Maf mediates the TGF-beta-dependent suppression of IL-22 production in T(H)17 cells. Nat Immunol. 2011;12:1238–45. https://doi.org/10.1038/ni.2134
Bauquet AT, Jin H, Paterson AM, Mitsdoerffer M, Ho IC, Sharpe AH, et al. The costimulatory molecule ICOS regulates the expression of c-Maf and IL-21 in the development of follicular T helper cells and TH-17 cells. Nat Immunol. 2009;10:167–75. https://doi.org/10.1038/ni.1690
Tanaka S, Suto A, Iwamoto T, Kashiwakuma D, Kagami S, Suzuki K, et al. Sox5 and c-Maf cooperatively induce Th17 cell differentiation via RORgammat induction as downstream targets of Stat3. J Exp Med. 2014;211:1857–74. https://doi.org/10.1084/jem.20130791
Minns D, Smith KJ, Alessandrini V, Hardisty G, Melrose L, Jackson-Jones L, et al. The neutrophil antimicrobial peptide cathelicidin promotes Th17 differentiation. Nat Commun. 2021;12:1285. https://doi.org/10.1038/s41467-021-21533-5
Veldhoen M, Hirota K, Westendorf AM, Buer J, Dumoutier L, Renauld JC, et al. The aryl hydrocarbon receptor links TH17-cell-mediated autoimmunity to environmental toxins. Nature. 2008;453:106–9. https://doi.org/10.1038/nature06881
Dang EV, Barbi J, Yang HY, Jinasena D, Yu H, Zheng Y, et al. Control of T(H)17/T(reg) balance by hypoxia-inducible factor 1. Cell. 2011;146:772–84. https://doi.org/10.1016/j.cell.2011.07.033
Shi LZ, Wang R, Huang G, Vogel P, Neale G, Green DR, et al. HIF1alpha-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J Exp Med. 2011;208:1367–76. https://doi.org/10.1084/jem.20110278
Esplugues E, Huber S, Gagliani N, Hauser AE, Town T, Wan YY, et al. Control of TH17 cells occurs in the small intestine. Nature. 2011;475:514–8. https://doi.org/10.1038/nature10228
Berod L, Friedrich C, Nandan A, Freitag J, Hagemann S, Harmrolfs K, et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat Med. 2014;20:1327–33. https://doi.org/10.1038/nm.3704
Chang SH, Chung Y, Dong C. Vitamin D suppresses Th17 cytokine production by inducing C/EBP homologous protein (CHOP) expression. J Biol Chem. 2010;285:38751–5. https://doi.org/10.1074/jbc.C110.185777
Krueger GG, Langley RG, Leonardi C, Yeilding N, Guzzo C, Wang Y, et al. A human interleukin-12/23 monoclonal antibody for the treatment of psoriasis. N. Engl J Med. 2007;356:580–92. https://doi.org/10.1056/NEJMoa062382
Kirkham BW, Lassere MN, Edmonds JP, Juhasz KM, Bird PA, Lee CS. et al. Synovial membrane cytokine expression is predictive of joint damage progression in rheumatoid arthritis: a two-year prospective study (the DAMAGE study cohort). Arthritis Rheum.2006;54:1122–31. https://doi.org/10.1002/art.21749.
Matusevicius D, Kivisäkk P, He B, Kostulas N, Ozenci V, Fredrikson S, et al. Interleukin-17 mRNA expression in blood and CSF mononuclear cells is augmented in multiple sclerosis. Mult Scler. 1999;5:101–4. https://doi.org/10.1177/135245859900500206
Duerr RH, Taylor KD, Brant SR, Rioux JD, Silverberg MS, Daly MJ, et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science. 2006;314:1461–3. https://doi.org/10.1126/science.1135245
Molet S, Hamid Q, Davoine F, Nutku E, Taha R, Pagé N, et al. IL-17 is increased in asthmatic airways and induces human bronchial fibroblasts to produce cytokines. J Allergy Clin Immunol. 2001;108:430–8. https://doi.org/10.1067/mai.2001.117929
Barczyk A, Pierzchala W, Sozanska E. Interleukin-17 in sputum correlates with airway hyperresponsiveness to methacholine. Respir Med. 2003;97:726–33. https://doi.org/10.1053/rmed.2003.1507
Torimoto K, Okada Y, Nakayamada S, Kubo S, Kurozumi A, Narisawa M, et al. Comprehensive immunophenotypic analysis reveals the pathological involvement of Th17 cells in Graves’ disease. Sci Rep. 2022;12:16880. https://doi.org/10.1038/s41598-022-19556-z
Heidt S, Segundo DS, Chadha R, Wood KJ. The impact of Th17 cells on transplant rejection and the induction of tolerance. Curr Opin Organ Transpl. 2010;15:456–61. https://doi.org/10.1097/MOT.0b013e32833b9bfb
Abadja F, Sarraj B, Ansari MJ. Significance of T helper 17 immunity in transplantation. Curr Opin Organ Transpl. 2012;17:8–14. https://doi.org/10.1097/MOT.0b013e32834ef4e4
Milner JD, Brenchley JM, Laurence A, Freeman AF, Hill BJ, Elias KM, et al. Impaired T(H)17 cell differentiation in subjects with autosomal dominant hyper-IgE syndrome. Nature. 2008;452:773–6. https://doi.org/10.1038/nature06764
Pène J, Chevalier S, Preisser L, Vénéreau E, Guilleux MH, Ghannam S, et al. Chronically inflamed human tissues are infiltrated by highly differentiated Th17 lymphocytes. J Immunol. 2008;180:7423–30. https://doi.org/10.4049/jimmunol.180.11.7423
Kotake S, Udagawa N, Takahashi N, Matsuzaki K, Itoh K, Ishiyama S, et al. IL-17 in synovial fluids from patients with rheumatoid arthritis is a potent stimulator of osteoclastogenesis. J Clin Invest. 1999;103:1345–52. https://doi.org/10.1172/JCI5703
Miranda-Carús ME, Benito-Miguel M, Balsa A, Cobo-Ibáñez T, Pérez de Ayala C, Pascual-Salcedo D, et al. Peripheral blood T lymphocytes from patients with early rheumatoid arthritis express RANKL and interleukin-15 on the cell surface and promote osteoclastogenesis in autologous monocytes. Arthritis Rheum. 2006;54:1151–64. https://doi.org/10.1002/art.21731
Sato K, Suematsu A, Okamoto K, Yamaguchi A, Morishita Y, Kadono Y, et al. Th17 functions as an osteoclastogenic helper T cell subset that links T cell activation and bone destruction. J Exp Med. 2006;203:2673–82. https://doi.org/10.1084/jem.20061775
Koenders MI, Lubberts E, van de Loo FA, Oppers-Walgreen B, van den Bersselaar L, Helsen MM, et al. Interleukin-17 acts independently of TNF-alpha under arthritic conditions. J Immunol. 2006;176:6262–9. https://doi.org/10.4049/jimmunol.176.10.6262
Koenders MI, Lubberts E, Oppers-Walgreen B, van den Bersselaar L, Helsen MM, Kolls JK, et al. Induction of cartilage damage by overexpression of T cell interleukin-17A in experimental arthritis in mice deficient in interleukin-1. Arthritis Rheum. 2005;52:975–83. https://doi.org/10.1002/art.20885
Lock C, Hermans G, Pedotti R, Brendolan A, Schadt E, Garren H, et al. Gene-microarray analysis of multiple sclerosis lesions yields new targets validated in autoimmune encephalomyelitis. Nat Med. 2002;8:500–8. https://doi.org/10.1038/nm0502-500
Ishizu T, Osoegawa M, Mei FJ, Kikuchi H, Tanaka M, Takakura Y, et al. Intrathecal activation of the IL-17/IL-8 axis in opticospinal multiple sclerosis. Brain. 2005;128:988–1002. https://doi.org/10.1093/brain/awh453
Kebir H, Kreymborg K, Ifergan I, Dodelet-Devillers A, Cayrol R, Bernard M, et al. Human TH17 lymphocytes promote blood-brain barrier disruption and central nervous system inflammation. Nat Med. 2007;13:1173–5. https://doi.org/10.1038/nm1651
Hernandez-Santos N, Gaffen SL. Th17 cells in immunity to Candida albicans. Cell Host Microbe. 2012;11:425–35. https://doi.org/10.1016/j.chom.2012.04.008
Ma CS, Chew GY, Simpson N, Priyadarshi A, Wong M, Grimbacher B, et al. Deficiency of Th17 cells in hyper IgE syndrome due to mutations in STAT3. J Exp Med. 2008;205:1551–7. https://doi.org/10.1084/jem.20080218
Conti HR, Gaffen SL. IL-17-mediated immunity to the opportunistic fungal pathogen candida albicans. J Immunol. 2015;195:780–8. https://doi.org/10.4049/jimmunol.1500909
Infante-Duarte C, Horton HF, Byrne MC, Kamradt T. Microbial lipopeptides induce the production of IL-17 in Th cells. J Immunol. 2000;165:6107–15. https://doi.org/10.4049/jimmunol.165.11.6107
Ye P, Rodriguez FH, Kanaly S, Stocking KL, Schurr J, Schwarzenberger P, et al. Requirement of interleukin 17 receptor signaling for lung CXC chemokine and granulocyte colony-stimulating factor expression, neutrophil recruitment, and host defense. J Exp Med. 2001;194:519–27. https://doi.org/10.1084/jem.194.4.519
Chung DR, Kasper DL, Panzo RJ, Chitnis T, Grusby MJ, Sayegh MH, et al. CD4 + T cells mediate abscess formation in intra-abdominal sepsis by an IL-17-dependent mechanism. J Immunol. 2003;170:1958–63. https://doi.org/10.4049/jimmunol.170.4.1958
Khader SA, Bell GK, Pearl JE, Fountain JJ, Rangel-Moreno J, Cilley GE, et al. IL-23 and IL-17 in the establishment of protective pulmonary CD4 + T cell responses after vaccination and during Mycobacterium tuberculosis challenge. Nat Immunol. 2007;8:369–77. https://doi.org/10.1038/ni1449
Rudner XL, Happel KI, Young EA, Shellito JE. Interleukin-23 (IL-23)-IL-17 cytokine axis in murine Pneumocystis carinii infection. Infect Immun. 2007;75:3055–61. https://doi.org/10.1128/IAI.01329-06
Huang W, Na L, Fidel PL, Schwarzenberger P. Requirement of interleukin-17A for systemic anti-Candida albicans host defense in mice. J Infect Dis. 2004;190:624–31. https://doi.org/10.1086/422329
Gaffen SL, Hernandez-Santos N, Peterson AC. IL-17 signaling in host defense against Candida albicans. Immunol Res. 2011;50:181–7. https://doi.org/10.1007/s12026-011-8226-x
Romani L. Immunity to fungal infections. Nat Rev Immunol. 2011;11:275–88. https://doi.org/10.1038/nri2939
Ogongo P, Tezera LB, Ardain A, Nhamoyebonde S, Ramsuran D, Singh A, et al. Tissue-resident-like CD4 + T cells secreting IL-17 control Mycobacterium tuberculosis in the human lung. J Clin Invest. 131, https://doi.org/10.1172/JCI142014 (2021).
Acosta-Rodriguez EV, Rivino L, Geginat J, Jarrossay D, Gattorno M, Lanzavecchia A, et al. Surface phenotype and antigenic specificity of human interleukin 17-producing T helper memory cells. Nat Immunol. 2007;8:639–46. https://doi.org/10.1038/ni1467
Eyerich K, Foerster S, Rombold S, Seidl HP, Behrendt H, Hofmann H, et al. Patients with chronic mucocutaneous candidiasis exhibit reduced production of Th17-associated cytokines IL-17 and IL-22. J Invest Dermatol. 2008;128:2640–5. https://doi.org/10.1038/jid.2008.139
Martonik D, Parfieniuk-Kowerda A, Rogalska, M, Flisiak R. The role of Th17 response in COVID-19. Cells. 10, https://doi.org/10.3390/cells10061550 (2021).
Nathan A, Beynor JI, Baglaenko Y, Suliman S, Ishigaki K, Asgari S, et al. Multimodally profiling memory T cells from a tuberculosis cohort identifies cell state associations with demographics, environment and disease. Nat Immunol. 2021;22:781–93. https://doi.org/10.1038/s41590-021-00933-1
Gosselin A, Monteiro P, Chomont N, Diaz-Griffero F, Said EA, Fonseca S, et al. Peripheral blood CCR4 + CCR6+ and CXCR3 + CCR6 + CD4 + T cells are highly permissive to HIV-1 infection. J Immunol. 2010;184:1604–16. https://doi.org/10.4049/jimmunol.0903058
Dubin PJ, Kolls JK. IL-23 mediates inflammatory responses to mucoid Pseudomonas aeruginosa lung infection in mice. Am J Physiol Lung Cell Mol Physiol. 2007;292:L519–528. https://doi.org/10.1152/ajplung.00312.2006
Romani L, Fallarino F, De Luca A, Montagnoli C, D'Angelo C, Zelante T, et al. Defective tryptophan catabolism underlies inflammation in mouse chronic granulomatous disease. Nature. 2008;451:211–5. https://doi.org/10.1038/nature06471
Crome SQ, Wang AY, Levings MK. Translational mini-review series on Th17 cells: function and regulation of human T helper 17 cells in health and disease. Clin Exp Immunol. 2010;159:109–19. https://doi.org/10.1111/j.1365-2249.2009.04037.x
Mitsdoerffer M, Lee Y, Jäger A, Kim HJ, Korn T, Kolls JK, et al. Proinflammatory T helper type 17 cells are effective B-cell helpers. Proc Natl Acad Sci USA. 2010;107:14292–7. https://doi.org/10.1073/pnas.1009234107
Patakas A, Benson RA, Withers DR, Conigliaro P, McInnes IB, Brewer JM, et al. Th17 effector cells support B cell responses outside of germinal centres. PLoS One. 2012;7:e49715 https://doi.org/10.1371/journal.pone.0049715
Ouyang W, Kolls JK, Zheng Y. The biological functions of T helper 17 cell effector cytokines in inflammation. Immunity. 2008;28:454–67. https://doi.org/10.1016/j.immuni.2008.03.004
Ouyang W, O’Garra A. IL-10 family cytokines IL-10 and IL-22: from basic science to clinical translation. Immunity. 2019;50:871–91. https://doi.org/10.1016/j.immuni.2019.03.020
Dudakov JA, Hanash AM, van den Brink MR. Interleukin-22: immunobiology and pathology. Annu Rev Immunol. 2015;33:747–85. https://doi.org/10.1146/annurev-immunol-032414-112123
Hirota K, Yoshitomi H, Hashimoto M, Maeda S, Teradaira S, Sugimoto N, et al. Preferential recruitment of CCR6-expressing Th17 cells to inflamed joints via CCL20 in rheumatoid arthritis and its animal model. J Exp Med. 2007;204:2803–12. https://doi.org/10.1084/jem.20071397
Annunziato F, Cosmi L, Santarlasci V, Maggi L, Liotta F, Mazzinghi B, et al. Phenotypic and functional features of human Th17 cells. J Exp Med. 2007;204:1849–61. https://doi.org/10.1084/jem.20070663
Brockmann L, Giannou AD, Gagliani N, Huber S. Regulation of TH17 cells and associated cytokines in wound healing, tissue regeneration, and carcinogenesis. Int J Mol Sci. 18, https://doi.org/10.3390/ijms18051033 (2017).
Zenewicz LA, Yancopoulos GD, Valenzuela DM, Murphy AJ, Karow M, Flavell RA. Interleukin-22 but not interleukin-17 provides protection to hepatocytes during acute liver inflammation. Immunity. 2007;27:647–59. https://doi.org/10.1016/j.immuni.2007.07.023
Zenewicz LA, Yancopoulos GD, Valenzuela DM, Murphy AJ, Stevens S, Flavell RA. Innate and adaptive interleukin-22 protects mice from inflammatory bowel disease. Immunity. 2008;29:947–57. https://doi.org/10.1016/j.immuni.2008.11.003
Wolk K, Witte E, Wallace E, Döcke WD, Kunz S, Asadullah K, et al. IL-22 regulates the expression of genes responsible for antimicrobial defense, cellular differentiation, and mobility in keratinocytes: a potential role in psoriasis. Eur J Immunol. 2006;36:1309–23. https://doi.org/10.1002/eji.200535503
Rutz S, Eidenschenk C, Ouyang W. IL-22, not simply a Th17 cytokine. Immunol Rev. 2013;252:116–32. https://doi.org/10.1111/imr.12027
Guglani L, Khader SA. Th17 cytokines in mucosal immunity and inflammation. Curr Opin HIV AIDS. 2010;5:120–7. https://doi.org/10.1097/COH.0b013e328335c2f6
Blaschitz C, Raffatellu M. Th17 cytokines and the gut mucosal barrier. J Clin Immunol. 2010;30:196–203. https://doi.org/10.1007/s10875-010-9368-7
Favre D, Lederer S, Kanwar B, Ma ZM, Proll S, Kasakow Z, et al. Critical loss of the balance between Th17 and T regulatory cell populations in pathogenic SIV infection. PLoS Pathog. 2009;5:e1000295. https://doi.org/10.1371/journal.ppat.1000295
Pallikkuth S, Micci L, Ende ZS, Iriele RI, Cervasi B, Lawson B, et al. Maintenance of intestinal Th17 cells and reduced microbial translocation in SIV-infected rhesus macaques treated with interleukin (IL)-21. PLoS Pathog. 2013;9:e1003471. https://doi.org/10.1371/journal.ppat.1003471
Fung TC, Artis D, Sonnenberg GF. Anatomical localization of commensal bacteria in immune cell homeostasis and disease. Immunol Rev. 2014;260:35–49. https://doi.org/10.1111/imr.12186
Bixler SL, Mattapallil JJ. Loss and dysregulation of Th17 cells during HIV infection. Clin Dev Immunol. 2013;2013:852418. https://doi.org/10.1155/2013/852418
Brenchley JM, Price DA, Schacker TW, Asher TE, Silvestri G, Rao S, et al. Microbial translocation is a cause of systemic immune activation in chronic HIV infection. Nat Med. 2006;12:1365–71. https://doi.org/10.1038/nm1511
Charles KA, Kulbe H, Soper R, Escorcio-Correia M, Lawrence T, Schultheis A, et al. The tumor-promoting actions of TNF-alpha involve TNFR1 and IL-17 in ovarian cancer in mice and humans. J Clin Invest. 2009;119:3011–23. https://doi.org/10.1172/JCI39065
Derhovanessian E, Adams V, Hähnel K, Groeger A, Pandha H, Ward S, et al. Pretreatment frequency of circulating IL-17 + CD4 + T-cells, but not Tregs, correlates with clinical response to whole-cell vaccination in prostate cancer patients. Int J Cancer. 2009;125:1372–9. https://doi.org/10.1002/ijc.24497
Kuang DM, Peng C, Zhao Q, Wu Y, Chen MS, Zheng L. Activated monocytes in peritumoral stroma of hepatocellular carcinoma promote expansion of memory T helper 17 cells. Hepatology. 2010;51:154–64. https://doi.org/10.1002/hep.23291
Muranski P, Boni A, Antony PA, Cassard L, Irvine KR, Kaiser A, et al. Tumor-specific Th17-polarized cells eradicate large established melanoma. Blood. 2008;112:362–73. https://doi.org/10.1182/blood-2007-11-120998
Martin-Orozco N, Muranski P, Chung Y, Yang XO, Yamazaki T, Lu S, et al. T helper 17 cells promote cytotoxic T cell activation in tumor immunity. Immunity. 2009;31:787–98. https://doi.org/10.1016/j.immuni.2009.09.014
Muranski P, Borman ZA, Kerkar SP, Klebanoff CA, Ji Y, Sanchez-Perez L, et al. Th17 cells are long lived and retain a stem cell-like molecular signature. Immunity. 2011;35:972–85. https://doi.org/10.1016/j.immuni.2011.09.019
Luckey CJ, Weaver CT. Stem-cell-like qualities of immune memory; CD4 + T cells join the party. Cell Stem Cell. 2012;10:107–8. https://doi.org/10.1016/j.stem.2012.01.011
Hong HS, Mbah NE, Shan M, Loesel K, Lin L, Sajjakulnukit P, et al. OXPHOS promotes apoptotic resistance and cellular persistence in TH17 cells in the periphery and tumor microenvironment. Sci Immunol. 2022;7:eabm8182. https://doi.org/10.1126/sciimmunol.abm8182
Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR, et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15:1016–22. https://doi.org/10.1038/nm.2015
Zou W, Restifo NP. T(H)17 cells in tumour immunity and immunotherapy. Nat Rev Immunol. 2010;10:248–56. https://doi.org/10.1038/nri2742
Veldhoen M, Hocking RJ, Atkins CJ, Locksley RM, Stockinger B. TGFbeta in the context of an inflammatory cytokine milieu supports de novo differentiation of IL-17-producing T cells. Immunity. 2006;24:179–89. https://doi.org/10.1016/j.immuni.2006.01.001
Bettelli E, Carrier Y, Gao W, Korn T, Strom TB, Oukka M, et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature. 2006;441:235–8. https://doi.org/10.1038/nature04753
Mangan PR, Harrington LE, O'Quinn DB, Helms WS, Bullard DC, Elson CO, et al. Transforming growth factor-β induces development of the TH17 lineage. Nature. 2006;441:231–4. https://doi.org/10.1038/nature04754
Pasare C, Medzhitov R. Toll pathway-dependent blockade of CD4 + CD25 + T cell-mediated suppression by dendritic cells. Science. 2003;299:1033–6. https://doi.org/10.1126/science.1078231
Qin H, Wang L, Feng T, Elson CO, Niyongere SA, Lee SJ, et al. TGF-β promotes Th17 cell development through inhibition of SOCS3. J Immunol. 2009;183:97–105.
Malhotra N, Robertson E, Kang J. SMAD2 is essential for TGF-beta-mediated Th17 cell generation. J Biol Chem. 2010;285:29044–8. https://doi.org/10.1074/jbc.C110.156745
Yoon J-H, Sudo K, Kuroda M, Kato M, Lee IK, Han JS, et al. Phosphorylation status determines the opposing functions of Smad2/Smad3 as STAT3 cofactors in TH17 differentiation. Nat Commun. 2015;6:7600. https://doi.org/10.1038/ncomms8600
Tanaka S, Jiang Y, Martinez GJ, Tanaka K, Yan X, Kurosaki T, et al. Trim33 mediates the proinflammatory function of Th17 cells. J Exp Med. 2018;215:1853–68. https://doi.org/10.1084/jem.20170779
Lu L, Wang J, Zhang F, Chai Y, Brand D, Wang X, et al. Role of SMAD and Non-SMAD signals in the development of Th17 and regulatory T cells. J Immunol. 2010;184:4295–306. https://doi.org/10.4049/jimmunol.0903418
Chang X, Liu F, Wang X, Lin A, Zhao H, Su B. The kinases MEKK2 and MEKK3 regulate transforming growth factor-beta-mediated helper T cell differentiation. Immunity. 2011;34:201–12. https://doi.org/10.1016/j.immuni.2011.01.017
Hahn JN, Falck VG, Jirik FR. Smad4 deficiency in T cells leads to the Th17-associated development of premalignant gastroduodenal lesions in mice. J Clin Investig. 2011;121:4030–42.
Zhang S, Zhang G, Wan YY. SKI and SMAD4 are essential for IL-21-induced Th17 differentiation. Mol Immunol. 2019;114:260–8. https://doi.org/10.1016/j.molimm.2019.07.029
Wang X, Ni L, Wan S, Zhao X, Ding X, Dejean A, et al. Febrile temperature critically controls the differentiation and pathogenicity of T helper 17 cells. Immunity. 2020;52:328–341:e325. https://doi.org/10.1016/j.immuni.2020.01.006
Sun Y, Liu X, Eaton EN, Lane WS, Lodish HF, Weinberg RA. Interaction of the ski oncoprotein with Smad3 regulates TGF-β signaling. Mol Cell. 1999;4:499–509. https://doi.org/10.1016/S1097-2765(00)80201-4
Tecalco-Cruz AC, Ríos-López DG, Vázquez-Victorio G, Rosales-Alvarez RE, Macías-Silva M. Transcriptional cofactors Ski and SnoN are major regulators of the TGF-β/Smad signaling pathway in health and disease. Signal Transduct Target Ther. 2018;3:15. https://doi.org/10.1038/s41392-018-0015-8
Deheuninck J, Luo K. Ski and SnoN, potent negative regulators of TGF-beta signaling. Cell Res. 2009;19:47–57. https://doi.org/10.1038/cr.2008.324
Patel DD, Kuchroo VK. Th17 cell pathway in human immunity: lessons from genetics and therapeutic interventions. Immunity. 2015;43:1040–51.
Gaffen SL, Jain R, Garg AV, Cua DJ. The IL-23–IL-17 immune axis: from mechanisms to therapeutic testing. Nat Rev Immunol. 2014;14:585–600.
Schwartz DM, Kanno Y, Villarino A, Ward M, Gadina M, O’Shea JJ. JAK inhibition as a therapeutic strategy for immune and inflammatory diseases. Nat Rev Drug Discov. 2017;16:843–62.
Leipe J, Grunke M, Dechant C, Reindl C, Kerzendorf U, Schulze-Koops H, et al. Role of Th17 cells in human autoimmune arthritis. Arthritis Rheumatism. 2010;62:2876–85.
Nakae S, Nambu A, Sudo K, Iwakura Y. Suppression of immune induction of collagen-induced arthritis in IL-17-deficient mice. J Immunol. 2003;171:6173–7.
Lubberts E, Koenders MI, Oppers-Walgreen B, van den Bersselaar L, Coenen-de Roo CJ, Joosten LA, et al. Treatment with a neutralizing anti‐murine interleukin‐17 antibody after the onset of collagen‐induced arthritis reduces joint inflammation, cartilage destruction, and bone erosion. Arthritis Rheumatism. 2004;50:650–9.
Yasuda K, Takeuchi Y, Hirota K. The pathogenicity of Th17 cells in autoimmune diseases. Semin Immunopathol. 2019;41:283–97. https://doi.org/10.1007/s00281-019-00733-8
Wu B, Zhang S, Guo Z, Bi Y, Zhou M, Li P, et al. The TGF-β superfamily cytokine Activin-A is induced during autoimmune neuroinflammation and drives pathogenic Th17 cell differentiation. Immunity. 2021;54:308–323.e306. https://doi.org/10.1016/j.immuni.2020.12.010
Li P, Guo Z, Wan YY. SKI expression suppresses pathogenic Th17 cell response and mitigates experimental autoimmune encephalomyelitis. Front Immunol. 2021;12:707899. https://doi.org/10.3389/fimmu.2021.707899
Choi G, Park YJ, Cho M, Moon H, Kim D, Kang CY, et al. A critical role for Th17 cell-derived TGF-β1 in regulating the stability and pathogenicity of autoimmune Th17 cells. Exp Mol Med. 2021;53:993–1004. https://doi.org/10.1038/s12276-021-00632-9
Das J, Ren G, Zhang L, Roberts AI, Zhao X, Bothwell AL, et al. Transforming growth factor β is dispensable for the molecular orchestration of Th17 cell differentiation. J Exp Med. 2009;206:2407–16. https://doi.org/10.1084/jem.20082286
Xu L, Kitani A, Fuss I, Strober W. Cutting edge: regulatory T cells induce CD4 + CD25-Foxp3- T cells or are self-induced to become Th17 cells in the absence of exogenous TGF-beta. J Immunol. 2007;178:6725–9. https://doi.org/10.4049/jimmunol.178.11.6725
Noack M, Miossec P. Th17 and regulatory T cell balance in autoimmune and inflammatory diseases. Autoimmun Rev. 2014;13:668–77. https://doi.org/10.1016/j.autrev.2013.12.004
Zhou X, Bailey-Bucktrout SL, Jeker LT, Penaranda C, Martínez-Llordella M, Ashby M, et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nat Immunol. 2009;10:1000–7. https://doi.org/10.1038/ni.1774
Komatsu N, Okamoto K, Sawa S, Nakashima T, Oh-hora M, Kodama T, et al. Pathogenic conversion of Foxp3+ T cells into TH17 cells in autoimmune arthritis. Nat Med. 2014;20:62–68. https://doi.org/10.1038/nm.3432
Huber S, Stahl FR, Schrader J, Lüth S, Presser K, Carambia A, et al. Activin a promotes the TGF-beta-induced conversion of CD4 + CD25- T cells into Foxp3+ induced regulatory T cells. J Immunol. 2009;182:4633–40. https://doi.org/10.4049/jimmunol.0803143
Annes JP, Munger JS, Rifkin DB. Making sense of latent TGFβ activation. J Cell Sci. 2003;116:217–24.
Lawrence DA. Latent-TGF-β: an overview. Mol Cell Biochem. 2001;219:163–70.
Acknowledgements
We thank Anthony Chen, Wei-chun Chou, Yongbin He, and Brandon Le for the critical reading and editing of the manuscript. We apologize to colleagues whose work we were unable to discuss and cite due to space limitations. This work was supported by the NIH (R01 AI160774, R01 AI123193, and R56 AG071256) and the National Multiple Sclerosis Society (RG-1802-30483 to YYW). The figures were created using BioRender.com.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Zhao, X. & Wan, Y.Y. Intricacies of TGF-β signaling in Treg and Th17 cell biology. Cell Mol Immunol 20, 1002–1022 (2023). https://doi.org/10.1038/s41423-023-01036-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-023-01036-7
- Springer Nature Limited
Keywords
This article is cited by
-
Th17/1 and ex-Th17 cells are detected in patients with polyarticular juvenile arthritis and increase following treatment
Pediatric Rheumatology (2024)
-
The dynamic shifts of IL-10-producing Th17 and IL-17-producing Treg in health and disease: a crosstalk between ancient "Yin-Yang" theory and modern immunology
Cell Communication and Signaling (2024)
-
Multiple treatments with human embryonic stem cell-derived mesenchymal progenitor cells preserved the fertility and ovarian function of perimenopausal mice undergoing natural aging
Stem Cell Research & Therapy (2024)
-
Reconstruction of the TGF-β signaling pathway of Fasciola gigantica
Parasitology Research (2024)
-
Immunological modulation in health and disease
Cellular & Molecular Immunology (2023)