Abstract
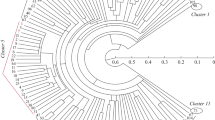
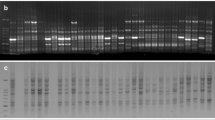
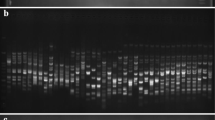
Genetic variation among five elite winter barley cultivars (H. vulgare L.) currently grown in Bulgaria was assessed at the molecular level using restriction fragment length polymorphism (RFLP) and randomly amplified polymorphic DNA (RAPD) markers. The present study sampled RFLPs in four well characterized multigene families in barley: the seed storage protein loci; the 18S, 5.8S and 26S ribosomal DNA loci; the loci coding for 5S ribosomal RNA and the loci coding subunit α of ATP-A complex in the mitochondrial genome. RFLPs were detected in three out of five investigated chromosomal loci in the barley cultivars studied. RAPD assay using arbitrary 10-base primers was applied to generate amplified length polymorphic markers in barley. Overall a total of 15 polymorphic phenotypes were found among the studied barley cultivars by using 11 out of 25 tested primers. All RAPDs were considered as dominant genetic markers except for two, where PCR and Southern blot analysis indicated the presence of codominant amplification products. Five RAPD polymorphisms in F1 and F2 progenies of the cross between Alpha and Obzor were inherited in Mendelian fashion. The determined values for the genetic variation proved a high genetic similarity among the tested cultivars. Genetic similarity (GS) calculated from RFLP and RAPD data ranged from 0.888 to 0.997 with a mean GS – 0.933.
Similar content being viewed by others
References
Allard, R.W., M.A. Saghai-Maroof, Q. Zhang & R.A. Jorgensen, 1990. Genetic and molecular organization of ribosomal DNA (rDNA) variants in wild and cultivated barley. Genetics 126: 743-751.
Appels, R., W.L. Gerlach, E.S. Dennis, H. Swift & W.J. Peacock, 1980. Molecular and chromosomal organisation of DNA sequences coding for the ribosomal RNAs in cereals. Chromosoma 78: 293-311.
Backes, G., A. Graner, B. Foroughi-Wehr, Fischbeck, G. Wenzel & A. Jahoor, 1995. Localization of quantitative trait loci (QTL) for agronomic important characters by the use of a RFLP map in barley (Hordeum vulgare L.). Theor Appl Genet 90: 294-302.
Beker, J. & M. Heun, 1995. Barley microsatellites: allele variation and mapping. Plant Mol Biol 27: 835-845.
Boer, P.M., J.E. McIntosh, M.W. Gray & L. Bonen, 1985. The wheat mitochondrial gene for apocytochrome b: Absence of prokaryotic ribosome binding site. Nucl Acids Res 13: 2281-2292.
Bonen, L., P.M. Boer & M.W. Gray, 1984. The wheat cytochrome oxidase subunit II gene has an intron insert and three radical amino acid changes relative to maize. EMBO J 3: 2531-2536.
Carlson, J.E., L.K. Tuisieram, J.C. Glaubitz, V.W.K. Luk, C. Kauffeldt & R. Rutledge, 1991. Segregation of random amplified polymorphic DNA markers in F1 progeny of conifers. Theor Appl Genet 83: 194-200.
Casas, M.A., E. Igartua, M.P. Valles & J.L. Molina-Cano, 1998. Genetic diversity of barley cultivars grown in Spain by RFLP, similarity and co ancestry coefficients. Plant Breed 117: 429-435.
Chvyrleva, C.D., A.K. Gazumyan & E.V. Ananiev, 1988. Organisation and the primary nucleotide sequence of the 5S rRNA gene in barley (H. vulgare). Genetika 24: 1830-1840.
Dahleen, L.S., 1997. Mapped clone sequences detecting differences among 28 North American barley cultivars. Crop Sci 37: 952-957.
Dawson, I.K., K.J. Chalmers, R. Waugh & W. Powell, 1993. Detection and analysis of genetic variation in Hordeum spontaneum population from Israel using RAPD markers. Mol Ecol 2: 151-159.
Forde, B.G., M. Kreis, M.B. Bahramian, J.M. Mattews, B.J. Miflin, R.D. Tompson, D. Bartels & R.B. Flavell, 1981. Molecular cloning and analysis of cDNA sequence derived from polyA+ RNA from barley endosperm: identification of B hordein related clones. Nucl Acids Res 9: 6689-6707.
Forde, B.G., M. Kreis, M. Williamson, P.R. Pring, J. Pywell, P.R. Shewry, N. Bunce & B.J. Milflin, 1985. Short tandem repeats shared by B and C hordein cDNA suggest a common evolutionary origin for two groups of storage protein genes. EMBO J 4: 9-15.
Gerlach, W.L. & J.R. Bedbrook, 1979. Cloning and characterisation of ribosomal DNA from wheat and barley. Nucl Acids Res 7: 1869-1885.
Gonzales, J.M. & E. Ferrer, 1993. Random amplified polymorphic DNA analysis in Hordeum spesies, Genome 36: 1029-1031.
Graner, A., A. Jahoor, G. Schondelmaier, H. Siedler, K. Pillen, G. Fishbeck, G. Wenzel & R.G. Herrmann, 1991. Construction of a RFLP map of barley. Theor Appl Genet 83: 250-256.
Graner, A., W.F. Ludwig & A.E. Melchinger, 1994. Relationships among European barley germplasm: II. Comparison of RFLP and pedigree data. Crop Sci 34: 1199-1205.
Heun, M., A.E. Kennedy, J.A. Anderson, N.L.V. Lapitan, M.E. Sorrells & S.D. Tanksley, 1991. Construction of a restriction fragment length polymorphism map for barley (Hordeum vulgare). Genome 34: 437-447.
Kleinhofs, A., A. Kilian, M.A. Saghai-Maroof, R.M. Biyashev, P. Hayes, F.Q. Chen, N. Lapitan, A. Fenwick, T.K. Blake, V. Kanazin, E. Ananiev, L., Dahleen, D. Kudrna, J. Bollinger, S.J. Knapp, B. Liu, M. Sorrells, M. Heun, J.D. Franckowiak, D. Hoffman, R. Skadsen & B.J. Steffenson, 1993. A molecular, isozyme and morphological map of the barley (Hordeum vulgare) genome. Theor Appl Genet 86: 705-712.
Kleinhofs, A. & A. Kilian, 1994. RFLP maps of barley. In: R.L. Philips & I.K. Vasil (Eds.), DNA-Based Markers in Plants, pp. 163-198. Kluwer Academic Publishers, Dordrecht.
Kolchinsky, A., V. Kanazin, E. Yakovleva, A. Gazumyan, C. Kole & E. Ananyev, 1990. 5S-RNA genes of barley are located on the second chromosome. Theor Appl Genet 80: 333-336.
Laurie, D.A., N. Pratchett, J.H. Bezant & J.W. Snape, 1995. RFLP mapping of five major genes and eight quantitative trait loci controlling flowering time in a winter × spring barley (Hordeum vulgare L.) cross. Genome 38: 575-585.
Loarce, Y., R. Gallego & E. Ferrer, 1996. A comparative analysis of the genetic relationships between rye cultivars using RFLP and RAPD markers. Euphytica 88: 107-115.
Manninen, O. & E., Nissila. 1997. Genetic diversity among Finnish six-rowed barley cultivars based on pedigree information and DNA markers. Hereditas 126: 87-93.
Martin, G.B., J.G.K. Williams & S.D. Tanksley, 1991. Rapid identification of markers linked to Pseudomonas resistance gene in tomato by using random primers and near isogenic lines. Proc Natl Acad Sci USA 88: 2336-2340.
Melchinger, A.E., A. Graner, M. Singh & M.M. Messmer, 1994. Relationships among European barley germplasm. I. Genetic diversity among winter and spring cultivars revealed by RFLPs. Crop Sci 34: 1191-1199.
Molnar, S.J, P.K. Gupta, G. Fedak & R. Wheatcroft, 1989. Ribosomal DNA repeat unit polymorphism in 25 Hordeum species. Theor Appl Genet 78: 387-392.
Molnar, S.J. & A. McKay, 1991. Restriction fragment analysis of ribosomal and hordein genes in eastern Canadian two-rowed barleys. Genome 34: 298-302.
Molnar, S.J. & A. McKay, 1995. Restriction fragment analysis of hordein genes in western Canadian two-rowed barleys. Can J Plant Sci 75: 191-193.
Nei, M. & W.H. Li, 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76: 5269-5273.
Pecchioni, N., A.M. Stanca, V. Terzi & L. Cattivelli, 1993. RFLP analysis of highly polymorphic loci in barley. Theor Appl Genet 85: 926-930.
Russell, J., J. Fuller, G. Yong, B. Thomas, G. Taramino, M. Macaulay, R. Waugh & W. Powell, 1997. Discrimination between barley genotypes using microsatellite markers. Genome 40: 442-450.
Saghai-Maroof, M.A., K.M. Soliman, R.A. Jorgensen & R.W. Allard, 1984. Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81: 8014-8018.
Saghai-Maroof, M.A., Q. Zhang & J. Chojecki, 1994. RFLPs in cultivated barley and their application in the evaluation of malting quality cultivars. Hereditas 121: 21-29.
Sambrook, J., E.F. Fritsch & T. Manniatis, 1989. Molecular Cloning: A Laboratory Manual, 2nd edn., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Shimron-Abarbanell, D. & A. Breiman, 1991. Comprehensive molecular characterization of tissue-culture-derived Hordeum marinum plants. Theor Appl Genet 83: 71-80.
Song, W. & R. Henry, 1995. Molecular analysis of the DNA polymorphism of wild barley (H. spontaneum) germplasm using the polymerase chain reaction. Gen Res Crop Evol 42: 273-281.
Spassova, M., F. Moneger, C.J. Leaver, P. Petrov, A. Atanassov, H.J.J. Nijkamp & J. Hille, 1994. Characterization and expression of the mitochondrial genome of a new type of cytoplasmic malesterile sunflower. Plant Mol Biol 26: 1819-1831.
Tinker, N.A., M.G. Fortin & D.E. Miller, 1993. Random amplified polymorphic DNA and pedigree relationships in spring barley. Theor Appl Genet 85: 976-984.
Weining, S. & P. Langridge, 1991. Identification and mapping polymorphism in cereals based on polymerase chain reaction. Theor Appl Genet 82: 209-216.
Williams, J.G.K, A.R. Kubelik, K.J. Livak, J.A. Rafalski & S.V. Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl Acids Res 18: 6531-6535.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Todorovska, E., Trifonova, A. & Atanassov, A. Genetic diversity among elite Bulgarian barley varieties evaluated by RFLP and RAPD markers. Euphytica 129, 325–336 (2003). https://doi.org/10.1023/A:1022205000732
Issue Date:
DOI: https://doi.org/10.1023/A:1022205000732