Abstract
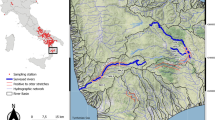
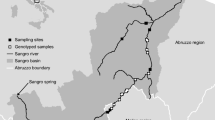
Non-invasive genetic techniques have proven to be cost-effective for monitoring and studying otter populations, largely due to the linearity of otter territories and the marking behavior of the species. After a severe decline, in the 60, the Eurasian otter is recovering in the Iberian Peninsula. However, the recovery pattern is not homogeneous and the species is still considered “Threatened” in many regions. During 2007–2010 a systematic non-invasive genetic sampling effort was carried out to determine the spatial distribution and to estimate the population size of an endangered otter population in northern Iberian Peninsula (Basque Country). Samples were identified to species level by sequencing a 226 bp mtDNA fragment prior to genotyping. Among the 132 obtained samples, 127 (98.4%) belonged to the study species, one sample was genetically identified as European mink (Mustela lutreola) and one as American mink (Neovison vison) while genetic species confirmation was not possible in the three remaining samples. These results provided novel and accurate data on species distribution, highlighting an overall increase of 25% in 10 x 10 UTM grids occupied by otter and a clear pattern of re-colonization upstream of the main rivers. All samples corresponding to otter were subsequently individually genotyped using a novel multiplex panel of 11 microsatellite markers and sexed by typing the sex-chromosome-related gene ZFX/ZFY. We obtained a complete individual genetic profile for 55 samples (genotyping success 43%), corresponding to 20 different individuals (11 females, 6 males, and 3 individuals of unknown gender). The mean otter density in occupied areas estimated to be 0.09 (0.06-0.12) individuals per river kilometer. The present study enabled us to obtain updated and relevant information about this elusive species’ distribution and population size, essential to define population status and to design successful and effective management and conservation programs.
Similar content being viewed by others
References
Aguilar, CM., Gómez-Gayubo, A., 2008. La nutria en La Rioja. In: López-Martín, J.M., Jiménez, J. (Eds.), La nutria en Espana. Veinte años de seguimiento de un mamífero amenazado. SECEM, Málaga, pp. 217–225.
Arrendal, J., Vila, C, Bjorklund, M., 2007. Reliability of noninvasive genetic census of otters compared to field censuses, Conserv Genet 8, 1097–1107.
Ben-David, M., Blundell, G.M., Kern, J.W., Maier, J.A.K., Brown, E.D., Jewett, S.C., 2005. Communication in river otters: creation of variable resource sheds for terrestrial communities, Ecology 86 (5), 1331–1345.
Beja-Pereira, A., Oliveira, R., Alves, P.C., Schwartz, M.K., Luikart, G., 2009. Advancing ecological understandings through technological transformations in noninvasive genetics, Mol. Ecol. Resour. 9 (5), 1279–1301.
Bonesi, L., Hale, M., Macdonald, D.W., 2013. Lessons for the use of non-invasive genetic sampling as a way to estimate Eurasian otter population size and sex-ratio, Acta Theriol. 58, 157–168.
Chanin, P., 1985. The Natural History of Otters. Croom Helm, Kent, pp. 179.
Caniglia, R., Fabbri, E., Galaverni, M., Milanesi, P., Randi, E., 2014. Non-invasive sampling and genetic variability, pack structure and dynamics in an expanding wolf population, J. Mammal. 95, 41–59.
Cassens, I., Tiedemann, R., Suchentrunk, F., Hartl, G.B., 2000. Mitochondrial variation in the European otter (Lutra lutra) and the use of spacial autocorrelation analysis in conservation.J, Hered. 90, 31–35.
Dallas, J.F., Bacon, P.J., Carss, D.N., Conroy, J.W.H., Green, R., Jefferies, D.J., Kruuk, H., Marshall, F., Piertney, S.B., Racey, P.A., 1999. Genetic diversity in the Eurasian otter Lutra lutra, in Scotland, Evidence from microsatellite polymorphism. Biol. J. Linnean. Soc. 68, 73–86.
Dallas, J.F., Carss, D.N., Marshall, F., Koepfli, K., Kruuk, H., Piertney, B., Bacon, P.J., 2000. Sex identification of the Eurasian otter Lutra lutra by PCRtyping of spraints, Conserv. Genet. 1, 181–183.
Dallas, J.F., Coxon, K.E., Sykes, T., Chanin, P.R.F., Marshall, F., Carss, D.N., Bacon, P., Piertney, S.B., Racey, P.A., 2003. Similar estimates of population genetic composition and sex ratio derived from carcasses and faeces of Eurasian otter Lutra lutra. Mol, Ecol. Notes 12, 275–282.
Dallas, J.F., Piertney, S.B., 1998. Microsatellite primers for the Eurasian otter, Mol. Ecol. 7, 1248–1251.
Davison, A., Chiba, S., 2003. Laboratory temperature variation is a previously unrecognized source of genotyping error during capillary electrophoresis, Mol. Ecol. Notes. 3 (2), 321–323.
Delibes, M., 1990. La Nutria (Lutra lutra) en España. ICONA, Serie Técnica, Madrid.
Diputación Foral de Álava, 2004. Orden Foral 880/2004, de 27 de octubre por la que se aprueba el Plan de Gestión de la Nutria, Lutra lutra (Linnaeus, 1758) en el Territorio Histórico de Álava. BOTHA no 136.
Effenberger, S., Suchentrunk, F., 1999. RFLP analysis of the mitochondrial DNA of otters (Lutra lutra) from Europe—implications for conservation of a flagship species, Biol. Conserv. 90, 229–234.
Eggert, L.S., Eggert, J.A., Woodruff, D.S., 2003. Estimating population sizes for elusive animals: the forest elephants of Kakum National Park Ghana, Mol. Ecol. Notes 12, 1389–1402.
Frantz, A., Roper, T.J., 2006. Simulations to assess the performance of different rarefaction methods in estimating population size using small datasets, Conserv. Genet. 7, 315–318.
Ferrando, A., Lecis, R., Domingo-Roura, X., Ponsà, M., 2008. Genetic diversity and individual identification of reintroduced otters (Lutra lutra) in north-eastern Spain by DNA genotyping of spraints, Conserv. Genet. 9, 129–139.
Ferrando, A., Ponsa, M., Marmi, J., Domingo-Roura, X., 2004. Eurasian otters Lutra lutra, have a dominant mtDNA haplotype from the Iberian Peninsula to Scandinavia, J. Hered. 95, 435–440.
Gómez-Moliner, B.J., Cabria, M.T., Rubines, J., Garin, I., Madeira, M.J., Elejalde, A., Palazón, S., 2004. PCR-RFLP identification of mustelid species: European mink (Mustela lutreola) American mink (M, vison) and polecat (M. putorius) by analysis of excremental DNA. J. Zool. 262(3), 311–316.
Gorman, M.L., Trowbridge, B.J., 1989. The role of odor in the social lives of carnivores. In: Gittleman, J.L. (Ed.), Carnivore Behavior, Ecology, and Evolution. Cornell University Press, Ithaca, New York, USA, pp. 57–88.
Hájková, P., Zemanová, B., Bryja, J., Hájek, B., Roche, K., Tkadlec, E., Zima, J., 2006. Factors affecting success of PCR amplification of microsatellite loci from otter faeces, Mol. Ecol. Notes 6, 559–562.
Hájková, P., Zemanová, B., Roche, K., Hájek, B., 2009. An evaluation of field and non invasive genetic methods for estimating Eurasian otter population size, Conserv. Genet. 10, 1667–1681.
Hernando, A., Martínez de Lecea, F., Illana, A., Bayona, J., Echegaray, J., 2005. Sondeo y evolución de la distribución de la nutria paleártica (Lutra lutra Linnaeus, 1758) en el País Vasco (N España), Galemys 1, 25–46.
Huang, C-C, Hsu, Y.-C., Lee, L.L., Li, S.-H., 2005. Isolation and characterization of tetramicrosatellite DNA markers in the Eurasian otter (Lutra lutra). Mol, Ecol. Notes 5, 314–316.
Hung, C.-M., Li, S.-H., Lee, L.-L., 2004. Faecal DNA typing determines the abundance and spacial organization of otters (Lutra lutra) along two stream systems in Kinmen, Anim. Conserv. 7, 301–311.
Ihaka, R., Gentleman, R., 1996. R: a language for data analysis and graphics. J. Comp. Graph. Stat. 5, 299–314.
Jones, O.R., Wang, J., 2010. COLONY: a program forparentage and sibship inference from multilocus genotype data, Mol. Ecol. Resour. 10 (3), 551–555.
Kalinowski, S.T., Wagner, A.P., Taper, M.L., 2006. ML-Relate: a computer program for maximum likelihood estimation of relatedness and relationship, Mol. Ecol. Notes 6, 576–579.
Kalz, B.Jewgenow, K., Fickel, J., 2006. Structure of an otter (Lutra lutra) population in Germany - results of DNA and hormone analyses from faecal samples, Mammal. Biol. 71 (6), 321–335.
Kauhala, K., 1996. Distributional history of the American mink (Mustela vison) in Finland with special reference to the trends in otter (Lutra lutra) populations, Ann. Zoolog. Fennici 33, 283–291.
Koelewijn, H.P., Pérez-Haro, M., Jansman, H.A.H., Boerwinkel, M.C, Bovernschen, D.R., 2010. The reintroduction of the Eurasian otter (Lutra lutra) into the Netherlands: hidden life revealed by noninvasive genetic monitoring, Conserv. Genet. 11, 601–614.
Kohn, M.H., York, E.C., Kamradt, D.A., Haught, G., Sauvajot, R.M., Wayen, R.K., 1999. Estimating population size by genotyping faeces, Proc. R. Soc. B. 266 (1420), 657–663.
Kranz, A., Polednik, L, Pinter, V., Parz-gollner, R., Fischotter, S., 2001. Distribution, status and conservation of otters in Lower Austria Wiss, Mitt. Niederösterr. Landesmuseum 14, 39–50.
Kruuk, H., 1992. Scent marking by otters (Lutra lutra): signaling the use of resources, Behav. Ecol. 3, 133–140.
Kruuk, H., 1995. Wild Otters Predation and Populations. Oxford Univ. Press, Oxford. Kruuk, H., 2006. Otters. Ecology Behavior and Conservation. Oxford University press, Oxford.
Lodé, T., 1993. The decline of otter Lutra lutra populations in the region of the pays de Loire, Western France, Biol. Conserv. 65 (1), 9–13.
López de Luzuriaga, J., Zuberogoitia, I., Zabala, J., 2008. La nutria en el País Vasco. In: López-Martín, J.M., Jiménez, J. (Eds.), La nutria en Espana. Veinte años de seguimiento de un mamífero amenazado. SECEM, Málaga, pp. 207–215.
López-Martín, J.M., Jiménez, J., 2008. La nutria en España. Veinte años de un mamífero amenazado. SECEM, Málaga.
Macdonald, S.M., Mason, C.F., 1983. The Otter Lutra lutra in Southern Italy, Biol. Conserv. 25 (2), 95–101.
Macdonald, S.M., Mason, C.F., 1994. Status and conservation need of the otter (Lutra lutra) in the western Paleartic, Council of Europe Press. Nat. Environ. 67, 1–54.
Mason, C.F., Macdonald, S.M., 2004. Growth in otter (Lutra lutra) populations in the UK as shown by long-term monitoring, Ambio 33, 148–152.
McDonald, R.A., Hara, K.O., Morrish, D.J., 2007. Decline of invasive alien mink (Mustela vison) is concurrent with recovery of native otters (Lutra lutra). Divers, Distrib. 13, 92–98.
Miller, C.R., Joyce, P., Waits, L.P., 2005. A new method for estimating the size of small populations from genetic mark-recapture data, Mol. Ecol. Notes 14, 1991–2005.
Mills, L., Citta, J., Lair, K., Schwartz, M., Tallmon, D., 2000. Estimating animal abundance using non-invasive DNA sampling: promise and pitfalls, J. Appl. Ecol. 10, 283–294.
Mucci, N., Arrendal, J., Ansorge, H., Bailey, M., Bodner, M., Delibes, M., Randi, E., 2010. Genetic diversity and landscape genetic structure of otter (Lutra lutra) populations in Europe, Conserv. Genet. 11, 583–599.
Mucci, N., Pertoldi, C, Madsen, A.B., Loeschcke, V., Randi, E., 1999. Extremely low mitochondrial DNA control-region sequence variation in the otter Lutra lutra population of Denmark, Hereditas 130, 331–336.
Mucci, N., Randi, E., 2007. Sex identification of Eurasian otter (Lutra lutra) noninvasive DNA samples using ZFX/ZFY sequences, Conserv. Genet. 8, 1479–1482.
O’Neill, D., Turnerm, P.D., O’Meara, D.B., Chadwick, E.A., Coffey, L., O’Reilly, C, 2013. Development of novel real-time TaqMan® PCR assays for the species and sex identification of otter (Lutra lutra) and their application to noninvasive genetic monitoring, Mol. Ecol. Resour. 13 (5), 877–883.
Palomo, L.J., Gisbert, J., 2002. Atlas de los mamíferos terrestres de Espan, Dirección General de Conservación de la Naturaleza-SECEM-SECEMU, Madrid, 564 pp. Park, H., Han, T., Kim, D., Min, M., Han, S., Kim, K., Lee, H., 2011. Individual identification and sex determination of Eurasian otters (Luta lutra) in Daegu city based on genetic analysis of otter spraints. Genes Genomics 33 (6), 653–657.
Parry, G.S., Bodger, O., McDonald, R.A., Forman, D.W., 2013. A systematic re-sampling approach to assess the probability of detecting otters Lutra lutra using spraint surveys on small lowland rivers, Ecol. Inform. 14, 64–70.
Pérez-Haro, M., Viñas, J., Mañnas, F., Batet, A., Ruiz-Olmo, J., Pla, C, 2005. Genetic variability in the complete mitochondrial control region of the Eurasian otter (Lutra lutra) in the Iberian Peninsula, Biol. J. Linn. Soc. 86 (4), 397–403.
Pompanon, F., Bonin, A., Bellemain, E., Taberlet, P., 2005. Genotyping errors: causes, consequences and solutions, Nat. Rev. Genet. 6(11), 847–849.
Prigioni, C, Remonti, L, Balestrieri, A., Sgrosso, S., Priore, G., Mucci, N., Randi, E., 2006. Estimation of European otter (Lutra lutra) population size by fecal DNA typing in southern Italy, J. Mammal. 87 (5), 855–858.
Prigioni, C., Balestrieri, A., Remonti, L., 2007. Decline and recovery in otter Lutra lutra populations in Italy, Mammal. Rev. 37 (1), 71–79.
Remonti, L., Prigioni, C., Balestrieri, A., Sgrosso, S., Priore, G., 2008. Distribution of a recolonising species may not reflect habitat suitability alone: the case of the Eurasian otter (Lutra lutra) in southern Italy, Wildl. Res. 35, 798–805.
Ruiz-González, A., Rubines, J., Berdión, O., Gómez-Moliner, B., 2008a. A non-invasive genetic method to identify the sympatric mustelids pine marten (Martes martes) and stone marten (Martes foina): preliminary distribution survey on the northern Iberian Peninsula. Eur. J. Wildl. Res. 54 (2), 253–261.
Ruiz-González, A., Ferrando, A., Pérez-Haro, M., Rubines, J., Madeira, M.J., Ponsá, M., Ruiz-Olmo, J., Romero, R., Gómez-Moliner, B.J., 2008b. Seguimientos moleculares de las poblaciones de nutria Lutra lutra (Linnaeus, 1758)a partir de muestreos no invasivos. In: López-Martín, J.M., Jiménez, J. (Eds.), La nutria en España. Veinte años de seguimiento de un mamífero amenazado. SECEM, Málaga, pp. 207–215.
Ruiz-González, A., Madeira, M.J., Randi, E., Urra, F., Gómez-Moliner, B.J., 2013. Non-invasive genetic sampling of sympatric marten species (Martes martes and Martes foina): assessing species and individual identification success rates on faecal DNAgenotyping, Eur. J. Wildl. Res 59 (3), 371–386.
Ruiz-Olmo, J., Delibes, M., 1998. La nutria en España ante el horizonte del año 2000. SECEM, Málaga.
Ruiz-Olmo, J., Loy, A., Cianfrani, C., Yoxon, P., Yoxon, G., De Silva, P.K., Roos, A., Bisther, M., Hájkova, P., Zemanová, B., 2008. Lutra lutra. In: IUCN 2013 (Ed.), IUCN Red List of Threatened Species. Version 2013.2., www.iucnredlist.org (downloaded 10.2.14).
Ruiz-Olmo, J., Batet, A., Mañas, F., Martínez-Vidal, R., 2011. Factors affecting otter (Lutra lutra) abundance and breeding success in freshwater habitats of the northeastern Iberian Peninsula, Eur. J. Wildl. Res. 57, 827–842.
Schwartz, M.K., Monfort, S.L., 2008. Genetic and endocrine tools for carnivore surveys. In: Lomg, R., MacKay, P., Ray, J., Zielinski, W. (Eds.), Noninvasive Survey Methods for North American Carnivores. Island Press, Washington, DC
Sulkava, R., 2007. Snow tracking: a relevant method for estimating otter Lutra lutra populations, Wildl. Biol. 13 (2), 208–218.
Taberlet, P., Luikart, G., 1999. Non-invasive genetic sampling and individual identification, Biol. J. Linn. Soc. 68, 41–55.
Valière, N., 2002. GIMLET: a computer program for analysing genetic individual identification data, Mol. Ecol Notes 2, 377–379.
Waits, L.P., Paetkau, D., 2005. Noninvasive genetic sampling tools for wildlife biologists: a review of applications and recommendations for accurate data collection, J. Wildl. Manag. 69, 1419–1433.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vergara, M., Ruiz-González, A., de Luzuriaga, J. et al. Individual identification and distribution assessment of otters (Lutra lutra) through non-invasive genetic sampling: Recovery of an endangered species in the Basque Country (Northern Spain). Mamm Biol 79, 259–267 (2014). https://doi.org/10.1016/j.mambio.2014.04.003
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.mambio.2014.04.003