Abstract
The biochar amendment plays a vital role in maintaining soil health largely due to its effects on soil microbial communities. However, individual cases and the variability in biochar properties are not sufficient to draw universal conclusions. The present study aimed to reveal how the biochar application affects soil microbial communities. Metadata of 525 ITS and 1288 16S rRNA sequencing samples from previous studies were reanalyzed and machine learning models were applied to explore the dynamics of soil microbial communities under biochar amendment. The results showed that biochar considerably changed the soil bacterial and fungal community composition and enhanced the relative abundances of Acidobacteriota, Firmicutes, Basidiomycota, and Mortierellomycota. Biochar enhanced the robustness of the soil microbial community but decreased the interactions between fungi and bacteria. The random forest model combined with tenfold cross-validation were used to predict biomarkers of biochar response, indicating that potentially beneficial microbes, such as Gemmatimonadetes, Microtrichales, Candidatus_Kaiserbacteria, and Pyrinomonadales, were enriched in the soil with biochar amendment, which promoted plant growth and soil nutrient cycling. In addition, the biochar amendment enhanced the ability of bacteria to biosynthesize and led to an increase in fungal nutrient patterns, resulting in an increase in the abundance and diversity of saprophytic fungi that enhance soil nutrient cycling. The machine learning model more accurately revealed how biochar affected soil microbial community than previous independent studies. Our study provides a basis for guiding the reasonable use of biochar in agricultural soil and minimizing its negative effects on soil microecosystem.
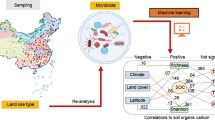
Graphical Abstract

Highlights
-
Biochar considerably affected the soil community composition of bacteria and fungi.
-
14 bacterial orders and 7 fungal families were enriched in biochar-amended soils.
-
Biochar enhanced positive interactions and the stability of soil community.
-
Biochar led to an increase in soil fungal nutrient patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Maintaining soil health is crucial for sustaining terrestrial life on Earth (Bardgett and Van Der Putten 2014). Progressive deterioration of soil conditions and poor nutrient use efficiency have severely limited agricultural productivity, posing a potential food safety risk (Singh et al. 2022; Xu et al. 2022b). To address the various issues limiting agricultural productivity, including the nutrient imbalance and inefficient water utilization, the design and functioning of agroecosystems need continuous optimization (Lal 2013). One potential solution is to return organic nutrients in plant residue, such as biochar, straw, or compost, back to the soil, which would improve the soil properties, especially the soil organic matter content.
For example, biochar, as a carbon-rich material produced by biomass pyrolysis under oxygen limited conditions, has potential as a soil amendment owing to its multiple agricultural and environmental returns (Li et al. 2018a, b), like improving soil fertility and structure (Ding et al. 2016), mitigating carbon emissions (Lehmann et al. 2021), immobilizing heavy metals (Lu et al. 2017), and adsorbing or degrading contaminants in farmland (Liu et al. 2018a; Zhu et al. 2017). Biochar is widely applied for soil improvement and environmental remediation. For instance, biochar can substantially reduce the content of cadmium in various plant systems, including soil rhizosphere, plant roots, and edible parts (Duan et al. 2023; Liu et al. 2018b). The biochar application during the growth stage of rice cultivation has been found to have a sustainable positive influence on soil properties and it has been observed to enhance rice yield (Liu et al. 2016).
The biochar application in soil can improve its biological properties by enhancing the functional activity of soil microbial community (Li et al. 2018a, b; Palansooriya et al. 2019). Soil microbial communities play a vital role in agricultural ecosystems by determining soil nutrient cycling, crop yield, and plant stress resistance (Hartmann & Six 2022). The pores in biochar provide a suitable habitat for soil microbes, facilitating their colonization and growth in the soil environment (Lehmann et al. 2011). Simultaneously, the biochar application leads to improvements in soil chemical properties like pH and dissolved organic carbon content, which in turn promote the proliferation of soil microbial communities (Farrell et al. 2013; Muhammad et al. 2014). However, the effects of biochar on soil microbial communities can be neutral or negative in some cases (Elzobair et al. 2016). The relationship between biochar and microorganisms, particularly the specific roles of biochar in soil microbial processes, has been the focus of many studies, but the mechanisms remain unclear (Palansooriya et al. 2019).
The effects on soil microbial communities under different biochar treatments have been investigated in many independent experiments with the fast development of high-throughput sequencing (HTS) technology in the last decade. However, contrasting results have been occasionally reported because of the heterogeneity of soils from different regions, the diversity of biochar properties, and the variation in experimental conditions. Therefore, identifying the key driving factors that regulate the soil microecosystem remains challenging due to the lack of consensus concerning soil microbiota responses to biochar amendment (Xu et al. 2021).
Traditional microbiological methods were often limited by environmental factors, sequencing tools, and analytical methods, making it difficult to analyze complex microbiomes more closely. Machine learning is becoming an efficient research tool with powerful training and prediction capabilities, and it has already been used successfully in precision medicine, compound design, and bioinformatics research (Ban et al. 2020; Zhang et al. 2022). Machine learning can also be used to advance microbiome research and further reflect potential community structure and sensitive microbial population structure related to soil properties and soil health. Raw sequencing data from public databases and the development of machine learning models for microbial ecological research provide the possibility to study the alteration rule of soil microbial communities (Lu et al. 2023). Machine learning has been applied by case studies to explore the correlations between soil microbial communities and environmental pollutants or plant pathogens (Palansooriya et al. 2022; Yuan et al. 2020; Zhang et al. 2022). However, how biochar amendment affects soil microbial communities has rarely been fully revealed by machine learning models. Therefore, this study aimed to : (i) investigate the responses of microbial communities (bacteria and fungi) to biochar amendment in agricultural soils through a global meta-analysis, (ii) predict the roles of soil microorganisms in biochar-amended soil using machine learning models, (iii) explore the relationship between biochar-mediated changes in soil microorganisms and the role of biochar in agriculture and the environment, and (iv) investigate the effects of biochar application on soil bacterial metabolic functions and fungal nutrient patterns.
2 Materials and methods
2.1 Data collection and description
Metadata were collected from 53 studies including 2225 samples, of which 1547 were bacterial samples and 678 were fungal samples, obtained by searching the Web of Science and the Sequence Read Archive (SRA) database for metadata on October 5, 2022. The keywords ‘Biochar soil microbial community,’ ‘Biochar soil microbiome,’ ‘Biochar bacteria,’ ‘Biochar fungal,’ and ‘Biochar soil structure’ were searched, and the results were compiled into a summary data file. The metadata sample covers twelve countries on five continents and mainly includes samples from China (Additional file 2: Tables S1 and S2). The biochar used in these experimental samples was either unmodified or added to other fertilizers. Straw, wood waste, green waste, and manure were used as raw materials for preparing the biochar (Additional file 2: Table S3). In this study, all the biochar was alkaline with a pH of 9.23 ± 1.09 (Additional file 2: Table S3). After integrating and grouping the metadata, soil biological samples treated with biochar were defined as the biochar group (1059 samples), and biological samples without any biochar addition were defined as the control group (756 samples). This dataset includes the 16S rRNA sequences of soils from 23 host plant species. Detailed information on the metadata, such as biochar type and crop species, is provided in Additional file 2: Tables S1 and S2.
2.2 Data processing
2.2.1 Processing of high-throughput sequencing metadata
The data of microbial HTS were imported into the QIIME 2 bioinformatics platform (v2021.08) for further analysis according to a standardized workflow (Bolyen et al. 2019). The detailed processing procedure was as follows: The raw data and the manifest file that contained the sample IDs, absolute path file, and direction of the reads were imported into QIIME 2, and the adapters and primer sequences were removed from the reads using Cutadapt (Martin 2011). The next workflow was performed using dada2, a plug-in installed in QIIME 2 for filtering, deletion, chimerization, double-end merging, noise reduction of sequences, and resolution of amplicon variants (Callahan et al. 2016). The length of the trimmed sample sequence when running dada2 depends on the sequencing quality of the sample. If the sequencing quality of the double-ended sample was poor, we attempted to run it again with the single-ended dada2 code to ensure that more reads were obtained after running dada2. The dada2 data-processing procedure generates a list of amplicon sequence variants (ASVs) and feature tables. The number of ASVs present in each sample was recorded. Representative sequences and ASVs were merged according to the merge instructions of QIIME2, and each representative sequence was classified using the SILVA database (v138.1, https://www.arb-silva.de/), and fungal ASVs were taxonomically classified using the UNITE database (v8.3, https://unite.ut.ee/). Characteristic sequences annotated as mitochondria or chloroplasts were excluded from analysis. Samples with fewer than 2000 feature sequence reads were also excluded (Zhang et al. 2022). A table of species characteristics was then generated. Finally, 412 samples were filtered, and the remaining 1813 samples were used for further analysis, including 1288 16S rRNA sequencing samples and 525 ITS rRNA sequencing samples (Additional file 2: Tables S1 and S2). The R package Tax4Fun2 (v1.1.5) was used to classify microorganisms into different metabolic function groups (Wemheuer et al. 2020). For fungal communities, FUNGuild (v0.2.0.9000) was used to classify microorganisms into different nutrient pattern groups (Nguyen et al. 2016).
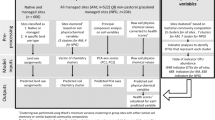
2.2.2 Construction and validation of machine learning models
To better distinguish the subtle changes in soil bacterial and fungal community structure from biochar applications, and to identify specific microbial taxa, three well-established machine learning methods including random forest (RF) (Liaw and Wiener 2002), logistic regression (LR), and support vector machine (SVM) (Cortes and Vapnik 1995) were used. The metadata were divided into a training set (75%) and a test set (25%) (Xu et al. 2022a). The performance of the classifier was evaluated by conducting tenfold cross-validation using the training set. The tenfold cross-validation is an important data training tool to solve the problem of model overfitting. It randomly divides the training set into 10 subsets, one of which is used for validation and the rest for training, repeating the process 10 iterations. With species level as the candidate feature, the optimal parameters of the model were selected for hyper-parameter tuning to ensure the accuracy of the model and reduce the degree of overfitting of the model. The results of model hyperparameterization showed that the random forest model was the most accurate in both bacterial and fungal community, with optimal parameters of mtry = 40, ntree = 800, and nodesize = 6 for bacterial communities and mtry = 20, ntree = 500, and nodesize = 7 for fungal communities (Additional file 1: Fig. S1 and S2). The best-performing classifiers were applied to a test set to independently verify their predictive ability at the phylum, order, family, genus, and species levels. Based on receiver-operating characteristic (ROC) curves and area under the curve (AUC) scores, the final best machine-learning model and the optimal taxonomic hierarchy of microorganisms were derived to distinguish the effects of biochar on soil microorganisms. An out-of-bag estimate of the error rate was used to assess the accuracy of each taxonomic levels and determine the optimal taxonomic levels for machine learning (Yuan et al. 2020). Several measures were implemented to prevent the model from overfitting. For example, microbial species with ambiguous classifications, such as those containing species names like 'uncultured', 'unclassified', and 'unidentified', were filtered out before a machine learning model combined with tenfold cross-validation was performed.
2.2.3 Statistical analysis and visualization
The α-diversity indexes (Chao1, Pielou evenness, Shannon and richness indices) were generated by using the R packages vegan and picante (v4.0.3). Permutational multivariate analysis of variance (PERMANOVA) (Adonis test, using Bray–Curtis transformed data, permutation = 999) was used to distinguish differences in the β-diversity of the biochar-treated and untreated soil microbial communities. Principal coordinate analysis (PCoA) data based on the Bray–Curtis distance matrix was performed by vegan (v4.0.3) and plotted using ggplot2 (v4.0.3). Global sample distribution and stacked and circos plots were generated using ggmap, ggplot2, and circlize software (v4.0.3). A parallel computational code was constructed based on the R package psych (version 2.3.6) to calculate the Spearman correlation coefficient of microbial communities. The detailed code is presented in the Data and Codes Availability Statement section of the article. Spearman correlation coefficients (|R|> 0.6, p < 0.05) for the species were used to construct microbial co-occurrence networks. The criterion for the Spearman correlation coefficient of the co-occurrence network for biomarkers was (|R|> 0.4, p < 0.05), as reported in previous studies (Yuan et al. 2020). Co-occurrence networks between biomarkers and soil microbial communities were visualized in R by calculating Spearman correlation coefficients and applying Cytoscape (version 3.9.1). Machine learning models were executed using the R packages caret (v6.0-94) and random forest (v4.7-1.1), and performance was evaluated using the R packages pROC (v1.18.0) and ROCR (v1.0-11). Annotation of bacterial metabolic functions: the R package Tax4fun2 (v1.1.5). Functional grouping of the fungi: the R package FUNGuild (v0.2.0.9000).
3 Results
3.1 Microbial community structure affected by biochar application
A total of 2225 HTS samples from 53 independent studies conducted in 12 countries (Additional file 1: Fig. S3A, Additional file 2: Tables S1 and S2) were collected, encompassing research on the effects of biochar on soil microbiota (Additional file 1: Fig. S3C, Additional file 2: Tables S1 and S2). The PCoA based on Bray–Curtis distances showed that the addition of biochar significantly affected the bacterial composition (PERMANOVA using the Adonis test: R2 = 0.004, p < 0.001; Fig. 1A). The relative abundances of Acidobacteriota and Firmicutes were higher in the biochar-amended soils, whereas those of Proteobacteria and Actinobacteriota were higher in the control soils (Fig. 1B). The biochar group had more unique genera in the soil bacterial community than the control group (Fig. 1C). However, the addition of biochar did not significantly change the α-diversity of the soil bacterial community, as measured by the Shannon and richness indices (p > 0.05, Kruskal–Wallis test; Fig. 1G, H). Significant difference was also found in the fungal composition between soils with and without biochar amendment (PERMANOVA using the Adonis test: R2 = 0.039, p < 0.001; Fig. 1D). Ascomycota was the main phylum constituting the fungal community in both the biochar and control groups. Compared to the control group, biochar treatment increased the relative abundances of Basidiomycota and Mortierellomycota and decreased that of Chytridiomycota and Olpidiomycota (Fig. 1E). Biochar produced more specific fungal genera than Control, similar to its effect on bacteria (Fig. 1F). Moreover, biochar treatment significantly lowered the Shannon and richness indices of soil fungal communities in the biochar group than in the control group (Kruskal–Wallis test, p < 0.01; Fig. 1I, J).
Effect of biochar application on the structure of soil bacterial and fungal microbial communities. A and D Principal coordinates analysis (PCoA) based on Bray–Curtis dissimilarity on community structure at genus level (A Bacteria; D Fungi). Statistical significance was evaluated via Adonis test. B and E The top 10 relative abundances of soil microbial community phylum level (B Bacteria; E Fungi). G–J represent the alpha diversity of soil microbial communities (G and H Bacteria; I and J Fungi) (Statistical significance was determined by Kruskal–Wallis test ns, not significant; ***p < 0.001)
3.2 Co-occurrence network among microbial communities under biochar application
Neutral models were constructed to clarify the potential importance of stochastic processes in microbial assembly following biochar application. The results showed that the relative contribution of the stochastic processes to the microbial community gradually decreased with increasing biochar content (Additional file 1: Fig. S4A, B), and this trend was more pronounced in the fungal community assembly than that of the bacteria (Additional file 1: Fig. S4C, D). Biochar changed the aggregation pattern of soil microbial communities, increasing the contribution of deterministic processes to community aggregation, suggesting that soil microbial aggregation is more substantially influenced by biochar than by stochastic factors.
To reveal the potential effects of biochar addition on soil bacterial and fungal communities, a microbial co-occurrence network was constructed based on Spearman's correlation coefficient (|R|> 0.6, p < 0.05) (Fig. 2). Biochar application reduced the number of nodes (biochar nodes: 527, control nodes: 561) (Additional file 1: Fig. S5; Additional file 2: Table S4) but increased the number of edges (biochar edges: 3107, control edges: 3098) (Additional file 1: Fig. S5; Additional file 2: Table S5), indicating that the application of biochar led to closer interactions between soil microbial communities (Fig. 2A, B). Based on the number of nodes and edge connections, Ascomycota, Proteobacteria, and Basidiomycota were the key taxa driving soil microbial community interactions, the proportions of which were reduced by biochar. However, the proportions of Firmicutes, Actinobacteriota, and Bacteroidota increased in biochar-amended soils (Fig. 2C). There was no significant difference in the degree of co-occurrence of network nodes between the biochar and control groups (p > 0.05, Kruskal–Wallis test); however, biochar increased the overall average degrees (average degrees: biochar 11.79, control 11.04) (Fig. 2D), indicating that biochar increases the mutualistic relationships among soil microorganisms. To compare the strength of the microbial network against disturbances in the biochar and control groups, a robustness analysis was conducted by gradually removing the nodes. Successive node removals caused a linear decrease in the natural connectivity of soil microbial communities in both biochar and control groups, with a more obvious decrease in that of the control group (Slope: − 0.08, R2: 0.95; Fig. 2E). This indicates that biochar reduced the natural connectivity and enhanced the anti-interference ability of the soil microbial communities.
Co-occurrence network analysis of microbial communities in the soils of biochar and control groups. A and B Co-occurrence network of soil microbial communities (A Biochar, B Control). C Proportion of the most dominant nodes in the co-occurrence network of soil microbial communities. D Degree comparison of nodes in co-occurrence networks of soil microbial communities (Kruskal–Wallis test; p > 0.05, ns). E Robustness assessment of soil microbial communities. The decreasing trend of natural connectivity is fitted with 80% node loss, and R2 and slope are shown in the figure. The smaller the absolute value of the slope is, the more stable the network is (Statistical significance was determined by a two-sided t-test, p < 0.001***)
3.3 Identification of biomarkers of soil microbial communities under biochar application
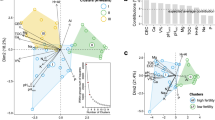
To determine whether the properties of soil bacterial and fungal communities could be used as biomarkers in the biochar and control groups, three machine learning models, RF, SVM, and LR, were constructed. To avoid overfitting the model to the limited training dataset, a tenfold cross-validation was performed to evaluate the prediction accuracy of the model based on different classification levels. The RF model had the best overall performance at different taxonomic levels for bacteria and fungi, with the highest prediction accuracy of 99.54% for the classification model constructed at the order level of bacterial community, and the prediction accuracy of nearly 100% for that at the family level of fungal community (Fig. 3A, D). The RF model performed well for both test and training set classifications (Additional file 1: Fig. S6). The bacterial and fungal AUCs reached 1.000 in the training set of the RF model, and the accuracy of the test set was also high at 0.812 for bacteria and 0.822 for fungi (Fig. 3B, E). These results show that the bacterial and fungal models can both accurately predict the effects of biochar on soil microbial communities. The most important 60 bacterial orders and 40 fungal families were identified as biomarkers using tenfold cross-validation error curves and simplified analysis (Fig. 3C, F; Additional file 2: Tables S6 and S7). Among the bacterial biomarkers, 14 bacterial orders were significantly enriched in biochar-amended soils, with Gemmatimonadetes, Microtrichales, Candidatus_Kaiserbacteria, and Pyrinomonadales being potentially beneficial (Kruskal–Wallis test, p < 0.05; Fig. 3G). Notably, among the fungal biomarkers, seven fungal families were enriched in biochar-amended soils, among which four belonged to Ascomycota phylum, and the abundance of Erythrobasidiaceae significantly increased (Kruskal–Wallis test, p < 0.05; Fig. 3H).
Effect of biochar application on the structure of soil bacterial and fungal microbial communities. A and D Out-of-bag (OOB) error curves were obtained by ten-fold cross-validation of different machine learning algorithms on different microbial species hierarchies to assess the performance of the predictive models (A bacteria; D fungi), with the corresponding microbial species taxonomic hierarchies in the horizontal coordinate. B and E AUC and ROC curves of different machine learning prediction models built on microbial communities (B Bacteria, E Fungi). C and F The errors of the tenfold cross-validation increase with the number of microorganisms, and the OOB error is calculated in this process to determine the number of biomarkers (C Bacterial order level, F Fungal family level). G and H The upper part of the graph shows the relative abundance of biomarkers, and the bottom graph shows the importance of biomarkers calculated by the random forest model, rearranged in descending order, and colored by taxonomic phylum. (G Bacteria, H Fungi. Statistical significance was determined by Kruskal–Wallis test, p < 0.05*)
3.4 Coordination of biomarkers
Co-occurrence networks were used to assess the cooperative features among biomarkers, and only those with significant correlations were shown (|R|> 0.4, p < 0.05) (Fig. 4; Additional file 2: Tables S8 and S9). The results showed that biochar made the interaction between the biomarkers substantially stronger than in the soil without biochar, and that even biomarkers with lower abundances could play a stronger linkage role. The application of biochar reduced the proportion of negatively related edges among the biomarkers, indicating that biochar promoted beneficial synergies in soil microbial communities. Notably, biochar made Aspergillus and Bacteroides more inhibitory to the synergistic ability of the fungal communities (Fig. 4C, D). In order to further explore the relationship between biomarkers and soil microorganisms, a co-occurrence network between biomarkers and soil microbial communities was also constructed. The results showed that although the number of nodes of biochar in the bacterial community (365) was less than that in the control group (383), the number of edges increased by as much as 242, which significantly increased the proportion of positive microbial community correlations (Biochar positive correlation edges ratio: 98.43%, negative correlation edges ratio: 1.57%; Control positive correlation edges ratio: 92.24%, negative correlation edges ratio: 7.76%) (Additional file 1: Fig. S7A, B). This phenomenon was more obvious in the co-occurrence network of fungal biomarkers and soil fungal communities. The biochar group had a total of 2923 edges, of which 98.97% were positively correlated, but only 320 nodes (Additional file 1: Fig. S7C). The control group had 2237 edges, of which 98.92% were positively correlated, and had more nodes (348) than the biochar group (Additional file 1: Fig. S7D). This further suggests that soil microbial community altered the relationship between collaborative relationships to promote better uptake of nutrients from biochar.
Co-occurrence network of biomarkers in soils of biochar and control groups. A and B show that biochar application is a Co-occurrence network for biomarkers of soil bacterial communities. The co-occurrence was constructed by screening 59 bacterial orders based on Spearman's correlation coefficient (|R|< 0.4, p < 0.05). C and D show that biochar application is a Co-occurrence network for biomarkers of soil fungal communities. 38 fungal families were screened in soils of biochar applied group based on Spearman's correlation coefficient (|R|< 0.4, p < 0.05), while 39 constructed networks in the control group
3.5 Effect of biochar on soil microbial functions
The 16S rRNA gene sequencing data were entered into the Kyoto Encyclopedia of Genes and Genomes (KEGG) database to evaluate the effects of biochar on bacterial function (Fig. 5A). In the present study, more than 37% of the sequences were enriched in organismal systems, followed by metabolism, which accounted for more than 28.9% of the sequences (Additional file 1: Fig. S8). However, there was no visible difference in the soil bacterial function between the biochar and control groups. Changes in the endophyte functions of soil fungal communities after biochar application were predicted using FUNGuild. Fungal endophytes were identified as probable and highly probable factors in this study. The results showed that the fungal trophic patterns could be divided into seven groups: symbiotrophs, saprotrophs, pathotrophs, pathotroph-saprotrophs, pathotroph-symbiotrophs, saprotroph-symbiotrophs, and pathotroph-saprotroph-symbiotrophs (Fig. 5B). Saprotrophs and saprotrophic symbiotrophs were the dominant trophic modes of soil fungal communities. Biochar increased the proportion of soil fungal endophytes in the trophic mode of saprotrophs and decreased the proportion of soil fungal endophytes in the saprotroph-symbiotrophs mode (Fig. 5C).
Bacterial metabolic function and fungal nutrient patterns in soils of biochar and control groups. A Circular heatmap demonstrating the different of relative abundance of soil bacterial functions in biochar and control groups. B Chord diagram demonstrating the composition of fungal trophic patterns. C Relative abundance of seven trophic modes of fungi
4 Discussion
Biochar has attracted heightened attention in recent years owing to its role in improving soil properties, as well as in mitigating climate change and environmental pollution (Agarwal et al. 2022; Hossain et al. 2020). The role of biochar in affecting soil microbial communities provides multiple agricultural and environmental benefits (Chen et al. 2019; Xu et al. 2023). In this study, machine learning was integrated with metadata to elucidate the mechanisms underlying the effects of biochar on soil microbial communities on a global scale. This investigation focused on assessing the specific impacts of biochar on the diversity, composition, and function of soil microbial communities.
4.1 Biochar affects soil microbial community structure and function
The α-diversity of soil fungal communities in the biochar group was significantly higher than that in the control group. However, biochar treatment did not cause statistically significant difference in the α-diversity of soil bacterial community, which was in accord with a previous study in which biochar had a greater effect on soybean inter-root soil fungal diversity than on bacterial diversity (Gao 2021). This may be due to the difference in carbon utilization of the major components of biochar by bacteria and fungi, with most fungi acting as decomposers in the soil and being more capable of utilizing the carbon composition of biochar than bacteria (Lehmann et al. 2011; Liu et al. 2022, 2017). This could be because the raw materials for biochar production are mostly composed of cellulose, hemicellulose, and lignin, and thus the prepared biochar often contains a certain amount of incomplete carbonized organic matter, which brings more benefits to the growth of saprophytic fungi than bacteria (Dai et al. 2021). The differences between biochar and control groups were significantly higher than those within groups (PERMANOVA: p < 0.001) for both bacterial and fungal communities, indicating that the biochar treatment does significantly change the soil microbial composition, which corroborates previous studies (Bello et al. 2021; Xu et al. 2023). Proteobacteria, Acidobacteriota, Actinobacteria, Ascomycota, Basidiomycota, and Mortierellomycota are the primary phyla found in agriculture soils. Biochar was previously found to upregulate certain beneficial bacterial phyla such as Acidobacteria, which play an important role in regulating soil community biogeochemical cycles (Kalam et al. 2020). Basidiomycota and Mortierellomycota were enriched in soils amended with biochar (Fig. 1). The fungal phylum Basidiomycota was a representative group of oligotrophic fungi, which plays a crucial role in the decomposition of complex organic matter in soils (Li et al. 2021; Yu et al. 2021). It breaks down the large molecular carbon compounds present in biochar into smaller molecules that are more easily absorbed by microorganisms and crops, thereby facilitating soil carbon nutrient cycling (Louca et al. 2018; Xu et al. 2021). Mortierellomycota is adapted to growing in nutrient-rich soil environments, which suggests that the application of biochar increases soil nutrients (Yu et al. 2020). Furthermore, biochar led to a considerable change in the trophic status of soil fungi and an obvious increase in the proportion of saprophytic fungi, among which the genus Lophiostoma contains a variety of beneficial endophytic bacteria that are present in biochar amended soils (Bai et al. 2020). Soil metabolites are important carbon sources for microbes (Swenson et al. 2018). The addition of biochar to oily sludge leads to an increase in the abundance of bacterial genes related to carbohydrate metabolism and xenobiotic biodegradation and metabolism (Shi et al. 2022). Bacterial carbon metabolism also increased after biochar was added to inter-root soils with polycyclic aromatic hydrocarbon contamination (Li et al. 2020). Our study also showed similar results: biochar led to changes in the soil bacterial functions including carbohydrate metabolism, and the global and overview maps increased by 0.03% and 0.10%, respectively (Fig. 5A). However, the effect of biochar on the bacterial metabolic function was not as great as expected, which may be due to the differences in the amount, source and properties of biochar. Fungal and bacterial communities were both affected by biochar treatment, with biochar affecting fungi to a greater extent in terms of α-diversity, community composition, species interactions, and shifts in trophic patterns.
4.2 Biochar promotes cooperation rather than competition among soil microbes
Co-occurrence network analysis provides a visualization of the interaction intensity between microbial communities and allows the evaluation of the soil micro-ecology stability (Ma et al. 2020). Biochar treatment increased the proportion of positive correlations and the number of edges in the microbial co-occurrence network, but decreased the number of nodes. This indicates that biochar promotes cooperation among microorganisms rather than competition to drive the uptake of biochar nutrients (Fig. 2A,B). Here we speculate that this phenomenon may be attributable to mutually beneficial interactions between fungal communities over a longer period of co-evolution (Zhang et al. 2014). This explanation was also confirmed in our study, as Ascomycota was a key taxonomy forming the soil microbial network in the biochar group, but with a downregulated proportion of soil nodes in the control group. This may be due to changes in the microbial pattern of resource acquisition and the trophic status of Ascomycota caused by the biochar. Biochar can alter the soil microenvironment, and the extent of this alteration depends largely on the characteristics of biochar. For example, the pH of biochar prepared at high temperatures is alkaline, and previous researchers have observed that high pH levels improve soil characteristics required for microbial colonization (Chen et al. 2017). However, certain microorganisms do not adapt to this improvement, which also leads to a decrease in their abundance in the soil environment. Actinomycetes prefer acidic and neutral pH environments, and higher pH biochar inevitably reduces the abundance of actinomycetes (Xu et al. 2021). The high porosity and high surface area of biochar provide a favorable habitat for soil microorganisms. Water molecules and soluble substances involved in microbial metabolism (such as acids, alcohols, aldehydes, ketones, and sugars) are stored in the mesopore and micropore of biochar. Therefore, when biochar is applied to a soil environment, it can regulate the abundance and activity of certain microorganisms that can take advantage of the physical properties of biochar (Palansooriya et al. 2019). Nutrients from biochar are also key factors in altering the structure and abundance of soil microbial communities, and increased soil phosphorus input due to biochar application leads to a subsequent decrease in fungal species (Nie et al. 2018).
In summary, the application of biochar in soil increased the activity, diversity, and abundance of soil microorganisms, and enhanced the ability of microorganisms to cooperate with each other. However, the physicochemical properties of biochar may inhibit the abundance of certain microorganisms, which make up only a small fraction of the overall microbial community. This results in diminished interactions of certain microorganisms in the soil environment.
4.3 Biochar enriches biomarkers with ecological functions
Machine learning further revealed the response patterns of the soil microbial communities to biochar application, considerably narrowing the search for potentially beneficial bacteria. Previous studies have successfully used RF models to identify biomarkers (Xu et al. 2022a; Yuan et al. 2020; Zhang et al. 2022). In the present study, RF also outperformed SVM and LR and identified 60 bacterial and 40 fungal biomarkers. Among them, 14 bacterial orders and 7 fungal families were enriched in biochar-amended soils, including potentially beneficial orders that promote plant growth and soil nutrient cycling, such as Gemmatimonadetes, Microtrichales, Candidatus_Kaiserbacteria, and Pyrinomonadales (Cai et al. 2020; Mujakić et al. 2022; Paulraj et al. 2022; Wu et al. 2022; Xu et al. 2023). Notably, biochar enriches methanogenic-related microorganisms, such as the S085 genera of Dehalococcoidia and Beggiatoales (Paoli et al. 2022; Saxton et al. 2016). Micrococcales is adapted to arid habitats, but its abundance decreased in biochar-applied soils (Sun et al. 2020). This indicates that biochar allows the soil to increase its water-holding capacity. Pirellulales lives in low-oxygen environments, and the pore structure of biochar allows more oxygen to circulate in the soil, leading to a decrease in the abundance of Pirellulales (Dedysh et al. 2020). Burkholderiales is widely reported to be a plant probiotic (Banerjee et al. 2018), and its abundance decreased in biochar-amended soils, yet it maintained a high relative abundance of 6.35%. Among the fungal biomarkers, Nectriaceae was enriched in biochar-amended soils in previous studies, where an increase in pH led to an increase in their abundance (Yang et al. 2019). There was also a strong response of Umbelopsidaceae to changes in soil pH (Rincón et al. 2015). Erythrobasidiaceae was significantly enriched in the biochar-amended soils (Adonis test: p < 0.001); however, its function has not been reported in detail or systematically (Peršoh et al. 2013). Helotiaceae includes many dark-isolated endophytes, a diverse group of filamentous ascomycetes commonly found in soils that act as saprophytic fungi to promote soil nutrient cycling (Detheridge et al. 2016; Herzog et al. 2019). The Helotiaceae family contains phytopathogenic bacteria that can cause serious plant diseases, such as dieback in ash trees (Griffiths et al. 2020). The biochar treatment greatly reduced the abundance of Helotiaceae, thereby reducing disease. Overall, it is considered that biochar has a positive influence on the soil micro-ecology in terms of its ability to improve soil nutrient accessibility and suppress plant diseases, which is mainly reflected in the enhanced positive interactions of soil microbes and the stability of soil community structure, as well as the reduced proportion of plant pathogens in the soil microbial community.
5 Conclusions
Through metadata analysis, our study showed that the application of biochar in agricultural land improves the soil microbial microenvironment, stimulates microbial activity, improves microbial interactions, and enhances microbial stability. Although large amounts of data could reduce the heterogeneity of individual experiments and help explain the rule of biochar shaping soil microorganisms, the decisive role of environmental factors, such as the physical and chemical characteristics of regional soil and certain structures of indigenous microorganisms, should be fully considered when carrying out agricultural production in a certain area. The biomarkers identified in this study have different ecological functions and their enrichment effects in biochar-amended soils require further study, especially the experimental verification should be strengthened in future work. In addition, the combination of biochar with key microorganisms may further promote soil improvement and plant growth.
Data and code availability
All metadata information of soil bacterial and fungal communities used in this study are collected from the SRA database and accession numbers are listed in Additional file 2 : Table S1 & S2. All analytical codes, visualization methods, and bioinformatics data used in this study are available online at https://github.com/users/leict0321/Biochar_script.
References
Agarwal H, Kashyap VH, Mishra A, Bordoloi S, Singh PK, Joshi NC (2022) Biochar-based fertilizers and their applications in plant growth promotion and protection. 3 Biotech 12:136. https://doi.org/10.1007/s13205-022-03195-2
Bai Y-C, Chang Y-Y, Hussain M, Lu B, Zhang J-P, Song X-B, Lei X-S, Pei D (2020) Soil chemical and microbiological properties are changed by long-term chemical fertilizers that limit ecosystem functioning. Microorganisms 8:694. https://doi.org/10.3390/microorganisms8050694
Ban Z, Yuan P, Yu F, Peng T, Zhou Q, Hu X (2020) Machine learning predicts the functional composition of the protein corona and the cellular recognition of nanoparticles. Proc Natl Acad Sci U S A 117:10492–10499. https://doi.org/10.1073/pnas.1919755117
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576. https://doi.org/10.1038/s41579-018-0024-1
Bardgett RD, Van Der Putten WH (2014) Belowground biodiversity and ecosystem functioning. Nature 515:505–511. https://doi.org/10.1038/nature13855
Bello A, Wang B, Zhao Y, Yang W, Ogundeji A, Deng L, Egbeagu UU, Yu S, Zhao L, Li D, Xu X (2021) Composted biochar affects structural dynamics, function and co-occurrence network patterns of fungi community. Sci Total Environ 775:145672. https://doi.org/10.1016/j.scitotenv.2021.145672
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Cai Y, Shen J-P, Di HJ, Zhang L-M, Zhang C, He J-Z (2020) Variation of soil nitrate and bacterial diversity along soil profiles in manure disposal maize field and adjacent woodland. J Soils Sediments 20:3557–3568. https://doi.org/10.1007/s11368-020-02679-z
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Chen J, Li S, Liang C, Xu Q, Li Y, Qin H, Fuhrmann JJ (2017) Response of microbial community structure and function to short-term biochar amendment in an intensively managed bamboo (Phyllostachys praecox) plantation soil: effect of particle size and addition rate. Sci Total Environ 574:24–33. https://doi.org/10.1016/j.scitotenv.2016.08.190
Chen L, Jiang Y, Liang C, Luo Y, Xu Q, Han C, Zhao Q, Sun B (2019) Competitive interaction with keystone taxa induced negative priming under biochar amendments. Microbiome 7:77. https://doi.org/10.1186/s40168-019-0693-7
Cortes C, Vapnik V (1995) Support-vector networks. Mach Learn 20:273–297. https://doi.org/10.1007/BF00994018
Dai Z, Xiong X, Zhu H, Xu H, Leng P, Li J, Tang C, Xu J (2021) Association of biochar properties with changes in soil bacterial, fungal and fauna communities and nutrient cycling processes. Biochar 3:239–254. https://doi.org/10.1007/s42773-021-00099-x
Dedysh SN, Kulichevskaya IS, Beletsky AV, Ivanova AA, Rijpstra WIC, Damsté JSS, Mardanov AV, Ravin NV (2020) Lacipirellula parvula gen. nov., sp. nov., representing a lineage of planctomycetes widespread in low-oxygen habitats, description of the family Lacipirellulaceae fam. nov. and proposal of the orders Pirellulales ord. nov., Gemmatales ord. nov. and Isosphaerales ord. nov. Syst Appl Microbiol 43:126050. https://doi.org/10.1016/j.syapm.2019.126050
Detheridge AP, Brand G, Fychan R, Crotty FV, Sanderson R, Griffith GW, Marley CL (2016) The legacy effect of cover crops on soil fungal populations in a cereal rotation. Agric Ecosyst Environ 228:49–61. https://doi.org/10.1016/j.agee.2016.04.022
Ding Y, Liu Y, Liu S, Li Z, Tan X, Huang X, Zeng G, Zhou L, Zheng B (2016) Biochar to improve soil fertility. A review. Agron Sustain Dev 36:36. https://doi.org/10.1007/s13593-016-0372-z
Duan Z, Chen C, Ni C, Xiong J, Wang Z, Cai J, Tan W (2023) How different is the remediation effect of biochar for cadmium contaminated soil in various cropping systems? A global meta-analysis. J Hazard Mater 448:130939. https://doi.org/10.1016/j.jhazmat.2023.130939
Elzobair KA, Stromberger ME, Ippolito JA, Lentz RD (2016) Contrasting effects of biochar versus manure on soil microbial communities and enzyme activities in an Aridisol. Chemosphere 142:145–152. https://doi.org/10.1016/j.chemosphere.2015.06.044
Farrell M, Kuhn TK, Macdonald LM, Maddern TM, Murphy DV, Hall PA, Singh BP, Baumann K, Krull ES, Baldock JA (2013) Microbial utilisation of biochar-derived carbon. Sci Total Environ 465:288–297. https://doi.org/10.1016/j.scitotenv.2013.03.090
Gao W (2021) Effect of biochar and charcoal-based fertilizer on bacterial and fungal communities in soybean inter-root soil. Ecol Environ Sci 30:205–212. https://doi.org/10.16258/j.cnki.1674-5906.2021.01.024
Griffiths SM, Galambao M, Rowntree J, Goodhead I, Hall J, O’Brien D, Atkinson N, Antwis RE (2020) Complex associations between cross-kingdom microbial endophytes and host genotype in ash dieback disease dynamics. J Ecol 108:291–309. https://doi.org/10.1111/1365-2745.13302
Hartmann M, Six J (2022) Soil structure and microbiome functions in agroecosystems. Nat Rev Earth Environ 4:4–18. https://doi.org/10.1038/s43017-022-00366-w
Herzog C, Hartmann M, Frey B, Stierli B, Rumpel C, Buchmann N, Brunner I (2019) Microbial succession on decomposing root litter in a drought-prone Scots pine forest. ISME J 13:2346–2362. https://doi.org/10.1038/s41396-019-0436-6
Hossain MZ, Bahar MM, Sarkar B, Donne SW, Ok YS, Palansooriya KN, Kirkham MB, Chowdhury S, Bolan N (2020) Biochar and its importance on nutrient dynamics in soil and plant. Biochar 2:379–420. https://doi.org/10.1007/s42773-020-00065-z
Kalam S, Basu A, Ahmad I, Sayyed RZ, El-Enshasy HA, Dailin DJ, Suriani NL (2020) Recent understanding of soil acidobacteria and their ecological significance: a critical review. Front Microbiol. https://doi.org/10.3389/fmicb.2020.580024
Lal R (2013) Food security in a changing climate. Ecohydrol Hydrobiol 13:8–21. https://doi.org/10.1016/j.ecohyd.2013.03.006
Lehmann J, Rillig MC, Thies J, Masiello CA, Hockaday WC, Crowley D (2011) Biochar effects on soil biota—a review. Soil Biol Biochem 43:1812–1836. https://doi.org/10.1016/j.soilbio.2011.04.022
Lehmann J, Cowie A, Masiello CA, Kammann C, Woolf D, Amonette JE, Cayuela ML, Camps-Arbestain M, Whitman T (2021) Biochar in climate change mitigation. Nat Geosci 14:883–892. https://doi.org/10.1038/s41561-021-00852-8
Li Y, Li Y, Chang SX, Yang Y, Fu S, Jiang P, Luo Y, Yang M, Chen Z, Hu S, Zhao M, Liang X, Xu Q, Zhou G, Zhou J (2018a) Biochar reduces soil heterotrophic respiration in a subtropical plantation through increasing soil organic carbon recalcitrancy and decreasing carbon degrading microbial activity. Soil Biol Biochem 122:173–185. https://doi.org/10.1016/j.soilbio.2018.04.019
Li Y, Hu S, Chen J, Müller K, Li Y, Fu W, Lin Z, Wang H (2018b) Effects of biochar application in forest ecosystems on soil properties and greenhouse gas emissions: a review. J Soils Sediments 18:546–563. https://doi.org/10.1007/s11368-017-1906-y
Li X, Yao S, Bian Y, Jiang X, Song Y (2020) The combination of biochar and plant roots improves soil bacterial adaptation to PAH stress: insights from soil enzymes, microbiome, and metabolome. J Hazard Mater 400:123227. https://doi.org/10.1016/j.jhazmat.2020.123227
Li H, Yang S, Semenov MV, Yao F, Ye J, Bu R, Ma R, Lin J, Kurganova I, Wang X, Deng Y, Kravchenko I, Jiang Y, Kuzyakov Y (2021) Temperature sensitivity of SOM decomposition is linked with a K-selected microbial community. Glob Change Biol 27:2763–2779. https://doi.org/10.1111/gcb.15593
Liaw A, Wiener M (2002) Classification and Regression by randomForest 2
Liu Y, Lu H, Yang S, Wang Y (2016) Impacts of biochar addition on rice yield and soil properties in a cold waterlogged paddy for two crop seasons. Field Crops Res 191:161–167. https://doi.org/10.1016/j.fcr.2016.03.003
Liu Y, Yao S, Wang Y, Lu H, Brar SK, Yang S (2017) Bio- and hydrochars from rice straw and pig manure: inter-comparison. Bioresour Technol 235:332–337. https://doi.org/10.1016/j.biortech.2017.03.103
Liu Y, Lonappan L, Brar SK, Yang S (2018a) Impact of biochar amendment in agricultural soils on the sorption, desorption, and degradation of pesticides: a review. Sci Total Environ 645:60–70. https://doi.org/10.1016/j.scitotenv.2018.07.099
Liu Y, Wang Y, Lu H, Lonappan L, Brar SK, He L, Chen J, Yang S (2018b) Biochar application as a soil amendment for decreasing cadmium availability in soil and accumulation in Brassica chinensis. J Soils Sediments 18:2511–2519. https://doi.org/10.1007/s11368-018-1927-1
Liu S, García-Palacios P, Tedersoo L, Guirado E, van der Heijden MGA, Wagg C, Chen D, Wang Q, Wang J, Singh BK, Delgado-Baquerizo M (2022) Phylotype diversity within soil fungal functional groups drives ecosystem stability. Nat Ecol Evol 6:900–909. https://doi.org/10.1038/s41559-022-01756-5
Louca S, Polz MF, Mazel F, Albright MBN, Huber JA, O’Connor MI, Ackermann M, Hahn AS, Srivastava DS, Crowe SA, Doebeli M, Parfrey LW (2018) Function and functional redundancy in microbial systems. Nat Ecol Evol 2:936–943. https://doi.org/10.1038/s41559-018-0519-1
Lu K, Yang X, Gielen G, Bolan N, Ok YS, Niazi NK, Xu S, Yuan G, Chen X, Zhang X, Liu D, Song Z, Liu X, Wang H (2017) Effect of bamboo and rice straw biochars on the mobility and redistribution of heavy metals (Cd, Cu, Pb and Zn) in contaminated soil. J Environ Manage Biogeochem Trace Elements Environ 186:285–292. https://doi.org/10.1016/j.jenvman.2016.05.068
Lu T, Xu N, Lei C, Zhang Q, Zhang Z, Sun L, He F, Zhou N-Y, Peñuelas J, Zhu Y-G, Qian H (2023) Bacterial biogeography in China and its association to land use and soil organic carbon. Soil Ecol Lett 5:230172. https://doi.org/10.1007/s42832-023-0172-8
Ma B, Wang Y, Ye S, Liu S, Stirling E, Gilbert JA, Faust K, Knight R, Jansson JK, Cardona C, Röttjers L, Xu J (2020) Earth microbial co-occurrence network reveals interconnection pattern across microbiomes. Microbiome 8:82. https://doi.org/10.1186/s40168-020-00857-2
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Muhammad N, Dai Z, Xiao K, Meng J, Brookes PC, Liu X, Wang H, Wu J, Xu J (2014) Changes in microbial community structure due to biochars generated from different feedstocks and their relationships with soil chemical properties. Geoderma 226–227:270–278. https://doi.org/10.1016/j.geoderma.2014.01.023
Mujakić I, Piwosz K, Koblížek M (2022) Phylum gemmatimonadota and its role in the environment. Microorganisms 10:151. https://doi.org/10.3390/microorganisms10010151
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nie C, Yang X, Niazi NK, Xu X, Wen Y, Rinklebe J, Ok YS, Xu S, Wang H (2018) Impact of sugarcane bagasse-derived biochar on heavy metal availability and microbial activity: a field study. Chemosphere 200:274–282. https://doi.org/10.1016/j.chemosphere.2018.02.134
Palansooriya KN, Li J, Dissanayake PD, Suvarna M, Li L, Yuan X, Sarkar B, Tsang DCW, Rinklebe J, Wang X, Ok YS (2022) Prediction of soil heavy metal immobilization by biochar using machine learning. Environ Sci Technol 56:4187–4198. https://doi.org/10.1021/acs.est.1c08302
Palansooriya KN, Wong JTF, Hashimoto Y, Huang L, Rinklebe J, Chang SX, Bolan N, Wang H, Ok YS (2019) Response of microbial communities to biochar-amended soils: a critical review. Biochar 1:3–22. https://doi.org/10.1007/s42773-019-00009-2
Paoli L, Ruscheweyh H-J, Forneris CC, Hubrich F, Kautsar S, Bhushan A, Lotti A, Clayssen Q, Salazar G, Milanese A, Carlström CI, Papadopoulou C, Gehrig D, Karasikov M, Mustafa H, Larralde M, Carroll LM, Sánchez P, Zayed AA, Cronin DR, Acinas SG, Bork P, Bowler C, Delmont TO, Gasol JM, Gossert AD, Kahles A, Sullivan MB, Wincker P, Zeller G, Robinson SL, Piel J, Sunagawa S (2022) Biosynthetic potential of the global ocean microbiome. Nature 607:111–118. https://doi.org/10.1038/s41586-022-04862-3
Paulraj S, Bhat R, Rajesh MK, Ramesh SV, Priya UK, Pandian RTP, Hegde V, Chowdappa P (2022) Metagenomic analysis reveals the predominance of Candidatus Patescibacteria in the rhizosphere of arecanut palms in yellow leaf disease (YLD) endemic areas of India. J Plant Crops. https://doi.org/10.25081/jpc.2022.v50.i1.7785
Peršoh D, Segert J, Zigan A, Rambold G (2013) Fungal community composition shifts along a leaf degradation gradient in a European beech forest. Plant Soil 362:175–186. https://doi.org/10.1007/s11104-012-1271-y
Rincón A, Santamaría-Pérez B, Rabasa SG, Coince A, Marçais B, Buée M (2015) Compartmentalized and contrasted response of ectomycorrhizal and soil fungal communities of Scots pine forests along elevation gradients in France and Spain. Environ Microbiol 17:3009–3024. https://doi.org/10.1111/1462-2920.12894
Saxton MA, Samarkin VA, Schutte CA, Bowles MW, Madigan MT, Cadieux SB, Pratt LM, Joye SB (2016) Biogeochemical and 16S rRNA gene sequence evidence supports a novel mode of anaerobic methanotrophy in permanently ice-covered Lake Fryxell, Antarctica. Limnol Oceanogr 61:S119–S130. https://doi.org/10.1002/lno.10320
Shi Y, Liu M, Li J, Yao Y, Tang J, Niu Q (2022) The dosage-effect of biochar on anaerobic digestion under the suppression of oily sludge: performance variation, microbial community succession and potential detoxification mechanisms. J Hazard Mater 421:126819. https://doi.org/10.1016/j.jhazmat.2021.126819
Singh H, Northup BK, Rice CW, Prasad PVV (2022) Biochar applications influence soil physical and chemical properties, microbial diversity, and crop productivity: a meta-analysis. Biochar 4:8. https://doi.org/10.1007/s42773-022-00138-1
Sun X, Zhang L, Pei J, Huang L-F (2020) Regulatory relationship between quality variation and environment of Cistanche deserticola in three ecotypes based on soil microbiome analysis. Sci Rep 10:6662. https://doi.org/10.1038/s41598-020-63607-2
Swenson TL, Karaoz U, Swenson JM, Bowen BP, Northen TR (2018) Linking soil biology and chemistry in biological soil crust using isolate exometabolomics. Nat Commun 9:19. https://doi.org/10.1038/s41467-017-02356-9
Wemheuer F, Taylor JA, Daniel R, Johnston E, Meinicke P, Thomas T, Wemheuer B (2020) Tax4Fun2: prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene sequences. Environ Microbiome 15:11. https://doi.org/10.1186/s40793-020-00358-7
Wu X, Rensing C, Han D, Xiao K-Q, Dai Y, Tang Z, Liesack W, Peng J, Cui Z, Zhang F (2022) Genome-resolved metagenomics reveals distinct phosphorus acquisition strategies between soil microbiomes. mSystems 7:e01107-e1121. https://doi.org/10.1128/msystems.01107-21
Xu W, Whitman WB, Gundale MJ, Chien C-C, Chiu C-Y (2021) Functional response of the soil microbial community to biochar applications. GCB Bioenergy 13:269–281. https://doi.org/10.1111/gcbb.12773
Xu N, Kang J, Ye Y, Zhang Q, Ke M, Wang Y, Zhang Z, Lu T, Peijnenburg WJGM, Penuelas J, Bao G, Qian H (2022a) Machine learning predicts ecological risks of nanoparticles to soil microbial communities. Environ Pollut 307:119528. https://doi.org/10.1016/j.envpol.2022.119528
Xu N, Zhao Q, Zhang Z, Zhang Q, Wang Y, Qin G, Ke M, Qiu D, Peijnenburg WJGM, Lu T, Qian H (2022b) Phyllosphere microorganisms: sources, drivers, and their interactions with plant hosts. J Agric FOOD Chem 70:4860–4870. https://doi.org/10.1021/acs.jafc.2c01113
Xu W, Xu H, Delgado-Baquerizo M, Gundale MJ, Zou X, Ruan H (2023) Global meta-analysis reveals positive effects of biochar on soil microbial diversity. Geoderma 436:116528. https://doi.org/10.1016/j.geoderma.2023.116528
Yang C, Liu N, Zhang Y (2019) Soil aggregates regulate the impact of soil bacterial and fungal communities on soil respiration. Geoderma 337:444–452. https://doi.org/10.1016/j.geoderma.2018.10.002
Yu H, Fan P, Hou J, Dang Q, Cui D, Xi B, Tan W (2020) Inhibitory effect of microplastics on soil extracellular enzymatic activities by changing soil properties and direct adsorption: an investigation at the aggregate-fraction level. Environ Pollut 267:115544. https://doi.org/10.1016/j.envpol.2020.115544
Yu H, Zhang Y, Tan W (2021) The “neighbor avoidance effect” of microplastics on bacterial and fungal diversity and communities in different soil horizons. Environ Sci Ecotechnology 8:100121. https://doi.org/10.1016/j.ese.2021.100121
Yuan J, Wen T, Zhang H, Zhao M, Penton CR, Thomashow LS, Shen Q (2020) Predicting disease occurrence with high accuracy based on soil macroecological patterns of Fusarium wilt. ISME J 14:2936–2950. https://doi.org/10.1038/s41396-020-0720-5
Zhang Y, Zhao Z, Dai M, Jiao N, Herndl GJ (2014) Drivers shaping the diversity and biogeography of total and active bacterial communities in the South China Sea. Mol Ecol 23:2260–2274. https://doi.org/10.1111/mec.12739
Zhang Z, Zhang Q, Cui H, Li Y, Xu N, Lu T, Chen J, Penuelas J, Hu B, Qian H (2022) Composition identification and functional verification of bacterial community in disease-suppressive soils by machine learning. Environ Microbiol 24:3405–3419. https://doi.org/10.1111/1462-2920.15902
Zhu X, Chen B, Zhu L, Xing B (2017) Effects and mechanisms of biochar-microbe interactions in soil improvement and pollution remediation: a review. Environ Pollut 227:98–115. https://doi.org/10.1016/j.envpol.2017.04.032
Acknowledgements
The authors thank all the researchers who uploaded the high-throughput sequencing data related to biochar studies.
Funding
This study was financially supported by the National Natural Science Foundation of China (42077090, 42377107) and the Key Science and Technology Research and Development Project of Hangzhou (202204T05).
Author information
Authors and Affiliations
Contributions
CL and YL designed the study. CL wrote the first manuscript and YL, HQ and TL contributed substantially to revision. CL contributed to the data collection, metadata analysis and visualization. YL and TL acquired funding for this project.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Handling editor: Jun Meng
Supplementary Information
Additional file 1:
The geographical region, country, and regional information of the high-throughput sequencing samples used in this study (Fig. S1); The suitability of Sloan’s neutral model for analyzing community assembly processes under biochar application (Fig. S2); Co-occurrence network analysis of microbial communities in biochar-treated and untreated soils (Fig. S3); The representation of random forest model in metadata collection (Fig. S4); Classification of bacterial metabolic function in biochar and control groups (Fig. S5); Co-occurrence network explains the relationship among biomarkers, fungal and bacterial community (Fig. S6); Bacteria communities based on hyperparameter tuning of a tenfold cross-validated machine learning algorithm (Fig. S7); Fungal communities based on hyperparameter tuning of a tenfold cross-validated machine learning algorithm (Fig. S8).
Additional file 2:
Metadata of 1548 16S rRNA gene and 678 ITS high throughput sequence data containing geographic information and Bio-project numbers used in this study (Tables S1 and S2); Physicochemical properties of biochar (Table S3); Co-occurrence network topological parameter of soil microbes in biochar and control groups (Tables S4 and S5); Biomarkers (bacteria and fungi) of soil microbial communities in biochar and control groups identified by random forest model (Tables S6 and S7); Co-occurrence network of bacterial and fungal biomarkers (Tables S8 and S9); Overview of the main microbiological phyla (Tables S10).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lei, C., Lu, T., Qian, H. et al. Machine learning models reveal how biochar amendment affects soil microbial communities. Biochar 5, 89 (2023). https://doi.org/10.1007/s42773-023-00291-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42773-023-00291-1