Abstract
Purpose of review
Hematopoietic stem cells (HSCs) drive blood-cell production (hematopoiesis). Out-competition of HSCs by malignant cells occurs in many hematologic malignancies like acute myeloid leukemia (AML). Through mathematical modelling, HSC dynamics and their impact on healthy blood cell formation can be studied, using mathematical analysis and computer simulations. We review important work within this field and discuss mathematical modelling as a tool for attaining biological insight.
Recent findings
Various mechanism-based models of HSC dynamics have been proposed in recent years. Key properties of such models agree with observations and medical knowledge and suggest relations between stem cell properties, e.g., rates of division and the temporal evolution of the HSC population. This has made it possible to study how HSC properties shape clinically relevant processes, including engraftment following an HSC transplantation and the response to different treatment.
Summary
Understanding how properties of HSCs affect hematopoiesis is important for efficient treatment of diseases. Mathematical modelling can contribute significantly to these efforts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To understand cell dynamics in a living organism, it is crucial that any tool for observation—be it an experimental, theoretical, or metaphorical tool—interferes as little as possible with the system being observed. The heat of a lamp or the disturbance of a needle might perturb the behavior of the biological system and lead to erroneous conclusions about it. In fact, it is well-known from fundamental physics that any observation perturbs the observed system, and hence that any experiment should minimize such unwanted perturbations [1]. Hence, it can be preferable to deduce biological insights, not only from direct measurements, but also indirectly based on non-invasive observations. One particular tool for such a task is mathematical modelling. By mathematically representing key components and mechanisms of a biological system, it is possible to carry out calculations and mathematical analysis, granting insight about the mathematical representation which may also hold for the biological system being modelled.
Simultaneously, mathematical modelling can be a tool for verifying if observed phenomena follow naturally from a particular proposed biological mechanism, or if some other explanation agrees better with observed data.
An area of research where mathematical modelling has been used recently is in understanding hematopoiesis, the process of blood cell production. Hematopoiesis arises from hematopoietic stem cells (HSCs), which are typically found within the bone marrow (BM). While the observation of blood cells and their abundance can be informative about hematopoiesis, the specific properties and dynamics of HSCs in their natural setting remain elusive. Mathematical modelling is one method for investigating HSCs via indirect observations from blood samples and for verifying or proposing novel ideas and hypotheses about the mechanisms underlying HSC regulation.
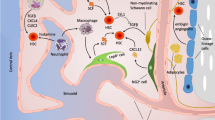
In this article, we give an overview of the role mathematical modelling has played in understanding the behavior of HSCs. Some of the approaches and concepts are illustrated in Fig. 1 for reference.
Mathematical modelling of hematopoietic stem cells covers a wide range of approaches and ideas. In the figure, some example concepts are illustrated. These illustrations do not reflect any particular model described in this paper but are only conceptual. Left panel: Starting from biological knowledge, the first step in mathematical modelling is to construct an abstract representation of the system considered. Such abstract representation could, as an example, be a diagram of cell behavior. A different abstraction is the compartment diagram, in which each group of cells is illustrated as a compartment, with arrows representing flows of cells from one compartment to another. In such diagrams, compartments do not need to correspond to biological compartments but can resolve more or less details of cellular sets or populations. Middle panel: There exist multiple mathematical or computational frameworks to represent a model. Common examples are systems of ordinary or stochastic differential equations, or individual-based/agent-based models, but a multitude of other frameworks has emerged. Right panel: Through analysis and simulation of a mathematical model, it is possible to investigate different aspects, and in turn provide potential insights into the real system being modelled. At the top, we illustrate how a model could investigate the effects of hypothetical medical interventions, while the bottom figure illustrates how differences in cell properties can yield different outcomes of some, here unspecified, outputs of interest
The Benefit of Mathematical Modelling
Mathematical modelling has long been used to investigate biological systems, particularly the human body. In a recent overview, Mackey and Maini [2] address the question “What has mathematics done for biology?” and discuss significant results about biology arising from mathematical investigations, with examples from diverse areas like pattern formation during embryonic development, atherosclerosis and neuro-oncology, among others.
In addition to the challenge of obtaining data without disturbing the biological system, as discussed above, ethical concerns may result in additional restrictions on which type of investigation is possible. Hence, mathematics is not simply a tool for making forecasts from existing data but can also contribute—as a “mathematical microscope”—to access otherwise hardly accessible properties of a biological system [3]. Using mathematical modelling, the relation between biological mechanisms and observations can be quantitatively understood, and novel ideas can be studied mathematically before doing costly experiments or clinical trials.
Most malignant hematological diseases are driven by (cancer) stem or progenitor cells, but only limited information about their dynamics can be derived from clinical measurements, e.g., bone marrow biopsies. Due to the discomfort imposed on the patient and potential risks, marrow biopsies are acquired at a lower frequency compared to blood samples. Mathematical modelling makes it possible to extract additional information from blood samples compared to current clinical practice. In particular, mathematical models can link peripheral blood cell counts to processes in the bone marrow and can, therefore, be used to make implications about stem and progenitor cell abundance and dynamics based on the blood samples which are routinely taken during follow-up examinations. This is clinically relevant since it illuminates the stem cell dynamics and their role in disease progression [4,5,6,7,8, 9••]. Similarly, mathematical models can extract complementary information about stem or progenitor cell kinetics by combining marrow and blood samples [8, 9••, 10, 11].
A broad range of quantitative methods in biology and medicine rely on mathematical models. However, most often the mathematical models are hidden in lab equipment or protocols of data analysis. Well-known examples encompass CT-, MR-, MRI-, ultrasound scanners, measurement of blood pressure by plethysmography and cuffs, administration of anesthesia [3, 12–15], or regression models establishing correlations between biomarkers and quantities of interest.
Mathematical Models of HSCs
In this section, we supplement recent reviews of mathematical modelling of stem cell function written by Stiehl and Marciniak-Czochra [16], and Brunetti, Mackey, and Craig [17].
In 1978, Mackey proposed a mathematical model [18] which inspired many of the mathematical models of blood cell formation that we describe in this article. The model proposed by Mackey describes the dynamics of a population of pluripotent stem cells, explicitly considering the cell cycle with HSCs shifting between a proliferating phase and a \({G}_{0}\) resting phase. Analyzing the model, the author connects changes in stem cell proliferation with the production of blood cells. The author focuses on a group of specific disorders characterized by periodic oscillations of mature blood cell counts. Mathematical models of this class of diseases are also reviewed by Dale and Mackey in 2015 [19]. The early paper by Mackey is an illustrative example of how mathematical modelling can be used to relate cell properties, e.g., the rate of apoptosis to dynamics of the entire pool of HSCs.
In a later paper [20], Mackey shows that the equivalent of the total human body weight worth of blood cells is produced about every 7 years in the typical human adult. In the same paper, the author relates an extended version of his previous model to data from an HSC-labelling experiment, estimating the differentiation rate of HSCs in humans to be between 0.01 and 0.02 days−1 and the apoptosis rate between 0.07 and 0.23 days−1 in a steady-state situation, i.e., in a situation where cell counts remain constant. In a similar effort to understand HSC processes, Abkowitz and colleagues [21–24] relate stochastic models of HSCs to in vivo data of murine, feline, and human origin, and are able to obtain estimates of, e.g., the replication rate of human HSCs. Furthermore, the authors suggest that the number of HSCs is comparable across different mammals, with estimates for humans observed in the blood, the success is onlyin the range of 11,400 to 22,400 [23]. Based on this work and [22], Catlin et al. [24] use 11,000 HSCs as an estimate for the steady-state size of the HSC pool, a number which has been used subsequently in several other models of HSC dynamics. Further examples of the application of mathematical models to infer cell kinetics based on labelling experiments in animals can be found in [25••] and [26].
Another early approach to model HSC dynamics is proposed in the work of Dingli and Michor [27]. In this model, hematopoiesis is described by a system of ordinary differential equations, in which one of the variables represents the pool of HSCs. The authors consider a rate of self-renewing production of HSCs, a rate of apoptosis as well as a rate of production of differentiated mature blood cells. The model is then extended for a population of leukemic stem cells (LSCs) with similar behavior, where division rates of the healthy HSCs and the LSCs are negatively regulated by the total number of stem cells. The resulting model captures how healthy hematopoiesis is suppressed by an increase in LSCs as hematologic malignancy progresses. The most significant result of the mathematical analysis is provided in the title: “Successful therapy must eradicate cancer stem cells” [27]. The high genetic similarity of diagnosis and relapse samples in acute myeloid leukemia, a severe blood cancer, supports this claim. Observations in sequencing studies are in line with the concept that the relapse is either triggered by cancer stem cells which have survived therapy or by malignant or pre-malignant stem cells which have acquired additional mutations [54]. This suggests that while some leukemia therapies may be successful at reducing the disease burden observed in the blood, the success is only temporary if they do not act on the population of HSCs and LSCs. This conclusion has later been confirmed by related models [6, 28]. In their later work, Dingli and coworkers propose models of hematopoiesis which account for multiple immature cell states [29, 30]. These models are used to predict the number of mitotic events linking the stem cell state to the mature cell compartment and to infer the phenotypic consequences of a mutation linked to oscillating blood cell counts.
Another approach to the mathematical modelling of HSCs is described in the model proposed by Roeder and Loeffler [31] and further investigated together with coworkers [32–34]. In the model, cells are assumed to move between two distinct growth environments, named GE-A and GE-Ω. In one of these abstract environments, GE-Ω, the cells are actively cycling and dividing while they are quiescent (non-dividing) in the other environment, GE-A. Each cell is characterized by its cycling status and its affinity to the GE-A environment. In contrast to other HSC models where a predefined pool of HSCs divides according to a model recapitulating the classical cell cycle phases, the model of Roeder and Loeffler [31] considers stemness and cycling activity as a property that emerges from cells dynamically switching between the two environments. The switching of a given cell between the environments is described as a function of its affinity to the GE-A environment and of the cell counts. Affinity is preserved or raised during quiescence while division degrades affinity. When affinity is sufficiently small, the cell is considered as terminally differentiated and unable to return to quiescence. Roeder and Loeffler [31] show through model simulations that the model is consistent with various experimental results from the literature, including those discussed by Abkowitz et al. [21]. Hence, the model provides a framework to understand experimental data, with the perspective that one should not only consider cell stemness as intrinsic to the cells but rather consider the interaction between HSCs and their environment. The notion of HSC reversibly exiting and re-entering quiescence, possibly adapting dynamically to external signalling, is in agreement with the notion of bone marrow stem cell niches, i.e., specific micro-environments supporting HSC quiescence and repair. A thorough review of the biological research on HSC niches is given by Wilson and Trumpp [35]. Trumpp, Essers, and Wilson [36] suggest that hematologic malignancies could be efficiently treated by combination therapy in which one drug stimulates the activation of quiescent HSCs while another drug then eradicates the newly activated non-quiescent HSCs. This is later investigated in the work of Glauche et al. [37] by adapting the model from [31], providing a mechanistic explanation behind the efficacy of combination therapy of chronic myeloid leukemia (CML).
A recent study by Ashcroft and colleagues models the binding of HSCs to HSC-specific niches in the bone marrow [38••]. In the model, HSCs detach from the niche and enter the peripheral blood at a certain rate. Upon returning to the bone marrow, the HSCs can reattach to unoccupied niches. Natural death of HSCs is assumed to occur more frequently in the peripheral blood, while only niche-bound HSCs divide. In the model, the majority of HSCs are assumed to be attached to a niche and quiescent, with a low number of unoccupied niches available in steady-state hematopoiesis. While Ashcroft et al. specifically relate the model to murine data, the model and related results should be valid for human HSCs as well. By using ordinary and stochastic differential equations, the authors investigate the prerequisites for clonal dominance in the stem cell niche. They conclude that clonal dominance in mice requires a selective advantage and cannot be the result of neutral drift. Furthermore, their model is used to investigate HSC dynamics after bone marrow transplantation. In this context, transplantation with and without prior emptying of the niches by chemotherapy (so-called preconditioning) is considered. Following the transplantation of HSCs without selective advantage into the blood stream, the engraftment of the transplanted cells in the bone marrow is limited by the number of unoccupied niches. The model can explain the experimental observation from Bhattacharya et al. [39, 40] that multiple smaller transplantations over a few days can lead to higher uptake of HSCs into the murine bone marrow compartment compared to a single-bolus transplant [38••]. The authors also use the model to predict the chimerism (abundance of donor-derived cells) after the transplantation of cells without selective advantage. Furthermore, Ashcroft et al. [38••] investigate how the probability of reconstitution after preconditioning depends on the transplanted stem cell dose. Special attention is paid to the scenario where only a single cell is transplanted. Experimental verification of such scenarios can be challenging, although not entirely impossible [41]. This is an example of an experiment that can easily be carried out in silico in a mathematical model. A related example is how patients, which in reality can only receive one treatment at a time, would have responded to a different treatment protocol or a different stem cell dose [42]. Other models of bone marrow transplantation include [43, 44] for the human and [45••] for the murine case.
Based on the observation that HSC counts are reduced in many acute myeloid leukemia (AML) patients [46], Stiehl et al. and Wang et al. [9••, 46] propose a model where HSCs and LSCs share a joint stem cell niche. This model is an extension of previous models which have been shown to recapitulate clinical blood sample data over time following bone marrow transplantation [42, 43] and the growth of leukemic cells [8, 9••, 10]. The model is given by a system of ordinary differential equations. Stem cell dynamics in the niche are coupled to the time evolution of progenitor, precursor, and mature cells as well as leukemic blasts. Clonal heterogeneity of HSCs and in turn the expansion of, e.g., a leukemic clone in the model of [38••] requires the presence of unoccupied niches and spontaneous detachment of HSCs from the niches. The model proposed in [9••, 46] considers a scenario where LSCs can actively dislodge HSCs from the niche. Since stem cells require the niche to maintain stemness properties, an invading clone has to conquer niche spaces to expand. Upon division of a LSC one of the two progeny occupies the niche of the parent, whereas the other attempts to dislodge an HSC from the niche. If this is successful, the dislodged HSC differentiates and the LSC maintains its stemness. In the opposite case, i.e., after a certain number of futile dislodgement attempts, the LSC differentiates. Similar assumptions hold for HSCs; however, to observe the expansion of leukemic cells, the probability that LSCs dislodge HSCs must outweigh the probability that HSCs dislodge LSCs. A leukemic clone with a sufficiently pronounced ability to dislodge HSCs is able to take over the niche, while the blood production might initially only show minor signs of the disease since it is transiently maintained by long-term progenitors. This model can explain the observation from [46] that AML patients with low HSC frequencies (in [46] defined as HSC frequencies below 15% of the median HSC frequency of healthy human individuals) have a poor survival. Furthermore, it is in line with the observation that HSC counts decrease before overt relapse of the AML [46]. Models assuming that HSCs and LSCs reside in separate niches and interact via systemic feedbacks that depend on the mature cell and leukemic blast counts cannot reproduce these observations [46]. A quantitative version of this model can be used to understand how the competition of stem cells for niche spaces impacts the clinical course of AML [9••, 11]. The authors develop a model-based risk stratification which uses HSC and blast counts at the time of AML diagnosis [9••] and show that this approach allows to refine the prognostic scoring of the patient cohort from [46].
In most of the models discussed above, the number of HSC niches is assumed to be independent of the dynamics of the HSC pool. To investigate the effect of feedback from HSCs to the niche-forming cells, [47] explicitly model the population of niche cells. Under the assumptions that niche cells undergo apoptosis in the absence of HSCs and that contact of HSCs to niche cells induces quiescence of HSCs but proliferation of niche cells, the model exhibits a homeostatic state. After perturbations, the model returns to homeostasis in a dampened oscillatory manner reminiscent of integral feedback control. The work of [47] illustrates that explicit modelling of the cells constituting the BM-niche can provide additional insights into the regulations of HSC dynamics.
Finally, recent work by the authors of this review aimed to model competition between HSC clones through niche occupation. Inspired by many of the models described above, a model was proposed in which competition for a limited pool of niches naturally leads to an expression of HSC fitness [48]. When two clones compete for the same niche space, e.g., in a situation where a leukemic clone has arisen due to mutations, the clone-specific fitness determines which of the two clones will eventually out-compete the other. In a scenario where the two clones have the exact same fitness, the clones will co-exist indefinitely. The model provides an expression of clonal fitness in terms of stem cell properties such as rates of proliferation, differentiation, and niche attachment. Combining this HSC model with a previous model of myeloproliferative neoplasms (MPNs) developed by Andersen et al. [49] suggests that differences in HSC fitness can explain both the disease-progression of MPNs as well as the efficacy of certain treatment protocols [50].
As approximations of reality models naturally have limitations. Besides limitations originating from the mathematical assumptions of the used modelling frameworks such as the high computational costs in the case of individual-based models or the neglect of random events in ordinary differential equation, there exist limitations coming from gaps in biological knowledge or from simplifying assumptions made during model design. One common source of limitations is the sparsity of in vivo data, especially in the human case. This sparsity originates from the inaccessibility of relevant quantities, such as kinetic parameters of stem cells, or the small number of follow-up time points. The sparsity of the data can lead to the unidentifiability of model parameters and, therefore, make it impossible to quantify important processes. Due to the sparsity of the data, it can also be challenging to distinguish between competing models. Since most human cancers require an instantaneous start of the therapy, longitudinal data of untreated cancers is practically unavailable. This makes it challenging to validate or to parameterize models of cancer dynamics. In most cases, invasive sampling techniques such as biopsies are required to extract the relevant information about a cancer. Since usually healthy individuals are not subjected to these invasive procedures, data on cancer evolution before clinical disease manifestation is rarely available. Therefore, a large part of our knowledge on tumor evolution is derived from experimental data or has been obtained by applying modelling or heuristic techniques to samples obtained at the time of diagnosis. Furthermore, important parts of our pathophysiological and pharmacological knowledge have been gained using animal or in vitro models, which do not necessarily reflect the human system [51, 52]. Although the latter may improve in the future due to advanced organoid or microfluidic systems, it has to be taken into account that human diseases are more complex than specific experimental systems [51, 52]. These limitations make it vital to validate models using real-world data and to be aware of the simplifying assumptions that have been applied during model development. In the context of personalized medicine, the applicability of models in clinical routine is further limited by the large inter-individual heterogeneity of the disease evolution, which is impacted by potentially unknown disease-specific factors (e.g., key mutations), patient-specific factors and environmental factors. Partially, this heterogeneity can be incorporated by the choice of individual model parameters for each patient [8, 9••, 11]. This, however, requires that the individualized parameters can be estimated based on patient data or derived from statistical or other approaches. Integration of mechanistic and artificial intelligence models is a promising direction to improve personalized medicine [53].
Conclusion
Through mathematical modelling, a range of significant insights into the otherwise hidden in vivo dynamics of HSCs has been obtained.
While cell division, apoptosis, and differentiation of individual cells are assumed to occur in response to a complicated system of regulatory processes, the average rates at which such events happen can often be estimated by relating mathematical models to experimental data. As an example, mathematical models can be used to understand how changes in such rates affect the production of blood cells. Such insight is particularly useful when investigating hematologic malignancies, where a disease may be caused by mutations which alter the rate at which a given biological process occurs. Mathematical modelling helps relate mutation-induced changes of stem cell properties to changes on the clinically accessible scale which are used to define disease entities. Furthermore, mathematical terminology and formalisms can substantially contribute to make more vaguely defined concepts rigorous whereby ambiguity may be avoided.
Naturally, there exist questions which are inaccessible to mathematical models and require in vivo experiments. Nevertheless, the methods of quantitative modelling provide a very useful starting point for investigating relevant hypotheses involving HSCs. Model selection techniques allow for falsifying of hypotheses by comparing model dynamics to available data. If a model of HSC dynamics does not agree with clinical or experimental data, at least one of the hypotheses underlying the model has to be incorrect.
The works reviewed here show different approaches to model HSC dynamics mathematically. Although some aspects of different models might be inconsistent with each other, the variety of proposed models that agree with experimental data illustrates the enigmatic nature of HSC behavior. Furthermore, it underlines an important aspect of mathematical modelling: Although a particular mathematical model might not describe all components of a biological system in detail, the model dynamics can still be sufficiently accurate to provide valuable insight and forecast how the system would behave under certain circumstances. Hence, the current state of mathematical modelling of HSC dynamics is a useful part of bio-medical research. Further work on current models and the development of new models will be necessary to understand and interpret novel experimental findings.
References
Papers of particular interest, published recently, have been highlighted as: •• Of major importance
Bohr N. The quantum postulate and the recent development of atomic theory1. Nature. 1928;121:580–90. https://doi.org/10.1038/121580a0.
Mackey MC, Maini PK. What has mathematics done for biology? Bull Math Biol. 2015;77:735–8. https://doi.org/10.1007/s11538-015-0065-9.
Ottesen JT. The Mathematical Microscope – Making the inaccessible accessible. BetaSys Syst. Biol. Regul. exocytosis Pancreat. β-cells, New York, NY: Springer New York; 2011, p. 97–118. https://doi.org/10.1007/978-1-4419-6956-9_6.
Pedersen RK, Andersen M, Knudsen TA, Sajid Z, Gudmand-Hoeyer J, Dam MJB, et al. Data-driven analysis of JAK2 V617F kinetics during interferon-alpha2 treatment of patients with polycythemia vera and related neoplasms. Cancer Med. 2020;9:2039–51. https://doi.org/10.1002/cam4.2741.
Sajid Z, Andersen M, Ottesen JT. System dynamics of cancer in erythropoiesis with multiple EPO feedbacks. Syst Dyn Rev. 2020;36:447–66. https://doi.org/10.1002/sdr.1670.
Ottesen JT, Pedersen RK, Sajid Z, Gudmand-Hoeyer J, Bangsgaard KO, Skov V, et al. Bridging blood cancers and inflammation: the reduced Cancitis model. J Theor Biol. 2019;465:90–108. https://doi.org/10.1016/j.jtbi.2019.01.001.
Ottesen JT, Andersen M. Potential of immunotherapies in treating hematological cancer-infection comorbidities—a mathematical modelling approach. Cancers (Basel). 2021;13:3789. https://doi.org/10.3390/cancers13153789.
Stiehl T, Baran N, Ho AD, Marciniak-Czochra A. Cell division patterns in acute myeloid leukemia stem-like cells determine clinical course: a model to predict patient survival. Cancer Res. 2015;75(6):940–9. https://doi.org/10.1158/0008-5472.CAN-14-2508.
•• Stiehl T, Wang W, Lutz C, Marciniak-Czochra A. Mathematical modeling provides evidence for niche competition in human AML and serves as a tool to improve risk stratification. Cancer Res. 2020;80:3983–92. https://doi.org/10.1158/0008-5472.CAN-20-0283. The authors consider a quantitative model of blood cell formation in acute myeloid leukemia patients. The model describes the competition of HSCs and LSCs for spaces in a joined stem cell niche with fixed capacity. The authors identify which parameters of LSCs impact on the course of the disease and develop a risk-stratification approach which combines the mathematical model with data of individual patients.
Stiehl T, Ho AD, Marciniak-Czochra A. Mathematical modeling of the impact of cytokine response of acute myeloid leukemia cells on patient prognosis. Sci Rep 2018:1–11. https://doi.org/10.1038/s41598-018-21115-4.
Stiehl T. Using mathematical models to improve risk-scoring in acute myeloid leukemia. Chaos An Interdiscip J Nonlinear Sci 2020;30:123150. https://doi.org/10.1063/5.0023830.
Große Ruse M, Søndergaard LR, Ditlevsen S, Damgaard M, Fuglsang S, Ottesen JT, et al. Absorption and initial metabolism of 75 Se- l -selenomethionine: a kinetic model based on dynamic scintigraphic data. Br J Nutr. 2015;114:1718–23. https://doi.org/10.1017/S000711451500344X.
Ottesen JT. Do not ask what mathematics can do for modelling. Teach. Learn. Math. Univ. Lev., Dordrecht: Kluwer Academic Publishers; n.d., p. 335–46. https://doi.org/10.1007/0-306-47231-7_30.
Ottesen JT, Olufsen MS, Larsen JK. Applied mathematical models in human physiology. Society for Industrial and Applied Mathematics; 2004. https://doi.org/10.1137/1.9780898718287.
Ottesen JT, Danielsen M. Mathematical modelling in medicine. IOS Press; 2000.
Stiehl T, Marciniak-Czochra A. How to characterize stem cells? Contributions from mathematical modeling. Curr Stem Cell Reports. 2019;5:57–65. https://doi.org/10.1007/s40778-019-00155-0.
Brunetti M, Mackey MC, Craig M. Understanding normal and pathological hematopoietic stem cell biology using mathematical modelling. Curr Stem Cell Reports. 2021;7:109–20. https://doi.org/10.1007/s40778-021-00191-9.
Mackey MC. Unified hypothesis for the origin of aplastic anemia and periodic hematopoiesis. Blood. 1978;51:941–56. https://doi.org/10.1182/blood.V51.5.941.941.
Dale DC, Mackey MC. Understanding, treating and avoiding hematological disease: better medicine through mathematics? Bull Math Biol. 2015;77:739–57. https://doi.org/10.1007/s11538-014-9995-x.
Mackey MC. Cell kinetic status of haematopoietic stem cells. Cell Prolif. 2001;34:71–83. https://doi.org/10.1046/j.1365-2184.2001.00195.x.
Abkowitz JL, Catlin SN, Guttorp P. Evidence that hematopoiesis may be a stochastic process in vivo. Nat Med. 1996;2:190–7. https://doi.org/10.1038/nm0296-190.
Abkowitz JL, Golinelli D, Harrison DE, Guttorp P. In vivo kinetics of murine hemopoietic stem cells. Blood. 2000;96:3399–405. https://doi.org/10.1182/blood.V96.10.3399.
Abkowitz JL, Catlin SN, McCallie MT, Guttorp P. Evidence that the number of hematopoietic stem cells per animal is conserved in mammals. Blood. 2002;100:2665–7. https://doi.org/10.1182/blood-2002-03-0822.
Catlin SN, Busque L, Gale RE, Guttorp P, Abkowitz JL. The replication rate of human hematopoietic stem cells in vivo. Blood. 2011;117:4460–6. https://doi.org/10.1182/blood-2010-08-303537.
•• Busch K, Klapproth K, Barile M, Flossdorf M, Holland-Letz T, Schlenner SM, et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature. 2015;518:542–6. https://doi.org/10.1038/nature14242. Combination of experimental work and mathematical modeling to provide quantitative insights in the dynamics of murine hematopoiesis under steady state conditions. The authors use inducible genetic labels and extract steady state HSC and progenitor cell kinetics (e.g., division and differentiation rates) by fitting a mathematical model to the observed time dynamics of label frequencies.
Stalmann USA, Ticconi F, Snoeren IAM, Li R, Gleitz HFE, Cowley GS, et al. Genetic barcoding systematically compares genes in del(5q) MDS and reveals a central role for CSNK1A1 in clonal expansion. Blood Adv. 2022;6:1780–96. https://doi.org/10.1182/bloodadvances.2021006061.
Dingli D, Michor F. Successful therapy must eradicate cancer stem cells. Stem Cells. 2006;24:2603–10. https://doi.org/10.1634/stemcells.2006-0136.
Andersen M, Hasselbalch HC, Kjær L, Skov V, Ottesen JT. Global dynamics of healthy and cancer cells competing in the hematopoietic system. Math Biosci. 2020;326:1–15. https://doi.org/10.1016/j.mbs.2020.108372.
Dingli D, Traulsen A, Pacheco JM. Compartmental architecture and dynamics of hematopoiesis. PLoS One 2007;2:e345. https://doi.org/10.1371/journal.pone.0000345.
Dingli D, Antal T, Traulsen A, Pacheco JM. Progenitor cell self-renewal and cyclic neutropenia. Cell Prolif. 2009;42:330–8. https://doi.org/10.1111/j.1365-2184.2009.00598.x.
Roeder I, Loeffler M. A novel dynamic model of hematopoietic stem cell organization based on the concept of within-tissue plasticity. Exp Hematol. 2002;30:853–61. https://doi.org/10.1016/S0301-472X(02)00832-9.
Roeder I, Kamminga LM, Braesel K, Dontje B, De Haan G, Loeffler M. Competitive clonal hematopoiesis in mouse chimeras explained by a stochastic model of stem cell organization. Blood. 2005;105:609–16. https://doi.org/10.1182/blood-2004-01-0282.
Roeder I, Horn M, Glauche I, Hochhaus A, Mueller MC, Loeffler M. Dynamic modeling of imatinib-treated chronic myeloid leukemia: functional insights and clinical implications. Nat Med. 2006;12:1181–4. https://doi.org/10.1038/nm1487.
Glauche I, Moore K, Thielecke L, Horn K, Loeffler M, Roeder I. Stem cell proliferation and quiescence - two sides of the same coin. PLoS Comput Biol. 2009;5:3–12. https://doi.org/10.1371/journal.pcbi.1000447.
Wilson A, Trumpp A. Bone-marrow haematopoietic-stem-cell niches. Nat Rev Immunol. 2006;6:93–106. https://doi.org/10.1038/nri1779.
Trumpp A, Essers M, Wilson A. Awakening dormant haematopoietic stem cells. Nat Rev Immunol. 2010;10:201–9. https://doi.org/10.1038/nri2726.
Glauche I, Horn K, Horn M, Thielecke L, Essers MAG, Trumpp A, et al. Therapy of chronic myeloid leukaemia can benefit from the activation of stem cells: simulation studies of different treatment combinations. Br J Cancer. 2012;106:1742–52. https://doi.org/10.1038/bjc.2012.142.
•• Ashcroft P, Manz MG, Bonhoeffer S. Clonal dominance and transplantation dynamics in hematopoietic stem cell compartments. PLOS Comput Biol 2017;13. https://doi.org/10.1371/journal.pcbi.1005803. This work illustrates the use of simple mathematical models of HSCs, by relating a model that is conceptually simple with experiments from the literature. Through mathematical analysis of the model, the authors investigate the outcomes of transplantation and the probability of successful reconstitution.
Bhattacharya D, Rossi DJ, Bryder D, Weissman IL. Purified hematopoietic stem cell engraftment of rare niches corrects severe lymphoid deficiencies without host conditioning. J Exp Med. 2006;203:73–85. https://doi.org/10.1084/jem.20051714.
Bhattacharya D, Czechowicz A, Ooi AGL, Rossi DJ, Bryder D, Weissman IL. Niche recycling through division-independent egress of hematopoietic stem cells. J Exp Med. 2009;206:2837–50. https://doi.org/10.1084/jem.20090778.
Osawa M, Hanada K, Hamada H, Nakauchi H. Long-term lymphohematopoietic reconstitution by a single CD34-low/negative hematopoietic stem cell. Science (80- ) 1996;273:242–5. https://doi.org/10.1126/science.273.5272.242.
Stiehl T, Ho AD, Marciniak-Czochra A. The impact of CD34+ cell dose on engraftment after SCTs: personalized estimates based on mathematical modeling. Bone Marrow Transplant. 2014;49:30–7. https://doi.org/10.1038/bmt.2013.138.
Marciniak-Czochra A, Stiehl T, Ho AD, Jäger W, Wagner W. Modeling of asymmetric cell division in hematopoietic stem cells—regulation of self-renewal is essential for efficient repopulation. Stem Cells Dev. 2009;18:377–86. https://doi.org/10.1089/scd.2008.0143.
Østby I, Rusten LS, Kvalheim G, Grøttum P. A mathematical model for reconstitution of granulopoiesis after high dose chemotherapy with autologous stem cell transplantation. J Math Biol. 2003;47:101–36. https://doi.org/10.1007/s00285-003-0198-6.
•• Manesso E, Teles J, Bryder D, Peterson C. Dynamical modelling of haematopoiesis: an integrated view over the system in homeostasis and under perturbation. J R Soc Interface 2013;10. https://doi.org/10.1098/rsif.2012.0817. The authors consider a detailed model of murine hematopoiesis with various feedback loops. Using data from literature in combination with an extensive simulation and optimization approach the authors establish a parameterization which is able to recapitulate blood cell dynamics after transplantation and hemorrhage.
Wang W, Stiehl T, Raffel S, Hoang VT, Hoffmann I, Poisa-Beiro L, et al. Reduced hematopoietic stem cell frequency predicts outcome in acute myeloid leukemia. Haematologica. 2017;102:1567–77. https://doi.org/10.3324/haematol.2016.163584.
Becker NB, Günther M, Li C, Jolly A, Höfer T. Stem cell homeostasis by integral feedback through the niche. J Theor Biol. 2019;481:100–9. https://doi.org/10.1016/j.jtbi.2018.12.029.
Pedersen RK, Andersen M, Stiehl T, Ottesen JT. Mathematical modelling of the hematopoietic stem cell-niche system: clonal dominance based on stem cell fitness. J Theor Biol 2021;518:110620. https://doi.org/10.1016/j.jtbi.2021.110620.
Andersen M, Sajid Z, Pedersen RK, Gudmand-Hoeyer J, Ellervik C, Skov V, et al. Mathematical modelling as a proof of concept for MPNs as a human inflammation model for cancer development. PLoS One 2017;12. https://doi.org/10.1371/journal.pone.0183620.
Pedersen RK, Andersen M, Knudsen TA, Skov V, Kjær L, Hasselbalch HC, et al. Dose‐dependent mathematical modeling of interferon‐α‐treatment for personalized treatment of myeloproliferative neoplasms. Comput Syst Oncol 2021;1. https://doi.org/10.1002/cso2.1030.
Mukherjee P, Roy S, Ghosh D, Nandi SK. Role of animal models in biomedical research: a review. Lab Anim Res. 2022;38(1):18.
Ingber DE. Is it time for reviewer 3 to request human organ chip experiments instead of animal validation studies? Adv Sci (Weinh). 2020;7(22):2002030.
Chakravarty K, Antontsev V, Bundey Y, Varshney J. Driving success in personalized medicine through AI-enabled computational modeling. Drug Discov Today. 2021;26(6):1459–65.
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS, Ritchey JK, Young MA, Lamprecht T, McLellan MD, McMichael JF, Wallis JW, Lu C, Shen D, Harris CC, Dooling DJ, Fulton RS, Fulton LL, Chen K, Schmidt H, Kalicki-Veizer J, Magrini VJ, Cook L, McGrath SD, Vickery TL, Wendl MC, Heath S, Watson MA, Link DC, Tomasson MH, Shannon WD, Payton JE, Kulkarni S, Westervelt P, Walter MJ, Graubert TA, Mardis ER, Wilson RK, DiPersio JF. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature. 2012;481(7382):506–10. https://doi.org/10.1038/nature10738.PMID:22237025;PMCID:PMC3267864.
Funding
This work was supported by the Fellowship “Personalized prediction of blood cancer progression using clinical data and mathematical modeling” from the Lundbeck Foundation (PI: TS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pedersen, R.K., Andersen, M., Stiehl, T. et al. Understanding Hematopoietic Stem Cell Dynamics—Insights from Mathematical Modelling. Curr Stem Cell Rep 9, 9–16 (2023). https://doi.org/10.1007/s40778-023-00224-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40778-023-00224-5