Abstract
Introduction
Neurotrophic tyrosine receptor kinase (NTRK) gene fusions occur in ~ 0.3% of all solid tumours but are enriched in some rare tumour types. Tropomyosin receptor kinase (TRK) inhibitors larotrectinib and entrectinib are approved as tumour-agnostic therapies for solid tumours harbouring NTRK fusions.
Methods
This study investigated the prevalence of NTRK fusions in Canadian patients and also aimed to help guide NTRK testing paradigms through analysis of data reported from a national clinical diagnostic testing program between September 2019 and July 2021.
Results
Of 1,687 patients included in the final analysis, NTRK fusions were detected in 0.71% (n = 12) of patients representing salivary gland carcinoma (n = 3), soft tissue sarcoma (n = 3), CNS (n = 3), and one in each of melanoma, lung, and colorectal cancer. All three salivary gland carcinomas contained ETV6-NTRK3 fusions. Thirteen (0.77%) clinically actionable incidental findings were also detected. Two of the 13 samples containing incidental findings were NTRK fusion-positive (GFOD1-NTRK2, FGFR3-TACC3 in a glioblastoma and AFAP1-NTRK2, BRAF c.1799T>A in a glioma). The testing algorithm screened most patient samples via pan-TRK immunohistochemistry (IHC), whereas samples from the central nervous system (CNS), pathognomonic cancers, and confirmed/ putative NTRK fusion-positive samples identified under research protocols were reflexed straight to next-generation sequencing (NGS).
Conclusion
These findings highlight the benefit and practicality of a diagnostic testing program to identify patients suitable for tumour-agnostic TRK inhibitor therapies, as well as other targeted therapies, due to clinically actionable incidental findings identified. Collectively, these findings may inform future guidance on selecting the appropriate testing approach per tumour type and on optimal NTRK testing algorithms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
A national clinical diagnostic testing program was established to address a gap in NTRK testing infrastructure in Canada. |
Out of 1,687 patients and 37 primary tumour types tested, the most commonly submitted tissue types for testing were colorectal cancer, lung/pleura, and soft tissue sarcoma. |
NTRK fusions were detected in 0.71% of patients. NTRK fusions were detected in salivary gland carcinomas, soft tissue sarcomas, primary CNS tumours, and one in each of melanoma, lung, and colorectal cancer. The most common fusion was ETV6-NTRK3, which was diagnosed for all three salivary gland carcinomas. |
Additional clinically actionable genomic findings were detected in 0.77% of patients. |
These findings may inform future diagnostic paradigms used to identify patients eligible for tumour-agnostic targeted TRK inhibitor therapy. |
1 Introduction
The neurotrophic tyrosine receptor kinase (NTRK) genes, NTRK1, NTRK2, and NTRK3, encode the tropomyosin receptor kinase (TRK) proteins TRKA, TRKB, and TRKC, respectively [1], which bind neurotrophins and act to regulate neuronal development and function [2]. NTRK genes are predominantly transcribed in the nervous system of adult tissues, and are also expressed during embryonic development [3]. A number of NTRK gene alternations, including point mutations, overexpression, amplifications, and fusions, have been previously described [4, 5]. In TRK fusion cancer, TRK fusion proteins exhibit the ability to remain constitutively active independent of a stimulating ligand and may cause cancer cells to grow [1, 4, 6]. The incidence of NTRK fusions in solid tumours overall is very low, occurring in only approximately 0.3% of all tumours [7,8,9]. Notably, NTRK fusions are common in select rare tumour types, such as secretory carcinomas of the breast and salivary gland in adults and infantile fibrosarcoma (IFS) in pediatrics, where reported fusion positivity rates often exceed 75% of cases [10]. Approximately 80 different NTRK fusion partners are currently known to exist across multiple tumour types [10].
As oncogenic drivers in multiple types of cancer, NTRK fusions have clinical utility as targets of tumour-agnostic TRK inhibitor therapies, in which response to therapy is independent of tumour/tissue histology [1]. Two TRK inhibitors are currently approved in countries around the world, including Canada, for patients with NTRK fusion-positive solid tumours: larotrectinib and entrectinib. Larotrectinib is approved for adult and pediatric (< 18 years) patients and entrectinib is approved for adult and pediatric (≥ 12 years) patients or adult patients only, depending on the country [11,12,13,14,15,16]. TRK inhibitor therapies have been issued marketing authorization with conditions in Canada, as they address a serious unmet patient medical need and/or demonstrate significant improvement in the benefit/risk profile over existing therapies [11, 12]. Indeed, both Canadian and international consensus statements recommend considering TRK inhibitors for NTRK fusion-positive cancers where there is high unmet need with no available satisfactory treatment options (e.g., surgery with high morbidity, excessive treatment toxicity, or low response rates) [17,18,19,20,21,22,23,24]. In clinical trials, larotrectinib was associated with an overall response rate (ORR) of 79.1%, median duration of response (DOR) of 35.2 months, median progression-free survival (PFS) of 28.3 months, and median overall survival (OS) of 44.4 months [6]. Entrectinib was associated with an ORR of 61.2%, median DOR of 20.0 months, median PFS of 13.8 months, and median OS of 33.8 months [25].
To identify patients eligible for TRK inhibitor therapy, strategies to detect the presence of an NTRK fusion include fluorescence in situ hybridization (FISH), reverse transcriptase polymerase chain reaction (RT-PCR), and next-generation sequencing (NGS) using DNA or RNA [26]. The advantages and limitations of the different diagnostic approaches have been extensively reviewed [26, 27]. Given the antibodies used in pan-TRK IHC do not discriminate between native and chimeric TRK proteins, IHC is used as a screening tool to triage patient samples with TRK protein expression to be subjected to molecular analyses to confirm the presence of NTRK fusion [26]. Multiple NGS strategies exist, including amplicon-based multiplex PCR, anchored multiplex PCR, and hybrid capture [28]. For example, the Oncomine Comprehensive Assay v3 (OCAv3; Thermo Fisher Scientific, Waltham, MA, USA) is a multiplex amplicon-based assay covering 51 fusion drivers. The FusionPlex® Lung panel (ArcherDX, Boulder, CO, USA) uses anchored multiplex PCR, and can identify novel fusion transcripts even if only one fusion partner is known [28, 29]. Considering the complexities of NTRK fusion identification, testing algorithms are variable in clinical practice [17, 19, 23, 26, 28, 30,31,32,33,34,35,36,37,38,39,40]. Canadian oncologists, endocrinologists, pathologists, and lab directors have generated consensus on testing and treatment algorithms [17, 20]. Canadian and global studies are also investigating the validity and suitability of NTRK fusion testing methods [30, 41, 42].
To address the gap in Canadian NTRK testing infrastructure revealed upon larotrectinib Health Canada approval, a centralized clinical testing program (FastTRK) was created to enable access to NTRK fusion clinical testing for patients with metastatic disease or who are at risk of severe morbidity with surgical resection and with no satisfactory treatment options. Here, we report the results of this testing program and provide a preliminary characterization of NTRK fusion prevalence in solid tumours among this Canadian patient population. The testing program aimed to provide insights on which tumour histologies contain NTRK fusions and to provide guidance on whom, when, and how to test for NTRK fusions in Canadian patients.
2 Materials and Methods
2.1 Study Design
This study was conducted as a retrospective analysis of data derived from submitted requisition forms and laboratory results from the Canadian FastTRK testing program between 16 September 2019 and 15 July 2021. Sponsored by Bayer Inc., the FastTRK testing program is a complimentary service for clinicians to determine whether their patients’ tumour harbours an NTRK fusion. The testing program sample testing algorithm is summarized in the Online Supplementary Material (OSM), Fig. 1.
Breakdown of tests and NTRK fusion results from the clinical diagnostic testing algorithm. a Flow of all patient samples obtained from the clinical diagnostic testing program, from receipt to NGS result. b Samples which underwent reverse reflex quality control to the alternative lab and findings. *Measurements in same patient, either same tumour site repeated due to insufficient sample or same patient, different tumour site (i.e. metastases). †Insufficient tissue with cancer cells, necrotic tissue. ‡Reflexed to KHSC from LifeLabs due to failed result by Archer due to RNA quality or quantity and reflexed to LifeLabs from KHSC due to negative result with OCAv3. CNS central nervous system, IHC immunohistochemistry, KHSC Kingston Health Sciences Centre, LL LifeLabs, NGS next-generation sequencing, NTRK neurotrophic tyrosine receptor kinase, TRK tropomyosin receptor kinase.
The testing program was performed through LifeLabs and KHSC to provide NTRK fusion testing services for Canadians. Up until 27 July 2020, LifeLabs offered testing to hospitals from Ontario (except Peterborough, Ottawa, and South East Local Health Integration Network), Manitoba, Saskatchewan, Alberta, British Columbia, Nunavut, Northwest Territories, and the Yukon. At program inception, KHSC testing coverage included South East Local Health Integration Network (Kingston General Hospital, Hotel Dieu Hospital (Kingston, ON), Providence Continuing Care Centre, Lennox and Addington County General Hospital, Brockville General Hospital, Perth and Smiths Falls Hospital, Quinte Healthcare (Belleville, Prince Edward County, Trenton, North Hastings hospitals)), and hospitals from Newfoundland, Nova Scotia, New Brunswick, and Prince Edward Island. Starting 27 July 2020, samples from hospitals in Peterborough (Ontario), Ottawa (Ontario), and Quebec were directed to and tested by KHSC.
The testing program accepted samples from all patients with solid tumours that were metastatic or when surgical resection was likely to result in severe morbidity, and for whom no satisfactory treatment options were available. Patients with commonly diagnosed tumour types, such as lung cancer and colorectal cancer, known to be negative for other oncogenic alterations through exclusionary testing, were considered appropriate candidates for this program. Only patients who provided consent for their personal information to be used for scientific research were included in this study analysis. Patients who did not consent for their personal information to be used for scientific research were still able to access testing through the program.
Physicians identified patients eligible for the program and submitted details in a downloadable requisition form for NTRK fusion testing from the http://www.FastTRK.ca website. Data used from submitted requisition forms included: patient sex, patient age range (0–6, 7–12, 13–18, 19–39, 40–65, 65+ years), tissue site, tumour type, previous treatment (chemotherapy, immunotherapy, biologic, oral tyrosine kinase inhibitor, surgery, radiation, other), originating institution (name, address), prior NTRK fusion testing performed, testing lab, and patient consent for scientific research. The requisition form also indicated that if the sample was being submitted for NGS that the sample have a viable tumor cellularity > 10%. The testing labs provided the following testing results: IHC result, percent cellularity, NTRK fusion NGS result, NTRK gene involved in fusion, 5’ fusion partner, and incidental genomic findings.
2.2 Testing Algorithm and Reporting
Commonly diagnosed tumour types (e.g., lung cancer, thyroid cancer, soft tissue sarcomas, colorectal cancer), which are known to harbour lower frequencies of NTRK fusions [43], were first subjected to a pan-TRK IHC screen. Positive and inconclusive/equivocal samples from IHC were reflexed for further molecular testing via NGS.
Tumour types known to frequently harbour NTRK fusions (i.e., pathognomonic cancers such as IFS, CMN, secretory carcinoma of the salivary gland, and salivary breast cancer) [43], or where native TRK protein is endogenously found, such as CNS tissues [3, 44], were reflexed straight to NGS without undergoing pan-TRK IHC. Despite the testing algorithm indicating that CNS samples would be reflexed straight to NGS, lab pathologists used clinical discretion to assay 12 CNS samples for pan-TRK IHC to inform their clinical assessments. If patients were confirmed to be putatively NTRK fusion-positive by either IHC or NGS through prior testing (before being submitted to FastTRK), samples were also reflexed straight to NGS. Research studies that identified samples as confirmed/putative NTRK positive used their own centre-specific IHC/NGS protocols.
All samples that were pan-TRK-positive via IHC and/or NGS-negative at KHSC, or failed to produce results by Archer at LifeLabs due to RNA quality/quantity, were subjected to “reverse reflex” to undergo further NGS testing at the other lab (i.e., LifeLabs/KHSC) for reassessment and validation.
Incidental findings of potential clinical interest to researchers and/or physicians were reported by the labs and shared with the requesting clinician. In this study analysis, clinically actionable findings were defined as OncoKB Therapeutic Levels of Evidence 1–4 [45, 46]. LifeLabs and KHSC initiated reporting on incidental findings starting 19 September 2019 and 1 June 2020, respectively.
2.3 Immunohistochemistry
2.3.1 Kingston Health Sciences Centre (KHSC)
Staining was performed on a BOND III autostainer (Leica Instruments, Wetzlar, Germany). Antigen retrieval at pH 9.0 was performed for 60 min. The primary pan-TRK antibody, EPR17341 (Abcam, Cambridge, UK) was diluted 1:100 in DAKO background-reducing diluent and incubated at 22 °C for 30 min. Detection was performed using a Bond Polymer Refine Detection kit and 3,3'-diaminobenzidine chromagen. On slide controls for IHC were appendix (cytoplasmic, membranous, and nuclear in peripheral nerves and ganglia, negative staining in epithelia) and cerebellum (membranous and cytoplasmic staining in neurons).
2.3.2 LifeLabs
Pan-TRK IHC was conducted using the VENTANA pan-TRK (EPR17341) Assay (Roche Diagnostics, Basel, Switzerland) per the manufacturer protocol with the following modifications: staining was performed on a Leica Bond III platform (Leica Instruments, Wetzlar, Germany). Antigen retrieval at pH 9.0 was performed for 20 min. Incubation time with the primary pan-TRK antibody (EPR17341) was at room temperature for 15 min. Detection was performed using Polymer Refine Detection System (Leica) and 3,3ʹ-diaminobenzidine chromagen. On slide controls for IHC was cerebellum (membranous and cytoplasmic staining in neurons).
2.3.3 Interpretation of TRK Positivity by Immunohistochemistry (IHC)
Since cases with NTRK fusions can show protein cytoplasmic, membranous, or nuclear expression of TRK proteins [47], both KHSC and LifeLabs defined pan-TRK positive cases as those with at least 10% of cancer cells showing moderate or strong staining in any of these cellular compartments. Obvious positives were not reviewed by another pathologist, but any equivocal cases were reviewed by at least one other pathologist.
2.4 Next-Generation Sequencing (NGS)
2.4.1 KHSC
Total nucleic acid was extracted from sectioned formalin-fixed paraffin-embedded tissue samples using the MagMAX FFPE RNA/DNA Ultra Kit, (Thermo Fisher Scientific, Waltham, MA, USA). Tissue samples were sectioned using sterile technique, laying 4-micron sections per slide onto uncharged, unbaked slides. Extraction was targeted specifically to the tumour area as indicated by examination of a hematoxylin phloxine saffron (HPS)-stained slide and identification by a pathologist. Concentration of DNA and RNA yield was assessed by the Qubit 2.0 (Life Technologies, Carlsbad, CA, USA), and 30 ng of RNA was used for reverse transcription with the Superscript IV VILO Mastermix with ezDNase. Library construction was done according to manufacturer’s instructions with the Oncomine Comprehensive Panel v3 primers (pool 1 + pool 2) using the Ion AmpliSeq Kit for Ion CHEF System (Thermo Fisher Scientific). Oncomine Comprehensive Assay v3 contains primers for 161 genes, with known partners for 51 fusion driver genes (Thermo Fisher Scientific). After amplification and pooling, each DNA and RNA library pool was assessed for concentration and integrity using the Ion Library Taqman Quantitation Kit, after which a 50 pM 80:20 DNA:RNA library pool was templated onto ION 540 Chips, using the Ion 540 Chef Kit (Thermo Fisher Scientific). Sequencing was performed on the Ion S5XL System, with 500 flows, and subsequent quality assessment for the run was completed using the Torrent Suite Software v5.6 August 2019, v5.12 February 2020 (Thermo Fisher Scientific). Obligatory run metrics were composed of mean RNA read length (> 60 bp), mean DNA read length (> 75 bp), mean raw accuracy (> 99%), and total sequencing reads (> 40,000,000). Further sample-specific quality assessment and analysis was completed using Ion Reporter software (v5.6 August 2019, v5.12 February 2020 (Thermo Fisher Scientific)), with a threshold of 500,000 minimum total mapped fusion reads and 3,000,000 mapped DNA reads required to pass the sample for interpretation and reporting. Individual fusion variant calls required > 20 fusion counts with ≥ 1% fusion counts/total mapped fusions, and individual DNA variant calls required a variant coverage of > 400 with a p-value of ≤ 0.00001 to be considered reportable.
2.4.2 LifeLabs
Formalin-fixed paraffin-embedded tissues were sectioned at 4 microns and fixed to charged, unbaked slides. Total nucleic acid was extracted from sample sections using ReliaPrep FFPE Total RNA Miniprep System (Promega, Madison, WI, USA). Extraction was targeted specifically to the tumour area as identified by a pathologist on an H&E (hematoxylin and eosin)-stained slide. RNA concentration was assessed on the Qubit 3.0 (Life Technologies) and 100–250 ng of RNA was combined with pre-aliquoted reverse transcription mix provided in the FusionPlex® Lung panel v1.0 kit (ArcherDx, Boulder, CO, USA). Library construction was carried out according to the manufacturer’s protocol using Anchored Multiplex PCR. The FusionPlex® Lung panel contains 163 unidirectional gene-specific primers targeting 14 clinically relevant genes, including 28 fusion targets across NTRK1, NTRK2, and NTRK3 exons. Input RNA quality was assessed with the PreSeq RNA QC assay (ArcherDx), and purified libraries were assessed prior to sequencing with the KAPA Library Quantification assay (Roche Sequencing, Basel, Switzerland). Libraries were compared to six known DNA standard dilutions and pooled to equimolar concentrations. FusionPlex® libraries were sequenced on an MiSeq NGS analyzer (Illumina, San Diego, CA, USA) using a standard v2 flow cell, and data were analyzed using Archer Suite Analysis v6.04 (ArcherDx) according to pre-defined quality metrics and RNA Fusion pipeline-classified NTRK fusions. A fusion was classified as detectable if it passed all defined metrics for unique reads per GSP2 (> 3), unique RNA start sites per GSP2 control (> 10), total fragments mapped (> 95%), median RNA fragment length > 200 bp, and passing sequencing reads (> 0.5 million).
2.5 Variables of Interest and Data Analysis
Primary variables of interest included the frequencies of patient demographics, tumour types, and NTRK fusion-positive samples in the clinical testing program. Secondary variables of interest included frequency of NTRK fusion-positivity per tumour type, frequency of clinically actionable findings, rate of pan-TRK positive but NGS-negative samples, and frequency of tumour types screened by each lab. Exploratory variables of interest included IHC/NGS turnaround time at LifeLabs and analysis to determine cost of identifying one patient with genomic findings.
Descriptive statistics were used to summarize study outcomes. Data were sorted and frequencies were determined using SPSS Statistics 26 (IBM, Armonk, NY, USA). Primary tumour types were input manually on the requisition form and grouped into categories prior to analysis as per previous studies [6, 48]. Carcinomas and adenocarcinomas were categorized histologically by tissue/organ site and rare tumours were given separate categories to highlight potential for NTRK fusions. The turnaround time calculation was performed in Microsoft Excel and excluded weekends but not holidays. Costing analysis was performed in Microsoft Excel using the average cost of IHC and NGS from both labs, which did not include set-up costs, shipping, or taxes.
3 Results
3.1 Patient and Sample Characteristics
Between the two labs selected for the testing program, LifeLabs and the Kingston Health Sciences Centre (KHSC), this study included 1862 patients, with 1687 included in the analysis after omitting repeat sample measurements (n = 52) and patients who did not consent to research (n = 123) (Fig. 1a). Analyzed samples were mostly from adult patients (> 18 years, 98.5%), with 91.2% of patients aged ≥ 40 years. Patient characteristics upon enrolment in the FastTRK program are listed in Table 1. Males and females were equally represented (53.5% and 46.5%, respectively). The study population included representation from all Canadian provinces except Newfoundland and Labrador, and the territories. Most samples in this study were obtained and handled by LifeLabs (87.3% vs. 12.7% by KHSC). The median percent cellularity of samples in the study was 70.0% (n = 1402, range 1–100%). Of the samples that underwent NGS and that had a percent cellularity reported by the pathologist, less than 0.5% of samples (n = 1) had a percent cellularity less than 10%. However, 14.9% (33/221) of all samples sent to NGS did not have a percent cellularity assigned. These data demonstrate that for the samples that went to NGS, the majority had a sufficient cellularity for NGS. As to be expected with patients meeting the inclusion criteria, 86.1% of patients had received previous treatment, with most (68.6%) having received one to two lines of therapy, most frequently chemotherapy (64.7%) and surgery (39.7%).
Samples encompassed 37 primary tumour types (Fig. 2), most commonly colorectal cancer (19.1%), lung/pleura (17.5%), soft tissue sarcoma (9.8%), thyroid (6.4%), and pancreas (6.1%). The distribution of tumour types was similar between labs, although colorectal cancer was the most common tumour type for LifeLabs and soft tissue sarcoma was the most common tumour type for KHSC (OSM Fig. 2). Three pathognomonic samples were tested, all of which were secretory carcinomas of the salivary gland.
Prevalence of tumour types analyzed in the clinical diagnostic testing program, % (n). Primary tumour types from both LifeLabs and KHSC (N=1687) combined were categorized similar to previous literature [6, 48]. Details are summarized in Supplementary Table 1. The “Other” category represents tumours with a frequency of <1% in the total sample pool. Prevalence by lab can be found in Supplementary Table 2 CUP cancer of unknown primary site, GI gastrointestinal, GIST gastrointestinal stromal tumour, KHSC Kingston Health Sciences Centre, PEComa perivascular epithelioid cell tumour, TRK tyrosine receptor kinase.
3.2 NTRK Fusion-Positive Samples
Of 1,687 samples analyzed, 12 (0.71%) were found to harbour NTRK fusions: three NTRK1 fusions (0.18%), three NTRK2 fusions (0.18%), and six NTRK3 fusions (0.35%). Details of NTRK fusion-positive samples are summarized in Table 2. Ten NTRK fusions were detected by LifeLabs and two were detected by KHSC. Nine samples submitted to the testing program were confirmed/putative pan-TRK-positive under research studies; however, none of these samples were confirmed to be NTRK fusion-positive via NGS and thus did not contribute to the frequency of NTRK fusions identified through this testing program. The reverse reflex pathway helped identify one additional NTRK fusion-positive sample (GFOD1-NTRK2). The OCAv3 panel does not have GFOD1 coverage, which therefore was missed by this kit. However, given the anchored multiplex PCR technology of ArcherDx’s FusionPlex panel, the identification of novel fusion transcripts with unknown 5’ partners such as GFOD1 was possible.
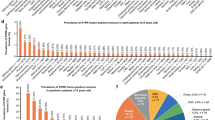
The most frequent fusion was ETV6-NTRK3 (n = 3, 25.0% of all NTRK fusions), which were all found in salivary gland tumours (two secretory and one adenocarcinoma). The following NTRK fusions were also detected: IRF2BP2-NTRK1, LMNA-NTRK1, TPM3-NTRK1, AFAP1-NTRK2, GFOD1-NTRK2, ERC1-NTRK3, SQSTM1-NTRK3, SPECC1L-NTRK3. The rate of NTRK fusion-positivity by age group in the total study population was: 19–39 years, 2.5%; 40–65 years, 0.8%; and 65+ years, 0.4%. All patients with NTRK fusions had at least one prior line of treatment: one prior line, 41.7%; two prior lines, 50.0%; and three prior lines, 8.3%. None of the patients identified had received oral tyrosine kinase inhibitors (data not shown). Central nervous system, soft tissue sarcoma, and salivary gland tumours harboured the most NTRK fusions overall (three each; Fig. 3a). On a per-tumour type basis, of all samples included in the program, melanoma (1/18, 5.6%), salivary gland (3/80, 3.8%), and CNS tumours (3/95, 3.2%) were most likely to harbour NTRK fusions (Fig. 3b). Of the 1,516 samples that underwent IHC screening, 8.0% (n = 121) were pan-TRK-positive or inconclusive and sent to NGS (Fig. 1a). The majority of samples that had pan-TRK positivity via IHC were confirmed negative for NTRK fusions by NGS (n = 70/81, 86.4%). However, of all samples screened by IHC, this corresponded to only 4.6% (n = 70/1516). The remaining samples that were pan-TRK-positive via IHC were categorized as NTRK fusion-positive (n = 6), NGS failure due to inadequate RNA quality (n = 4), and NGS not performed (n = 1). Of the 12 CNS samples assayed by pan-TRK IHC, two samples had insufficient RNA quality/quantity and one sample was pan-TRK negative, resulting in nine samples for analysis. Samples that were pan-TRK-positive but NGS-negative for NTRK fusion were most abundant in lung/pleura (n = 5/6, 83.3%), salivary gland (n = 15/18, 83.3%), soft tissue sarcoma (n = 16/20, 80.8%), and CNS (n = 7/9, 77.8%) tumours (Fig. 3c).
Breakdown of tumour types of NTRK fusion-positive samples in the clinical diagnostic testing program. a Prevalence of NTRK fusion-positive samples by tumour type (N=12). b Frequency of NTRK fusion-positive sample per tumour type of all samples in the clinical diagnostic testing program (N=1687). c Percentage of samples which were pan-TRK positive, NGS-negative for NTRK fusion by tumour type (N=81). Only tumour types containing >5 samples meeting the criteria are shown. *Spindle cell tumour (n=1) and MPNST (n=2); †Astrocytoma, glioma, and grade IV glioblastoma; ‡Secretory carcinoma (n=2) and unspecified (n=1). CUP cancer of unknown primary, GI gastrointestinal, GIST gastrointestinal stromal tumour, MPNST malignant peripheral nerve sheath tumour, NTRK neurotrophic tyrosine receptor kinase, TRK tropomyosin receptor kinase.
3.3 Incidental Findings
Of all 260 samples analyzed by NGS (221, plus the 39 samples that underwent reverse reflex), 13 (5.0%) contained clinically actionable incidental findings, as defined by OncoKB Levels of Evidence 1–4 [45, 46]. Clinically actionable incidental findings, patient demographics, and tumour clinical characteristics are summarized in Table 3. A complete list of incidental findings reported by the labs can be found in OSM Table 2 with samples containing clinically actionable findings highlighted in grey. Two of the 13 samples containing incidental findings were NTRK fusion-positive (GFOD1-NTRK2, FGFR3-TACC3 in a glioblastoma and AFAP1-NTRK2, BRAF c.1799T>A in a glioma). Epidermal growth factor receptor (EGFR) was the most frequently altered gene, with three (23.1%) alterations. Other incidental findings included point mutations in oncogenes PIK3CA, BRAF and fusions of ALK, RET, FGFR2, FGFR3 and NRG1. Samples with clinically actionable findings were predominantly from CNS (23.1%) and lung/pleural (15.3%) tumours. All patients with clinically actionable findings had received one, two, or three (7.7%, 69.2%, and 23.1%, respectively) previous lines of treatment. None of the identified patients had received oral tyrosine kinase inhibitors (data not shown).
3.4 Operational Outcomes
Overall, NTRK fusions and/or clinically actionable findings were uncovered in 23 samples, which accounts for 10.4% of those sent to NGS and 1.4% of all samples (Fig. 1a). Half of the samples with NTRK fusions were reflexed straight to NGS (Table 2). Notably, the independent parallel sequencing confirmation (reverse reflex) process/quality control mechanism helped identify one additional NTRK fusion (Fig. 1b), which was identified by LifeLabs upon a negative result with OCAv3 NGS at KHSC.
Data on report turnaround times were provided by LifeLabs. The median turnaround time on the H&E report was 1.0 workday (SD 0.8) and 7.6 workdays (SD 2.5) for the NGS report. Overall, the total cost to analyze the samples in the clinical diagnostic testing program (including 1,516 pan-TRK IHC assays performed and 260 (n = 221, Fig. 1a and n = 39, Fig. 1b) NGS samples tested) was $395,350.00 CAD (not including set-up costs, shipping, prior testing costs, or taxes). Since 12 patients (1/141) were identified as having NTRK fusions, the cost to identify one NTRK fusion patient was $32,945.83 CAD. The cost to identify one patient with any genomic finding (NTRK fusion and incidental) reported by the labs in this program was $15,814.00 CAD.
4 Discussion
We conducted an observational study on the prevalence of NTRK fusions detected in solid tumours through a Canadian clinical diagnostic testing program (FastTRK). To our knowledge, this is the first systematic characterization of the prevalence of NTRK fusions among the Canadian patient population with advanced solid tumours. These findings highlight the benefit and practicality of a diagnostic testing program to identify patients suitable for tumour-agnostic TRK inhibitor therapies like larotrectinib and entrectinib, as well as other targeted therapies, due to clinically actionable incidental findings identified. Collectively, these findings may inform future guidance on selecting the appropriate testing approach per tumour type and on optimal NTRK testing algorithms.
The study identified an overall NTRK fusion-positivity rate of 0.71% (n = 12/1,687), which is more than double previously identified rates of approximately 0.3% in recent studies [7,8,9]. Although the sample size of this study is smaller than the 11,500 to > 295,000 included in these previous studies, differences in testing algorithms and patient population characteristics may underly the different NTRK fusion-positivity rates. The findings from this observational study are also derived from two facilities that used different pan-TRK IHC and NGS methods, so heterogeneity in NTRK fusion detection should also be considered. The pan-TRK IHC screen used in this study may have helped improve the rate of NTRK fusion detection. A recent small-scale study (n = 127) also observed a considerably higher NTRK fusion-positivity rate of 3.1% following enrichment for tumours negative for other driver mutations or overexpressing TRK [38]. In addition, a positive result by IHC in combination with NTRK fusion-positivity provides greater confidence that the fusion mRNA is translated into chimeric protein (in tissues without native TRK expression). Lack of chimeric (fusion) protein expression (due to out-of-frame fusions) was hypothesized as a possible cause of larotrectinib non-response in the pivotal clinical trial [48]. Overall, a pan-TRK IHC screen may be a valuable tool to triage patients for NGS testing where routine NGS-screening is not feasible.
The inclusion of primary CNS tumours in this study, unlike in the Rosen et al. study [8], may have also contributed to the higher NTRK fusion-positivity rate considering three NTRK fusions were detected in this tumour type. The reflex of CNS pathognomonic samples straight to NGS may have also help improved NTRK fusion detection, considering half of the NTRK fusions were identified through this method. Indeed, two of the three NTRK-fusion positive secretory carcinomas of the salivary gland were reflexed straight to NGS as pathognomonic samples. In the healthcare setting, it therefore may be more practical to perform NGS testing directly in patients with these tumour types who are eligible for targeted treatment.
The prevalence rates of NTRK fusions that were observed in salivary gland tumours (3.8%) and soft tissue sarcomas (1.8%) are similar to rates seen in other studies, ranging from 2.6 to 5.3% and 1.2 to 1.5%, respectively [7, 8]. The detection of NTRK fusions in secretory salivary gland carcinoma in this study is also consistent with the literature [49]. The soft tissue sarcomas identified as harbouring NTRK fusions in this study included a spindle cell tumour and two malignant peripheral nerve sheath tumours (MPNST). Malignant peripheral nerve sheath tumours were previously included in pivotal TRK inhibitor clinical trial populations [48, 50]. The recent classification of soft tissue and bone tumours recognizes two tumour classifications with a high prevalence of NTRK fusions: IFS/congenital mesoblastic nephroma (CMN) and NTRK-rearranged spindle cell neoplasms [51, 52]. The NTRK-rearranged spindle cell neoplasms include soft tissue sarcomas with spindle cells in a long or short fascicular growth or a haphazard arrangement, described as morphology akin to IFS or MPNST [52, 53]. Thus, the NTRK fusion-positive MPNST identified in this study may represent a distinct entity of the NTRK-rearranged spindle cell neoplasm, which may mimic MPNST [53, 54].
Although the observed NTRK fusion prevalence of 3.2% in CNS tumours in this study is considerably higher than the 0.1–0.4% seen in the recent Westphalen et al. study [7], NTRK fusions have been observed at rates of 1.1–2.6% in glioblastomas [55,56,57,58,59], 2.3% in anaplastic astrocytomas [59], and 0.4–4.3% in low-grade gliomas [59,60,61]. The identification of NTRK2 fusions in primary CNS tumours in this study also reflects the literature [62]. Similarly, although the NTRK fusion rate in cutaneous or mucosal melanoma is usually < 1.0% [63], it was higher (4.8%) in this study, albeit with only one positive sample identified. For common tumour types like lung and colorectal cancers, our observed NTRK fusion-positivity rates (0.3% for both) are comparable to rates of < 1.0% seen in a recent systemic review [43].
The most frequent NTRK fusion in our dataset was ETV6-NTRK3 (n = 3, 25.0% of all NTRK fusions), consistent with the literature [7, 9]. As these were all found in salivary gland tumours (two of three confirmed to be of secretory origin), this is in line with previous reports of ETV6-NTRK3 being characteristic of secretory salivary gland tumours [64]. These findings support and expand on previous knowledge of NTRK fusion partners with our observations of IRF2BP2-NTRK1 in lung, AFAP1-NTRK2 in CNS, ERC1-NTRK3 in melanoma, and SQSTM1-NTRK3 and SPECC1L-NTRK3 in soft tissue sarcoma [65]. To the best of our knowledge, the GFOD1-NTRK2 fusion (identified here in glioblastoma) has not yet been reported.
Although the clinical testing program was designed for the detection of NTRK fusions to help identify patients eligible for TRK inhibitor therapy, the added benefit of this centralized testing program was the possible detection of other targetable genomic alterations through NGS. Indeed, in addition to the 12 NTRK fusion-positive tumours, 13 tumours were identified that contained clinically actionable incidental findings. These findings encompass many oncogenic pathways, including point mutations in EGFR, PIK3CA, and BRAF, and fusions of ALK, RET, FGFR2, FGFR3, and NRG1. Although NTRK fusions are generally mutually exclusive from other oncogenic drivers, two of the 12 NTRK fusion-positive samples had clinically actionable incidental findings [7, 8, 66, 67]. The two exceptions were GFOD1-NTRK2 and AFAP1-NTRK2, which co-occurred with FGFR3-TACC3 fusion and BRAF 1799T>A (V600E), respectively, both in CNS tumours. Indeed, primary CNS tumours are known to harbour concurrent alterations [7, 68,69,70], which is in line with our observation of both NTRK fusions and incidental findings in only CNS samples. Ultimately, the detection of clinically actionable incidental findings through the testing program provided additional information to the treating clinician.
Considering operational outcomes of the clinical diagnostic testing program, the observed turnaround times of 1.00 and 7.63 days for the IHC and NGS reports, respectively, exceeded expectations of report delivery times, despite the impact of the COVID-19 pandemic on personnel/administrative duties. Occasional delays may have occurred due to the time required to answer technical questions; however, the expected turnaround time was 2–4 business days for IHC and up to 14 business days for NGS. It is important to note the lab that provided the turnaround data (LifeLabs) is a private lab, whereas KHSC is an academic health science centre funded by the Ontario Ministry of Health, thus the turnaround times may differ depending on the lab. The testing program currently operates as a privately funded program, so low sample volumes likely contributed to a higher cost of testing. Our estimated cost of $32,945 CAD to detect one NTRK fusion patient should be considered a high estimate; however, it would presumably decrease through program expansion to more Canadian centres. Further, costing analysis is based on Bayer’s agreements with LifeLabs and KHSC, and may not accurately reflect the cost to the Canadian public healthcare system to conduct the same testing.
4.1 Limitations
This study has certain limitations:
Submitted samples: There were a number of factors that influenced the diversity and proportion of histologies submitted. (1) This national clinical diagnostic testing program was open to samples from all patients with solid tumours that were metastatic or when surgical resection was likely to result in severe morbidity, and for whom no satisfactory treatment options were available. It was primarily the treating physician or assigned pathologist who would have used their clinical judgement to decide which patients qualified for and accessed this testing program. (2) The patients selected for this program could have been driven by the specific testing pathways for each tissue type, which is unique to each Canadian province and often hospital. (3) Physicians were asked to submit samples for testing for any histology type, which were negative for other known oncogenic drivers or mutations (although this was not always the case), thus introducing possible enrichment for NTRK fusion-positive samples. (4) The amount of sample needed for testing may have prevented testing uptake from select histologies. (5) The current study was also dependent on the accuracy of information entered into the requisition form. Taken together, the clinical testing program had no control on what histologies were submitted for testing, and coupled with a relatively small patient population tested, the results may not reflect the true prevalence of TRK fusion tumours across Canada and precluded statistical hypothesis-based analyses in the current study.
Diagnostic considerations: Although this study was not sufficiently powered nor designed to explore the overall sensitivity and specificity of pan-TRK IHC, the current study results suggest a higher rate of pan-TRK-positive but NGS-negative samples than previously reported for tumour types known to have non-specific or physiological staining by pan-TRK IHC, such as soft tissue sarcomas, and CNS samples that did not follow the pre-defined algorithm [9, 32]. The rate of pan-TRK positive but NGS-negative samples in this analysis was similar to another study with a false positivity rate of 96.8%, which also selected samples for NGS based on pan-TRK IHC and, thus, the tested samples were enriched for tumour types with non-specific/physiological pan-TRK staining [38]. It should also be noted that pan-TRK IHC is known to have limited sensitivity for chimeric TRKC proteins arising from NTRK3 fusions [9], and, therefore, NTRK fusion-positive samples could have been missed. As the diagnostic, prognostic and predictive biomarker testing continues to rapidly evolve for locally advanced and metastatic solid tumours, the utility of upfront NGS will continue to expand, therefore improving the systematic and efficient identification of patients with rare actionable driver mutations. Overall, although reflective of multi-centre testing programs, the two facilities conducting testing in the current study used variable approaches to IHC (lab-based pan-TRK IHC developed by KHSC versus the pan-TRK Assay used by LifeLabs) and NGS (Oncomine Comprehensive Panel v3 used by KHSC versus FusionPlex® Lung panel v1.0), thus this heterogeneity in protocols may have impacted rates of NTRK fusion detection. The reverse reflex pathway helped identify one additional NTRK fusion-positive sample (GFOD1-NTRK2). Although the practicality of implementing such quality control methods in clinical practice still needs to be determined, this method could help improve detection of NTRK fusions in some contexts.
Future research should include a larger Canadian population to corroborate and expand upon these findings, as well as the impact of testing on patient outcomes, such as PFS and OS. While the study was not designed to collect data on treatment decisions, the impact on subsequent therapy choice could be the subject of future study.
4.2 Conclusions
This observational study summarizes the outcomes of a national NTRK fusion tumour clinical testing program designed to identify patients eligible for TRK inhibitor therapy. The study population included considerable patient diversity in terms of age, geographic representation, and tumour type, and thus provides valuable insights into the prevalence of NTRK fusions in a real-world Canadian setting. The algorithm used in this clinical testing program may also provide guidance on strategies to improve detection of NTRK fusions in clinical practice.
References
Cocco E, Scaltriti M, Drilon A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol. 2018;15:731–47.
Reichardt LF. Neurotrophin-regulated signalling pathways. Philos Trans R Soc Lond B Biol Sci. 2006;361:1545–64.
Barbacid M, Lamballe F, Pulido D, Klein R. The trk family of tyrosine protein kinase receptors. Biochim Biophys Acta. 1991;1072:115–27.
Vaishnavi A, Le AT, Doebele RC. TRKing down an old oncogene in a new era of targeted therapy. Cancer Discov. 2015;5:25–34.
Okamura R, Boichard A, Kato S, Sicklick JK, Bazhenova L, Kurzrock R. Analysis of JCO. Precis Oncol 2018;2018.
Hong DS, DuBois SG, Kummar S, Farago AF, Albert CM, Rohrberg KS, van Tilburg CM, Nagasubramanian R, Berlin JD, Federman N, Mascarenhas L, Geoerger B, Dowlati A, Pappo AS, Bielack S, Doz F, McDermott R, Patel JD, Schilder RJ, Tahara M, Pfister SM, Witt O, Ladanyi M, Rudzinski ER, Nanda S, Childs BH, Laetsch TW, Hyman DM, Drilon A. Larotrectinib in patients with TRK fusion-positive solid tumours: a pooled analysis of three phase 1/2 clinical trials. Lancet Oncol. 2020;21:531–40.
Westphalen CB, Krebs MG, Le Tourneau C, Sokol ES, Maund SL, Wilson TR, Jin DX, Newberg JY, Fabrizio D, Veronese L, Thomas M, de Braud F. Genomic context of NTRK1/2/3 fusion-positive tumours from a large real-world population. NPJ Precis Oncol. 2021;5:69.
Rosen EY, Goldman DA, Hechtman JF, Benayed R, Schram AM, Cocco E, Shifman S, Gong Y, Kundra R, Solomon JP, Bardelli A, Scaltriti M, Drilon A, Iasonos A, Taylor BS, Hyman DM. TRK fusions are enriched in cancers with uncommon histologies and the absence of canonical driver mutations. Clin Cancer Res. 2020;26:1624–32.
Gatalica Z, Xiu J, Swensen J, Vranic S. Molecular characterization of cancers with NTRK gene fusions. Mod Pathol. 2019;32:147–53.
Hsiao SJ, Zehir A, Sireci AN, Aisner DL. Detection of tumor NTRK gene fusions to identify patients who may benefit from tyrosine kinase (TRK) inhibitor therapy. J Mol Diagn. 2019;21:553–71.
Vitrakvi product monograph. Bayer Inc. (2021).
Rozlytrek product monograph. Hoffmann-La Roche Limited. (2021).
Vitrakvi prescribing information. Bayer Inc. (2018).
Vitrakvi summary of product characteristics. Bayer Inc. (2019).
Rozyltrek Prescribing information. Hoffmann-La Roche Limited. (2019).
Rozyltrek summary of product characteristics. Hoffmann-La Roche Limited. (2020).
Bebb DG, Banerji S, Blais N, Desmeules P, Gill S, Grin A, Feilotter H, Hansen AR, Hyrcza M, Krzyzanowska M, Melosky B, Noujaim J, Purgina B, Ruether D, Simmons CE, Soulieres D, Torlakovic EE, Tsao MS. Canadian consensus for biomarker testing and treatment of TRK fusion cancer in adults. Curr Oncol. 2021;28:523–48.
Awada A, Berghmans T, Clement PM, Cuppens K, De Wilde B, Machiels JP, Pauwels P, Peeters M, Rottey S, Van Cutsem E. Belgian expert consensus for tumor-agnostic treatment of NTRK gene fusion-driven solid tumors with larotrectinib. Crit Rev Oncol Hematol. 2022;169: 103564.
Lim KHT, Kong HL, Chang KTE, Tan DSW, Tan IBH, Mohamad F, Soh SY, Pang BN, Soo RA, Choo SP, Hsieh WS, Aung L. Recommended testing algorithms for NTRK gene fusions in pediatric and selected adult cancers: consensus of a Singapore Task Force Asia. Pac J Clin Oncol. 2021;18:394.
Perreault S, Chami R, Deyell RJ, El Demellawy D, Ellezam B, Jabado N, Morgenstern DA, Narendran A, Sorensen PHB, Wasserman JD, Yip S. Canadian consensus for biomarker testing and treatment of TRK fusion cancer in pediatric patients. Curr Oncol. 2021;28:346–66.
Hanna NH, Robinson AG, Temin S, Baker S, Brahmer JR, Ellis PM, Gaspar LE, Haddad RY, Hesketh PJ, Jain D, Jaiyesimi I, Johnson DH, Leighl NB, Moffitt PR, Phillips T, Riely GJ, Rosell R, Schiller JH, Schneider BJ, Singh N, Spigel DR, Tashbar J, Masters G. Therapy for stage IV non-small-cell lung cancer with driver alterations: ASCO and OH (CCO) joint guideline update. J Clin Oncol. 2021;39:1040–91.
Orbach D, Sparber-Sauer M, Laetsch TW, Minard-Colin V, Bielack SS, Casanova M, Corradini N, Koscielniak E, Scheer M, Hettmer S, Bisogno G, Hawkins DS, Ferrari A. Spotlight on the treatment of infantile fibrosarcoma in the era of neurotrophic tropomyosin receptor kinase inhibitors: international consensus and remaining controversies. Eur J Cancer. 2020;137:183–92.
Marchiò C, Scaltriti M, Ladanyi M, Iafrate AJ, Bibeau F, Dietel M, Hechtman JF, Troiani T, López-Rios F, Douillard JY, Andrè F, Reis-Filho JS. ESMO recommendations on the standard methods to detect NTRK fusions in daily practice and clinical research. Ann Oncol. 2019;30:1417–27.
Yoshino T, Pentheroudakis G, Mishima S, Overman MJ, Yeh KH, Baba E, Naito Y, Calvo F, Saxena A, Chen LT, Takeda M, Cervantes A, Taniguchi H, Yoshida K, Kodera Y, Kitagawa Y, Tabernero J, Burris H, Douillard JY. JSCO-ESMO-ASCO-JSMO-TOS: international expert consensus recommendations for tumour-agnostic treatments in patients with solid tumours with microsatellite instability or NTRK fusions. Ann Oncol. 2020;31:861–72.
Demetri GD, De Braud F, Drilon A, Siena S, Patel MR, Cho BC, Liu SV, Ahn MJ, Chiu CH, Lin JJ, Goto K, Lee J, Bazhenova L, John T, Fakih M, Chawla SP, Dziadziuszko R, Seto T, Heinzmann S, Pitcher B, Chen D, Wilson TR, Rolfo C. Updated integrated analysis of the efficacy and safety of entrectinib in patients with. Clin Cancer Res. 2022;28:2196.
Penault-Llorca F, Rudzinski ER, Sepulveda AR. Testing algorithm for identification of patients with TRK fusion cancer. J Clin Pathol. 2019;72:460–7.
Zito Marino F, Pagliuca F, Ronchi A, Cozzolino I, Montella M, Berretta M, Errico ME, Donofrio V, Bianco R, Franco R. NTRK fusions, from the diagnostic algorithm to innovative treatment in the era of precision medicine. Int J Mol Sci. 2020;21:3718.
Park HJ, Baek I, Cheang G, Solomon JP, Song W. Comparison of RNA-based next-generation sequencing assays for the detection of NTRK gene fusions. J Mol Diagn. 2021;23:1443–51.
Heydt C, Wölwer CB, Velazquez Camacho O, Wagener-Ryczek S, Pappesch R, Siemanowski J, Rehker J, Haller F, Agaimy A, Worm K, Herold T, Pfarr N, Weichert W, Kirchner T, Jung A, Kumbrink J, Goering W, Esposito I, Buettner R, Hillmer AM, Merkelbach-Bruse S. Detection of gene fusions using targeted next-generation sequencing: a comparative evaluation. BMC Med Genom. 2021;14:62.
De Winne K, Sorber L, Lambin S, Siozopoulou V, Beniuga G, Dedeurwaerdere F, D’Haene N, Habran L, Libbrecht L, Van Huysse J, Weynand B, Wouters K, Pauwels P, Zwaenepoel K. Immunohistochemistry as a screening tool for NTRK gene fusions: results of a first Belgian ring trial. Virchows Arch. 2021;478:283–91.
Pfarr N, Kirchner M, Lehmann U, Leichsenring J, Merkelbach-Bruse S, Glade J, Hummel M, Stögbauer F, Lehmann A, Trautmann M, Kumbrink J, Jung A, Dietmaier W, Endris V, Kazdal D, Evert M, Horst D, Kreipe H, Kirchner T, Wardelmann E, Lassen U, Büttner R, Weichert W, Dietel M, Schirmacher P, Stenzinger A. Testing NTRK testing: wet-lab and in silico comparison of RNA-based targeted sequencing assays. Genes Chromosom Cancer. 2020;59:178–88.
Solomon JP, Linkov I, Rosado A, Mullaney K, Rosen EY, Frosina D, Jungbluth AA, Zehir A, Benayed R, Drilon A, Hyman DM, Ladanyi M, Sireci AN, Hechtman JF. NTRK fusion detection across multiple assays and 33,997 cases: diagnostic implications and pitfalls. Mod Pathol. 2020;33:38–46.
Solomon JP, Hechtman JF. Detection of NTRK fusions: merits and limitations of current diagnostic platforms. Cancer Res. 2019;79:3163–8.
Garrido P, Hladun R, de Álava E, Álvarez R, Bautista F, López-Ríos F, Colomer R, Rojo F. Multidisciplinary consensus on optimising the detection of NTRK gene alterations in tumours. Clin Transl Oncol. 2021;23:1529–41.
Hechtman JF. NTRK insights: best practices for pathologists. Mod Pathol. 2021.
Bormann Chung C, Lee J, Barritault M, Bringuier PP, Xu Z, Huang WY, Beharry A, Castillo J, Christiansen J, Lin JC, Sheffield BS. Evaluating targeted next-generation sequencing assays and reference materials for NTRK fusion detection. J Mol Diagn. 2022;24:18–32.
Rohrberg KS, Lassen U. Detecting and targeting NTRK fusions in cancer in the era of tumor agnostic oncology. Drugs. 2021;81:445–52.
de González AK, Mansukhani MM, Fernandes H, Hsiao SJ. Pan-tumor screening for NTRK gene fusions using pan-TRK immunohistochemistry and RNA NGS fusion panel testing. Cancer Genet. 2022;262–263:47–52.
Sholl L, Zheng M, Nardi V, Hornick J. Predictive “biomarker piggybacking”: an examination of reflexive pan-cancer screening with pan-TRK immunohistochemistry. Histopathology. 2021;79:260–4.
Wong D, Yip S, Sorensen PH. Methods for identifying patients with tropomyosin receptor kinase (TRK) fusion cancer. Pathol Oncol Res. 2020;26:1385–99.
Tsao MS, Torlakovic E, Stockley T, Lo B. In: Proceedings of the association for molecular pathology annual meeting, virtual. ST07.
Kirchner M, Glade J, Lehmann U, Merkelbach-Bruse S, Hummel M, Lehmann A, Trautmann M, Kumbrink J, Jung A, Dietmaier W, Endris V, Kazdal D, Ploeger C, Evert M, Horst D, Kreipe H, Kirchner T, Wardelmann E, Büttner R, Weichert W, Dietel M, Schirmacher P, Stenzinger A, Pfarr N. NTRK testing: first results of the QuiP-EQA scheme and a comprehensive map of NTRK fusion variants and their diagnostic coverage by targeted RNA-based NGS assays. Genes Chromosom Cancer. 2020;59:445–53.
Forsythe A, Zhang W, Phillip Strauss U, Fellous M, Korei M, Keating K. A systematic review and meta-analysis of neurotrophic tyrosine receptor kinase gene fusion frequencies in solid tumors. Ther Adv Med Oncol. 2020;12:1758835920975613.
Wadhwa S, Nag TC, Jindal A, Kushwaha R, Mahapatra AK, Sarkar C. Expression of the neurotrophin receptors Trk A and Trk B in adult human astrocytoma and glioblastoma. J Biosci. 2003;28:181–8.
Center MSKC. OncoKB therapeutic level of evidence V2, https://www.oncokb.org/levels
Chakravarty D, Gao J, Phillips SM, Kundra R, Zhang H, Wang J, Rudolph JE, Yaeger R, Soumerai T, Nissan MH, Chang MT, Chandarlapaty S, Traina TA, Paik PK, Ho AL, Hantash FM, Grupe A, Baxi SS, Callahan MK, Snyder A, Chi P, Danila D, Gounder M, Harding JJ, Hellmann MD, Iyer G, Janjigian Y, Kaley T, Levine DA, Lowery M, Omuro A, Postow MA, Rathkopf D, Shoushtari, AN, Shukla N, Voss M, Paraiso E, Zehir A, Berger MF, Taylor BS, Saltz LB, Riely GJ, Ladanyi M, Hyman DM, Baselga J, Sabbatini P, Solit DB, Schultz N. OncoKB: a precision oncology knowledge base JCO. Precis Oncol. 2017;2017.
Hechtman JF, Benayed R, Hyman DM, Drilon A, Zehir A, Frosina D, Arcila ME, Dogan S, Klimstra DS, Ladanyi M, Jungbluth AA. Pan-Trk immunohistochemistry is an efficient and reliable screen for the detection of NTRK fusions. Am J Surg Pathol. 2017;41:1547–51.
Drilon A, Laetsch TW, Kummar S, DuBois SG, Lassen UN, Demetri GD, Nathenson M, Doebele RC, Farago AF, Pappo AS, Turpin B, Dowlati A, Brose MS, Mascarenhas L, Federman N, Berlin J, El-Deiry WS, Baik C, Deeken J, Boni V, Nagasubramanian R, Taylor M, Rudzinski ER, Meric-Bernstam F, Sohal DPS, Ma PC, Raez LE, Hechtman JF, Benayed R, Ladanyi M, Tuch BB, Ebata K, Cruickshank S, Ku NC, Cox MC, Hawkins DS, Hong DS, Hyman DM. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N Engl J Med. 2018;378:731–9.
Skalova A, Michal M, Simpson RH. Newly described salivary gland tumors. Mod Pathol. 2017;30:S27–43.
Doebele RC, Drilon A, Paz-Ares L, Siena S, Shaw AT, Farago AF, Blakely CM, Seto T, Cho BC, Tosi D. Entrectinib in patients with advanced or metastatic NTRK fusion-positive solid tumours: integrated analysis of three phase 1–2 trials. Lancet Oncol. 2020;21:271–82.
Davis J, Bahrami A. In: WHO classification of tumours of soft tissue and bone (ed WHO Classification of Tumours Editorial Board). pp. 119–21. 2020.
Suurmeijer A. In: WHO classification of tumours of soft tissue and bone (ed WHO Classification of Tumours Editorial Board). pp. 287–9. 2020.
Surrey LF, Davis JL. NTRK-Rearranged soft tissue neoplasms: a review of evolving diagnostic entities and algorithmic detection methods. Cancer Genet. 2022;260–261:6–13.
Suurmeijer AJH, Dickson BC, Swanson D, Zhang L, Sung YS, Cotzia P, Fletcher CDM, Antonescu CR. A novel group of spindle cell tumors defined by S100 and CD34 co-expression shows recurrent fusions involving RAF1, BRAF, and NTRK1/2 genes. Genes Chromosom Cancer. 2018;57:611–21.
Frattini V, Trifonov V, Chan JM, Castano A, Lia M, Abate F, Keir ST, Ji AX, Zoppoli P, Niola F, Danussi C, Dolgalev I, Porrati P, Pellegatta S, Heguy A, Gupta G, Pisapia DJ, Canoll P, Bruce JN, McLendon RE, Yan H, Aldape K, Finocchiaro G, Mikkelsen T, Privé GG, Bigner DD, Lasorella A, Rabadan R, Iavarone A. The integrated landscape of driver genomic alterations in glioblastoma. Nat Genet. 2013;45:1141–9.
Shah N, Lankerovich M, Lee H, Yoon JG, Schroeder B, Foltz G. Exploration of the gene fusion landscape of glioblastoma using transcriptome sequencing and copy number data. BMC Genom. 2013;14:818.
Zheng Z, Liebers M, Zhelyazkova B, Cao Y, Panditi D, Lynch KD, Chen J, Robinson HE, Shim HS, Chmielecki J, Pao W, Engelman JA, Iafrate AJ, Le LP. Anchored multiplex PCR for targeted next-generation sequencing. Nat Med. 2014;20:1479–84.
Kim J, Lee Y, Cho HJ, Lee YE, An J, Cho GH, Ko YH, Joo KM, Nam DH. NTRK1 fusion in glioblastoma multiforme. PLoS ONE. 2014;9:e91940.
Ferguson SD, Zhou S, Huse JT, de Groot JF, Xiu J, Subramaniam DS, Mehta S, Gatalica Z, Swensen J, Sanai N, Spetzler D, Heimberger AB. Targetable gene fusions associate with the IDH wild-type astrocytic lineage in adult gliomas. J Neuropathol Exp Neurol. 2018;77:437–42.
Zhang J, Wu G, Miller CP, Tatevossian RG, Dalton JD, Tang B, Orisme W, Punchihewa C, Parker M, Qaddoumi I, Boop FA, Lu C, Kandoth C, Ding L, Lee R, Huether R, Chen X, Hedlund E, Nagahawatte P, Rusch M, Boggs K, Cheng J, Becksfort J, Ma J, Song G, Li Y, Wei L, Wang J, Shurtleff S, Easton J, Zhao D, Fulton RS, Fulton LL, Dooling DJ, Vadodaria B, Mulder HL, Tang C, Ochoa K, Mullighan CG, Gajjar A, Kriwacki R, Sheer D, Gilbertson RJ, Mardis ER, Wilson RK, Downing JR, Baker SJ, Ellison DW, Project SJCSRHWUPCG. Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat Genet. 2013;45:602–12.
Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C. The landscape of kinase fusions in cancer. Nat Commun. 2014;5:4846.
Doz F, van Tilburg CM, Geoerger B, Højgaard M, Øra I, Boni V, Capra M, Chisholm J, Chung HC, DuBois SG, Gallego-Melcon S, Gerber NU, Goto H, Grilley-Olson JE, Hansford JR, Hong DS, Italiano A, Kang HJ, Nysom K, Thorwarth A, Stefanowicz J, Tahara M, Ziegler DS, Gavrilovic IT, Norenberg R, Dima L, De La Cuesta E, Laetsch TW, Drilon A, Perreault S. Efficacy and safety of larotrectinib in TRK fusion-positive primary central nervous system tumors. Neuro Oncol. 2021;24:997.
Forschner A, Forchhammer S, Bonzheim I. NTRK gene fusions in melanoma: detection, prevalence and potential therapeutic implications. J Dtsch Dermatol Ges. 2020;18:1387–92.
Le X, Baik C, Bauman J, Gilbert J, Brose MS, Grilley-Olson JE, Patil T, McDermott R, Raez LE, Johnson JM, Shen L, Tahara M, Ho AL, Norenberg R, Dima L, Brega N, Drilon A, Hong DS. Larotrectinib treatment for patients with TRK fusion-positive salivary gland cancers. Oncologist. 2022.
Rudzinski ER, Hechtman J, Roy-Chowdhuri S, Rudolph M, Lockwood CM, Silvertown J, Wierzbinska J, Shen K, Norenberg R, Nogai H, Hong DS, Drilon A, Laetsch TW. Diagnostic testing approaches for the identification of patients with TRK fusion cancer prior to enrollment in clinical trials investigating larotrectinib. Cancer Genet. 2022;260–261:46–52.
Bazhenova L, Lokker A, Snider J, Castellanos E, Fisher V, Fellous M, Nanda S, Zong J, Keating K, Jiao X. TRK fusion cancer: patient characteristics and survival analysis in the real-world setting. Targ Oncol. 2021;16:389–99.
Bridgewater J, Jiao X, Parimi M, Flach C, Stratford J, Kamburov A, Zong J. Prognosis and molecular characteristics of patients with TRK fusion cancer in the 100,000 Genomes Project Proceedings of the American Association for Cancer Research Annual Meeting 2021;Abstract 394
Torre M, Vasudevaraja V, Serrano J, DeLorenzo M, Malinowski S, Blandin AF, Pages M, Ligon AH, Dong F, Meredith DM, Nasrallah MP, Horbinski C, Dahiya S, Ligon KL, Santi M, Ramkissoon SH, Filbin MG, Snuderl M, Alexandrescu S. Molecular and clinicopathologic features of gliomas harboring NTRK fusions. Acta Neuropathol Commun. 2020;8:107.
Schram AM, Jonsson P, Drilon A, Bale TA, Hechtman JF, Benayed R, Hanusch B, Young RJ, Grommes C, Ku N, Kaley T, Hyman DM, Taylor BS. Genomic heterogeneity underlies mixed response to tropomyosin receptor kinase inhibition in recurrent glioma. JCO Precis Oncol. 2018;2.
Shepherd DJ, Miller TE, Forst DA, Jones P, Nardi V, Martinez-Lage M, Stemmer-Rachamimov A, Gonzalez RG, Lafrate AJ, Ritterhouse LL. Mosaicism for receptor tyrosine kinase activation in a glioblastoma involving both PDGFRA amplification and NTRK2 fusion. Oncologist. 2021;26:919–24.
Acknowledgements
The authors acknowledge Bahiyyih Schmalenberg (Bayer Inc., Mississauga, ON, Canada) for project management support; Chrisoula Giannaris (Bayer Inc.), Vadim Bernard-Gauthier (Bayer Inc.), Shurjeel Choudhri (Bayer Inc.), Edward Feijoo-Olaya (Bayer Inc.), Brandon Levac (Bayer Inc.), and Jason Lee (Bayer Inc.) for technical guidance; the LifeLabs Histology Department; Andrea Brumwell and Danyelle Liddle (MEDUCOM Health Inc., Guelph, ON, Canada) for data analysis and medical writing services. The authors also acknowledge the oncologists, pathologists, geneticists, and laboratories that contributed to the data submitted in this work.
Author information
Authors and Affiliations
Contributions
Joshua D. Silvertown, Laura Semenuk, Colleen Knapp, Jillan Jaynes, Doreen Berg, David Berman, Harriet Feilotter, Connie Lisle, Nabodita Kaul, Josianne Lachappelle, Leslie Richardson, Marsha Speevak, Ronald Carter and Timothy Feltis conceived the project, implemented and managed program, were involved in study design, management and data analysis. Joshua D. Silvertown wrote the study protocol. Haya Sarras reviewed study design, supported data interpretation and oversaw of data acquisition and revision of intellectual content. All authors were involved in writing, reviewing, and editing the manuscript, approved the final version of the manuscript, and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding author
Ethics declarations
Funding
This research was funded by Bayer Inc.
Conflicts of interest
Harriet Feilotter received funding from Bayer Inc. to support implementation of the FastTRK program and received honoraria from Bayer Inc. for speaking engagements. David Berman received compensation as a member of the Scientific Advisory Board for Acrivon Therapeutics. Marsha Speevak is employed as a consultant for LifeLabs. Joshua D. Silvertown and Haya Sarras are employed by Bayer Inc. Connie Lisle and Doreen Berg were previously employed by LifeLabs. Nabodita Kaul, Josianne Lachappelle, Leslie Richardson, Ronald Carter, and Timothy Feltis are currently employed by LifeLabs. Laura Semenuk, Colleen Knapp, Jillan Jaynes, David Berman, and Harriet Feilotter are currently employed by Kingston Health Sciences Centre.
Ethics approval
This study was conducted in accordance with the Helsinki Declaration of 1975 and with the approval of Veritas IRB Inc.
Consent to participate
Written informed consent was obtained from all subjects involved in the study.
Consent for participation
Not applicable.
Data availability
Availability of the data underlying this publication will be determined later according to Bayer’s commitment to the EFPIA/PhRMA Principles for Responsible Clinical Trial Data Sharing. This pertains to scope, time point and process of data access. As such, Bayer commits to sharing, upon request from qualified scientific and medical researchers, anonymized patient-level clinical trial data, study-level clinical trial data and protocols from clinical trials in patients for medicines and indications approved in the United States (US) and European Union (EU) as necessary for conducting legitimate research. This applies to data on new medicines and indications that have been approved by the EU and US regulatory agencies on or after January 1, 2014. Interested researchers can use http://www.clinicalstudydatarequest.com to request access to anonymized patient-level data and supporting documents from clinical studies to conduct further research that can help advance medical science or improve patient care. Information on the Bayer criteria for listing studies and other relevant information is provided in the study sponsors section of the portal. Data access will be granted to anonymized patient-level data, protocols and clinical study reports after approval by an independent scientific review panel. Bayer is not involved in the decisions made by the independent scientific review panel. Bayer will take all necessary measures to ensure that patient privacy is safeguarded.
Code availability
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Silvertown, J.D., Lisle, C., Semenuk, L. et al. Prevalence of NTRK Fusions in Canadian Solid Tumour Cancer Patients. Mol Diagn Ther 27, 87–103 (2023). https://doi.org/10.1007/s40291-022-00617-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-022-00617-y