Abstract
Despite a vaccine being available, human papillomavirus virus (HPV)-driven cancers remain the ninth most prevalent cancers globally. Current therapies have significant drawbacks and often still lead to poor prognosis and underwhelming survival rates. With gene therapy becoming more available in the clinic, it poses a new front for therapeutic development. A characteristic of HPV-driven cancers is the ability to encode oncoproteins that aberrate normal p53 function without mutating this tumour-suppressor gene. The HPV E6 oncoprotein degrades p53 to allow the HPV-driven carcinogenic process to proceed. This review aimed to investigate the use of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) gene-editing technology and how it may be used to overcome HPV-mediated silencing of p53 by hyper-expressing the p53 promoter. Increasing p53 bioavailability may have promising potential as a therapy and has been a goal in the context of HPV-driven cancers. Clinical trials and proof-of-concept pre-clinical work have shown positive outcomes and tumour death when p53 levels are increased. Despite previous successes of RNA-based medicines, including the knockout of HPV oncogenes, the use of CRISPR activation is yet to be investigated as a promising potential therapy. This short review summarises key developments on attempts that have been made to increase p53 expression in the context of HPV cancer therapy, but leaves open the possibility for other cancers bearing a p53 wild-type gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Restoring p53 may be a key clinical factor for the treatment of HPV cancers. |
CRISPR-based gene therapy may be a promising treatment modality for HPV-driven cancers |
1 Introduction
High-risk human papillomavirus (HPV) types 16 and 18 are the causative agents for most HPV-driven cancers. Despite there being an effective vaccine against HPV, HPV cancers still rank ninth for the most prevalent cancer type in the world [1]. HPV is implicated in over 99% of cervical cancer cases, with HPV16 alone accounting for 70.8% of those [2]. Oropharyngeal squamous cell carcinomas are also caused by persistent HPV infection [3], an aetiology strongly associated with HPV type 16 [4]. Indeed, the global incidence of HPV-associated cancers has not significantly improved since 2008, with these cancers being especially prevalent in lower income countries [5]. With the global incidence being strongly associated with several external carcinogenic and geographic factors, its seemingly upward trend is hard to reverse.
Current standard therapies, including chemotherapy, surgery, or both, have not improved survival rates in the past 2 decades, suggesting that there is a need for more novel treatment approaches. The carcinogenic process in HPV cancers is largely attributed to the E6 and E7 oncoproteins encoded by HPV [6]. A notable interaction that exists is the ability for HPV to dysregulate wild-type p53 function, via the E6 oncoprotein [7]. p53 is a tumour-suppressor protein and its inhibition results in cells losing control of the mitotic cell cycle at the G2/M checkpoint. This works synergistically with E7, which inactivates the phosphorylated retinoblastoma protein (pRb), taking the brakes off the cell cycle and thereby allowing cells to proliferate uncontrollably [8,9,10]. Mutations or dysregulations having the same effect on p53 are highly selected for in other cancers [11], including lung, colon, invasive ductal breast, pancreatic and high-grade ovarian serous carcinoma [12]. Coining the term “guardian of the genome”, p53 has versatile roles as a tumour-suppressor gene.
With gene therapy becoming more available in the clinic and the advances of the Clustered Regularly Interspaced Short Palindromic Repeats-associated Cas 9 (CRISPR/Cas)-based systems, it gives rise to new avenues for gene editing-based therapies. Indeed, therapeutic editing using CRISPR technology to correct p53 mutations have been proposed for a few human diseases including Li-Fraumeni syndrome (see review by Mirgayazova et al. [13]). However, in this review, we aimed to address the possibility of overcoming wild-type p53 dysregulation, as commonly seen in HPV-driven cancers. Here, we review the aetiology of HPV-driven cancers with previous efforts to reverse this, and why overcoming HPV oncogene-mediated silencing of p53 using genetic manipulation is a potential treatment option.
2 The Physiological Function of p53
p53 can be activated via two distinct pathways. Firstly, the onset of the DNA damage response (DDR) causes the serine/threonine protein ataxia-telangiectasia mutated (ATM) and ATM-Rad3 (ATR) signal kinase pathways to stabilise and activate downstream checkpoint kinases (CHK). ATM-Chk2 is responsible for detection of predominantly double-strand breaks, whereas ATR-Chk1 is broader and may be activated by general DNA stressors, such as stress on the replication fork during replication [14]. Once ATM-Chk2 and ATR-Chk1 are activated, these kinases phosphorylate p53 (Fig. 1). Secondly, hyperproliferative signals, which can be caused by transcription factors such as E2F, trigger prompt activation of the alternate reading frame product (ARF). Other oncogenic insults such as c-myc and k-ras are also able to prompt stabilisation of ARF. ARF are a class of GTP proteins that activate the cell cycle, and when dephosphorylated into GDP, block entrance into the cell cycle. Specifically, ARF blocks the E3 ubiquitin ligase activity of the minute double murine 2 protein (MDM2), so that p53 is not subjected to proteasomal degradation through the tagging process of ubiquitination [15]. MDM is a gene that encodes E3 ubiquitin-ligases, which are involved in the p53 negative feedback loop, degrading p53 rapidly when there are no signals for DNA stabilisation, such as in DNA damage.
There is also a dimerization partner, RB-like, E2F and multi-vulval class B (DREAM) complex, known as the p53-p21-DREAM induction pathway, that regulates the progression from G2 into the mitotic phase of the cell cycle [9]. DREAM is a large dimerization complex that recruits several genes that all regulate the cell cycle, notably pRb/E2F. Normal cells conform to a set number of cell divisions, defined as Hayflick’s limit [16], by regulating apoptosis and cellular senescence. Otherwise, cells are transformed and cancerous. Apoptosis can occur from p53 activating pro-apoptotic signallers such as PUMA and NOXA, which then act at the mitochondria, where apoptosis largely occurs [17, 18]. Finally, all this activity that is initiated by p53 leads to cellular senescence—p53-mediated degradation and purposeful cessation of the cell cycle. A quiescent stage is a hallmark at which the cells are considered to no longer respond to growth factor signalling.
As one of the functions of p53 is p53-dependent apoptosis, its functions need to be tightly regulated. Ubiquitination is a form of post-translational modification, which results in p53 being directed for proteasomal degradation via the 26S proteasome [19, 20]. This process is otherwise normal and is achieved through MDM2. Another class of MDM proteins is MDM4, which targets the p53 transcription start site, slowing levels of expression [21]. Collectively, these two prevent p53 from inducing expression of several down-stream proteins including PUMA and NOXA, so that p53-mediated apoptosis is regulated.
3 The Role of p53 in Cancer
Naturally p53 is a transcriptional regulator, and outside the context of cancer may regulate processes for other developmental pathways. As discussed, p53 has potent anti-tumour effects. Consequently, p53 is one of the most mutated genes associated with cancer [22, 23]. There are a few ways in which p53 mutations may arise. Germline p53 mutants can be inherited, leading to the development of Li-Fraumeni or Li-Fraumeni-like syndrome [24]. This is the early development of cancer at a young age, coupled with poor prognosis as the cancers are of high risk, generally differing from the tissue of origin, making initial detection and treatment even more challenging [25]. Missense mutations can occur across multiple codons that are found within the DNA-binding domain, leading to aberrated binding and subsequent function of p53 [26]. Finally, viral proteins can mitigate the effects of p53, not by mutation, but rather by dysregulation. Viral proteins are able to target p53, allowing constant induction into the DNA replicative phase and cell cycle, bypassing its checkpoints and avoiding apoptosis [25]. Mutations in p53 are involved in, but not exclusive to, ovarian, oesophageal, colorectal, head and neck, larynx, and lung cancers, as well as primary leukemia, testicular cancer and malignant melanoma [27]. Other cancer types may aberrate, regulate or inactivate p53 to promote malignancy, but not necessarily mutate it. Some examples of this include sarcomas [28], myelomas [29] and HPV-driven cancers. The modes of p53 dysregulation including deletions, methylation, mutations, microRNAs (miRNAs), isoforms, and regulators in cancers have been reviewed elsewhere in detail [29]. Overall, p53 deficiency or mutations are strongly selected for during the evolution of a cancer cell.
4 p53 Dysfunction in Human Papillomavirus (HPV) Cancers
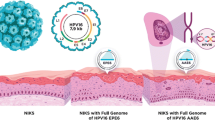
As part of the HPV life cycle, HPV encodes two oncoproteins, E6 and E7. Persistent HPV infection is required for the development of cancer [30, 31]. HPV first infects the deeper basal layer, where it also remains in a low copy number, making it harder to be detect by the immune system, remaining latent [32]. Viral load then steadily increases when progressing towards malignancy, where expression of E6 and E7 then concurrently increase, eventually becoming an invasive cancer spreading through to the more superficial suprabasal cell layer [32]. Only when integration of HPV has occurred do E6 and E7 oncoproteins become active, causing cells to progress towards malignancy [6]. Similarities to this may be drawn with that of the lytic lifecycle of a phage, where it remains latent until eventually producing and shedding more viral proteins, causing it to progress invasively. Indeed, HPV infection itself is not sufficient to induce cancer and requires further genetic mistakes to occur.
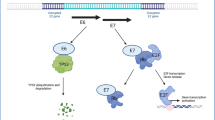
E7 functions as a transcriptional regulator [33] by binding to Rb protein and alleviating it from the transcription factor, E2F, subsequently leading to the entry from a quiescent G0 stage into a replicative S-phase. This hyperproliferative signal triggers p53 stabilisation. Although the E6 oncoprotein is able to bind to residual amino acids on a ubiquitin-protein ligase (UBE3A) known as E6AP [7], binding causes residual changes to the enzyme substrate complex, causing E6AP to bind surrounding proteins, including p53 [7, 34]. Through cross-linking of amino acid residues, p53 is placed right in the catalytic centre where it is ubiquitinated and eventually degraded by the 26S proteasome [7] (Fig. 2).
5 Previous Efforts to Reverse p53 Deficiency in HPV Cancers
There are several ways that p53 expression can be upregulated or bioavailability increased [35]. This has been shown to be successful, in the context of cancers still bearing a wild-type p53 [36]. Targeting p53 aggregates have also been proposed as a potential therapy, as common protein aggregates can lead to p53 clearance in cells [37]. Inhibiting the aggregation process has led to elevated levels of normal p53 functions [38, 39]. Finally, there are several chemical compounds that have been used to increase p53 levels in cervical cancer treatments [35]. Other indirect ways include inhibition of MDM, which is involved in the negative feedback of p53. To this effect, the therapeutic potential of targeting MDM to re-engage p53 activity with MDM2-specific inhibitors has been tested in both pre-clinical and clinical settings for a range of malignancies with a low frequency of p53 mutations, such as haematological malignancies, and indeed a number of these molecules are under clinical evaluation for acute myeloid lymphoma and multiple myeloma [40].
Previous studies have explored ways in which wild type (WT) p53 function can be restored or increased within the context of HPV cancers (Table 1). One successful example is a p53 gene therapy drug, gendicine, which was approved by the China Food and Drugs Administration (CFDA) in 2003. Gendicine is a human recombinant adenovirus that expresses a WT p53 protein, commonly used in combinational therapies within a clinical setting [41]. Expressing a WT p53 protein in combinational therapy has been the most common way in which p53 levels are increased and has certainly been shown to induce great therapeutic benefit when used in a combinational therapy setting. A clinical trial has compared the use of an adenoviral vector expressing p53 when used in combination of neoadjuvant chemotherapy and found that combinational therapy resulted in tumour regression almost comparable to that in a cisplatin, vinblastine, and bleomycin (PVB) treatment group [42]. Other pre-clinical studies have also explored the in vitro use of an adenoviral vector expressing p53 (rAd-p53) [43], and others have ventured into the delivery of a WTp53 plasmid using nanoparticles in HeLa cells, an HPV-positive (+) cervical cancer cell line [44]. Inhibiting proteasomal degradation of p53 using bortezomib and siRNA targeting of E6 and E7 also increased p53 levels in HPV+ head and neck squamous cell carcinoma (HNSCC) cells, and hence could serve as a dual-targeting approach [45]. miRNAs are also involved in p53 regulation through ubiquitin ligases. Indeed, miR-375 overexpression combined with radiotherapy increases p53-dependent apoptosis in HeLa cells [46]. Curcumin is a derivative of turmeric and when combined with paclitaxel, an existing chemotherapeutic agent, was shown to increase p53 levels in HPV+ cervical cancer cells [47]. Celecoxib is a non-steroidal inhibitor of COX-2 and targets anti- and pro-p53 networks, resulting in a net upregulation of p53 in a range of HPV+ cancer cell types, including patient-derived tumours [48]. RITA, a class of drug that activates p53 and its apoptotic-dependent pathways, has been shown to induce p53 and p53-apoptotic-dependent proteins in vitro and HPV+ cervical cancer tumour suppression in vivo [49]. Finally, reactivating p53 by targeting pathways or interrupting oncoprotein function has also been successful in restoring some p53 function [50, 51]. These studies in particular reveal a lot about the interaction that the oncoproteins have at the molecular level, through p53 interaction or surrounding proteins that affect p53 function. All these interventions have been shown to increase p53 expression and restore the activity of p53-dependent apoptotic pathways to culminate in tumour cell death. Finally, any form of E6 silencing or knockout using siRNA- or CRISPR-based technologies would also result in subsequent increase of p53 expression [52].
6 Using CRISPRa to Hyperactivate p53 in HPV Cancers
CRISPR activation (CRISPRa) is a candidate therapeutic tool for HPV-driven cancers. In general, the CRISPR system consists of a gRNA, also known as CRISPR RNA (crRNA), allowing for efficient Watson-Crick base pairing to the complementary DNA. Inclusion of a tracrRNA helps direct and complex the gRNA with the Cas9 endonuclease [53]. CRISPRa is a variant of CRISPR that allows activation of endogenous genes through promoters [54]. It consists of a “deficient” cas9 (dCas9), whereby the nuclease’s activity is diminished through mutating its domains, but still allowing for gRNA/Cas9 complexing to occur [55]. When a deficient Cas9 variant is fused with heterologous activator domains, endogenous genes may be hyper-expressed based on what the gRNA targets [53]. Using activator domains such as a construct of VP64-p65-Rta amplifies the response when hyper-expressing endogenous genes [53].
CRISPRa has been adopted as a tool for multiplexed activation of endogenous genes as an immunotherapy (MAEGI) [56]. The aim is to increase the amount of tumour-associated antigens using CRISPRa, allowing for more robust binding CD8+ T cells, thus inducing a stronger adaptive immune response [56]. CRISPRa has also been used to screen and study pro-oncogenes by activating them [57], revealing information on cancer pathways and vulnerabilities for future treatments. Outside these methods of screening, this system has not been used to increase bioavailability of tumour suppressor proteins, such as p53, let alone been used within the context of treating cancers. Given that HPV cancers retain a WT form of p53, reversing the HPV-mediated degradation of p53 and subsequent function by hyper-expressing p53 is an attractive proposal.
This poses the future possibility of utilising such a CRISPRa/dCas9 mechanism with a gRNA intended to target a p53 promoter. Having a gRNA that targets a p53 transcriptional activator, with the utilisation of a dCas9 fused with activator domains, could then hyper-express p53 and potentially overcome HPV-mediated p53 degradation. The question remains then whether the anticipated effects of p53 hyper-expression would be advantageously therapeutic, resulting in the reduction of tumour burden and overall disease. Indeed, a recent study provided proof-of-concept data that increasing p53 bioavailability using fenofibrate, which belongs to a class of drugs prescribed to manage dyslipidaemias, resulted in the alteration of HPV+ head and neck tumour microenvironment in vivo and loss of HPV+ cancer cell proliferation [58]. After all, it is well known that the introduction of p53 into the tumour microenvironment has anti-tumour effects resulting in cellular senescence [58,59,60] and p53-dependent apoptosis, causing cancer cell killing [58, 60,61,62]. A proposed design could utilise the well-known CRISPR doxycycline inducible system known as the tet-on/off system [58, 63], which is an approach our lab is currently testing in an effort to hyper-express p53. It is hypothesised that healthy cells remain untouched and that only tumour cells will be killed [62]. This is because the onset of oncogenic stressors and DNA damage are major contributors to the stabilisation of p53 [64, 65], likely to otherwise be absent in normal cells where p53 is regulated by MDM2. Only transformed cells that have DNA damage, or hyper-proliferative signals, such as caused by E7, would stabilise p53. Even if p53 were to be expressed in normal cells, its own negative feedback loop would kick in and rapidly degrade excess p53, further rationalising p53 being a good candidate for hyper-expression in HPV cancers.
7 Potential Challenges and Conclusions
In the context of HPV cancers, a switch from MDM2-p53 binding to E6-mediated degradation of p53 is a further hallmark of HPV+ cancers [4, 7]. It is a possibility that this will prove to be problematic as the increased levels of p53 will then need to compete with not only the E6 oncoprotein but also its own negative feedback loop involving the class of MDM proteins [20, 21]. The E7 oncoprotein has also been shown to play a role in p53 aberration to some extent. The p53-p21-DREAM is a target for the E7 oncoprotein and is directly involved in the p53-p21 activation pathway [10]. Therefore, increasing p53 expression alone may not be enough to overcome this and may require a combined approach, such as using standard chemotherapy [41, 42]. It has also been observed in yeast that hyper-expression of p53 leads to the formation of prions [66]. Whilst this is not yet proven in mammalian cell culture, it is a possibility that despite initial hyper-expression of p53, which may lead to tumour death, the formation of prions may cause a paradoxical induction of another cancer.
Directly targeting HPV E6 and E7 oncoproteins using RNA-based methods including siRNA, miRNA and CRISPR has been previously utilized to target HPV-driven cancers (see recent review by Salinas-Montalvo et al. [67]). Indeed, the use of CRISPR technology to delete HPV E7 has been shown to be successful in killing almost 100% of tumours in in vivo HPV+ cervical cancer models. [61]. As effective as this may be, it is likely that combination therapy of targeting HPV oncogenes and increasing p53 bioavailability can provide more efficacious therapeutic outcomes. Indeed, the co-delivery of a plasmid expressing p53 and the CRISPR-based targeting of the E7 oncoprotein led to the inhibition of HPV+ tumour growth and ultimately reversed the effects of HPV carcinogenesis in transgenic mice [60].
HPV-related cancers are becoming increasingly more prevalent, despite the vaccine being largely protective, especially for HPV+ head and neck cancers [68,69,70]. It is too early to draw conclusions of the impact of the HPV vaccine, which was initially designed for cervical cancer, on the incidence of other HPV-driven cancers as longitudinal studies to address this are currently ongoing. With poor prognosis and survival rates, HPV cancer patients need better therapy options. Despite being relatively new, gene therapy is making its way to the clinic in various forms. Reintroduction of p53 into the tumour microenvironment by simply adding a recombinant or pharmacologically targeting MDM has proven protective. This reflects the potent anti-tumour effects when p53 is present and why so many cancer types develop in the presence of p53 mutations or aberrations.
In conclusion, there are a lot of possibilities with the advent of gene-manipulating technologies such as CRISPRa, not just for hyper-expressing p53 for HPV-driven cancers, but also for other cancers carrying a WT p53 gene.
References
Sung H, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49.
Serrano B, et al. Human papillomavirus genotype attribution for HPVs 6, 11, 16, 18, 31, 33, 45, 52 and 58 in female anogenital lesions. Eur J Cancer. 2015;51(13):1732–41.
Castellsagué X, et al. HPV involvement in head and neck cancers: comprehensive assessment of biomarkers in 3680 patients. JNCI J Natl Cancer Inst. 2016;108(6):403.
Ndiaye C, et al. HPV DNA, E6/E7 mRNA, and p16INK4a detection in head and neck cancers: a systematic review and meta-analysis. Lancet Oncol. 2014;15(12):1319–31.
Arbyn M, et al. Estimates of incidence and mortality of cervical cancer in 2018: a worldwide analysis. Lancet Glob Health. 2020;8(2):e191–203.
Yeo-Teh NSL, Ito Y, Jha S. High-risk human papillomaviral oncogenes E6 and E7 target key cellular pathways to achieve oncogenesis. Int J Mol Sci. 2018;19(6):1706.
Sailer C, et al. Structural dynamics of the E6AP/UBE3A-E6-p53 enzyme-substrate complex. Nat Commun. 2018;9(1):4441.
Engeland K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 2018;25(1):114–32.
Fischer M, et al. The p53–p21-DREAM-CDE/CHR pathway regulates G2/M cell cycle genes. Nucleic Acids Res. 2016;44(1):164–74.
Fischer M, et al. Human papilloma virus E7 oncoprotein abrogates the p53–p21-DREAM pathway. Sci Rep. 2017;7(1):2603.
Levine AJ. p53: 800 million years of evolution and 40 years of discovery. Nat Rev Cancer. 2020;20(8):471–80.
My Cancer Genome. https://www.mdpi.com/2073-4425/11/6/704/htm#B1-genes-11-00704. Accessed 4 Mar 2022.
Mirgayazova R, et al. Therapeutic editing of the TP53 gene: is CRISPR/Cas9 an option? Genes. 2020;11(6):704.
Maréchal A, Zou L. DNA damage sensing by the ATM and ATR kinases. Cold Spring Harbor Perspect Biol. 2013;5(9):a012716.
Ozenne P, et al. The ARF tumor suppressor: structure, functions and status in cancer. Int J Cancer. 2010;127(10):2239–47.
Hayflick L, Moorhead PS. The serial cultivation of human diploid cell strains. Exp Cell Res. 1961;25(3):585–621.
Aubrey BJ, et al. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death Differ. 2018;25(1):104–13.
Bieging KT, Mello SS, Attardi LD. Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer. 2014;14(5):359–70.
Brooks CL, Gu W. p53 regulation by ubiquitin. FEBS Lett. 2011;585(18):2803–9.
Hou H, Sun D, Zhang X. The role of MDM2 amplification and overexpression in therapeutic resistance of malignant tumors. Cancer Cell Int. 2019;19(1):216.
Yu D-H, et al. Targeting MDMX for cancer therapy: rationale, strategies, and challenges. Front Oncol. 2020;10:1389.
Mendiratta G, et al. Cancer gene mutation frequencies for the US population. Nat Commun. 2021;12(1):5961.
Richardson RB. p53 mutations associated with aging-related rise in cancer incidence rates. Cell Cycle (Georgetown, Tex). 2013;12(15):2468–78.
Gargallo P, et al. Li–Fraumeni syndrome heterogeneity. Clin Transl Oncol. 2020;22(7):978–88.
Brychtová V, Hrabal V, Vojtěšek B. Oncogenic viral protein interactions with p53 Family Proteins. Klin Onkol. 2019;32(Supplementum 3):72–7.
Baugh EH, et al. Why are there hotspot mutations in the TP53 gene in human cancers? Cell Death Differ. 2018;25(1):154–60.
Olivier M, Hollstein M, Hainaut P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol. 2010;2(1):a001008–a001008.
Sciot R. MDM2 amplified sarcomas: a literature review. Diagnostics (Basel, Switzerland). 2021;11(3):496.
Herrero AB, et al. Molecular mechanisms of p53 deregulation in cancer: an overview in multiple myeloma. Int J Mol Sci. 2016;17(12):2003.
McLaughlin-Drubin ME, Münger K. The human papillomavirus E7 oncoprotein. Virology. 2009;384(2):335–44.
Hoffman SR, et al. Patterns of persistent HPV infection after treatment for cervical intraepithelial neoplasia (CIN): a systematic review. Int J Cancer. 2017;141(1):8–23.
Shukla S, et al. Physical state and copy number of high risk human papillomavirus type 16 DNA in progression of cervical cancer. Indian J Med Res. 2014;139(4):531–43.
Songock WK, Kim S-M, Bodily JM. The human papillomavirus E7 oncoprotein as a regulator of transcription. Virus Res. 2017;231:56–75.
Beaudenon S, Huibregtse JM. HPV E6, E6AP and cervical cancer. BMC Biochem. 2008;9(1):S4.
Zhao X, et al. Therapeutic potential of p53 reactivation in cervical cancer. Crit Rev Oncol Hematol. 2021;157:103182.
Tisato V, et al. MDM2/X inhibitors under clinical evaluation: perspectives for the management of hematological malignancies and pediatric cancer. J Hematol Oncol. 2017;10(1):133.
Kanapathipillai M. Treating p53 mutant aggregation-associated cancer. Cancers (Basel). 2018;10(6):154.
Chen Z, et al. Polyarginine and its analogues inhibit p53 mutant aggregation and cancer cell proliferation in vitro. Biochem Biophys Res Commun. 2017;489(2):130–4.
Chen Z, Kanapathipillai M. Inhibition of p53 mutant peptide aggregation in vitro by cationic osmolyte acetylcholine chloride. Protein Pept Lett. 2017;24(4):353–7.
Sanz G, et al. Inhibition of p53 inhibitors: progress, challenges and perspectives. J Mol Cell Biol. 2019;11(7):586–99.
Zhang WW, et al. The first approved gene therapy product for cancer Ad-p53 (Gendicine): 12 years in the clinic. Hum Gene Ther. 2018;29(2):160–79.
Xiao J, et al. Efficacy of recombinant human adenovirus-p53 combined with chemotherapy for locally advanced cervical cancer: a clinical trial. Oncol Lett. 2017;13(5):3676–80.
Liu YG, Zheng XL, Liu FM. The mechanism and inhibitory effect of recombinant human P53 adenovirus injection combined with paclitaxel on human cervical cancer cell HeLa. Eur Rev Med Pharmacol Sci. 2015;19(6):1037–42.
Han H, et al. Inhibition of cell proliferation and migration through nucleobase-modified polyamidoamine-mediated p53 delivery. Int J Nanomed. 2018;13:1297–311.
Li C, Johnson DE. Liberation of functional p53 by proteasome inhibition in human papilloma virus-positive head and neck squamous cell carcinoma cells promotes apoptosis and cell cycle arrest. Cell Cycle. 2013;12(6):923–34.
Song L, et al. miR-375 modulates radiosensitivity of HR-HPV-positive cervical cancer cells by targeting UBE3A through the p53 pathway. Med Sci Monit Int Med J Exp Clin Res. 2015;21:2210–7.
Dang Y-P, et al. Curcumin improves the paclitaxel-induced apoptosis of HPV-positive human cervical cancer cells via the NF-κB-p53-caspase-3 pathway. Exp Ther Med. 2015;9(4):1470–6.
Saha B, et al. Restoration of tumor suppressor p53 by differentially regulating pro- and anti-p53 networks in HPV-18-infected cervical cancer cells. Oncogene. 2012;31(2):173–86.
Zhao CY, et al. Rescue of p53 function by small-molecule Rita in cervical carcinoma by blocking E6-mediated degradation. Can Res. 2010;70(8):3372.
Caicedo-Granados E, et al. Wild-type p53 reactivation by small-molecule Minnelide™ in human papillomavirus (HPV)-positive head and neck squamous cell carcinoma. Oral Oncol. 2014;50(12):1149–56.
de Bakker T, et al. Restoring p53 function in head and neck squamous cell carcinoma to improve treatments. Front Oncol. 2022;11:799993.
Butz K, et al. siRNA targeting of the viral E6 oncogene efficiently kills human papillomavirus-positive cancer cells. Oncogene. 2003;22(38):5938–45.
Sontheimer EJ, Barrangou R. The bacterial origins of the CRISPR genome-editing revolution. Hum Gene Ther. 2015;26(7):413–24.
Dominguez AA, Lim WA, Qi LS. Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nat Rev Mol Cell Biol. 2016;17(1):5–15.
Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol. 2014;32(4):347–55.
Wang G, et al. Multiplexed activation of endogenous genes by CRISPRa elicits potent antitumor immunity. Nat Immunol. 2019;20(11):1494–505.
Wangensteen KJ, et al. Combinatorial genetics in liver repopulation and carcinogenesis with a in vivo CRISPR activation platform. Hepatology. 2018;68(2):663–76.
O’Neill WQ, et al. Repositioning fenofibrate to reactivate p53 and reprogram the tumor-immune microenvironment in HPV+ head and neck squamous cell carcinoma. Cancers. 2022;14(2):282.
Lujambio A, et al. Non-cell-autonomous tumor suppression by p53. Cell. 2013;153(2):449–60.
Xiong J, et al. Co-delivery of p53 restored and E7 targeted nucleic acids by poly (beta-amino ester) complex nanoparticles for the treatment of HPV related cervical lesions. Front Pharmacol. 2022;13:826771.
Jubair L, Fallaha S, McMillan NAJ. Systemic delivery of CRISPR/Cas9 targeting HPV oncogenes is effective at eliminating established tumors. Mol Ther. 2019;27(12):2091–9.
Dabiri Y, et al. p53-dependent anti-proliferative and pro-apoptotic effects of a Gold(I) N-heterocyclic carbene (NHC) complex in colorectal cancer cells. Front Oncol. 2019;9:438.
Das AT, Tenenbaum L, Berkhout B. Tet-On Systems For Doxycycline-inducible Gene Expression. Curr Gene Ther. 2016;16(3):156–67.
Shieh S-Y, et al. DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell. 1997;91(3):325–34.
Zhang Y, Xiong Y, Yarbrough WG. ARF promotes MDM2 Degradation and stabilizes p53: ARF-INK4a locus deletion impairs both the Rb and p53 tumor suppression pathways. Cell. 1998;92(6):725–34.
Park S-K, et al. Tumor suppressor protein p53 expressed in yeast can remain diffuse, form a prion, or form unstable liquid-like droplets. iScience. 2021;24(1):102000.
Salinas-Montalvo AM, et al. RNA-based gene targeting therapies for human papillomavirus driven cancers. Cancer Lett. 2021;523:111–20.
Ferreira DA, et al. A “hit-and-run” affair—a possible link for cancer progression in virally driven cancers. Biochim Biophys Acta Rev Cancer. 2021;1875(1):188476.
Kostov S, et al. A case of human papillomavirus infection and vulvar cancer in a young patient—“hit and run” theory. Gynecol Oncol Rep. 2021;36:100760.
Rossi C, Vanhomwegen C, Laurent F. HPV vaccination in boys and men: update and recommendations. Rev Med Brux. 2018;39(4):352–8.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this article. Open access funding was part of the CAUL transformative agreement.
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and material
Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
Code availability
Not applicable.
Author contributions
Conceptualization: Adi Idris, Nigel McMillan; writing – original draft preparation: Yusuf Idres; literature search: Yusuf Idres; writing – review and editing: Adi Idris, Nigel McMillan; supervision: Adi Idris, Nigel McMillan.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Idres, Y.M., McMillan, N.A.J. & Idris, A. Hyperactivating p53 in Human Papillomavirus-Driven Cancers: A Potential Therapeutic Intervention. Mol Diagn Ther 26, 301–308 (2022). https://doi.org/10.1007/s40291-022-00583-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-022-00583-5