Abstract
The remarkable advance in gene editing technology presents unparalleled opportunities for transforming medicine and finding cures for hereditary diseases. Human trials of clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein-9 nuclease (Cas9)-based therapeutics have demonstrated promising results in disrupting or deleting target sequences to treat specific diseases. However, the potential of targeted gene insertion approaches, which offer distinct advantages over disruption/deletion methods, remains largely unexplored in human trials due to intricate technical obstacles and safety concerns. This paper reviews the recent advances in preclinical studies demonstrating in vivo targeted gene insertion for therapeutic benefits, targeting somatic solid tissues through systemic delivery. With a specific emphasis on hemophilia as a prominent disease model, we highlight advancements in insertion strategies, including considerations of DNA repair pathways, targeting site selection, and donor design. Furthermore, we discuss the complex challenges and recent breakthroughs that offer valuable insights for progressing towards clinical trials.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Progress in utilizing in vivo targeted gene insertion, primarily clustered regularly interspaced short palindromic repeats/(CRISPR)/CRISPR-associated protein-9 nuclease (Cas9)-based, for treating inherited diseases has been demonstrated in preclinical hemophilia studies. |
Technical advancements in targeted gene insertion include strategies to enhance desired DNA repair pathways, developing universal target sites/locations and expression strategies, and promoting the survival of edited cells. Outcomes highlight efficiency, safety, and areas for improvement. |
We explore challenges in current technologies and the potential impact of novel advances such as lipid nanoparticles delivery and ex vivo gene editing on technology advancement and clinical trial drug development. |
1 Introduction
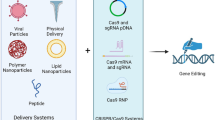
Genome editing technology has revolutionized biomedical research and promoted the development of novel biomedicines [1]. Engineered nucleases can recognize a preselected sequence in a genome and generate double-strand breaks (DSBs) with high specificity in various cell and organism models [2]. Zinc-finger nucleases (ZFNs) [3] and transcription activator-like effector nucleases (TALENs) [4] were early established tools that rely on reprogrammed DNA-binding protein motifs to target a specific sequence. The clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein-9 nuclease (Cas9) system has emerged later as a complex of a non-specific Cas9 nuclease and site-specific single guide RNA(s) (sgRNA) [5]. Complementary base-pairing between sgRNA and its target DNA sequence enables the Cas9/sgRNA complex to recognize and cleave the DNA at a preselected site [6]. The CRISPR/Cas9 system has surpassed ZFNs and TALENs owing to its remarkably enhanced specificity and flexibility in reprogramming sgRNA, becoming the most popular tool for genome manipulation [1].
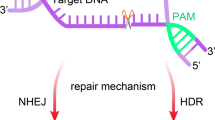
The genomic DSBs introduced by engineered nucleases trigger immediate cellular responses and evoke DNA repair via multiple intrinsic pathways. Non-homologous end joining (NHEJ) [7, 8], microhomology-mediated end joining (MMEJ) [9], and single-strand annealing (SSA) pathways [10] mediate error-prone repair processes, which predispose to indels at the DSB sites, whereas homology-directed repair (HDR) enables accurate repairing of a DNA lesion based on homology sequences from sister chromatid or exogenous templates [11]. By selectively favoring a particular DNA repair pathway and the corresponding repair outcomes, different gene editing strategies can achieve different forms of genome manipulation [12]. Gene/enhancer disruption, intron/exon deletion, mutation correction, and targeted gene insertion, have all been reported, in preclinical models, to produce therapeutic benefits [13]. Among these advances, gene insertion entails the integration of a large sequence into the host genome to restore normal function. This versatile strategy produces curative benefits regardless of the type, quantity, and location of disease-causing mutations [14]. However, the requirement of a donor DNA for targeted gene insertion poses significant technical hurdles, hampering its development for therapeutic application.
Therapeutic gene editing through in vivo delivery of the engineered nucleases has achieved great success using recombinant adeno-associated virus (AAV) [15], a vector system demonstrating excellent safety profiles in both preclinical and clinical studies [16, 17] and enabling the approval of eight gene therapy drugs by regulatory authorities (Table 1). AAV-delivered ZFNs for targeted gene insertion were first tested in humans (Table 2). Clinical trials for SB-913, SB-318, and SB-FIX were conducted in 2017–2019, inserting a healthy copy of gene into the albumin (ALB) locus to treat mucopolysaccharidosis (MPS) type I [18], MPS type II [19] and hemophilia B [20], respectively. Although the safety of AAV-ZFNs in humans was largely confirmed, the trials failed to demonstrate the desired therapeutic effectiveness [21]. Further investigations speculated the requirement of much higher AAV inputs for gene insertion therapy, which are prohibitively expensive and associated with high risks of toxicity.
Subsequent clinical studies for CRISPR/Cas9-based therapeutic gene editing were carried out with extra caution (Table 2). Exagamglogene autotemcel (Exa-cel) [CTX001] was designed for ex vivo genome editing in hematopoietic stem and progenitor cells (HSPCs) to alleviate sickle cell disease (SCD) and transfusion-dependent β-thalassemia (TDT) by disrupting the BCL11A enhancer [22] (Fig. 1). In vivo treatment using AAV-delivered CRISPR/Cas9 was initiated with a retinal injection of EDIT-101 for treating hereditary blindness Leber congenital amaurosis type 10 (LCA10). It co-delivers two sgRNAs with Cas9 to delete the intronic region containing the IVS26 mutation, and therefore rescues CEP290 function in photoreceptor cells [23]. Exa-cel, showcasing robust efficacy and safety in phase III clinical trials, was submitted to the European Medicines Agency (EMA) and United States (US) Food and Drug Administration (FDA) for regulatory approval in January 2023 [24]. It received marketing authorization in the UK on 16 November 2023, and subsequently gained approval from the FDA in the US on 8 December 2023 [25]. Meanwhile, the phase I/II clinical trial of EDIT-101 has revealed clinically significant improvement of visual acuity in 3/24 participants [26] (Fig. 1). These achievements have prompted CRISPR-mediated targeted gene insertion for disease therapy, which has not been tested in humans.
Overview of CRISPR drugs tested in humans. Left: CTX-001 is an ex vivo gene editing in HSPs, for the treatment of SCD and TDT by disrupting the BCL11A enhancer. Right upper: EDIT-101 is an in vivo treatment for LCA10 by deleting the CEP290 intronic region containing IVS26 mutation, which is delivered by AAV5 and administrated through retinal injection. Right lower: NTLA-2001 and NTLA-2002 are both liver-targeted in vivo editing systemically delivered by LNP. NTLA-2001 is intended for the treatment of ATTR by disrupting the TTR gene, while NTLA-2002 targets HAE by disrupting the KLKB1 gene. The status and year of clinical trials are shown. Lower: Schematics of gene editing strategies applied. Gene/enhancer disruption via NHEJ introduces indels at the target site (blue). Exon/intron deletion employs two sgRNAs to delete the disease-causing mutation (reddish). The labeled color of CRISPR drugs indicates the type of gene editing strategies applied. (Created with BioRender.com.) SCD sickle cell disease, TDT transfusion-dependent β-thalassemia, LCA10 Leber congenital amaurosis type 10, ATTR transthyretin amyloidosis, HAE hereditary angioedema, AAV adeno-associated virus, LNP lipid nanoparticle, CRISPR/Cas9 clustered regularly interspaced short palindromic repeat-associated 9 nucleases, indels insertions and deletions, NHEJ non-homologous end joining, HSPs hematopoietic stem cells, sgRNAs single guide RNAs
Currently, ongoing research conducted in animal models has exhibited robust therapeutic potential of gene insertion strategies. Hemophilia A and B, caused by the deficiency of blood coagulation factor VIII (FVIII) and IX (FIX), respectively, are among the most intensively investigated disease models [27]. In this review, we focus on the relevant preclinical studies that explore in vivo targeting strategies for achieving efficient somatic knock-in of therapeutic payloads via systemic administration. Technical components of different targeting strategies, new findings, remaining challenges, and potential solutions with implications in clinical translation will be discussed.
2 Strategies for Targeted DNA Insertion for in vivo Therapy via Systemic Delivery
Various targeting strategies relying on DSB-activated intrinsic repair pathways, including HDR, NHEJ, MMEJ and SSA, have been reported to introduce DNA insertion at specific loci. Each repair pathway has distinct advantages and limitations for targeted insertions, while donor designs can selectively enhance the targeting specificity and minimize non-specific and potential adverse effects.
2.1 Homology-Directed Repair (HDR)-Based Gene Insertion as a Classical and Broadly Used Targeting Strategy
HDR accurately corrects or removes a DNA lesion through synthesizing new DNA based on existing homology sequences [28]. Targeted gene insertion via the HDR mechanism requires donors carrying flanking homology arms of approximately 0.6–1.4 kb [29,30,31,32,33,34,35]. The long homologies facilitate the HDR repair of not only site-specific DSBs induced by a nuclease but also endogenous DNA lesions within a large region, which can lead to low frequency gene insertion in the absence of nuclease.
Targeted gene insertion via HDR repair is a widely adopted strategy to introduce therapeutic payloads in preclinical models. The compact size of the human F9 gene (hF9, 1.4 kb) allows the inclusion of long homology arms within the limited capacity of AAV vectors (< 4.7 kb), which therefore renders hemophilia B as a desirable model for HDR knock-in studies. In vivo hF9 insertion via HDR-based methods has been broadly achieved through systemic AAV administration of either ZFNs [30, 31] or CRISPR/Cas9 systems [32,33,34], demonstrating therapeutic benefits in both neonatal and adult mice with hemophilia B. The plasma hFIX levels ranged from 3 to 23% of normal in most studies targeting the F9 locus [30,31,32, 34], while Wang et al. observed notably higher plasma hFIX levels by targeting the mAlb locus, which reached 40% of normal in adult mice and 120% of normal in neonates [33]. Among these studies, the insertion rates of the hF9 transgene in somatic livers varied from 1 to 3.8% in adult mice, and from 11 to 16.1% in neonates, primarily due to donor designs and target loci involved, whereas on-target DNA cleavage was detected at much higher levels, with indel rates ranging from 34 to 47% [30, 32, 33]. Researchers have also explored nuclease-free gene correction/insertion, aiming to bypass the use of nucleases and minimize DNA damage; however, the insertional efficiency was relatively low. Barzel et al. delivered a single AAV8 vector carrying hF9 donor and flanking homology sequences to the Alb locus into hemophilia B mice without any nuclease, and detected only around 0.5% of Alb alleles carrying hF9 in both neonatal and adult mice [29]. In the study by Li et al. targeting the hF9 locus in humanized mice, administration of AAV8 hF9 donor in the absence of a nuclease failed to produce detectable hFIX [30].
In addition to Cas9, alternative Cas families also hold considerable promise in mediating in vivo gene insertion. The Cas12a (Cpf1) family offers a distinct advantage of producing staggered DSB ends, making it well-suited for precise and long homology-based HDR insertion [36, 37]. Recently, investigations have further highlighted a compact AsCas12f (422 amino acids) [38,39,40]. Hino et al. utilized a hyperactive variant enAsCas12f to facilitate HDR-based hF9 knock-in at the mAlb 3′ untranslated region (UTR) locus in hemophilia B neonatal mice using a single AAV vector, yielding over 40% of normal FIX activity [38].
In vivo gene insertion in livers via systemic delivery has also been applied to treat other diseases beyond hemophilia B. In 2015, Sharma et al. reported HDR-based gene insertion at the mAlb site as a versatile platform for therapeutic gene knock-in to treat liver metabolic diseases, including Fabry and Gaucher diseases, and Hurler and Hunter’s syndromes (also known as MPS type I or II, respectively) [41]. Meanwhile, in situ correction of disease genes via HDR-based insertion at the original loci has also shown therapeutic efficacy in addressing Crigler–Najjar disease, Ornithine transcarbamylase (OTC) deficiency, and Hereditary tyrosinemia type 1 (HT-1) diseases [42,43,44,45,46,47,48].
Collectively, these advances supported HDR-mediated insertion as a promising approach to treat a broad spectrum of diseases; nevertheless, the insertion rates attained are still limited, restricting it to treating diseases with low therapeutic thresholds [33]. Furthermore, the requirement of long homology arms in the donor imposes constraint on AAV packaging capacity, confining the application of HDR-based insertion to small transgene sequences. More investigations are thus warranted to enhance the insertional efficiency and reduce the length of homology, without sacrificing insertion precision.
2.2 Targeted Insertion via Non-homologous End Joining (NHEJ) Provides Distinct Alternatives
The NHEJ pathway functions particularly in repairing DSBs rather than other DNA lesions [7]; hence, a nuclease-induced site-specific DSB is essential for NHEJ-based targeted gene insertion. Unlike HDR, NHEJ repairs DSBs by directly ligating the broken DNA ends in a homology-independent and template-free manner [8]. The NHEJ donors are therefore devoid of homology arms and are presented in a linear form to expose the DNA ends [49, 50]. Remarkably, NHEJ insertion exhibited much higher efficiency than HDR-based strategy in cellular assays [49] due to the rapid and dominant nature of NHEJ-based DSB repair in mammalian species [51].
Despite the error-prone and nondirectional features of NHEJ repair, targeted insertion mediated by NHEJ emerges as a distinct alternative to the HDR strategy for therapeutic application, owing to its high efficiency and homology-independent flexibility [50]. In 2016, Suzuki et al. provided the first demonstration of NHEJ-mediated gene insertion in rodents [50]. This study also established a useful design termed homology-independent targeted integration (HITI), which can significantly enhance forward integration by reconstructing the sgRNA target sites and permitting re-cleavages upon reverse integration [50].
NHEJ-based insertion has been extensively employed in various in vivo disease models, particularly favoring the knock-in of larger genes such as hF8, as the limited capacity of AAV precludes the HDR vector design. Chen et al. used two AAV8 vectors to deliver SaCas9/sgRNA targeting the mAlb locus and an NHEJ donor carrying B-domain deleted hF8 (BDD-F8), a truncated gene of 4.4 kb to encode functional hFVIII [52]. The hemophilia A mice receiving intervention showed long-term and dose-dependent production of hFVIII, with the plasma hFVIII level and activity reaching around 34% and 13% of the normal, respectively. Droplet digital PCR (ddPCR) analysis detected the insertion of BDD-F8 at the mAlb locus in 0.2–0.3% liver DNA [52]. Meanwhile, Zhang et al. also performed effective somatic knock-in of BDD-F8 in hemophilia A mice using three AAV8 vectors to carry SpCas9, sgRNA targeting mAlb, and the BDD-F8 NHEJ donor. Stable plasma hFVIII was robustly detected within 1 month after the treatment, demonstrating 100–200% of normal activity [53].
Targeted insertion of the small hF9 gene via NHEJ is technically less challenging and is often used as trailblazer to explore new Cas9 systems, target sites, or donor designs to address unsolved issues. He et al. used triple AAV8 delivery of SpCas9, sgRNA, and the NHEJ hF9 donor targeting the mAlb 3′UTR to treat hemophilia B mice and achieved liver-specific hF9 knock-in via systemic administration [54]. They demonstrated a significant synergy yielded by the high Cas9 expression, active sgRNAs, and hyperactive hFIX Padua variant, which substantially reduced the AAV input doses required by approximately 100-fold in both neonatal and adult mice. RNA-seq analysis detected the mAlb-hF9 chimeric transcripts produced from desired hF9 insertion, at around 0.49% and 0.08% mAlb mRNA in the adult and neonatal mice receiving the lowest effective AAV doses. Lee et al. demonstrated hF9 knock-in using a less commonly used Cas9 from Campylobacter jejuni (CjCas9) and by targeting the APOC3 transgene locus, showing that a bidirectional AAV-trap donor resulted in much higher hFIX production at the insertion rate of approximately 3%, as both forward and reverse insertions could support functional expression (55). In another study, Chen et al. employed the HITI strategy to treat hemophilia B in a rat model and achieved a notable increase in forward hF9 insertion by 7.5-fold [56].
2.3 Other Repair Mechanisms Involved in Gene Insertion
The MMEJ pathway requires 1–16 nt homology flanking the DSB for repair [9]. DSB repair via the MMEJ mechanism is also an error-prone process and mostly contributes to larger deletions that often co-exist with the indels produced by NHEJ [57].
Studies have reported the potential of MMEJ to mediate large gene insertion at DSB sites by adopting linearized donors flanked by 20-bp microhomology sequences [58, 59]. However, the insertion of MMEJ donors was often supported by both MMEJ and NHEJ mechanisms, and the evident MMEJ insertion was only observed in mouse embryonic stem cells and fetus, suggesting its activity may be cell context-dependent [60]. Using AAV donors and ex vivo cell models, Fu et al. found that the MMEJ pathway competed with NHEJ and HDR in mediating the insertions at nuclease-induced DSBs at a relatively slower process [61]. Zhao et al. utilized a recombinant donor design that featured one microhomology arm and one long homology arm, which facilitated precise insertion in multiple cell models by employing both MMEJ and HDR repair [62]. To date, in vivo gene knock-in via MMEJ was only tested via electroporation and hydrodynamic injection of plasmids [60, 63]. Additional research is needed to examine the efficacy of AAV-delivered MMEJ-mediated in vivo knock-in, since the efficient and precise insertion circumventing the need of long homologous arms is appealing for AAV delivery.
Compared with MMEJ, SSA repair involves longer homology and leads to larger deletions [64]. The potential of SSA in gene knock-in has primarily been validated in single-stranded DNA (ssDNA)-mediated targeted insertion [65]. Co-delivery of Cas9/sgRNA and single-stranded oligonucleotides (ssODN; <200 nt) has previously led to the successful correction of HBB and CYBB mutations in human HSPCs [66, 67] and the efficient knock-in in mouse and rat embryos [68], which supported ssDNA donors as an attractive option because short homology arms at 30–100 nt can mediate effective insertion [69, 70]. To test the potential of ssDNA for somatic gene knock-in, Guan et al. delivered naked 120 nt ssODN and Cas9/sgRNA plasmids in adult mice by hydrodynamic injection, which indeed corrected a point mutation in the F9 gene, yet showing a low rate at only 0.56% due to plasma instability of ssODN [71]. Recently, long ssDNA (lssDNA) donors were reported to mediate gene-size knock-in and demonstrated high efficacy and low cytotoxicity in mouse embryos and multiple cell models [72,73,74]. New designs and modifications of ssDNA have further improved its transfection efficiency and in vivo stability [72, 75, 76], actively promoting new research. However, costly production and unstable delivery persist as unsolved obstacles, especially to the in vivo targeted insertion via systemic delivery.
3 Choices of Insertion Site and Expression Strategies
3.1 Maximizing Therapeutic Effect and Versatility by Insertion Locus Selection
The transgene expression and therapeutic benefits produced from a gene insertion greatly depend on the choice of integration locus (Table 3). Inserting a therapeutic sequence into the original locus to functionally replace the defective sequence was first tested in hemophilia B mice. Ohmori et al. [34] and Wang et al. targeted the endogenous mF9 locus [32], while Li et al. targeted hF9 transgene locus in humanized mice [30], for the integration of mF9 or hF9 exons 2-8. These insertions all achieved stable plasma FIX production and hemostasis correction. However, the efficacy of this substitutive insertion approach relies on the unscathed transcriptional activity of the target loci and hence cannot benefit diseased subjects who carry mutations in promotors or regulatory regions.
Alternatively, general target loci with high transcription activity and tissue specificity become attractive as versatile platforms for therapeutic insertion. The Alb locus is one of the most popular target sites owing to its exclusive and high expression in the liver. The proof-of-concept study by Sharma et al. in 2015 generated successful transgene knock-in at mAlb intron-1 for the treatment of hemophilia A and B and other liver metabolic diseases [41]. The versatility and efficacy of targeting other sites within the Alb locus were also investigated by using hemophilia A or B models [29, 33, 52,53,54, 56]. Thus far, no reduction of ALB production has been reported in mice carrying an insertion at Alb locus [33], but the validity of the concern is yet to be verified.
More recently, the APOC3 locus in humanized mice was assessed for hF9 insertion, producing 300 ng/mL plasma hFIX after administration of AAVs carrying CjCas9/sgRNA and a bidirectional hF9 donor [55]. APOC3 was selected because of its liver specificity and minimal safety risks, as decreased APOC3 expression is clinically asymptomatic or even beneficial for cholesterol modulation in the human body [55]. In 2023, Lee et al. also directed hF9 knock-in into Serpinc1 locus, which encodes antithrombin (AT), the most highly expressed anticoagulant factor. Marked improvement in coagulation activity was observed, attributed to the combinational effect of hFIX production and AT decrease [77]. Albeit showing promising results, careful control of AT suppression is necessary to prevent thrombotic adverse events.
Transgene knock-in at safe harbor loci such as Rosa26 in mice or AAVS1 in humans have rarely been examined for therapeutic insertion due to the requisite for exogenous promoters, which limited the donor delivery via AAV, and, meanwhile, may result in unexpected gene activation with risk for tumorigenesis [31, 50].
3.2 Balancing High Expression and Low Disturbance by Selecting a Target Site
Early research favored intronic targeting to avoid frameshift caused by NHEJ-produced indels. The F9 exon 2-8 from human/mouse has been inserted into the intron 1 of multiple loci, such as mF9 [34], hF9 [30] and mAlb [33, 41]; each produced plasma FIX and ameliorated hemophilia B symptoms in mice (Fig. 2). In these studies, a synthetic splice acceptor (SA) site was included in the donors to connect the integrated transgene sequence with the upstream exon 1, yielding desirable mRNA and protein [30, 33, 34, 41]. The mALB-hFIX hybrid proteins carried a short sequence from endogenous loci at the N-terminal but fully represented FIX activities for therapeutic benefits [33, 41]. It is noteworthy that the F9 and Alb exon 1 encode signal peptides, making the intron 1 an ideal target site for the expression of secretory proteins like FIX, FVIII, or some metabolic enzymes in liver, but may not be suitable for proteins with cell-autonomous functions.
Selection of insertion sites and expression strategies. Engineered nucleases, including ZFN and CRISPR/Cas9, have been systemically delivered by AAV for in vivo transgene insertion in mouse models. Intron 1, exon 2, intron 13, and exon 14 are intensively used insertion sites. SA is typically included in the donor when targeting intron 1, to connect the GOI with the upstream exon 1. Targeting exon 2 results in the fusion of GOI with exon 1, generating a chimeric protein. When targeting intron 13, a compensated exon 14 sequence is contained in the donor to avoid impacting endogenous gene expression level. Auxiliary 2A is also added to segregate two proteins. Stop codon and downstream 3′UTR at exon 14 are also tested in combination with 2A-GOI or IRES-GOI for bicistronic expression, respectively. (Created with BioRender.com.) AAV adeno-associated virus, ZFN zinc-finger nuclease, CRISPR/Cas9 clustered regularly interspaced short palindromic repeat-associated 9 nucleases, SA splicing acceptor, GOI gene of interest, 2A self-cleaving peptide, IRES internal ribosome entry site, UTR untranslated region
Recent studies also evaluated mAlb intron 13 for the targeted insertion of BDD-F8 using NHEJ approaches, recording plasma hFVIII production and hemostasis correction in hemophilia A mice [52, 53]. To avoid altering endogenous ALB protein, NHEJ donors were specially designed to compensate mAlb exon 14 coding region and carry auxiliary 2A sequences to separate ALB and hFVIII proteins. Using a hemophilia B rat model, Chen et al. demonstrated successful integration of hF9 Padua into the endogenous rat Alb intron 13 through an SpCas9/sgRNA-induced NHEJ-based HITI targeting approach [56] (Fig. 2).
Targeting sites in exons has been explored in parallel. Wang et al. inserted hF9 exon 2–8 into the endogenous mF9 exon 2 (Fig. 2), producing a chimeric mFIX-hFIX protein detectable in mouse plasma and with therapeutic activity [32]. Barzel et al. and De Caneva et al. targeted stop codon of mAlb at exon 14 to insert 2A-hF9 [29] and 2A-UGT1A1 [48], respectively, by applying HDR-based methods to precisely position the insertions. Via adopting a NHEJ-based strategy, He et al. specifically integrated hF9 at mAlb exon 14 but in the 3′UTR downstream of the stop codon [54] (Fig. 2). The donor contained an internal ribosome entry site (IRES) to translate hF9 separately from mAlb. Compared with the targeting at mAlb stop codon, insertion at the 3′UTR region provided greater flexibility in choosing a highly active sgRNA target site, which reached a higher efficiency and achieved therapeutic effects with much lower AAV doses [54].
The targeted insertions in adult mouse liver were generally detected in the vicinity of 0.5–3.8%, whereas the indels at DSB sites often reached much higher levels (12–50.3%) in both HDR- and NHEJ-based insertion studies [33, 54]. The high indel rates at target sites, driven by the intrinsic potency of NHEJ regardless of targeting strategies, underscore the value of intronic or 3′UTR-based targeting strategies in avoiding massive disruption of the coding sequences at target loci. Nonetheless, extra precautions are still necessary due to the presence of genomic regulatory elements.
3.3 Donor Design and Expression Strategies
Promoterless donors targeting a well-characterized locus, such as Alb, have drawn considerable attention in the field of gene insertion therapy (Table 3). Including a promoterless sequence serves the dual purpose of reducing donor size and bypassing exogenous promoters, which mitigates the risk of interfering with endogenous transcription, gene silencing [78], and unintended activation of oncogenes by random integration [79].
Ectopic SA is often used to link the promoterless transgene to the upstream endogenous exons upon intronic targeting [30, 33, 41], while self-cleaving 2A peptides and IRES discovered in viruses are widely used to introduce bicistronic expression [80] (Fig. 2).
The 2A sequences usually encode 18–22 amino acids and mediate simultaneous expression of two proteins through ribosomal skipping during translation [81]. The upstream protein carries the majority of 2A peptide residues at the C terminus, while the downstream protein contains one extra proline at the N terminus [82]. The small size of 2A sequences (54–66 bp) is ideal for AAV delivery, and the efficiency of 2A peptides in mediating bicistronic expression is typically high [80], especially P2A, which reached nearly 100% in a previous study [83]. Preclinical studies employing either HDR- or NHEJ-based insertion strategies have harnessed 2A sequences to separate transgenes from endogenous coding sequences, thereby achieving successful expression and therapeutic benefits [29, 48, 52, 53].
IRES is an RNA element that can recruit ribosomes to initiate translation from the internal region of mRNA [84], obviating the need for in-frame connection of two ORFs. It can therefore tolerate indels and serve as a suitable element for NHEJ insertion [85]. IRES-mediated bicistronic expression is less efficient, generally resulting in 20–50% expression of the upstream gene [86]. Nevertheless, in cases in which a target locus with high transcription activity (such as Alb) was selected, a superior level of plasma hFIX could still be produced with a low input AAV dose using IRES [54]. To date, the efficiencies of bicistronic expression, as well as the side products and their functional implications, were less thoroughly investigated in preclinical studies, warranting further examination in the future.
4 Remaining Challenges and Potential Solutions
4.1 Challenges Arising from Adeno-Associated Virus-Based Delivery
At present, in vivo gene insertion therapies validated in preclinical models have primarily relied on the AAV system for systemic delivery, and hence they face the same hurdles as AAV gene augmentation therapy [87]. The barriers include pre-existing immunity in the human population [88], limited payload capacity [89, 90], toxicity at high dose [91, 92], and difficulty in mass production [17]. In addition to the extensive research on advancing AAV technologies [93], studies on targeted gene insertion also explored the potential to address the obstacles arising from high AAV dosage via improved targeting designs or other delivery tools [94]. Through combining optimal settings, He et al. demonstrated robust and functional in vivo hF9 insertion with significantly reduced AAV input despite delivering SpCas9, sgRNAs, and donor using three separate AAV vectors [54]. Lee et al. alternatively combined viral and non-viral vectors for in vivo hF9 knock-in and revealed the possibility of replacing AAV for Cas9/sgRNA delivery with lipid nanoparticles (LNPs), which reduced the total AAV dose required [77, 95].
4.2 Relatively Low Efficiency of Somatic Gene Knock-In
In vivo gene insertion targeting somatic tissues including the liver could only achieve modest efficiency (approximately 1–3% of target alleles) with current technologies and tools, thus restricting the preclinical demonstration to diseases with low therapeutic thresholds, such as hemophilia. Although technological innovations can further enhance targeting efficiency, the extent of improvement may be limited by unidentified intrinsic factors. The evidence presented in the dosage analysis by He et al. suggested that the knock-in of hF9 in mouse livers may reach a plateau level, with approximately 4.66% of functional insertions detected in adult mice and 0.96% observed in neonates [54].
Another intriguing avenue to boost the therapeutic potential of somatic knock-in lies in the in vivo selective expansion of target cells. Mature hepatocytes with genetic advantages can expand substantially in vivo under selective conditions, as naturally manifested in HT1 mice with fumarylacetoacetate hydrolase (FAH) deficiency and Wilson disease (WD) mice with P-type copper-transporting ATPase (ATP7B) deficiency [96, 97]. Nygaard et al. developed a universal selection system by applying a chemical inhibitor to replicate FAH deficiency conditions, while delivering hF9 donor with short hairpin RNA (shRNA) against upstream enzymes to ameliorate the cytotoxicity specifically in targeted cells [98]. Indeed, hepatocytes with hF9 knock-in significantly repopulated in liver, and the plasma hFIX levels increased by 10- to 1000-fold after selective expansion, robustly reaching therapeutic levels. Recent studies then used acetaminophen (APAP) to induce cytochrome P450 (CYP)-dependent hepatotoxicity and co-delivered sgRNA against CYP-cofactor (Cypor) to render growth advantage in target cells, which displayed over 30-fold expansion of transgene-bearing hepatocytes in both CRISPR-mediated [99] and cleavage-free insertions [100].
4.3 Tissue Specificity of Transgene Expression and Somatic Genome Editing
Tissue-specific transgene expression is preferred but is challenging in AAV-delivered gene augmentation. While naturally occurring or engineered capsids may result in distinct tissue tropism [101], the features differ considerably between preclinical models and human patients, and exclusive specificity to certain tissues or cell types has not been conferred [102]. Markedly, targeted insertion of promoterless sequences restrains transgene expression under the control of endogenous target loci [49], which provides an excellent solution to introduce tissue/cell type-specific expression. Targeting hF9 integration at either mAlb or Apoc3 locus secured hepatocyte-specific expression in mouse [29, 30, 33, 41, 48, 52,53,54,55,56].
Moreover, tissue specificity arises as a significant challenge when implementing systemic administration of AAVs carrying nucleases for in vivo gene editing. To address this issue, Li et al. and Anguela et al. controlled ZFN activities by liver-specific ApoE/hAAT1 promoter and constricted hF9 knock-in and expression in liver, despite lacking direct verification of tissue-specific gene editing [30, 31]. He et al. and Lee et al. applied liver-specific LP1 and TBG promoters, respectively, and achieved confined Cas9 expression in mouse hepatocytes and liver-specific genome editing [54, 55]. These observations indicate the feasibility of cell-type specific genome editing, which could support a broad range of applications with enhanced safety profiles.
Owing to the privilege of germline cells in resisting AAV infection [103], vertical transmission of editing events through germlines have not been perceived, although in vivo gene editing and insertion via systemic AAV-delivery were widely reported in various somatic tissues [54, 104]. However, the possibility of germline infection by novel AAV capsids cannot be eliminated, which highlights the importance of confining editing activity to disease-relevant tissues/cells for new gene editing or insertion therapy.
4.4 Off-Target Effect
Off-targeting remains a pivotal concern for gene editing-related applications. Extensive efforts have been dedicated to minimizing the risk of unintended targeting, including advancements in in silico prediction, restriction on Cas9 activity duration, and further engineering of Cas9 protein and sgRNA [105]. Lee et al. effectively shortened the Cas9 expression window by employing LNP delivery instead of AAV [77, 95]. Additionally, high-fidelity Cas9 variants, such as HypaCas9 [106] and Cas9-HF1 [107], have been rationally designed, albeit with compromised activity to give way to specificity [108]. The chemical modifications of sgRNA have also been evidenced to enhance specificity through incorporating 2′-O-methyl-3′-phosphonoacetate (MP), bridged nucleic acids, or locked nucleic acids [109]. Optimizing sgRNA length such as addition of two guanine nucleotides [110] and truncation of 2–3 bp at the 5′ end could also diminish off-target potentials [111].
4.5 Unintended Integration and Editing Outcomes
Despite efforts to channel the repair process towards a specific pathway, endogenous mechanisms may competitively participate in repairing nuclease-induced DSBs and mediating gene insertion, resulting in diverse yet unintended modifications at the genomic target site. The prevalence of on-target indels regardless of targeting strategies was broadly confirmed. For instance, the study by Li et al. on HDR-based hF9 insertion revealed up to 45% indels yielded by NHEJ repair at the DSB site [30], while the study by He et al. on NHEJ-based hF9 insertion detected up to 42% indels, at a comparable level [54]. Subsequent studies by Sharma et al. and Wang et al. detected NHEJ-mediated donor integration events, despite the presence of HDR donors with homology arms [33, 41]. Additional undesired editing outcomes, including large deletions, chromosome rearrangement, and truncated or concatemerized integration, have also been reported [112,113,114]. Although new methods have been actively developed [114, 115], full-spectrum detection of these heterogenous and often unpredictable editing outcomes based on the current deep sequencing platforms remains technically challenging. Subsequent research is compulsory to minimize their occurrence and thoroughly evaluate the functional implications.
5 New Advances for Gene Editing Therapy
5.1 Lipid Nanoparticles Provide an Alternative Vehicle to Introduce In vivo Gene Insertion
LNP is another promising liver-targeted vehicle to deliver CRISPR/Cas9 through systemic administration, due to its primary uptake by hepatocytes mediated by the ApoE receptor [116]. Compared with recombinant AAV vectors, LNPs have a relatively short expression window and low immunogenicity, and offer advantages in potency, payload capacity and design flexibility [117] (Fig. 3).
Features of AAV and LNP delivery platforms and challenges for in vivo gene insertion. AAV and LNP have emerged as promising delivery vehicles for in vivo gene insertion. AAV exhibits a broad range of tissue tropisms, while LNP naturally targets mainly the liver despite recent advancements in other tissue targeting. AAV delivers the DNA vector, in which Cas9 and sgRNA are packaged separately or in the same vector. The donor was delivered by AAV DNA vector. LNP facilitates the encapsulation of mRNA or proteins to deliver Cas9/sgRNA as mRNA or RNP. LNPs also allow the delivery of dsODN donor, while large donor encapsulation remains a challenge. AAV-mediated in vivo gene insertion enables specific expression owing to the locus-specific insertion of promoterless donor and restriction of Cas9 expression. In contrast, LNP lacks expression specificity due to the difficulty in large donor delivery and the lack of restriction by cell type/tissue-specific promoters. The remaining challenges are shown in grey and are marked with red crosses. (Created with BioRender.com.) AAV adeno-associated virus, LNP lipid nanoparticle, Cas9 clustered regularly interspaced short palindromic repeat-associated 9 nuclease, sgRNA single guide RNA, RNP ribonucleoprotein, dsODN double-stranded oligodeoxynucleotide
The most common in vivo application of LNP-packed Cas9 mRNA/sgRNA is gene disruption in the liver. Han et al. targeted the Serpinc1 gene to lower AT expression in hemophilia mice, resulting in significantly reduced plasma AT and enhanced thrombosis. Moreover, in vivo bioluminescence imaging and indels analysis confirmed the predominant occurrence of editing in the liver [118]. Similarly, Qiu et al. disrupted the Angptl3 locus in mice and observed reduced serum angiopoietin-like 3 (ANGPTL3) and blood lipoprotein levels for more than 100 days after a single dosing [119]. These advances have prompted clinical tests of LNP-mediated gene disruption. NTLA-2001 and NTLA-2002 were devised as remedies for transthyretin amyloidosis (ATTR) and hereditary angioedema (HAE) by disrupting the TTR and KLKB1 genes, respectively [120, 121] (Fig. 1). According to the data released thus far, NTLA-2001 has successfully reduced serum TTR levels in all tested subjects with ATTR [122], while the HAE patients receiving NTLA-2002 were recorded with robust decline in plasma kallikrein levels and HAE attack rates [123].
To date, LNP-Cas9-mediated in vivo gene insertion was only attempted in combination with AAV or dsODN donor, given the challenges in delivering long DNA sequences via LNPs. Yin et al. first combined LNP-Cas9 mRNA/sgRNA and AAV donor for in vivo knock-in of Fah gene in HT1 mice, which achieved gene correction in > 6% of hepatocytes and ameliorated disease symptoms [95]. Another example is the study by Lee et al., in which coadministration of AAV donor and LNP-Cas9 complex showed efficient in vivo hF9 knock-in in hemophilia mice [77]. Co-delivery of LNP-Cas9 and dsODN donor (54 bp) was also conducted by Samanta et al. to correct the p.R83C mutation in mice with glycogen storage disease type-Ia (GSD-Ia), which produced approximately 3.6 ± 0.8% of normal G6Pase-α activity [124].
In summary, the well-demonstrated efficiency and safety profiles of LNP for in vivo gene editing in both preclinical and clinical trials have illustrated its potential in future gene insertion therapy.
5.2 Ex vivo Gene Editing and Potential of Cell Therapy for Hemophilia Treatment
Ex vivo gene therapy comprises the extraction of particular autologous cells from patients, followed by culturing, expansion, editing and selection in vitro, and eventual transplantation back to the patients. This approach avoids the toxicity associated with AAV systemic administration and non-specific tissue targeting [125]. Successful ex vivo gene insertions have been accomplished in hematologic cell types, achieving the highest efficiency through coupling AAV6 HDR-donor with Cas9/sgRNA ribonucleoprotein (RNP) [126]. Notably, AAV6-RNP delivery has been widely used to introduce the targeted insertion of engineered chimeric antigen receptor (CAR) into human T cells [127], and the generated CAR-T cells have been actively tested in clinical trials [128].
Engineering B cells for cell therapy represents a nascent therapeutic strategy for genetic diseases, including hemophilia [129]. Primary B cells can undergo differentiation into plasma cells, rendering them suitable for long-term expression of secretory factors and feasible for multiple infusions. Therefore, B-cell therapy provides incomparable flexibility in adjusting treatment according to patient responses. Furthermore, unlike engineered HSPCs and CAR T cells, autologous B-cell transplants can be efficiently engrafted without lymphodepletion, granting B-cell-based therapy a unique advantage by obviating the requirement for toxic preconditioning, which is a customary practice in emerging ex vivo approaches targeting HSCs for hemophilia treatment [130, 131]. In 2018, Hung et al. applied AAV6 HDR-donor alongside Cas9/sgRNA RNP to insert MND promoter-driven hF9 Padua at the CCR5 locus in human primary B cells and successfully generated secretory hFIX with coagulation activity [132]. Recently, Liu et al. further demonstrated the successful infusion of the engineered B cells into immunodeficient mice, revealing rapid engraftment and stable FIX production for up to 20 weeks [133]. Two studies identified the IgH intronic region between JH segments and Eμ enhancer as a promising site for targeted insertion in primary human B cells, leading to successful production of antigen-specific antibodies [134, 135]. In conclusion, ex vivo therapy offers distinct advantages and can potentially serve as a treatment option for addressing hemophilia, particularly as a compensatory approach when in vivo therapy falls short.
6 Conclusion
The past decade has witnessed the breakthrough of gene editing tools as well as the development of ZFN and CRISPR/Cas9-based in vivo gene editing, which opens new avenues for the treatment of inherited genetic diseases. Recent preclinical advances have focused on optimizing targeted gene insertion by leveraging distinct targeting mechanisms, donor designs, and insertion sites, resulting in promising efficacy and safety profiles. While insertion approaches feature unique advantages and achieved primary success in treating conditions such as hemophilia using animal models, several challenges need to be addressed prior to their transition to human clinical applications. The implementation of LNP delivery and ex vivo gene editing may offer feasible directions for addressing these concerns but demands more comprehensive and deliberate considerations, especially on practicality. Despite these challenges, the ongoing research efforts and continuous improvements in related studies suggest targeted gene insertion as a potent constituent of gene editing therapies and a potential candidate in the clinical treatment of genetic diseases.
References
Doudna JA. The promise and challenge of therapeutic genome editing. Nature. 2020;578(7794):229–36.
Gaj T, Gersbach CA, Barbas CF 3rd. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013;31(7):397–405.
Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD. Genome editing with engineered zinc finger nucleases. Nat Rev Genet. 2010;11(9):636–46.
Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol. 2013;14(1):49–55.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816–21.
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157(6):1262–78.
Lieber MR. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem. 2010;79:181–211.
Davis AJ, Chen DJ. DNA double strand break repair via non-homologous end-joining. Transl Cancer Res. 2013;2(3):130–43.
Wang H, Xu X. Microhomology-mediated end joining: new players join the team. Cell Biosci. 2017;7:6.
Bhargava R, Onyango DO, Stark JM. Regulation of single-strand annealing and its role in genome maintenance. Trends Genet. 2016;32(9):566–75.
Morrical SW. DNA-pairing and annealing processes in homologous recombination and homology-directed repair. Cold Spring Harb Perspect Biol. 2015;7(2): a016444.
Xue C, Greene EC. DNA repair pathway choices in CRISPR-Cas9-mediated genome editing. Trends Genet. 2021;37(7):639–56.
Li G, Li X, Zhuang S, Wang L, Zhu Y, Chen Y, et al. Gene editing and its applications in biomedicine. Sci China Life Sci. 2022;65(4):660–700.
Rezazade Bazaz M, Dehghani H. From DNA break repair pathways to CRISPR/Cas-mediated gene knock-in methods. Life Sci. 2022;295: 120409.
Wang D, Zhang F, Gao G. CRISPR-based therapeutic genome editing: strategies and in vivo delivery by AAV vectors. Cell. 2020;181(1):136–50.
Samulski RJ, Muzyczka N. AAV-mediated gene therapy for research and therapeutic purposes. Annu Rev Virol. 2014;1(1):427–51.
He X, Urip BA, Zhang Z, Ngan CC, Feng B. Evolving AAV-delivered therapeutics towards ultimate cures. J Mol Med (Berl). 2021;99(5):593–617.
Harmatz P, Lau HA, Heldermon C, Leslie N, Foo CWP, Vaidya SA, et al. EMPOWERS: a phase 1/2 clinical trial of SB-318 ZFN-mediated in vivo human genome editing for treatment of MPS I (Hurler syndrome). Mol Genet Metab. 2019;126(2):S68.
Muenzer J, Prada CE, Burton B, Lau HA, Ficicioglu C, Foo CWP, et al. CHAMPIONS: a phase 1/2 clinical trial with dose escalation of SB-913 ZFN-mediated in vivo human genome editing for treatment of MPS II (Hunter syndrome). Mol Genet Metab. 2019;126(2):S104.
Harmatz P, Prada CE, Burton BK, Lau H, Kessler CM, Cao L, et al. First-in-human in vivo genome editing via AAV-zinc-finger nucleases for mucopolysaccharidosis I/II and hemophilia B. Mol Ther. 2022;30(12):3587–600.
Sheridan C. Sangamo’s landmark genome editing trial gets mixed reception. Nat Biotechnol. 2018;36(10):907–8.
Frangoul H, Altshuler D, Cappellini MD, Chen YS, Domm J, Eustace BK, et al. CRISPR-Cas9 gene editing for sickle cell disease and beta-thalassemia. N Engl J Med. 2021;384(3):252–60.
Maeder ML, Stefanidakis M, Wilson CJ, Baral R, Barrera LA, Bounoutas GS, et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat Med. 2019;25(2):229–33.
CRISPR Therapeutics provides business update and reports first quarter 2023 Financial Results [press release]. 8 May 2023.
Hoy SM. Exagamglogene autotemcel: first approval. Mol Diagn Ther. Epub 17 Jan 2024. https://doi.org/10.1007/s40291-024-00696-z
Editas medicine announces clinical data demonstrating proof of concept of EDIT-101 from Phase 1/2 BRILLIANCE Trial [press release]. 17 November 2022.
Mancuso ME, Mahlangu JN, Pipe SW. The changing treatment landscape in haemophilia: from standard half-life clotting factor concentrates to gene editing. Lancet. 2021;397(10274):630–40.
Yang H, Ren S, Yu S, Pan H, Li T, Ge S, et al. Methods favoring homology-directed repair choice in response to CRISPR/Cas9 induced-double strand breaks. Int J Mol Sci. 2020;21(18):6461.
Barzel A, Paulk NK, Shi Y, Huang Y, Chu K, Zhang F, et al. Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature. 2015;517(7534):360–4.
Li H, Haurigot V, Doyon Y, Li T, Wong SY, Bhagwat AS, et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature. 2011;475(7355):217–21.
Anguela XM, Sharma R, Doyon Y, Miller JC, Li H, Haurigot V, et al. Robust ZFN-mediated genome editing in adult hemophilic mice. Blood. 2013;122(19):3283–7.
Wang L, Yang Y, Breton CA, White J, Zhang J, Che Y, et al. CRISPR/Cas9-mediated in vivo gene targeting corrects hemostasis in newborn and adult factor IX-knockout mice. Blood. 2019;133(26):2745–52.
Wang Q, Zhong X, Li Q, Su J, Liu Y, Mo L, et al. CRISPR-Cas9-mediated in vivo gene integration at the albumin locus recovers hemostasis in neonatal and adult hemophilia B mice. Mol Ther Methods Clin Dev. 2020;18:520–31.
Ohmori T, Nagao Y, Mizukami H, Sakata A, Muramatsu S-I, Ozawa K, et al. CRISPR/Cas9-mediated genome editing via postnatal administration of AAV vector cures haemophilia B mice. Sci Rep. 2017;7(1):4159.
Lisjak M, De Caneva A, Marais T, Barbon E, Biferi MG, Porro F, et al. Promoterless gene targeting approach combined to CRISPR/Cas9 efficiently corrects hemophilia B phenotype in neonatal mice. Front Genome Ed. 2022;4: 785698.
Li P, Zhang L, Li Z, Xu C, Du X, Wu S. Cas12a mediates efficient and precise endogenous gene tagging via MITI: microhomology-dependent targeted integrations. Cell Mol Life Sci CMLS. 2020;77(19):3875–84.
Dai X, Park JJ, Du Y, Na Z, Lam SZ, Chow RD, et al. Massively parallel knock-in engineering of human T cells. Nat Biotechnol. 2023;41(9):1239–55.
Hino T, Omura SN, Nakagawa R, Togashi T, Takeda SN, Hiramoto T, et al. An AsCas12f-based compact genome-editing tool derived by deep mutational scanning and structural analysis. Cell. 2023;186(22):4920–35.
Wu Z, Zhang Y, Yu H, Pan D, Wang Y, Wang Y, et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat Chem Biol. 2021;17(11):1132–8.
Wu T, Liu C, Zou S, Lyu R, Yang B, Yan H, et al. An engineered hypercompact CRISPR-Cas12f system with boosted gene-editing activity. Nat Chem Biol. 2023;19(11):1384–93.
Sharma R, Anguela XM, Doyon Y, Wechsler T, DeKelver RC, Sproul S, et al. In vivo genome editing of the albumin locus as a platform for protein replacement therapy. Blood. 2015;126(15):1777–84.
Yang Y, Wang L, Bell P, McMenamin D, He Z, White J, et al. A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice. Nat Biotechnol. 2016;34(3):334–8.
Ou L, DeKelver RC, Rohde M, Tom S, Radeke R, St Martin SJ, et al. ZFN-mediated in vivo genome editing corrects murine hurler syndrome. Mol Ther. 2019;27(1):178–87.
Krooss SA, Dai Z, Schmidt F, Rovai A, Fakhiri J, Dhingra A, et al. Ex vivo/in vivo gene editing in hepatocytes using “all-in-one” CRISPR-adeno-associated virus vectors with a self-linearizing repair template. Iscience. 2020;23(1):100764.
Wang L, Yang Y, Breton C, Bell P, Li M, Zhang J, et al. A mutation-independent CRISPR-Cas9-mediated gene targeting approach to treat a murine model of ornithine transcarbamylase deficiency. Sci Adv. 2020;6(7):eaax5701.
Ibraheim R, Tai PWL, Mir A, Javeed N, Wang J, Rodriguez TC, et al. Self-inactivating, all-in-one AAV vectors for precision Cas9 genome editing via homology-directed repair in vivo. Nat Commun. 2021;12(1):6267.
Zhang QS, Tiyaboonchai A, Nygaard S, Baradar K, Major A, Balaji N, et al. Induced liver regeneration enhances CRISPR/Cas9-mediated gene repair in tyrosinemia type 1. Hum Gene Ther. 2021;32(5–6):294–301.
De Caneva A, Porro F, Bortolussi G, Sola R, Lisjak M, Barzel A, et al. Coupling AAV-mediated promoterless gene targeting to SaCas9 nuclease to efficiently correct liver metabolic diseases. JCI Insight. 2019;5(15).
He X, Tan C, Wang F, Wang Y, Zhou R, Cui D, et al. Knock-in of large reporter genes in human cells via CRISPR/Cas9-induced homology-dependent and independent DNA repair. Nucl Acids Res. 2016;44(9): e85.
Suzuki K, Tsunekawa Y, Hernandez-Benitez R, Wu J, Zhu J, Kim EJ, et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature. 2016;540(7631):144–9.
Mao Z, Bozzella M, Seluanov A, Gorbunova V. DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells. Cell Cycle. 2008;7(18):2902–6.
Chen H, Shi M, Gilam A, Zheng Q, Zhang Y, Afrikanova I, et al. Hemophilia A ameliorated in mice by CRISPR-based in vivo genome editing of human factor VIII. Sci Rep. 2019;9(1):16838.
Zhang JP, Cheng XX, Zhao M, Li GH, Xu J, Zhang F, et al. Curing hemophilia A by NHEJ-mediated ectopic F8 insertion in the mouse. Genome Biol. 2019;20(1):276.
He X, Zhang Z, Xue J, Wang Y, Zhang S, Wei J, et al. Low-dose AAV-CRISPR-mediated liver-specific knock-in restored hemostasis in neonatal hemophilia B mice with subtle antibody response. Nat Commun. 2022;13(1):7275.
Lee JH, Oh HK, Choi BS, Lee HH, Lee KJ, Kim UG, et al. Genome editing-mediated knock-in of therapeutic genes ameliorates the disease phenotype in a model of hemophilia. Mol Ther Nucl Acids. 2022;29:551–62.
Chen X, Niu X, Liu Y, Zheng R, Yang L, Lu J, et al. Long-term correction of hemophilia B through CRISPR/Cas9 induced homology-independent targeted integration. J Genet Genom. 2022;49(12):1114–26.
Sfeir A, Symington LS. Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway? Trends Biochem Sci. 2015;40(11):701–14.
Yao X, Wang X, Liu J, Hu X, Shi L, Shen X, et al. CRISPR/Cas9—mediated precise targeted integration in vivo using a double cut donor with short homology arms. EBioMedicine. 2017;20:19–26.
Nakade S, Tsubota T, Sakane Y, Kume S, Sakamoto N, Obara M, et al. Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9. Nat Commun. 2014;5:5560.
Yao X, Wang X, Hu X, Liu Z, Liu J, Zhou H, et al. Homology-mediated end joining-based targeted integration using CRISPR/Cas9. Cell Res. 2017;27(6):801–14.
Fu YW, Dai XY, Wang WT, Yang ZX, Zhao JJ, Zhang JP, et al. Dynamics and competition of CRISPR-Cas9 ribonucleoproteins and AAV donor-mediated NHEJ, MMEJ and HDR editing. Nucl Acids Res. 2021;49(2):969–85.
Zhao Z, Shang P, Sage F, Geijsen N. Ligation-assisted homologous recombination enables precise genome editing by deploying both MMEJ and HDR. Nucl Acids Res. 2022;50(11): e62.
Aida T, Nakade S, Sakuma T, Izu Y, Oishi A, Mochida K, et al. Gene cassette knock-in in mammalian cells and zygotes by enhanced MMEJ. BMC Genom. 2016;17(1):979.
Vu TV, Das S, Nguyen CC, Kim J, Kim JY. Single-strand annealing: Molecular mechanisms and potential applications in CRISPR-Cas-based precision genome editing. Biotechnol J. 2022;17(7).
Kan Y, Ruis B, Takasugi T, Hendrickson EA. Mechanisms of precise genome editing using oligonucleotide donors. Genome Res. 2017;27(7):1099–111.
DeWitt MA, Magis W, Bray NL, Wang T, Berman JR, Urbinati F, et al. Selection-free genome editing of the sickle mutation in human adult hematopoietic stem/progenitor cells. Sci Transl Med. 2016;8(360):360ra134.
De Ravin SS, Li L, Wu X, Choi U, Allen C, Koontz S, et al. CRISPR-Cas9 gene repair of hematopoietic stem cells from patients with X-linked chronic granulomatous disease. Sci Transl Med. 2017;9(372).
Renaud J-B, Boix C, Charpentier M, De Cian A, Cochennec J, Duvernois-Berthet E, et al. Improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR-Cas9 nucleases. Cell Rep. 2016;14(9):2263–72.
Yoshimi K, Kunihiro Y, Kaneko T, Nagahora H, Voigt B, Mashimo T. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat Commun. 2016;7:10431.
Shao Y, Guan Y, Wang L, Qiu Z, Liu M, Chen Y, et al. CRISPR/Cas-mediated genome editing in the rat via direct injection of one-cell embryos. Nat Protoc. 2014;9(10):2493–512.
Guan Y, Ma Y, Li Q, Sun Z, Ma L, Wu L, et al. CRISPR/Cas9-mediated somatic correction of a novel coagulator factor IX gene mutation ameliorates hemophilia in mouse. EMBO Mol Med. 2016;8(5):477–88.
Shy BR, Vykunta VS, Ha A, Talbot A, Roth TL, Nguyen DN, et al. High-yield genome engineering in primary cells using a hybrid ssDNA repair template and small-molecule cocktails. Nat Biotechnol. 2023;41(4):521–31.
Gu B, Posfai E, Rossant J. Efficient generation of targeted large insertions by microinjection into two-cell-stage mouse embryos. Nat Biotechnol. 2018;36(7):632–7.
Quadros RM, Miura H, Harms DW, Akatsuka H, Sato T, Aida T, et al. Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol. 2017;18(1):92.
Han W, Li Z, Guo Y, He K, Li W, Xu C, et al. Efficient precise integration of large DNA sequences with 3′-overhang dsDNA donors using CRISPR/Cas9. Proc Natl Acad Sci U S A. 2023;120(22): e2221127120.
Richardson CD, Ray GJ, DeWitt MA, Curie GL, Corn JE. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat Biotechnol. 2016;34(3):339–44.
Lee JH, Han JP, Song DW, Lee GS, Choi BS, Kim M, et al. In vivo genome editing for hemophilia B therapy by the combination of rebalancing and therapeutic gene knockin using a viral and non-viral vector. Mol Ther Nucleic Acids. 2023;32:161–72.
Bhagwan JR, Collins E, Mosqueira D, Bakar M, Johnson BB, Thompson A, et al. Variable expression and silencing of CRISPR-Cas9 targeted transgenes identifies the AAVS1 locus as not an entirely safe harbour. F1000Res. 2019;8:1911.
Miller DG, Petek LM, Russell DW. Adeno-associated virus vectors integrate at chromosome breakage sites. Nat Genet. 2004;36(7):767–73.
Shaimardanova AA, Kitaeva KV, Abdrakhmanova II, Chernov VM, Rutland CS, Rizvanov AA, et al. Production and application of multicistronic constructs for various human disease therapies. Pharmaceutics. 2019;11(11):580.
Liu Z, Chen O, Wall JBJ, Zheng M, Zhou Y, Wang L, et al. Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci Rep. 2017;7(1):2193.
Donnelly MLL, Luke G, Mehrotra A, Li X, Hughes LE, Gani D, et al. Analysis of the aphthovirus 2A/2B polyprotein “cleavage” mechanism indicates not a proteolytic reaction, but a novel translational effect: a putative ribosomal “skip.” J Gen Virol. 2001;82(Pt 5):1013–25.
Kim JH, Lee SR, Li LH, Park HJ, Park JH, Lee KY, et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE. 2011;6(4): e18556.
Lee KM, Chen CJ, Shih SR. Regulation mechanisms of viral IRES-driven translation. Trends Microbiol. 2017;25(7):546–61.
Zhang C, He X, Kwok YK, Wang F, Xue J, Zhao H, et al. Homology-independent multiallelic disruption via CRISPR/Cas9-based knock-in yields distinct functional outcomes in human cells. BMC Biol. 2018;16(1):151.
Mizuguchi H, Xu ZL, Ishii-Watabe A, Uchida E, Hayakawa T. IRES-dependent second gene expression is significantly lower than cap-dependent first gene expression in a bicistronic vector. Mol Ther. 2000;1(4):376–82.
Wang D, Tai PWL, Gao G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat Rev Drug Discov. 2019;18(5):358–78.
Rapti K, Grimm D. Adeno-associated viruses (AAV) and host immunity—a race between the hare and the hedgehog. Front Immunol. 2021;12: 753467.
Grieger JC, Samulski RJ. Packaging capacity of adeno-associated virus serotypes: impact of larger genomes on infectivity and postentry steps. J Virol. 2005;79(15):9933–44.
Wu Z, Yang H, Colosi P. Effect of genome size on AAV vector packaging. Mol Ther. 2010;18(1):80–6.
Kishimoto TK, Samulski RJ. Addressing high dose AAV toxicity—’one and done’ or “slower and lower”? Expert Opin Biol Ther. 2022;22(9):1067–71.
Lek A, Wong B, Keeler A, Blackwood M, Ma K, Huang S, et al. Death after high-dose rAAV9 gene therapy in a patient with duchenne’s muscular dystrophy. N Engl J Med. 2023;389(13):1203–10.
Li C, Samulski RJ. Engineering adeno-associated virus vectors for gene therapy. Nat Rev Genet. 2020;21(4):255–72.
Taha EA, Lee J, Hotta A. Delivery of CRISPR-Cas tools for in vivo genome editing therapy: Trends and challenges. J Control Release. 2022;342:345–61.
Yin H, Song C-Q, Dorkin JR, Zhu LJ, Li Y, Wu Q, et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol. 2016;34(3):328–33.
Overturf K, Al-Dhalimy M, Tanguay R, Brantly M, Ou CN, Finegold M, et al. Hepatocytes corrected by gene therapy are selected in vivo in a murine model of hereditary tyrosinaemia type I. Nat Genet. 1996;12(3):266–73.
Padula A, Spinelli M, Nusco E, Bujanda Cundin X, Capolongo F, Campione S, et al. Genome editing without nucleases confers proliferative advantage to edited hepatocytes and corrects Wilson disease. JCI Insight. 2023;8(21).
Nygaard S, Barzel A, Haft A, Major A, Finegold M, Kay MA, et al. A universal system to select gene-modified hepatocytes in vivo. Sci Transl Med. 2016;8(342):342ra79.
Vonada A, Tiyaboonchai A, Nygaard S, Posey J, Peters AM, Winn SR, et al. Therapeutic liver repopulation by transient acetaminophen selection of gene-modified hepatocytes. Sci Transl Med. 2021;13(597).
Tiyaboonchai A, Vonada A, Posey J, Pelz C, Wakefield L, Grompe M. Self-cleaving guide RNAs enable pharmacological selection of precise gene editing events in vivo. Nat Commun. 2022;13(1):7391.
Srivastava A. In vivo tissue-tropism of adeno-associated viral vectors. Curr Opin Virol. 2016;21:75–80.
Westhaus A, Cabanes-Creus M, Dilworth KL, Zhu E, Salas Gomez D, Navarro RG, et al. Assessment of pre-clinical liver models based on their ability to predict the liver-tropism of adeno-associated virus vectors. Hum Gene Ther. 2023;34(7–8):273–88.
Rajasekaran S, Thatte J, Periasamy J, Javali A, Jayaram M, Sen D, et al. Infectivity of adeno-associated virus serotypes in mouse testis. BMC Biotechnol. 2018;18(1):70.
Nelson CE, Wu Y, Gemberling MP, Oliver ML, Waller MA, Bohning JD, et al. Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat Med. 2019;25(3):427–32.
Guo C, Ma X, Gao F, Guo Y. Off-target effects in CRISPR/Cas9 gene editing. Front Bioeng Biotechnol. 2023;11:1143157.
Chen JS, Dagdas YS, Kleinstiver BP, Welch MM, Sousa AA, Harrington LB, et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature. 2017;550(7676):407–10.
Kleinstiver BP, Pattanayak V, Prew MS, Tsai SQ, Nguyen NT, Zheng Z, et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature. 2016;529(7587):490–5.
Li T, Yang Y, Qi H, Cui W, Zhang L, Fu X, et al. CRISPR/Cas9 therapeutics: progress and prospects. Signal Transduct Target Ther. 2023;8(1):36.
Kocak DD, Josephs EA, Bhandarkar V, Adkar SS, Kwon JB, Gersbach CA. Increasing the specificity of CRISPR systems with engineered RNA secondary structures. Nat Biotechnol. 2019;37(6):657–66.
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S, et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 2014;24(1):132–41.
Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol. 2014;32(3):279–84.
Liu M, Zhang W, Xin C, Yin J, Shang Y, Ai C, et al. Global detection of DNA repair outcomes induced by CRISPR-Cas9. Nucl Acids Res. 2021;49(15):8732–42.
Canaj H, Hussmann JA, Li H, Beckman KA, Goodrich L, Cho NH, et al. Deep profiling reveals substantial heterogeneity of integration outcomes in CRISPR knock-in experiments. bioRxiv. 2019:841098.
Higashitani Y, Horie K. Long-read sequence analysis of MMEJ-mediated CRISPR genome editing reveals complex on-target vector insertions that may escape standard PCR-based quality control. Sci Rep. 2023;13(1):11652.
Lackner M, Helmbrecht N, Paabo S, Riesenberg S. Detection of unintended on-target effects in CRISPR genome editing by DNA donors carrying diagnostic substitutions. Nucl Acids Res. 2023;51(5): e26.
Witzigmann D, Kulkarni JA, Leung J, Chen S, Cullis PR, van der Meel R. Lipid nanoparticle technology for therapeutic gene regulation in the liver. Adv Drug Deliv Rev. 2020;159:344–63.
Kazemian P, Yu SY, Thomson SB, Birkenshaw A, Leavitt BR, Ross CJD. Lipid-nanoparticle-based delivery of CRISPR/Cas9 genome-editing components. Mol Pharm. 2022;19(6):1669–86.
Han JP, Kim M, Choi BS, Lee JH, Lee GS, Jeong M, et al. In vivo delivery of CRISPR-Cas9 using lipid nanoparticles enables antithrombin gene editing for sustainable hemophilia A and B therapy. Sci Adv. 2022;8(3):eabj6901.
Qiu M, Glass Z, Chen J, Haas M, Jin X, Zhao X, et al. Lipid nanoparticle-mediated codelivery of Cas9 mRNA and single-guide RNA achieves liver-specific in vivo genome editing of Angptl3. Proc Natl Acad Sci USA. 2021;118(10).
Gillmore JD, Gane E, Taubel J, Kao J, Fontana M, Maitland ML, et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N Engl J Med. 2021;385(6):493–502.
Seitzer J. NTLA-2002: CRISPR/Cas9-mediated gene knockout of KLKB1 to treat hereditary angioedema. J Allergy Clin Immunol. 2021;147(2):AB147.
Intellia and regeneron announce initial data from the cardiomyopathy arm of ongoing phase 1 study of NTLA-2001, an investigational CRISPR therapy for the treatment of transthyretin (ATTR) amyloidosis [press release]. 16 September 2022.
Intellia therapeutics presents new interim data from first-in-human study of NTLA-2002 for the treatment of hereditary angioedema (HAE) at the American College of Allergy, Asthma & Immunology 2022 Annual Scientific Meeting [press release]. 12 November 2022.
Samanta A, George N, Arnaoutova I, Chen HD, Mansfield BC, Hart C, et al. CRISPR/Cas9-based double-strand oligonucleotide insertion strategy corrects metabolic abnormalities in murine glycogen storage disease type-Ia. J Inherit Metab Dis. 2023;46(6):1147–58.
Li Y, Glass Z, Huang M, Chen ZY, Xu Q. Ex vivo cell-based CRISPR/Cas9 genome editing for therapeutic applications. Biomaterials. 2020;234: 119711.
Dever DP, Bak RO, Reinisch A, Camarena J, Washington G, Nicolas CE, et al. CRISPR/Cas9 beta-globin gene targeting in human haematopoietic stem cells. Nature. 2016;539(7629):384–9.
Ren J, Liu X, Fang C, Jiang S, June CH, Zhao Y. Multiplex genome editing to generate universal CAR T cells resistant to PD1 inhibition. Clin Cancer Res. 2017;23(9):2255–66.
Lin H, Cheng J, Mu W, Zhou J, Zhu L. Advances in universal CAR-T cell therapy. Front Immunol. 2021;12: 744823.
Edelstein J, Fritz M, Lai SK. Challenges and opportunities in gene editing of B cells. Biochem Pharmacol. 2022;206: 115285.
Doering CB, Gangadharan B, Dukart HZ, Spencer HT. Hematopoietic stem cells encoding porcine factor VIII induce pro-coagulant activity in hemophilia A mice with pre-existing factor VIII immunity. Mol Ther. 2007;15(6):1093–9.
Pavani G, Laurent M, Fabiano A, Cantelli E, Sakkal A, Corre G, et al. Ex vivo editing of human hematopoietic stem cells for erythroid expression of therapeutic proteins. Nat Commun. 2020;11(1):3778.
Hung KL, Meitlis I, Hale M, Chen CY, Singh S, Jackson SW, et al. Engineering protein-secreting plasma cells by homology-directed repair in primary human B cells. Mol Ther. 2018;26(2):456–67.
Liu H, Singh S, Mullen T, Bullock C, Keegan S, Patterson T, et al. Development of an ex vivo precision gene engineered B cell medicine that produces active and sustained levels of FIX for the treatment of hemophilia B. Blood. 2023;142(Suppl 1):463.
Moffett HF, Harms CK, Fitzpatrick KS, Tooley MR, Boonyaratanakornkit J, Taylor JJ. B cells engineered to express pathogen-specific antibodies protect against infection. Sci Immunol. 2019;4(35).
Hartweger H, McGuire AT, Horning M, Taylor JJ, Dosenovic P, Yost D, et al. HIV-specific humoral immune responses by CRISPR/Cas9-edited B cells. J Exp Med. 2019;216(6):1301–10.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was supported by funds provided by the Research Grants Council of Hong Kong (14116719 and 14115520 to BF; T13-605/18-W to NI), and in part by the Children’s Thalassaemia Foundation in Hong Kong (CTFHK 2018/02 to BF), the Children’s Cancer Foundation in Hong Kong (CCF-PORG 2023.10 to BF), and the Health@InnoHK Program launched by the Innovation Technology Commission of the Hong Kong SAR, China. SZ received graduate studentships from the Chinese University of Hong Kong.
Conflicts of interest
Zhenjie Zhang, Siqi Zhang, Hoi Ting Wong, Dali Li, and Bo Feng declare they have no conflicts of interest that might be relevant to the contents of this manuscript.
Availability of data and materials
Not applicable.
Ethics approval
Not applicable.
Informed consent
Not applicable.
Author contributions
ZZ, SZ, and HW wrote the manuscript. DL and BF revised the manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Zhang, Z., Zhang, S., Wong, H.T. et al. Targeted Gene Insertion: The Cutting Edge of CRISPR Drug Development with Hemophilia as a Highlight. BioDrugs 38, 369–385 (2024). https://doi.org/10.1007/s40259-024-00654-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40259-024-00654-5