Abstract
The CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 genome editing system has been a major technological breakthrough that has brought revolutionary changes to genome editing for therapeutic and diagnostic purposes and precision medicine. With the advent of the CRISPR/Cas9 system, one of the critical limiting factors has been the safe and efficient delivery of this system to cells or tissues of interest. Several approaches have been investigated to find delivery systems that can attain tissue-targeted delivery, lowering the chances of off-target editing. While viral vectors have shown promise for in vitro, in vivo and ex vivo delivery of CRISPR/Cas9, their further clinical applications have been restricted due to shortcomings including limited cargo packaging capacity, difficulties with large-scale production, immunogenicity and insertional mutagenesis. Rapid progress in nonviral delivery vectors, including the use of lipid, polymer, peptides, and inorganic nanoparticle-based delivery systems, has established nonviral delivery approaches as a viable alternative to viral vectors. This review will introduce the molecular mechanisms of the CRISPR/Cas9 gene editing system, current strategies for delivering CRISPR/Cas9-based tools, an overview of strategies for overcoming off-target genome editing, and approaches for improving genome targeting and tissue targeting. We will also highlight current developments and recent clinical trials for the delivery of CRISPR/Cas9. Finally, future directions for overcoming the limitations and adaptation of this technology for clinical trials will be discussed.
Graphical Abstract

Similar content being viewed by others
Introduction
Genome editing has brought revolutionary changes in gene therapy research by elucidating biological mechanisms, improving disease modelling, implementing gene therapies, and developing precision medicines [1]. So far, gene editing has shown potential utility to treat single gene-linked conditions such as Huntington’s disease, cystic fibrosis, and sickle cell anaemia [2, 3]. The capacity has also been demonstrated in treating more complex polygenic conditions, such as cancers, heart disease, and neurological disorders; however, this usage is limited by difficulties in editing the gene of interest in multiple cells at differing sites [4]. Traditionally genome editing has used zinc nuclease fingers (ZFN) and transcription-activator-like effector nucleases (TALENS). The use of these tools required substantial time and effort to ensure that their design targeted specific genomic sequences of interest, and thus limited their use. The development of the clustered regularly interspaced short palindromic repeats (CRISPR), and the CRISPR-associated protein 9 (CRISPR/Cas9) system for genome editing has provided a cost effective and programable tool for precise genome editing, addressing issues with ZFN and TALENS [5].
The CRISPR/Cas9 system was first discovered as an adaptive immune effector system in bacteria [6]. The CRISPR/Cas9 system is composed of CRISPR association system (Cas) genes organized in an operon, which encodes nucleases that complex with single guide RNA (sgRNA), which is composed of a transverse acting CRIPSR RNA strand (tracrRNA) and CRIPSR RNA strand (crRNA) [7]. The Cas9 protein is an RNA-guided DNA endonuclease, which is a powerful gene editing tool. It acts as biomolecular scissors to cut the targeted DNA at a specific location specified by the sgRNA. The CRISPR/Cas9 complex randomly scans DNA in the cells through three-dimensional collisions to search for protospacer adjacent motifs (PAMs), which are short specific sequences that allows for further binding of the Cas9 nuclease to the DNA (Fig. 1). If no PAM site is detected, then Cas9 rapidly disassociates from the DNA [8,9,10,11]. If a PAM site is encountered, the CRISPR/Cas9 complex is bought into contact with the DNA strand and unwinds the first 10–12 nucleotides adjacent to the PAM sequence, allowing for the formation of a DNA-RNA hybrid. If complementary DNA binding occurs to the sgRNA’s first 8–12 nucleotides, referred to as the “seed region”, confirmational changes of Cas9 allow for its HNH nuclease domain to cleave the primary DNA stand, while the RuvC domain of the Cas9 cleaves the secondary DNA strand [8,9,10,11]. The first 13 nucleotides upstream of the PAM sequence are the most important regarding sgRNA and DNA binding, with mismatches in this “seed” region leading to failed binding and cleavage. In contrast, mismatches are more tolerated in regions downstream of the seed region, and thus can result in off-target cleavage [12].
Overview of the Cas9 endonuclease. TracrRNA is attached to 3′ end of crRNA and acts an anchor to the Cas9 endonuclease, this combined RNA molecule is known as a single guide RNA (sgRNA). Cas9 scans potential target DNA for the appropriate protospacer adjacent motif (PAM). When the protein finds the PAM, a confirmation change occurs leading to unwinding of the DNA allowing for interaction between the crRNA and DNA. The RuvC and HNH domains then cut the target DNA if complementary binding occurs between the guide and seed region
Genome editing using CRISPR/Cas9 results in permanent genomic changes in the affected cells. Typically, therapeutic gene editing in somatic cells specifically targets the inherited gene, strictly limiting the effect to the individual patients, without being passed down to future generations. In contrast, gene editing in germline cells may induce changes that can be inherited by future generations, sparking ethical concerns with its usage [10]. Notably, in 2019, a highly controversial illegal gene editing activity on human embryos violated ethics and aroused public concerns [13, 14]. A crucial challenge for applying CRISPR/Cas9 technology is off-target editing in host cells with editing occurring at undesired genome sites. Efforts have been made to optimize CRISPR/Cas9 systems and sgRNA sequences to limit off-target events [15, 16]. Eradication of off-target events is challenging, especially in larger genomes such as humans, with magnitudes higher proportion of off-target mutations occurring in humans than in simpler genomes [17]. Another challenge is gene-editing efficiency. Editing efficiency is dictated not just by the Cas and sgRNA complex, but also by the efficiency of the delivery methods. Additionally, tissue and cell-specific delivery of CRISPR/Cas9 systems remains a key challenge as well, and the successful targeted delivery will hinge many potential therapeutic applications [17].
To date, several CRISPR/Cas9 gene editing systems have been translated into clinical trials. The first clinical trial of using the CRISPR/Cas9-based DNA repair pathway was initially introduced in 2016 in China, where CRISPR/Cas9 mediated knockout of PD1 was performed to reactivate T cells in a patient with lung cancer [18, 19]. The United States Food and Drug Administration (FDA) has also listed more than 30 clinical trials of CRISPR/Cas9 to treat different genetic disorders, including carcinomas, sickle cell anaemia, inflammatory disorders and retinal disorder [20]. In this review, we will discuss the recent advances in the development of CRISPR/Cas9 gene editing tools and diverse strategies for its delivery. We will discuss the challenges and limitations of these systems and will highlight therapeutic applications. We will also shed light on the clinical translation of promising gene editing therapeutic technologies.
Approaches for Cas9 delivery
The delivery of sgRNA and Cas9 endonuclease into host cells remains a crucial challenge for gene editing. Their delivery into cells currently relies on two key aspects; firstly, which form the Cas9 system takes (plasmid encoding the Cas9 endonuclease (pDNA) and sgRNA, Cas9 mRNA plus sgRNA, or recombinant Cas9 protein and sgRNA forming a ribonucleotide protein (RNP)) and secondarily, the method to deliver the CRISPR/Cas9 system [21,22,23,24]. The selection of which system and delivery method is used will determine the time before editing occurs, and the likelihood of off-targeting events occurring, the choice of CRISPR/Cas9 system will also affect which delivery method is employed, for example Cas9 RNP will require a delivery construct which is capable of complexing with negatively charged protein complexes [15, 21].
In comparison to nucleic acid delivery, where nucleic acids (e.g., DNA, mRNA, siRNA, miRNA or antisense RNA) only need to be delivered as a single species to be effective, CRISPR/Cas9 systems require that the targeting sgRNA is complexed to the Cas9 endonuclease protein to achieve its biological functions [25]. Currently, three different approaches are applied to deliver CRISPR/Cas9 complexes with sgRNA (Fig. 2). CRISPR/Cas9 can be delivered as genetically encoded material (virally encoded, plasmid DNA (pDNA) or mRNA or as ribonucleoprotein (RNP)-based (Cas9 protein complexed with sgRNA) system [21]. Among these, pDNA is the most widely used form for the CRISPR/Cas9 system due to its stability, ease of construction and flexibility of plasmid preparation [26, 27]. Once pDNA is successfully delivered into a cell’s nucleus, it can efficiently undergo transcription and translation to express the desired Cas9 endonuclease (Fig. 2). Detectable genomic insertion/deletion (Indel) events from Cas9 endonuclease expression through pDNA delivery can take 24–48 h to appear, as unlike the other processes, DNA encoded Cas9 endonuclease requires both transcription and translation to occur to generate functional Cas9/sgRNA complexes. In addition, as pDNA can replicate in the host, with basal expression of Cas9 which may increase the chance of off-target gene editing. Furthermore, pDNA can also potentially undergo undesirable stable integration into the host genome, which may lead to long-term expression of Cas9, increasing the likelihood of off-target events occurring [28, 29].
Delivery of CRISPR/Cas9 in the form of pDNA, mRNA, and RNP. pDNA will converge into mRNA pathway, and mRNA will converge into the RNP pathway. pDNA steps: Successful delivery of pDNA results in the plasmid being transported to the nucleus, where transcription can occur. Transcribed sgRNA and Cas9 mRNA is exported from the nucleus to the cytoplasm. mRNA steps: mRNA is translated to yield the folded Cas9 endonuclease, which then interacts with sgRNA to form the Cas9 RNP complex. Cas9 RNP steps: the RNP complex is imported into the nucleus, where genome editing occurs
In comparison to plasmid DNA, the Cas9 mRNA is simpler to introduce and quicker to express, as this form circumvents the requirement for nuclear delivery for transcription of DNA and can commence mRNA translation instantly [25, 30]. Additionally, this approach alleviates the plasmid-associated risk of insertional mutagenesis and reduces chances of off-target effect due to short time presence in the cell and reverse transcription to Cas9 DNA does not occur. However, mRNA is unstable compared to plasmid DNA and is highly susceptible to RNase-mediated degradation.
The use of purified RNP Cas9 is a straightforward form of the system that has attracted much research interest (Fig. 2). Gene editing using this system is the most rapid due to its bypassing of transcription and translation steps, with indel events detected from 1 hr of delivery. RNP-based approaches also have reduced off-target effects when compared to the pDNA system due to the short-lived transient presence of the Cas9 endonuclease. However, due to the high molecular weight of Cas9 protein and the additional requirement to be connected with sgRNA and delivery difficulties due to unable to cross the cell membrane, loading and delivering protein cargo is challenging, therefore, requires more optimization to formulate into nanoparticles that can load the protein [31, 32].
Strategies for CRISPR/Cas9 delivery
A significant bottleneck for in vivo applications of CRISPR/Cas9 systems is the availability of an efficient delivery vehicle. Currently CRIPSR/Cas9 delivery systems have limited effectiveness at bypassing biological barriers (e.g. systemic degradation, targeted intracellular delivery, endosomal escape, nuclear localization) required for successful delivery [33]. Therefore, efforts have conducted to identify safe, less immunogenic, biocompatible, and efficient delivery systems for CRISPR/Cas9. Such delivery systems are broadly classified into two types — viral and nonviral delivery systems.
Viral vectors are commonly used for in vivo delivery as they can incorporate therapeutic genes into their genome or encapsulate genetic materials to facilitate their intracellular delivery [34]. Commonly used viral vectors include Adeno associated virus (AAV), adenoviruses, and lentiviruses [35]. However, the applications of these vectors are restricted due to the limited package size of the vector limiting use of CRISPR/Cas9 constructs to smaller variants systems, as well as concerns with immunogenicity, which limits their ability to be used multiple times in the same host. Viral vectors are also associated with insertional mutagenesis, tumorigenesis, and other associated safety issues [36]. These shortcomings have led researchers to explore safer and less immunogenic alternatives for delivering CRISPR/Cas9 components.
The nonviral approach used for CRISPR/Cas9 delivery can be split between physical delivery and carrier-mediated delivery (Table 1). The most common physical methods are reliable but lack specificity and scalability. Carrier-mediated delivery can provide beneficial characteristics of scalability and specificity, biodegradablity, high packaging capacity, ease of fabrication, and biostability [37]. Many nonviral carriers have been investigated in different studies for CRISPR/Cas9 delivery (Table 1) Among all, lipid-based, polymer-based, peptide-based, and inorganic nanocarriers are the most widely studies delivery systems.
Physical methods for nonviral CRISPR/Cas9 delivery
Physical nonviral gene delivery methods are less complex than other non-viral approaches for the delivery of CRISPR/Cas9 cassettes. They employ physical forces to facilitate the intracellular delivery of CRISPR/Cas9 components though the disruption of host cellular and nuclear membranes. Despite their laboratory success, in vivo applications of physical methods are still restricted due to their technical limitation of scalability and administration skill [38]. Physical delivery methods include electroporation, microinjection, and hydrodynamic injection. Among these, electroporation is a widely used method for both in vitro and ex vivo delivery [33, 39]. It employs electrical currents to stimulate the formation and opening of pores in cell membranes, which permits the internalisation of the CRISPR/Cas9 cargo. Ex vivo electroporation approaches have been used in several clinical trials for CRISPR/Cas9 therapeutic delivery. For example, autologous CD34+ cells edited with CRISPR/Cas9 targeting the BCL11A enhancer were administered back to donor patients by electroporation for the treatment of transfusion-dependent β-thalassemia (TDT) and sickle cell disease (SCD) (Clinical trial number: NCT03745287; NCT03655678) [40]. Despite its success in transferring these gene-editing tools into clinical trials; this method is still limited in ex vivo applications and the requirement of in vivo application without prior isolation of cells for electroporation in the process of further development [41].
Microinjection is one of the most used techniques to deliver CRISPR/Cas9 into cells. It involves injecting CRISPR/Cas9 into cells via the use of a microneedle under microscopic visualisation conditions. This technique has challenges relating to its intensive technical requirements to inject a single cell by hand, and scalability, thus is still confined to animal models and excludes it from in vivo human application [33, 41]. However, direct delivery through injecting the CRISPR/Cas9 therapeutic system has the advantage of delivering all form of CRISPR/Cas9 system regardless of their overall size of the delivery approach, e.g., protein. For example, in a study by Ma et al. [42], this technique was efficiently applied in knocking out four genes (ApoE, B2m, Prf1, and Prkdc) in rats in one-step, by co-injection of Cas9 mRNA and sgRNAs into zygotes. In a recent study, Crispo et al. [43] successfully delivered sgRNA along with Cas9 protein specific for ovine myostatin (MSTN) through microinjection into the cytoplasm of ovine zygotes to disrupt the MSTN genomic sequence in the whole embryo. This approach resulted in successfully knocking out of MSTN in ship. The intensive nature of this system limits its usage in vivo delivery as only single cells can be efficiently transfected.
While microinjection and electroporation are limited to use in in vitro or ex vivo cultured cells, hydrodynamic injection has already been applied for in vivo delivery of CRISPR/Cas9 tools. Hydrodynamic injection involves the injection of a large volume of solution containing the CRISPR/Cas9 system into animal's bloodstream [21]. This causes a sudden increase in hydrodynamic pressure, which temporarily improves the permeability of endothelial and parenchymal cells, allowing for the CRISPR/Cas9 payload to enter cells in many tissues including the heart, liver, kidneys, lungs, and muscles [21]. For example, Zhen et al. successfully applied the hydrodynamic injection technique to deliver CRISPR/Cas9 pDNA targeting the coding region of the hepatitis B surface antigen (HBsAg) into hepatitis B virus (HBV)-positive mice through tail vein injection [44]. Immunohistochemical results demonstrated almost no HBsAg-positive cells in the liver tissue of HBV infected mice. However, this technically simple method is only available for in vivo applications due to the requirement of hydrodynamic pressure which is not applicable for in vitro applications. Furthermore, hydrodynamic delivery is quite traumatic to the organism, commonly resulting in cardiac dysfunction, liver expansion, and elevated blood pressure [45].
Lipid nanoparticle-based nonviral CRISPR/Cas9 delivery
Nanoparticles (NPs) are materials with nano-scale dimensions (e.g., 1–100 nm), which have unique biological properties due their surface properties and size [46]. NPs can be formulated or self-assembled and are broadly used for drug delivery. Lipid nanoparticle (LNP)-based delivery systems have been widely adopted for gene transfection. These systems commonly feature cationic phospholipids, which complex with and condense DNA, and improve their cellular uptake. Current research is focused on improving the properties of LNPs with respect to cell-penetration, endosome escape, reducing toxicity, degradation, and improving long-term storage stability [47, 48].
Due to the large size and negative charge of the Cas9 RNP complex, they cannot be readily transported through negatively charged mammalian cell membranes [21]. A key delivery system to address this issue is the application of cationic lipids to condense the anionic-charged cargo via electrostatic interactions into LNPs, which can facilitate endocytosis through the cell membrane [49, 50]. The condensation of these payloads provides protection from degradation. An optimal equilibrium between the complexation level required for endocytosis and the ability of the CRISPR/Cas9 system to be released at its site of action is required for efficient transfection. The use of LNPs instead of traditional viral delivery systems has been shown to lead to less immune responses because LNPs do not containing any viral-sourced immunogenic components. To date, they are being broadly utilized for in vitro, ex vivo, and in vivo applications [51].
LNP-based delivery systems provide the possibility for highly efficient and stable nonviral CRISPR/Cas9 delivery. For example, Wang and colleagues [52] used a library of bioreducible LNP-based delivery systems to form Cas9/sgRNA complexes. In this work three lipids (amines with disulphide bond and 14-carbon hydrophobic tail) were formulated with cholesterol, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE) and C16-PEG2000-ceramide which showed high gene knockout efficiency (> 70%) in human embryonic kidney (HEK) cells. Lipofectamine is a family of phospholipid formulations, which have been developed for optimal delivery of different oligonucleotides or proteins, including DNA (Lipofectamine™ 2000, 3000, and LTX), mRNA (Lipofectamine MessengerMAX™) and RNPs (Lipofectamine CRISPRMAX™) [53, 54]. Lipofectamine 3000 is a commercially available LNP-based system for gene delivery, which is used as a gold standard for pDNA-based CRISPR/Cas9 delivery research [55]. This cationic lipid-based system can associate with and cross anionic cell membranes to deliver genetic materials into cells [53, 54]. However, due to issues with rapid plasma clearance, immunogenicity and toxicity, Lipofectamine formulations are generally not applicable for clinical purposes [56, 57]. Functional modification of lipids can further improve the transfection efficiency and reduce the toxicity of cationic liposomes. For example, Zhang et al. [58] have developed a polymer polyethylene glycol-modified lipid nanocarrier (LPLNP) for in vitro delivery of cas9 Plasmid / sgRNA into A375 cells. The system efficiently condensed and encapsulated Cas9/sgPLK-1 plasmids to form a core–shell structure (PLNP/DNA) and achieved 47.4% transfection efficiency in A375 cells. In vivo application through intra-tumoral injection of as9/sgPLK-1 plasmids into melanoma tumor-bearing mice significantly downregulated Polo-like kinase 1 (PLK-1) protein and suppressed tumor growth (> 67%).
Polymer-based systems for CRISPR/Cas9 delivery
Polymers can form highly versatile molecular complexes with CRISPR/Cas9 systems, which can be functionalized to incorporate a variety of components to improve cell/tissue targeting, cell uptake and endosome escape [59,60,61]. In contrast to cationic lipids, cationic polymer carriers show chemical diversity and functional potential, providing more choices for flexible structural designs and may directly complex with CRISPR/Cas9 to improve its delivery characteristics [62]. Polyethylenimine (PEI) is a hydrophilic cationic polymers with linear or branched structures, and varying molecular weights, which can complex with and condense negatively charged DNA via electrostatic interactions and provides pH buffering capabilities that assist with endosomal escape [63, 64]. Ryu et al. [65] used 25-kDa branched PEI (bPEI-25 k) to deliver CRISPR/Cas9 plasmids to Neuro2a cells, with the polymer/plasmid complex demonstrating > 70% transfection efficiency, and > 20% of Nuero2a cells demonstrated Indels in the targeted genes, which is compatible to commercial reagents Lipofectamine 2000. Unfortunately, PEI exhibits significant toxicity due to its high density of cationic charge, which restricts its in vivo and clinical applications [66]. New polymers or combinations of polymers with less toxicity are currently under development for CRISPR/Cas9 system delivery [67]. Recently, O’Keeffe Ahern et al. [68] have successfully used a highly branched poly(β-amino ester) polymer to deliver an RNP-based CRISPR/Cas9 system to HEK293 and human recessive dystrophic epidermolysis bullosa (REDB) cell lines with high efficiency (15–20% of HEK293 cells demonstrated indels in the targeted genes, > 40% of REDB cells demonstrated indels in the targeted genes) and high cell viability (72% for HEK293) after transfection.
Polymers can also be incorporated into cationic LNP-based systems to deliver CRISPR/Cas9 cargo and to enhance in vivo stability [27]. Polyethylene glycol (PEG) is a family of neutral biocompatible polymers with high hydrophilicity and low toxicity [69, 70]. In many studies, PEG demonstrated reduced interactions with biological components, increased the colloidal stability of the delivery systems, and reduced uptake of the particles by the reticuloendothelial system, improving persistence [70,71,72]. Finn et al. [73] used the lipid myristoyl diglyceride-conjugated to PEG (DMG-PEG) in a LNP-based CRISPR mRNA/sgRNA delivery system to edit the transthyretin (TTR) gene in the liver, which achieved high in vivo efficiency (97% reduction in serum protein levels) in vivo for at least 1 year. The DMG-PEG complex has been demonstrated to be safe in humans, and is widely utilized in the Moderna mRNA-1273 COVID-19 vaccine elasomeran (Spikevax) [74]. Zhang and colleagues [75] used PEGylated chitosan to deliver Cas9 pDNA/sgRNA in vitro, providing a 15% transfection efficiency at low pH (6.5–6.8) in HEK293 cell line. PEGylated LNPs are promising for CRISPR/Cas9 delivery, but high levels of PEGylation may hinder interactions between phospholipids and cell membranes, blocking their cellular uptake. PEGylation of neutral and anionic LNPs has also been investigated. However, these systems demonstrate lower levels of cellular uptake, which is further reduced through PEGylation, hence, limiting its applications for these systems [71].
Peptide-based systems for CRISPR/Cas9 delivery
Peptide-based delivery systems have shown great potential for the delivery of CRISPR/Cas9 gene editing systems as cationic peptides have the potential to complex with more cargo than viral systems and facilitate intracellular delivery with reduced immune or cytotoxic responses. Cell-penetrating peptides (CPPs) are short peptides that can directly interact with cell membranes allowing for the intracellular delivery of therapeutic cargos through cell membranes and, therefore successfully used to facilitate intracellular delivery of therapeutic cargos [76].
Possibilities of CPP-mediated delivery of Cas9 RNPs have been explored in several studies where covalent approaches were used to conjugate the Cas9 endonuclease and CPP [76,77,78]. However, these studies revealed that the covalent approach for CPP-driven RNP delivery required large amounts of Cas9 and multiple incubation steps with target cells, resulting in low gene editing efficiencies and ultimately reduced therapeutic efficacy [78]. Thus, to increase gene editing efficiency, optimization of the RNP-CPP delivery approach has been required. Lostale-Seijo et al. [76] developed a non-covalent strategy for delivering a complex of the Cas9 RNP with an amphipathic CPP PT24 (PT24/Cas9 RNP). The complex was formed by a hydrazone bond formation between a cationic peptide scaffold PT24 and a hydrophobic aldehyde tail. This work reported that the complexes that were formed only required a single incubation step for efficient delivery of the Cas9 RNP cargo while demonstrating good efficiency and low toxicity. Similarly, Del’Guidice et al. [79] have used an amphiphilic peptide 6His-CM18-PTD4 to deliver functional transcription factor HoxB4, CRISPR/Cas9, and Cpf1 RNP complexes into the hard-to-transfect primary natural killer (NK) cells. This peptide is composed of a hexahistidine fused with the endosomolytic peptide CM18 and the CPP PTD4. The authors reported that co-incubation of the 6HisCM18-PTD4 peptide with spCas9 and/or asCpf1 CRISPR RNPs achieved robust gene editing in less than 2 min. When the same procedure of co-incubating with the transcription factor HoxB4 was investigated, transcriptional regulation was also achieved [79]. In another in vitro study, PEGylated NPs (P-HNPs) based on the cationic α-helical polypeptide poly(γ-4-((2-(piperidin-1-yl)ethyl)aminomethyl)benzyl-L-glutamate) (PPABLG) for the delivery of CRISPR/Cas9 expression plasmid and sgRNA were investigated [80]. This study demonstrated that P-HNP mediated delivery of a CRISPR/Cas9 plasmid/sgRNA targeting the polo-like kinase 1 (Plk1) gene resulted in a 35% gene deletion in HeLa cells,reduced the Plk1 protein level by 66.7%, suppressed tumor growth by > 71%, and prolonged animal survival to 60% within 60 days [80]. Recently, Gustafsson et.al [78]. investigated the capacity of RNA-delivering CPP, PepFect14 (PF14), to deliver a Cas9 RNP using a non-covalent interaction between the sgRNA and Cas9 RNP. The RNP-CPP complexes demonstrated high levels of efficiency (i.e., up to 80% gene editing in HEK293T cells), comparable to the commercial lipofectamine reagents RNAiMAX and CRISPRMax™. The possibility of using other cationic peptides like dendrons or dendrimers have also been evaluated as potential carriers for CRISPR/Cas9. In a recent study, Zamolo et al. [81] tested several dendron systems and identified one peptide dendrimer Z22 ((rl)8(krl)4(krl)2kk(C18)c) as a potential peptide-based career for CRISPR/Cas9 pDNA delivery with low immunogenicity and toxicity.
Inorganic nanoparticle-based delivery systems
Inorganic nanoparticles, including metal NPs, metal–organic frameworks, carbon nanotubes, and mesoporous silica nanoparticles, have also been used to facilitate the delivery of CRISPR/Cas9 systems [82, 83]. Due to being chemically inert, gold NPs (AuNPS) gained much interest in being used as a delivery tool, which also does not trigger host’s immune system. With high efficiency and low toxicity, gold NPs are considered as a safer alternative for the delivery of gene editing toolboxes. Lee and collaborators [84] injected gold NPs complexed with CRISPR/Cas9 system in vivo, and showed that there are no immune responses towards the complexes. Gold NPs have been successfully used in many in vitro, ex vivo, and in vivo applications [85]. Generally, the size and in-cell retention time of gold NPs(AuNp) are critical for their efficiency [86, 87]. Shahbazi et al. [88] designed an ex vivo colloidal gold NP-based CRISPR/Cas9 pDNA delivery system for treating hard-to-transfect haematopoietic stem and progenitor cells (HSPCs). The treated HSPCs demonstrated efficient gene editing with low toxicity, with greater than 80% cell viability observed after transfection. Wang and collaborators [89] used PEG-lipid/AuNPs/Cas9-sgPlk-1 (LACP) system to knock out the Plk-1 gene and down regulate Plk-1 protein expression in a melanoma model (subcutaneously implanted A375 cells). This Cas9-sgPlk-1 plasmid delivery achieved 65% down-regulation of Plk-1 protein compared to the PBS control. Mesoporous silica nanoparticles represent another inorganic nanoparticulate delivery platform that has been investigated for the delivery of nucleic acid-based molecules. These particles have a large internal surface area and high cargo loading capacity, which have demonstrated the potential to deliver CRISPR/Cas9 in several studies. Furthermore, silica nanoparticles can be readily chemically modified, including with lipids and polymers for improving encapsulation of CRISPR/Cas9 systems. Most of the silica based CRISPR/Cas9 delivery systems are mesoporous, formulated with tetraethyl orthosilicate (TEOS) and conjugated with lipid and polymer. For example, in a study conducted by Wang et al. [90], stimuli-responsive silica nanoparticles (SNPs) demonstrated the capacity to efficiently deliver CRISPR/Cas9 cargoes. In vivo studies showed that subretinally injected SNPs conjugated with all-trans-retinoic acid (ATRA) and intravenously injected glutathione-responsive SNPs conjugated with N-acetylgalactosamine effectively delivered Cas9 mRNA and RNP to murine retinal pigment epithelium (RPE) cells and liver cells with good biocompatibility and efficient gene editing capability (1.3-fold higher gene knockdown efficiency than lipofectamine 2000 and CRISPRMAX [90].
The nanoscale zeolitic imidazolate framework (ZIF) is a biomimetic metal–organic framework, which enables effective and cell-specific delivery of the CRISPR/Cas9 gene editing system [91]. Different cells showed different selectivity of the cancer cell membrane coated CRISPR/Cas9 in ZIF delivery, which allows for targeted delivery of gene editing tools, which good potential for future applications [91]. ZIF-8 is a cage-like coordination compound with rhombic dodecahedron crystals [92]. Alsaiari et al. [93] initially used ZIF-8 to deliver the Cas9 plasmid/sgRNA system in 2018, and the delivery system showed good biocompatibility, and loading capacity (17%) for both plasmid and sgRNA and gene knockdown efficiency (37% over 4 days) in Chinese hamster ovary (CHO) cells.
Targeted delivery for CRISPR/Cas9
Target-oriented delivery of gene therapeutics is an exciting approach for successful gene therapy. Gene editing using the CRISPR/Cas9 systems results in permanent genomic changes in the affected cells within an organism [21]. A crucial challenge for applying CRISPR/Cas9 technology is reduction of off-target editing events in host cells, which have been readily demonstrated in mice embryos and adult human cells [103, 104]. Two types of off-target effects are associated with CRISPR/Cas9-mediated gene editing. Several studies have demonstrated that the CRISPR/Cas9 protein can tolerate small mismatches between the sgRNA and genomic DNA, allowing for cleavage at off target sites [104,105,106]. This could lead to lethal genetic mutations, genotoxicity, and carcinogenesis. Several approaches have been applied to redesign sgRNA through the use of chemical modifications and engineering to reduce or prevent off-target gene editing [15]. Several studies reported that optimising the GC content (40–60%), shortening the length (18–20 bp), dual nicking of sgRNA and Cas9, and incorporation of 2′-O-methyl-3′-phosphonoacetate in the sgRNA backbone can increase target sequence specificity and decrease off-target effects [107]. Additionally, the development of alternative or improved Cas9 variants also provide a means to reduce off-target effects.
Predicting the genomic sites that are susceptible to CRISPR/Cas9 off-target activity is primarily determined using in silico modelling of the homology between the sgRNA sequence and genome, while considering increased mismatch tolerability with the increasing distance from the sgRNA “seed” region [108]. In vitro screening is still used to verify target-specificity and off-target effects. Currently, the determination of off-target effects is still being refined to give more accurate predictions, as well as the designing of sgRNAs with high targeting specificity with low off-target effects [109]. DNA site selection is vital to limit off-target events and requires the selection of a unique target sequence. Research into the rational design of sgRNA has led to the development of a variety of criteria and design algorithms [110]. Despite significant efforts in sgRNA design, experimental testing is still required, with many predicted sgRNAs showing little to no target site activity [111, 112]. Tolerability of mismatches between sgRNA and target DNA sequences has led to the development of engineered Cas9 proteins to reduce off-target effects. One such example involves the inactivation of a cleavage domain to create a Cas9 variant that can only break one DNA strand, known as Cas9 nickase (nCas9) [113]. Using a pair of the nCas9 to flank the target sequence with one binding to the forward DNA sequence and the other to the reverse sequence, means that double stranded breaks only occur were both nCas9 molecules are bound in flanking locations to each other. Thus, off-target cleavage only results in a single-strand nick, which is quickly repaired by DNA ligase. This system has shown a 50% reduction in off-target effects with a minimal decrease in on-target cleavage compared to conventional Cas9 editing [17, 114, 115].
In addition to off-target genome cleavage events, targeting CRISPR-mediated gene editing to specific cells or tissues within an organism may be required. This has led to the development of approaches that promote cell and tissue targeting for CRISPR/Cas9 delivery through modifications to nonviral vectors with targeting moieties, such as peptides, aptamers and antibodies for receptor or antigen-targeting delivery [116]. Various liposome and lipidated nanoparticles formulations have been used to allow for targeted delivery into specific tissues or cancer cells based upon the biophysical properties of the formulations. Here, we cover some recently developed targeted delivery approaches for CRISPR/Cas9 delivery; these approaches can be categorized into ligand-mediated active targeting and passive targeting. Ligand-mediated active targeting involves the use of a targeting moieties, such as peptides, antibodies, and nucleic acid aptamers, that specifically target overexpressed receptors on tumor cells, or antigens. The targeting agent serves as a “homing device” that directs the delivery system specifically to the cells of interest. This approach is highly specific and can selectively target the desired cells. Passive targeting, on the other hand, relies on the physicochemical properties of the delivery system, such as its size, charge, or lipophilicity, and pathophysiological condition of the targeting site; for example, leaky vasculature and impaired lymphatic drainage are distinctive factors of the microenvironment of solid tumors [117, 118]. In the following section a detailed overview of both approaches for CRISPR/Cas9 delivery, with specific examples, are discussed.
Active targeting
Aptamers are short single-stranded nucleic acid sequences which conform to a unique 3-dimensional structure; screening of aptamers has shown some 3-dimensional structures that can bind to specific target molecules, such as proteins or small molecules, with high affinity and specificity. Large libraries of aptamers can be rapidly synthesized and screened, are biocompatible, and readily modifiable to improve their stability under physiological conditions. They can be generated through a process called in vitro selection or systematic evolution of ligands by exponential enrichment (SELEX), which involves the iterative selection and amplification of aptamer sequences that bind to the target of interest [119]. Aptamers have widely implemented in delivery systems and have the capacity to selectively interact with a wide range of molecules, including receptors on the surface of cells, enzymes, and other biomolecules. In one study, cationic liposomes were modified by incorporating RNA aptamers to allow for the targeted delivery of the Cas9 pDNA/sgRNA system to prostate tumor cells expressing the prostate-specific membrane antigen. This aptamer-liposome-CRISPR/Cas9 system demonstrated significant cell-specific binding and was used to effectively knock out the survival gene, polo-like kinase 1 (PK1) in both in vitro and in vivo studies [120]. In another study, Zhuang et al. [121] have developed a valency-controlled tetrahedral DNA nanostructures (TDNs) conjugated to specific cancer cell specific DNA aptamers (TLS11a aptamer) and loaded it on a extracellular vessel surface via cholesterol anchoring for specific cell targeting (e.g., HepG2 cells). In vitro, in vivo, and ex vivo studies demonstrated successful delivery of CRISPR/Cas9 RNPs and significant downregulation of GFP expression and WNT10B in cultured HepG2 cells and human primary liver cancer-derived organoids, as well as xenograft tumor models.
The use of peptides as targeting ligands for non-viral delivery systems is an attractive option because of several favourable characteristics and has been studied in many gene delivery studies. Peptides can be readily synthesized and can be engineered to be conjugated with non-viral delivery vectors, demonstrate low toxicity, and have the potential to specifically target many different cells and tissue types in a target-specific manner [122]. Due to their ease of incorporation into delivery systems and cancer specific receptor-targeting capacity, they have been incorporated into non-viral delivery systems to allow for targeted delivery into cancer cells. For example, the iRGD peptide was used as a targeting ligand for liposome-templated hydrogel nanoparticles (LHNPs) to selectively deliver CRISPR/Cas9 to U87 cells (in vitro) and tumors in the brain (in vivo). The iRGD modified LHNPs demonstrated higher efficiency than the commercial agent Lipofectamine 2000 for delivering Cas9 protein/sgRNA in cell culture. When (PLK1) targeted CRISPR/Cas9 protein was delivered using iRGD modified LHNPs, inhibition of tumor growth and improved survival of tumor-bearing mouse were observed [123]. In addition to receptor targeting, peptides can also be used to target cargoes to the nucleus. Nuclear localization signals (NLS) are generally short peptides, which facilitate the nuclear transport of proteins from the cytoplasm into the nucleus [124]. A multiple-functionalized targeting delivery system was developed for efficient delivery of Cas9/sgRNA plasmids into tumors. Protamine decorated by nuclear location signal peptide (CPKKKRKV) and the nucleon targeted AS1411 aptamer with specific affinity for nucleolin in the tumor cell membrane was used to the deliver Cas9/sgRNA plasmids. With the aid of the NLS and nucleon targeted aptamer, this multi-factionalized vector specifically delivered the plasmid to the nuclei of HeLa cells and demonstrated high gene editing efficiency by knocking out the protein tyrosine kinase 2 (PTK2) gene to down-regulate focal adhesion kinase (FAK) [125].
Antibody-mediated delivery of CRISPR/Cas9 systems has recently become increasingly examined for their ability to provide high selectivity towards species that are exposed on the surface of cells. Plasmid and RNP CRISPR/Cas9 have been delivered via the use of antibodies that are directly conjugated to the CRIPSR/Cas9 RNP. In a recent study, antibodies Trastuzumab or Pertuzumab, as Fab fragments that are targeting HER2, were conjugated to the Cas9 protein to deliver Cas9/sgRNA RNPs to SKBR3 breast cancer cells, which overexpress the HER2 receptor. This Fab-conjugated Cas9/sgRNA RNP demonstrated target-specific cellular uptake and gene editing [126]. Guo et al. [127] engineered a triple-negative breast cancer (TNBC)-targeted nanolipogel system (tNLG) featuring an ICAM1 antibody that is covalently conjugated to DOPC-DSPE-PEG for the delivery of a CRISPR/Cas9 plasmid. In vivo application of this system demonstrated a greater than 81% knockout of Lipocalin 2 (Lcn2) in TNBC tumour tissues, which resulted in significant tumour growth suppression. Antibodies provide a highly-specific targeting approach with low immunogenicity concerns. Antibody orientation is required to be controlled to ensure targeting properties of the antibody are functional. This requires strict configuration and chemical functionalization protocols to be observed, limiting the development of antibody-mediated delivery of CRIPSR/Cas9 systems. An exciting approach to overcome this is the one-step synthesis of a metal-organic framework (MOFs) antibody-drug delivery system, which pairs antibodies with a metal framework that can incorporate the cargo of interest. This metal framework preserves the targeting capability of the antibody and provides a system to load the cargo without requiring synthesis consideration for antibody orientation. This holds much potential for targeted CRISPR Cas9 delivery [128].
Exosomes are extracellular lipid vesicles that are released by cells to mediate specific cell-to-cell communication [129]. Exosomes can provide a biocompatible, cell-targeted system with the potential to load various cargos for in vivo delivery [130] Cell targeting with exosomes occurs through a variety of mechanisms. A key mechanism is through the presence of certain proteins, such as tetraspanins, associated with the exosome membrane, which can interact with proteins on the target cell's surface. Additionally, exosomes also have specific lipids and nucleic acids on their surface, which can also interact with the target cell surface. Engineered cancer-derived exosomes have been successfully used to target ovarian tumors to deliver CRISPR/Cas9 pDNA resulting in the induction of apoptosis [131]. The use of exosomes holds great promise, but it is currently in its infancy. Understanding the mechanisms that allow for specific cell targeting and delivery is still broad and requires further exploration for each cell-derived and engineered exosome. Due to their cell-derived nature, the current production of exosomes is limited to small scales, and their usage for CRISPR/Cas9 delivery is an emerging area of research [27].
Passive targeting
Unique features of tumor pathophysiology have drawn significant interest as potential targets for tumor-targeted delivery of therapeutics. Tumor growth has been associated with the formation of acidic microenvironment around solid tumor with the pH range of 6.0-–6.5 [132]. Considering this, pH-responsive tumor-targeted nucleic acid delivery systems have been developed, which rely on changes in their delivery characteristics in the acidic tumor microenvironment. For example, Tang et al. [133] have developed a multistage delivery nanoparticle (MDNP) with pH-responsive tumor targeting. This system features a core–shell structure, composed of a cationic polyplex of CRISPR/Cas9 pDNA and phenylboronic acid (PBA) modified low molecular weight polyethyleneimine, while the shell is formed by the polymer 2,3-dimethylmaleic anhydride (DMMA)-modified poly(ethylene glycol)-b-polylysine (mPEG113-b-PLys100/DMMA). When exposed to an acidic microenvironment, the polymer shell DMMA groups rapidly decompose, causing the polymer to convert from an anionic to a cationic surface charge. The cationic polymer then dissociates from the cationic core, exposing the core and leading to tumor accumulation of these exposed cationic cores. Along with the cationic charge of the core, the PBA groups on the core structure bind with sialic acid, which is overexpressed in cancer cells resulting in enhanced cellular internalisation of the core structure into the cells. Systemic administration of this MDNP containing CRISPR/Cas9 (MDNP/dCas9-miR-524) to tumor-bearing mice resulted effective upregulation of miR-524 in tumor, resulting in tumor growth retardation [134].
Organ targeted or specific tissue targeted delivery of CRISPR/Cas9 system is another attractive strategy for targeted delivery of gene editing therapeutics. Several studies have utilized this tissue specific targeting strategy for CRISPR/Cas9 delivery. In many in vivo delivery, lipidated nanoparticles were found to accumulate in the liver, therefore, they have been further investigated as potential liver targeted systems for CRISPR/Cas9 delivery [135]. Studies have also focused on adapting LNPs for targeting other organ (such as kidney, spleen), with the adaption of the systems occurring via alteration of lipid compositions to modify the physicochemical properties such as size, shape, charge, or surface chemistry of the nanoparticles to allow for selectively distribution to the target organ. The interactions between a LNPs and tissue specific proteins in physiological environment are theorized to be responsible for the accumulation of LNPs in certain tissues [136]. Being inspired with this strategy, Cheng et al. developed a CRISPR/Cas9 delivery approach which exemplifies organ targeting known as Selective Organ Targeting (SORT) [137]. This SORT approach is a promising technique which uses various combinational approaches of LNPs with the addition of a optimized ratio of a cationic lipid supplemental molecules (e.g., dioleoyl-3-trimethylammonium propane) which exhibits specific biophysical properties and results in the LNPs exhibiting specificity and effectively mediated CRISPR/Cas9 delivery to specific organs including; lungs, spleen, and liver depending on ratios used [137]. A key advantage is that this system can target the numerous relevant cell types in the targeted organ and successfully deliver CRISPR/Cas9 system in vivo (e.g., epithelial cells, endothelial cells, B cells). Tissue-specific LNP delivery of the CRISPR/Cas9 system has also been used in another in vivo study to rescue dystrophin expression in Duchenne muscular dystrophy mice [138].
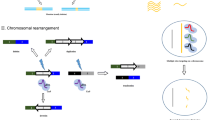
Overall, targeted delivery system for CRISPR/Cas9 system mediated genome editing is an emerging area of great interest, Fig. 3. summarizes key methods which have commonly been adopted for targeted delivery of the CRISPR/Cas9 system in many studies.
Summary of methods for targeted CRISPR/Cas9 delivery. A Cargo loading shows different forms of the CRISPR/Cas9 system which can be loading into non-targeted delivery vectors, with the examples being liposomes, polymer nanoparticles, and peptide. B Modifications for targeting is achieved through attaching with a targeting moity (such as a peptide, antibody, or aptamer) that can selectively bind to a receptor/cell surface protein/sugar and allow for active targeted cellular delivery. Modification with tissue specific promoter can direct a delivery system to a cell of interest. C Passive targeting can be divided into tumor and organ targeting. Tumor targeting occurs via the use of confirmational changes of the delivery systems under acidic environment of tumor micro-environment. Organ targeting uses a lipid nanoparticle approach in using the biophysical (siz, and charge) properties of the formulations resulting in accumulation in a specific organ, e.g., liver, lung, and spleen depending on formulation used, leading to targeted delivery
Translation to the clinic
Although the development of CRISPR/Cas9-associated therapeutics have progressed rapidly over the past decade, these techniques have not been widely investigated in human clinical trials due to various issues (e.g., difficulties with tissue-targeted delivery and the potential for off-target gene editing). Currently, more than 30 clinical trials are listed in the NIH clinicaltrials.gov register [20], which indicate the use of CRISPR/Cas9 system as a therapeutic intervention, with the number of active clinical trials slowly increasing each year. These trials include treatments for various genetic disorders, including cancers, disorders of the eye, metabolic diseases, and infections [20]. For specific examples, readers are directed to Table 2. Many of these trials are in the early stages, and thus the possibility of any of these approaches proving successful in phase III trials, and being approved by drug regulatory bodies, will still be several years away.
The clinical translation of CRISPR/Cas9 gene editing technologies relies on the careful selection of each component (e.g., the Cas9 sequence/enzyme used; the type of guide RNA used, and whether this will be a single sequence or a pool; whether a protein, mRNA, or DNA-based approach will be used; and what type of delivery approach will be used) to maximize tissue or cell-specific delivery, uptake and gene editing, and to minimize the likelihood of off-target gene editing [144]. As can be seen in Table 2, the vast majority of CRISPR/Cas9 human clinical trials have used ex vivo approaches, where specific cells are removed from a patient, enriched, subjected to gene editing, and then reintroduced into the patient. This process means that more complex tissue/cell-targeted delivery approaches (such as those needed for in vivo approaches) are not required, potentially increasing the gene editing efficiency and reducing the potential for toxicity, as only the enriched cells are subjected to the gene editing approach. The ex vivo approaches mostly used stem cells and immune-responsive cells. For instance, the program cell death (PD-1) protein, a membrane of immune cells, is targeted and knocked out by CRISPR/Cas9 technology for treating several tumors and cancer cells. PD-1 downregulates T cell activity and suppresses the immune system's antitumor activity. Clinical trials are investigating the potential of ex vivo CRISPR/Cas9-mediated PD1 knockout technique in humans for different solid tumors, liver cancer ((NCT04417764), lung cancer (NCT02793856), and prostate cancer (NCT03525652) [139]. However, ex vivo approaches are only applicable where the cells that are being targeted for gene editing can be extracted from the patient, which is not always possible. Therefore, in vivo approaches are still required despite their increased complexity. The application of in vivo gene editing therapeutic delivery is gradually stepping forward to clinical translation. NIH Clinical trial has listed a few in vivo applications of the CRISPR/Cas9 mediated therapeutic delivery (Examples in Table 2). NTLA-2001 is currently in phase I clinical trial that is used or treating transthyretin amyloidosis. Intravenous administration of NTLA-2001 composed of lipid nanoparticle encapsulating messenger RNA for Cas9 protein and a single guide RNA targeting T transthyretin (TTR) resulted in knocking out of TTR protein and thus led to decreased concentration (> 50% depending on dose concentration) of TTR protein in serum with few adverse effect [56]. Compared to ex vivo application, in vivo delivery of gene editing therapeutics is more challenging due to the concern of tissue specificity, selectivity, off-targeting, and safety; thus, robust optimisation and strategical application are essential. Progressively these clinical trials will establish the therapeutic potential for these ongoing gene engineered therapeutics and will also provide insight for future research on genome editing tool in a wide range of genetic disorders.
Conclusion
The CRISPR/Cas9 system has been heralded as revolutionary for the treatment of genetic disorders. This system has allowed for more targeted and accessible gene editing than ever before. In vivo therapeutic use of this system is limited by the lack of specific and effective delivery systems. Viral vectors have shown vast potential for the delivery of CRISPR/Cas9. However, the immunogenicity and limited ability to carrying large genetic cargoes has reduced their applications. Alternatively, nonviral delivery systems provide promising stability and delivery efficiency with in vitro, ex vivo, and in vitro applications, and each of them comes with advantages and disadvantages. Most delivery systems are based on nanoparticles, which could be merged to gather the benefits and reduce the drawbacks. The emergence of targeted delivery systems primarily depends on DNA aptamer or peptide ligands to facilitate cell-specific delivery. These emerging systems still require further research to establish efficacy and avoid cytotoxicity or considerations. The reduction of genomic off-targeting has been a critical issue in implementation of gene editing therapeutics. Improved rational design of sgRNA, truncation of sgRNA sequence and determination of Cas variant have been proven as simple and effective ways to mitigate the risk of off-targeting events. Overall, there has been exciting progress in CRISPR/Cas9-based gene editing therapy, which has translated several therapeutics into clinical trials.
However, continuous developments are required for the mode of delivery and tailoring of the systems as most therapeutic deliveries are still based on ex vivo approaches and in vivo applications stills require more optimisation in this area. Tissue-specific and selective organ-targeted deliveries are the breakthroughs that have been advent in recent years. Inorganic nanoparticles like carbon nanotubes and bare mesoporous silica nanoparticles are also emerging as exciting platforms for CRISPR/Cas9 delivery.
Data availability
Data generation and sharing are not applicable to this article as no datasets were generated or analyzed during the current study.
References
Pickar-Oliver A, Gersbach CA. The next generation of CRISPR-Cas technologies and applications. Nat Rev Mol Cell Biol. 2019;20:490–507. https://doi.org/10.1038/s41580-019-0131-5.
Genetic Alliance. District of Columbia Department of Health Understanding genetics: a district of Columbia guide for patients and health professionals, in: Genetic Alliance Monographs and Guides. Washington (DC): Genetic Alliance; 2010.
Kotagama OW, Jayasinghe CD, Abeysinghe T. Era of genomic medicine: a narrative review on CRISPR technology as a potential therapeutic tool for human diseases. Biomed Res Int. 2019;2019:1369682. https://doi.org/10.1155/2019/1369682.
Gillet J-P, Macadangdang B, Fathke RL, Gottesman MM, Kimchi-Sarfaty C. The development of gene therapy: from monogenic recessive disorders to complex diseases such as cancer. In: Gene Therapy of Cancer. 2009.
Gupta RM, Musunuru K. Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J Clin Invest. 2014;124:4154–61. https://doi.org/10.1172/JCI72992.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21. https://doi.org/10.1126/science.1225829.
Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565–75. https://doi.org/10.1046/j.1365-2958.2002.02839.x.
Jiang F, Doudna JA. CRISPR-Cas9 structures and mechanisms. Annu Rev Biophys. 2017;46:505–29. https://doi.org/10.1146/annurev-biophys-062215-010822.
Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature. 2014;507:62–7. https://doi.org/10.1038/nature13011.
Esvelt KM, Mali P, Braff JL, Moosburner M, Yaung SJ, Church GM. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat Methods. 2013;10:1116–21. https://doi.org/10.1038/nmeth.2681.
Szczelkun MD, Tikhomirova MS, Sinkunas T, Gasiunas G, Karvelis T, Pschera P, Siksnys V, Seidel R. Direct observation of R-loop formation by single RNA-guided Cas9 and Cascade effector complexes. Proc Natl Acad Sci U S A. 2014;111:9798–803. https://doi.org/10.1073/pnas.1402597111.
Jinek M, Jiang F, Taylor DW, Sternberg SH, Kaya E, Ma E, Anders C, Hauer M, Zhou K, Lin S, Kaplan M, Iavarone AT, Charpentier E, Nogales E, Doudna JA. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science. 2014;343:1247997. https://doi.org/10.1126/science.1247997.
Greely HT. CRISPR’d babies: human germline genome editing in the “He Jiankui affair.” J Law Biosci. 2019;6:111–83. https://doi.org/10.1093/jlb/lsz010.
Krimsky S. Ten ways in which He Jiankui violated ethics. Nat Biotechnol. 2019;37:19–20. https://doi.org/10.1038/nbt.4337.
Naeem M, Majeed S, Hoque MZ, Ahmad I. Latest developed strategies to minimize the off-target effects in CRISPR-Cas-mediated genome editing. Cells. 2020. https://doi.org/10.3390/cells9071608.
Tycko J, Myer VE, Hsu PD. Methods for optimizing CRISPR-Cas9 genome editing specificity. Mol Cell. 2016;63:355–70. https://doi.org/10.1016/j.molcel.2016.07.004.
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S, Kim JS. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 2014;24:132–41. https://doi.org/10.1101/gr.162339.113.
Liu Q. World-first phase I clinical trial for CRISPR-Cas9 PD-1-edited T-cells in advanced nonsmall cell lung cancer. Glob Med Genet. 2020;7:73–4. https://doi.org/10.1055/s-0040-1721451.
Cyranoski D. Chinese scientists to pioneer first human CRISPR trial. Nature. 2016;535:476–7. https://doi.org/10.1038/nature.2016.20302.
U.S. National Library of Medicine. Clinicaltrials.gov. 2022. https://clinicaltrials.gov/.
Lino CA, Harper JC, Carney JP, Timlin JA. Delivering CRISPR: a review of the challenges and approaches. Drug Deliv. 2018;25:1234–57. https://doi.org/10.1080/10717544.2018.1474964.
Angelov B, Angelova A, Filippov SK, Narayanan T, Drechsler M, Štěpánek P, Couvreur P, Lesieur S. DNA/Fusogenic Lipid Nanocarrier Assembly: Millisecond Structural Dynamics. J Phys Chem Lett. 2013;4:1959–64. https://doi.org/10.1021/jz400857z.
Angelov B, Garamus VM, Drechsler M, Angelova A. Structural analysis of nanoparticulate carriers for encapsulation of macromolecular drugs. J Mol Liq. 2017;235:83–9. https://doi.org/10.1016/j.molliq.2016.11.064.
Mukherjee D, Hasan MN, Ghosh R, Ghosh G, Bera A, Prasad SE, Hiwale A, Vemula PK, Das R, Pal SK. Decoding the Kinetic Pathways toward a Lipid/DNA Complex of Alkyl Alcohol Cationic Lipids Formed in a Microfluidic Channel. J Phys Chem B. 2022;126:588–600. https://doi.org/10.1021/acs.jpcb.1c07263.
Qiu M, Glass Z, Xu Q. Nonviral nanoparticles for CRISPR-based genome editing: Is it just a simple adaption of what have been developed for nucleic acid delivery? Biomacromol. 2019;20:3333–9. https://doi.org/10.1021/acs.biomac.9b00783.
Xu Y, Li Z. CRISPR-Cas systems: Overview, innovations and applications in human disease research and gene therapy. Comput Struct Biotechnol J. 2020;18:2401–15. https://doi.org/10.1016/j.csbj.2020.08.031.
Duan L, Ouyang K, Xu X, Xu L, Wen C, Zhou X, Qin Z, Xu Z, Sun W, Liang Y. Nanoparticle delivery of CRISPR/Cas9 for genome editing. Front Genet. 2021;12:673286. https://doi.org/10.3389/fgene.2021.673286.
Kim S, Kim D, Cho SW, Kim J, Kim JS. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 2014;24:1012–9. https://doi.org/10.1101/gr.171322.113.
Glass Z, Lee M, Li Y, Xu Q. Engineering the delivery system for CRISPR-based genome editing. Trends Biotechnol. 2018;36:173–85. https://doi.org/10.1016/j.tibtech.2017.11.006.
Jiang C, Mei M, Li B, Zhu X, Zu W, Tian Y, Wang Q, Guo Y, Dong Y, Tan X. A non-viral CRISPR/Cas9 delivery system for therapeutically targeting HBV DNA and pcsk9 in vivo. Cell Res. 2017;27:440–3. https://doi.org/10.1038/cr.2017.16.
DeWitt MA, Corn JE, Carroll D. Genome editing via delivery of Cas9 ribonucleoprotein. Methods. 2017;121–122:9–15. https://doi.org/10.1016/j.ymeth.2017.04.003.
Tang X, Wang Z, Zhang Y, Mu W, Han X. Non-viral nanocarriers for CRISPR-Cas9 gene editing system delivery. Chem Eng J. 2022;435:135116. https://doi.org/10.1016/j.cej.2022.135116.
Yip BH. Recent advances in CRISPR/Cas9 delivery strategies. Biomolecules. 2020;10:839. https://doi.org/10.3390/biom10060839.
Robbins PD, Ghivizzani SC. Viral vectors for gene therapy. Pharmacol Ther. 1998;80:35–47. https://doi.org/10.1016/s0163-7258(98)00020-5.
Asmamaw Mengstie M. Viral vectors for the in vivo delivery of CRISPR components: Advances and challenges. Front Bioeng Biotechnol. 2022. https://doi.org/10.3389/fbioe.2022.895713.
David RM, Doherty AT. Viral vectors: The road to reducing genotoxicity. Toxicol Sci. 2017;155:315–25. https://doi.org/10.1093/toxsci/kfw220.
Lostalé-Seijo I, Montenegro J. Synthetic materials at the forefront of gene delivery. Nat Rev Chem. 2018;2:258–77. https://doi.org/10.1038/s41570-018-0039-1.
Li L, Hu S, Chen X. Non-viral delivery systems for CRISPR/Cas9-based genome editing: Challenges and opportunities. Biomaterials. 2018;171:207–18. https://doi.org/10.1016/j.biomaterials.2018.04.031.
Young JL, Dean DA. Electroporation-mediated gene delivery. Adv Genet. 2015;89:49–88. https://doi.org/10.1016/bs.adgen.2014.10.003.
Frangoul H, Altshuler D, Cappellini MD, Chen YS, Domm J, Eustace BK, Foell J, de la Fuente J, Grupp S, Handgretinger R, Ho TW, Kattamis A, Kernytsky A, Lekstrom-Himes J, Li AM, Locatelli F, Mapara MY, de Montalembert M, Rondelli D, Sharma A, Sheth S, Soni S, Steinberg MH, Wall D, Yen A, Corbacioglu S. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. N Engl J Med. 2021;384:252–60. https://doi.org/10.1056/NEJMoa2031054.
Lesueur LL, Mir LM, Andre FM. Overcoming the specific toxicity of large plasmids electrotransfer in primary cells in vitro. Mol Ther Nucleic Acids. 2016;5:e291. https://doi.org/10.1038/mtna.2016.4.
Ma Y, Shen B, Zhang X, Lu Y, Chen W, et al. Heritable multiplex genetic engineering in rats using CRISPR/Cas9. PLoS ONE. 2014;9(3):e89413. https://doi.org/10.1371/journal.pone.0089413
Crispo M, Mulet AP, Tesson L, Barrera N, Cuadro F, dos Santos-Neto PC, Nguyen TH, Crénéguy A, Brusselle L, Anegón I, Menchaca A. Efficient generation of myostatin knock-out sheep using CRISPR/Cas9 technology and microinjection into zygotes. PLoS ONE. 2015;10:e0136690. https://doi.org/10.1371/journal.pone.0136690.
Zhen S, Hua L, Liu YH, Gao LC, Fu J, Wan DY, Dong LH, Song HF, Gao X. Harnessing the clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated Cas9 system to disrupt the hepatitis B virus. Gene Ther. 2015;22:404–12. https://doi.org/10.1038/gt.2015.2.
Bonamassa B, Hai L, Liu D. Hydrodynamic gene delivery and its applications in pharmaceutical research. Pharm Res. 2011;28:694–701. https://doi.org/10.1007/s11095-010-0338-9.
Khan I, Saeed K, Khan I. Nanoparticles: properties, applications and toxicities. Arab J Chem. 2019;12:908–31. https://doi.org/10.1016/j.arabjc.2017.05.011.
Evers MJW, Kulkarni JA, van der Meel R, Cullis PR, Vader P, Schiffelers RM. State-of-the-art design and rapid-mixing production techniques of lipid nanoparticles for nucleic acid delivery. Small Methods. 2018. https://doi.org/10.1002/smtd.201700375.
Allen TM, Cullis PR. Liposomal drug delivery systems: From concept to clinical applications. Adv Drug Deliv Rev. 2013;65:36–48. https://doi.org/10.1016/j.addr.2012.09.037.
Kazemian P, Yu SY, Thomson SB, Birkenshaw A, Leavitt BR, Ross CJD. Lipid-nanoparticle-based delivery of CRISPR/Cas9 genome-editing components. Mol Pharm. 2022;19:1669–86. https://doi.org/10.1021/acs.molpharmaceut.1c00916.
Felgner PL, Gadek TR, Holm M, Roman R, Chan HW, Wenz M, Northrop JP, Ringold GM, Danielsen M. Lipofection: a highly efficient, lipid-mediated DNA-transfection procedure. Proc Natl Acad Sci U S A. 1987;84:7413–7. https://doi.org/10.1073/pnas.84.21.7413.
Tassler S, Dobner B, Lampp L, Ziolkowski R, Malinowska E, Wolk C, Brezesinski G. DNA delivery systems based on peptide-mimicking cationic lipids-the effect of the co-lipid on the structure and DNA binding capacity. Langmuir. 2019;35:4613–25. https://doi.org/10.1021/acs.langmuir.8b04139.
Wang M, Zuris JA, Meng F, Rees H, Sun S, Deng P, Han Y, Gao X, Pouli D, Wu Q, Georgakoudi I, Liu DR, Xu Q. Efficient delivery of genome-editing proteins using bioreducible lipid nanoparticles. Proc Natl Acad Sci U S A. 2016;113:2868–73. https://doi.org/10.1073/pnas.1520244113.
Dalby B, Cates S, Harris A, Ohki EC, Tilkins ML, Price PJ, Ciccarone VC. Advanced transfection with Lipofectamine 2000 reagent: primary neurons, siRNA, and high-throughput applications. Methods. 2004;33:95–103. https://doi.org/10.1016/j.ymeth.2003.11.023.
Zhao M, Yang H, Jiang X, Zhou W, Zhu B, Zeng Y, Yao K, Ren C. Lipofectamine RNAiMAX: an efficient siRNA transfection reagent in human embryonic stem cells. Mol Biotechnol. 2008;40:19–26. https://doi.org/10.1007/s12033-008-9043-x.
Yu X, Liang X, Xie H, Kumar S, Ravinder N, Potter J. de Mollerat du Jeu X, Chesnut JD, Improved delivery of Cas9 protein/gRNA complexes using lipofectamine CRISPRMAX. Biotechnol Lett. 2016;38:919–29. https://doi.org/10.1007/s10529-016-2064-9.
Gillmore JD, Gane E, Taubel J, Kao J, Fontana M, Maitland ML, Seitzer J, O’Connell D, Walsh KR, Wood K, Phillips J, Xu Y, Amaral A, Boyd AP, Cehelsky JE, McKee MD, Schiermeier A, Harari O, Murphy A, Kyratsous CA, Zambrowicz B, Soltys R, Gutstein DE, Leonard J, Sepp-Lorenzino L, Lebwohl D. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N Engl J Med. 2021;385:493–502. https://doi.org/10.1056/NEJMoa2107454.
Liang X, Potter J, Kumar S, Zou Y, Quintanilla R, Sridharan M, Carte J, Chen W, Roark N, Ranganathan S, Ravinder N, Chesnut JD. Rapid and highly efficient mammalian cell engineering via Cas9 protein transfection. J Biotechnol. 2015;208:44–53. https://doi.org/10.1016/j.jbiotec.2015.04.024.
Zhang L, Wang P, Feng Q, Wang N, Chen Z, Huang Y, Zheng W, Jiang X. Lipid nanoparticle-mediated efficient delivery of CRISPR/Cas9 for tumor therapy. NPG Asia Mater. 2017;9:e441. https://doi.org/10.1038/am.2017.185.
Jhaveri AM, Torchilin VP. Multifunctional polymeric micelles for delivery of drugs and siRNA. Front Pharmacol. 2014;5:77. https://doi.org/10.3389/fphar.2014.00077.
Mariano A, Lubrano C, Bruno U, Ausilio C, Dinger NB, Santoro F. Advances in cell-conductive polymer biointerfaces and role of the plasma membrane. Chem Rev. 2022;122:4552–80. https://doi.org/10.1021/acs.chemrev.1c00363.
Fox LJ, Richardson RM, Briscoe WH. PAMAM dendrimer - cell membrane interactions. Adv Colloid Interface Sci. 2018;257:1–18. https://doi.org/10.1016/j.cis.2018.06.005.
Duan L, Ouyang K, Xu X, Xu L, Wen C, Zhou X, Qin Z, Xu Z, Sun W, Liang Y. Nanoparticle delivery of CRISPR/Cas9 for genome editing. Front Gen. 2021. https://doi.org/10.3389/fgene.2021.673286.
Hall A, Lachelt U, Bartek J, Wagner E, Moghimi SM. Polyplex evolution: understanding biology, optimizing performance. Mol Ther. 2017;25:1476–90. https://doi.org/10.1016/j.ymthe.2017.01.024.
Benjaminsen RV, Mattebjerg MA, Henriksen JR, Moghimi SM, Andresen TL. The possible "proton sponge " effect of polyethylenimine (PEI) does not include change in lysosomal pH. Mol Ther. 2013;21:149–57. https://doi.org/10.1038/mt.2012.185.
Ryu N, Kim MA, Park D, Lee B, Kim YR, Kim KH, Baek JI, Kim WJ, Lee KY, Kim UK. Effective PEI-mediated delivery of CRISPR-Cas9 complex for targeted gene therapy. Nanomedicine. 2018;14:2095–102. https://doi.org/10.1016/j.nano.2018.06.009.
Fahira AI, Amalia R, Barliana MI, Gatera VA, Abdulah R. Polyethyleneimine (PEI) as a polymer-based co-delivery system for breast cancer therapy. Breast Cancer (Dove Med Press). 2022;14:71–83. https://doi.org/10.2147/BCTT.S350403.
Zuckermann M, Hovestadt V, Knobbe-Thomsen CB, Zapatka M, Northcott PA, Schramm K, Belic J, Jones DT, Tschida B, Moriarity B, Largaespada D, Roussel MF, Korshunov A, Reifenberger G, Pfister SM, Lichter P, Kawauchi D, Gronych J. Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling. Nat Commun. 2015;6:7391. https://doi.org/10.1038/ncomms8391.
O’Keeffe Ahern J, Lara-Saez I, Zhou D, Murillas R, Bonafont J, Mencia A, Garcia M, Manzanares D, Lynch J, Foley R, Xu Q, Sigen A, Larcher F, Wang W. Non-viral delivery of CRISPR-Cas9 complexes for targeted gene editing via a polymer delivery system. Gene Ther. 2022;29:157–70. https://doi.org/10.1038/s41434-021-00282-6.
Reddy MSB, Ponnamma D, Choudhary R, Sadasivuni KK. A comparative review of natural and synthetic biopolymer composite scaffolds. Polymers (Basel). 2021. https://doi.org/10.3390/polym13071105.
Suk JS, Xu Q, Kim N, Hanes J, Ensign LM. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv Drug Deliv Rev. 2016;99:28–51. https://doi.org/10.1016/j.addr.2015.09.012.
Zhen S, Li X. Liposomal delivery of CRISPR/Cas9. Cancer Gene Ther. 2020;27:515–27. https://doi.org/10.1038/s41417-019-0141-7.
Li SD, Huang L. Nanoparticles evading the reticuloendothelial system: role of the supported bilayer. Biochim Biophys Acta. 2009;1788:2259–66. https://doi.org/10.1016/j.bbamem.2009.06.022.
Finn JD, Smith AR, Patel MC, Shaw L, Youniss MR, van Heteren J, Dirstine T, Ciullo C, Lescarbeau R, Seitzer J, Shah RR, Shah A, Ling D, Growe J, Pink M, Rohde E, Wood KM, Salomon WE, Harrington WF, Dombrowski C, Strapps WR, Chang Y, Morrissey DV. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep. 2018;22:2227–35. https://doi.org/10.1016/j.celrep.2018.02.014.
Cabanillas B, Novak N, Akdis CA. The form of PEG matters: PEG conjugated with lipids and not PEG alone could be the specific form involved in allergic reactions to COVID-19 vaccines. Allergy. 2022;77:1658–60. https://doi.org/10.1111/all.15187.
Zhang H, Bahamondez-Canas TF, Zhang Y, Leal J, Smyth HDC. PEGylated chitosan for nonviral aerosol and mucosal delivery of the CRISPR/Cas9 system in vitro. Mol Pharm. 2018;15:4814–26. https://doi.org/10.1021/acs.molpharmaceut.8b00434.
Lostale-Seijo I, Louzao I, Juanes M, Montenegro J. Peptide nanostructures for ribonucleoprotein cell membrane transport and gene edition. Chem Sci. 2017;8:7923–31. https://doi.org/10.1039/c7sc03918b.
Dasari BC, Cashman SM, Kumar-Singh R. Reducible PEG-POD/DNA nanoparticles for gene transfer in vitro and in vivo: Application in a mouse model of age-related macular degeneration. Mol Ther Nucleic Acids. 2017;8:77–89. https://doi.org/10.1016/j.omtn.2017.06.004.
Gustafsson O, Rädler J, Roudi S, Lehto T, Hällbrink M, Lehto T, Gupta D, Andaloussi SE, Nordin JZ. Efficient peptide-mediated in vitro delivery of Cas9 RNP. Pharmaceutics. 2021;13:878.
Del’Guidice T, Lepetit-Stoffaes J-P, Bordeleau L-J, Roberge J, Théberge V, Lauvaux C, Barbeau X, Trottier J, Dave V, Roy D-C, Gaillet B, Garnier A, Guay D. Membrane permeabilizing amphiphilic peptide delivers recombinant transcription factor and CRISPR-Cas9/Cpf1 ribonucleoproteins in hard-to-modify cells. PLoS ONE. 2018;13:e0195558. https://doi.org/10.1371/journal.pone.0195558.
Wang HX, Song Z, Lao YH, Xu X, Gong J, Cheng D, Chakraborty S, Park JS, Li M, Huang D, Yin L, Cheng J, Leong KW. Nonviral gene editing via CRISPR/Cas9 delivery by membrane-disruptive and endosomolytic helical polypeptide. Proc Natl Acad Sci U S A. 2018;115:4903–8. https://doi.org/10.1073/pnas.1712963115.
Zamolo SJ, Darbre T, Reymond J-L. Transfecting tissue models with CRISPR/Cas9 plasmid DNA using peptide dendrimers. Chem Commun. 2020;56:11981–4. https://doi.org/10.1039/D0CC04750C.
Dunbar T, Tsakirpaloglou N, Septiningsih EM, Thomson MJ. Carbon nanotube-mediated plasmid DNA delivery in rice leaves and seeds. Int J Mol Sci. 2022. https://doi.org/10.3390/ijms23084081.
Garcia-Fernandez A, Vivo-Llorca G, Sancho M, Garcia-Jareno AB, Ramirez-Jimenez L, Barber-Cano E, Murguia JR, Orzaez M, Sancenon F, Martinez-Manez R. Nanodevices for the efficient codelivery of CRISPR-Cas9 editing machinery and an entrapped cargo: a proposal for dual anti-inflammatory therapy. Pharmaceutics. 2022. https://doi.org/10.3390/pharmaceutics14071495.
Lee K, Conboy M, Park HM, Jiang F, Kim HJ, Dewitt MA, Mackley VA, Chang K, Rao A, Skinner C, Shobha T, Mehdipour M, Liu H, Huang WC, Lan F, Bray NL, Li S, Corn JE, Kataoka K, Doudna JA, Conboy I, Murthy N. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat Biomed Eng. 2017;1:889–901. https://doi.org/10.1038/s41551-017-0137-2.
Yigit MV, Medarova Z. In vivo and ex vivo applications of gold nanoparticles for biomedical SERS imagingi. Am J Nucl Med Mol Imaging. 2012;2:232–41.
Ho LWC, Chan CKW, Han R, Lau YFY, Li H, Ho YP, Zhuang X, Choi CHJ. Mammalian cells exocytose alkylated gold nanoparticles via extracellular vesicles. ACS Nano. 2022;16:2032–45. https://doi.org/10.1021/acsnano.1c07418.
Wang SH, Lee CW, Chiou A, Wei PK. Size-dependent endocytosis of gold nanoparticles studied by three-dimensional mapping of plasmonic scattering images. J Nanobiotechnology. 2010;8:33. https://doi.org/10.1186/1477-3155-8-33.
Shahbazi R, Sghia-Hughes G, Reid JL, Kubek S, Haworth KG, Humbert O, Kiem HP, Adair JE. Targeted homology-directed repair in blood stem and progenitor cells with CRISPR nanoformulations. Nat Mater. 2019;18:1124–32. https://doi.org/10.1038/s41563-019-0385-5.
Wang P, Zhang L, Zheng W, Cong L, Guo Z, Xie Y, Wang L, Tang R, Feng Q, Hamada Y, Gonda K, Hu Z, Wu X, Jiang X. Thermo-triggered release of CRISPR-Cas9 system by lipid-encapsulated gold nanoparticles for tumor therapy. Angew Chem Int Ed Engl. 2018;57:1491–6. https://doi.org/10.1002/anie.201708689.
Wang Y, Shahi PK, Wang X, Xie R, Zhao Y, Wu M, Roge S, Pattnaik BR, Gong S. In vivo targeted delivery of nucleic acids and CRISPR genome editors enabled by GSH-responsive silica nanoparticles. J Control Release. 2021;336:296–309. https://doi.org/10.1016/j.jconrel.2021.06.030.
Alyami MZ, Alsaiari SK, Li Y, Qutub SS, Aleisa FA, Sougrat R, Merzaban JS, Khashab NM. Cell-type-specific CRISPR/Cas9 delivery by biomimetic metal organic frameworks. J Am Chem Soc. 2020;142:1715–20. https://doi.org/10.1021/jacs.9b11638.
Xie H, Liu X, Huang Z, Xu L, Bai R, He F, Wang M, Han L, Bao Z, Wu Y, Xie C, Gong Y. Nanoscale zeolitic imidazolate framework (ZIF)-8 in cancer theranostics: current challenges and prospects. Cancers (Basel). 2022. https://doi.org/10.3390/cancers14163935.
Alsaiari SK, Patil S, Alyami M, Alamoudi KO, Aleisa FA, Merzaban JS, Li M, Khashab NM. Endosomal escape and delivery of CRISPR/Cas9 genome editing machinery enabled by nanoscale zeolitic imidazolate framework. J Am Chem Soc. 2018;140:143–6. https://doi.org/10.1021/jacs.7b11754.
Raveux A, Vandormael-Pournin S, Cohen-Tannoudji M. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote. Sci Rep. 2017;7:42661. https://doi.org/10.1038/srep42661.
Kim YS, Kim GR, Park M, Yang SC, Park SH, Won JE, Lee JH, Shin HE, Song H, Kim HR. Electroporation of AsCpf1/RNP at the zygote stage is an efficient genome editing method to generate knock-out mice deficient in leukemia inhibitory factor. Tissue Eng Regen Med. 2020;17:45–53. https://doi.org/10.1007/s13770-019-00225-8.
Niola F, Dagnæs-Hansen F, Frödin M. In Vivo Editing of the Adult Mouse Liver Using CRISPR/Cas9 and Hydrodynamic Tail Vein Injection. Methods Mol Biol. 2019;1961:329–41. https://doi.org/10.1007/978-1-4939-9170-9_20.
Guan Y, Ma Y, Li Q, Sun Z, Ma L, Wu L, Wang L, Zeng L, Shao Y, Chen Y, Ma N, Lu W, Hu K, Han H, Yu Y, Huang Y, Liu M, Li D. CRISPR/Cas9-mediated somatic correction of a novel coagulator factor IX gene mutation ameliorates hemophilia in mouse. EMBO Mol Med. 2016;8:477–88. https://doi.org/10.15252/emmm.201506039.
Wei T, Cheng Q, Min Y-L, Olson EN, Siegwart DJ. Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing. Nat Commun. 2020;11:3232. https://doi.org/10.1038/s41467-020-17029-3.
Han JP, Kim M, Choi BS, Lee JH, Lee GS, Jeong M, Lee Y, Kim E-A, Oh H-K, Go N, Lee H, Lee KJ, Kim UG, Lee JY, Kim S, Chang J, Lee H, Song DW, Yeom SC. In vivo delivery of CRISPR-Cas9 using lipid nanoparticles enables antithrombin gene editing for sustainable hemophilia A and B therapy. Sci Adv. 2022;8:eabj6901. https://doi.org/10.1126/sciadv.abj6901.
Wang P, Zhang L, Xie Y, Wang N, Tang R, Zheng W, Jiang X. Genome editing for cancer therapy: delivery of Cas9 protein/sgRNA plasmid via a gold nanocluster/lipid core-shell nanocarrier. Adv Sci (Weinh). 2017;4:1700175. https://doi.org/10.1002/advs.201700175.
Zhang L, Wang L, Xie Y, Wang P, Deng S, Qin A, Zhang J, Yu X, Zheng W, Jiang X. Triple-targeting delivery of CRISPR/Cas9 to reduce the risk of cardiovascular diseases. Angew Chem Int Ed. 2019;58:12404–8. https://doi.org/10.1002/anie.201903618.
Xu X, Koivisto O, Liu C, Zhou J, Miihkinen M, Jacquemet G, Wang D, Rosenholm JM, Shu Y, Zhang H. Effective delivery of the CRISPR/Cas9 system enabled by functionalized mesoporous silica nanoparticles for GFP-tagged paxillin knock-in. Adv Ther. 2021;4:2000072. https://doi.org/10.1002/adtp.202000072.
Yang X, Tang Q, Jiang Y, Zhang M, Wang M, Mao L. Nanoscale ATP-responsive zeolitic imidazole framework-90 as a general platform for cytosolic protein delivery and genome editing. J Am Chem Soc. 2019;141:3782–6. https://doi.org/10.1021/jacs.8b11996.
Uddin F, Rudin CM, Sen T. CRISPR gene therapy: applications, limitations, and implications for the future. Front Oncol. 2020;10:1387. https://doi.org/10.3389/fonc.2020.01387.
Zhang XH, Tee LY, Wang XG, Huang QS, Yang SH. Off-target effects in CRISPR/Cas9-mediated genome engineering. Mol Ther Nucleic Acids. 2015;4:e264. https://doi.org/10.1038/mtna.2015.37.
Atkins A, Chung CH, Allen AG, Dampier W, Gurrola TE, Sariyer IK, Nonnemacher MR, Wigdahl B. Off-target analysis in gene editing and applications for clinical translation of CRISPR/Cas9 in HIV-1 therapy. Front Genome Ed. 2021;3:673022. https://doi.org/10.3389/fgeed.2021.673022.
Ryan DE, Taussig D, Steinfeld I, Phadnis SM, Lunstad BD, Singh M, Vuong X, Okochi KD, McCaffrey R, Olesiak M, Roy S, Yung CW, Curry B, Sampson JR, Bruhn L, Dellinger DJ. Improving CRISPR-Cas specificity with chemical modifications in single-guide RNAs. Nucleic Acids Res. 2018;46:792–803. https://doi.org/10.1093/nar/gkx1199.
Anderson KR, Haeussler M, Watanabe C, Janakiraman V, Lund J, Modrusan Z, Stinson J, Bei Q, Buechler A, Yu C, Thamminana SR, Tam L, Sowick MA, Alcantar T, O’Neil N, Li J, Ta L, Lima L, Roose-Girma M, Rairdan X, Durinck S, Warming S. CRISPR off-target analysis in genetically engineered rats and mice. Nat Methods. 2018;15:512–4. https://doi.org/10.1038/s41592-018-0011-5.
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK, Sander JD. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol. 2013;31:822–6. https://doi.org/10.1038/nbt.2623.
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Sullender M, Ebert BL, Xavier RJ, Root DE. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol. 2014;32:1262–7. https://doi.org/10.1038/nbt.3026.
Haeussler M, Schönig K, Eckert H, Eschstruth A, Mianné J, Renaud J-B, Schneider-Maunoury S, Shkumatava A, Teboul L, Kent J, Joly J-S, Concordet J-P. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 2016;17:148. https://doi.org/10.1186/s13059-016-1012-2.
Xu H, Xiao T, Chen CH, Li W, Meyer CA, Wu Q, Wu D, Cong L, Zhang F, Liu JS, Brown M, Liu XS. Sequence determinants of improved CRISPR sgRNA design. Genome Res. 2015;25:1147–57. https://doi.org/10.1101/gr.191452.115.
Liu Y, Zhou C, Huang S, Dang L, Wei Y, He J, Zhou Y, Mao S, Tao W, Zhang Y, Yang H, Huang X, Chi T. A Cas-embedding strategy for minimizing off-target effects of DNA base editors. Nat Commun. 2020;11:6073. https://doi.org/10.1038/s41467-020-19690-0.
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 2013;154:1380–9. https://doi.org/10.1016/j.cell.2013.08.021.
Mali P, Aach J, Stranges PB, Esvelt KM, Moosburner M, Kosuri S, Yang L, Church GM. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol. 2013;31:833–8. https://doi.org/10.1038/nbt.2675.
Alghuthaymi MA, Ahmad A, Khan Z, Khan SH, Ahmed FK, Faiz S, Nepovimova E, Kuca K, Abd-Elsalam KA, Exosome/liposome-like nanoparticles: New carriers for CRISPR genome editing in plants. Int J Mol Sci. 2021;22. https://doi.org/10.3390/ijms22147456.
Begum AA, Toth I, Hussein WM, Moyle PM. Advances in targeted gene delivery. Curr Drug Deliv. 2019;16:588–608. https://doi.org/10.2174/1567201816666190529072914.
Attia MF, Anton N, Wallyn J, Omran Z, Vandamme TF. An overview of active and passive targeting strategies to improve the nanocarriers efficiency to tumour sites. J Pharm Pharmacol. 2019;71:1185–98. https://doi.org/10.1111/jphp.13098.
Zhu G, Chen X. Aptamer-based targeted therapy. Adv Drug Deliv Rev. 2018;134:65–78. https://doi.org/10.1016/j.addr.2018.08.005.
Zhen S, Takahashi Y, Narita S, Yang YC, Li X. Targeted delivery of CRISPR/Cas9 to prostate cancer by modified gRNA using a flexible aptamer-cationic liposome. Oncotarget. 2017;8:9375–87. https://doi.org/10.18632/oncotarget.14072.
Zhuang J, Tan J, Wu C, Zhang J, Liu T, Fan C, Li J, Zhang Y. Extracellular vesicles engineered with valency-controlled DNA nanostructures deliver CRISPR/Cas9 system for gene therapy. Nucleic Acids Res. 2020;48:8870–82. https://doi.org/10.1093/nar/gkaa683.
Borner K, Kienle E, Huang LY, Weinmann J, Sacher A, Bayer P, Stullein C, Fakhiri J, Zimmermann L, Westhaus A, Beneke J, Beil N, Wiedtke E, Schmelas C, Miltner D, Rau A, Erfle H, Krausslich HG, Muller M, Agbandje-McKenna M, Grimm D. Pre-arrayed Pan-AAV peptide display libraries for rapid single-round screening. Mol Ther. 2020;28:1016–32. https://doi.org/10.1016/j.ymthe.2020.02.009.
Chen Z, Liu F, Chen Y, Liu J, Wang X, Chen AT, Deng G, Zhang H, Liu J, Hong Z, Zhou J. Targeted Delivery of CRISPR/Cas9-Mediated Cancer Gene Therapy via Liposome-Templated Hydrogel Nanoparticles. Adv Funct Mater. 2017. https://doi.org/10.1002/adfm.201703036.
Lu J, Wu T, Zhang B, Liu S, Song W, Qiao J, Ruan H. Types of nuclear localization signals and mechanisms of protein import into the nucleus. Cell Commun Signal. 2021;19:60. https://doi.org/10.1186/s12964-021-00741-y.
Liu B-Y, He X-Y, Xu C, Ren X-H, Zhuo R-X, Cheng S-X. Peptide and aptamer decorated delivery system for targeting delivery of Cas9/sgRNA plasmid to mediate antitumor genome editing. ACS Appl Mater Interfaces. 2019;11:23870–9. https://doi.org/10.1021/acsami.9b05772.
Ubiparipovic S, Christ D, Rouet R. Antibody-mediated delivery of CRISPR-Cas9 ribonucleoproteins in human cells. Protein Eng Des Sel. 2022;35:011. https://doi.org/10.1093/protein/gzac011.
Guo P, Yang J, Huang J, Auguste DT, Moses MA. Therapeutic genome editing of triple-negative breast tumors using a noncationic and deformable nanolipogel. Proc Natl Acad Sci U S A. 2019;116:18295–303. https://doi.org/10.1073/pnas.1904697116.
Alt K, Carraro F, Jap E, Linares-Moreau M, Riccò R, Righetto M, Bogar M, Amenitsch H, Hashad RA, Doonan C, Hagemeyer CE, Falcaro P. Self-assembly of oriented antibody-decorated metal-organic framework nanocrystals for active-targeting applications. Adv Mater. 2022;34:2106607. https://doi.org/10.1002/adma.202106607.
Zhang Y, Liu Y, Liu H, Tang WH. Exosomes: biogenesis, biologic function and clinical potential. Cell Biosci. 2019;9:19. https://doi.org/10.1186/s13578-019-0282-2.
Choi H, Choi Y, Yim HY, Mirzaaghasi A, Yoo JK, Choi C. Biodistribution of exosomes and engineering strategies for targeted delivery of therapeutic exosomes. Tissue Eng Regen Med. 2021;18:499–511. https://doi.org/10.1007/s13770-021-00361-0.
Kim SM, Yang Y, Oh SJ, Hong Y, Seo M, Jang M. Cancer-derived exosomes as a delivery platform of CRISPR/Cas9 confer cancer cell tropism-dependent targeting. J Control Release. 2017;266:8–16. https://doi.org/10.1016/j.jconrel.2017.09.013.
Boedtkjer E, Pedersen SF. The acidic tumor microenvironment as a driver of cancer. Annu Rev Physiol. 2020;82:103–26. https://doi.org/10.1146/annurev-physiol-021119-034627.
Tang Q, Liu J, Jiang Y, Zhang M, Mao L, Wang M. Cell-selective messenger RNA delivery and CRISPR/Cas9 genome editing by modulating the interface of phenylboronic acid-derived lipid nanoparticles and cellular surface sialic acid. ACS Appl Mater Interfaces. 2019;11(50):46585–90. https://doi.org/10.1021/acsami.9b17749. Epub 2019 Dec 9 PMID: 31763806.
Liu Q, Zhao K, Wang C, Zhang Z, Zheng C, Zhao Y, Zheng Y, Liu C, An Y, Shi L, Kang C, Liu Y. Multistage delivery nanoparticle facilitates efficient CRISPR/dCas9 activation and tumor growth suppression in vivo. Adv Sci (Weinh). 2019;6:1801423. https://doi.org/10.1002/advs.201801423.
Zhang YN, Poon W, Tavares AJ, McGilvray ID, Chan WCW. Nanoparticle-liver interactions: cellular uptake and hepatobiliary elimination. J Control Release. 2016;240:332–48. https://doi.org/10.1016/j.jconrel.2016.01.020.
Dilliard SA, Cheng Q, Siegwart DJ. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc Natl Acad Sci. 2021;118:e2109256118. https://doi.org/10.1073/pnas.2109256118.
Cheng Q, Wei T, Farbiak L, Johnson LT, Dilliard SA, Siegwart DJ. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR-Cas gene editing. Nat Nanotechnol. 2020;15:313–20. https://doi.org/10.1038/s41565-020-0669-6.
Wei T, Cheng Q, Min YL, Olson EN, Siegwart DJ. Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing. Nat Commun. 2020;11(1):3232. https://doi.org/10.1038/s41467-020-17029-3.
Cheng H, Zhang F, Ding Y. CRISPR/Cas9 delivery system engineering for genome editing in therapeutic applications. Pharmaceutics. 2021. https://doi.org/10.3390/pharmaceutics13101649.
Modarai SR, Kanda S, Bloh K, Opdenaker LM, Kmiec EB. Precise and error-prone CRISPR-directed gene editing activity in human CD34+ cells varies widely among patient samples. Gene Ther. 2021;28:105–13. https://doi.org/10.1038/s41434-020-00192-z.
Hu Z, Yu L, Zhu D, Ding W, Wang X, Zhang C, Wang L, Jiang X, Shen H, He D, Li K, Xi L, Ma D, Wang H. Disruption of HPV16-E7 by CRISPR/Cas system induces apoptosis and growth inhibition in HPV16 positive human cervical cancer cells. Biomed Res Int. 2014;2014:612823. https://doi.org/10.1155/2014/612823.
Xu L, Wang J, Liu Y, Xie L, Su B, Mou D, Wang L, Liu T, Wang X, Zhang B, Zhao L, Hu L, Ning H, Zhang Y, Deng K, Liu L, Lu X, Zhang T, Xu J, Li C, Wu H, Deng H, Chen H. CRISPR-edited stem cells in a patient with HIV and acute lymphocytic leukemia. N Engl J Med. 2019;381:1240–7. https://doi.org/10.1056/NEJMoa1817426.
Bruno B, Wasch R, Engelhardt M, Gay F, Giaccone L, D’Agostino M, Rodriguez-Lobato LG, Danhof S, Gagelmann N, Kroger N, Popat R, Van de Donk N, Terpos E, Dimopoulos MA, Sonneveld P, Einsele H, Boccadoro M. European Myeloma Network perspective on CAR T-Cell therapies for multiple myeloma. Haematologica. 2021;106:2054–65. https://doi.org/10.3324/haematol.2020.276402.
Foley RA, Sims RA, Duggan EC, Olmedo JK, Ma R, Jonas SJ. Delivering the CRISPR/Cas9 system for engineering gene therapies: Recent cargo and delivery approaches for clinical translation. Front Bioeng Biotechnol. 2022. https://doi.org/10.3389/fbioe.2022.973326.
Acknowledgements
Figures were created using BioRender.com.
Funding
This research was supported by Australian Research Council (ARC Project number: DP100101271).
Author information
Authors and Affiliations
Contributions
A.A.B., F.S, and C.C.D formulated the conception and design. The first draft of the manuscript was written by F.S. and A.A.B and all authors critically revised and commented on previous versions of the manuscript. C.C.D and F.S. have prepared all the figures. A.A.B, F.S, and P.M.M has edited the revised version. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This paper is a review article without any in vitro or in vivo test.
Consent for publication
All authors agree to publish this review article to this journal.
Competing interests
All authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sinclair, F., Begum, A.A., Dai, C.C. et al. Recent advances in the delivery and applications of nonviral CRISPR/Cas9 gene editing. Drug Deliv. and Transl. Res. 13, 1500–1519 (2023). https://doi.org/10.1007/s13346-023-01320-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13346-023-01320-z