Abstract
High-energy diffraction microscopy (HEDM) combined with in situ mechanical testing is a powerful nondestructive technique for tracking the evolving microstructure within polycrystalline materials during deformation. This technique relies on a sophisticated analysis of X-ray diffraction patterns to produce a three-dimensional reconstruction of grains and other microstructural features within the interrogated volume. However, it is known that HEDM can fail to identify certain microstructural features, particularly smaller grains or twinned regions. Characterization of the identical sample volume using high-resolution surface-specific techniques, particularly electron backscatter diffraction (EBSD), can not only provide additional microstructure information about the interrogated volume but also highlight opportunities for improvement of the HEDM reconstruction algorithms. In this study, a sample fabricated from undeformed “low solvus, high refractory” nickel-based superalloy was scanned using HEDM. The volume interrogated by HEDM was then carefully characterized using a combination of surface-specific techniques, including epi-illumination optical microscopy, zero-tilt secondary and backscattered electron imaging, scanning white light interferometry, and high-precision EBSD. Custom data fusion protocols were developed to integrate and align the microstructure maps captured by these surface-specific techniques and HEDM. The raw and processed data from HEDM and serial sectioning have been made available via the Materials Data Facility (MDF) at https://doi.org/10.18126/4y0p-v604 for further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
This data descriptor article covers a 3D microstructural imaging experiment intended to push the limits of electron backscatter diffraction (EBSD) and high energy diffraction microscopy (HEDM), particularly via improved orientation resolution in 3D-EBSD and investigation of data collection and reconstruction parameters for HEDM. Previous works [1,2,3,4] have shown that HEDM, compared to EBSD, can fail to correctly identify smaller grains or twinned regions in a sample. While at least one 3D-to-3D study has been done for synchrotron-based diffraction contrast tomography [5], there is little quantitative analysis of this issue in 3D for HEDM due to a lack of “ground truth” 3D datasets for comparison. Toward this end, we collected a high-quality 3D-EBSD dataset from a volume of interest (VOI) that was initially examined using HEDM. The angular resolution of 3D-EBSD was significantly improved by employing spherical harmonic indexing and empirical pattern center fitting, while the 3D reconstruction accuracy was improved by employing several secondary imaging techniques, including epi-illumination optical microscopy, zero-tilt backscattered electron imaging, and scanning white light interferometry. Furthermore, the HEDM data from the VOI were collected with a wide range of acquisition parameters so that the impact of both collection and reconstruction parameters could be investigated vis-à-vis the high-resolution 3D-EBSD dataset. In addition, a major goal of this study was to develop and document a set of detailed data fusion protocols so that the microstructure maps produced by surface-specific and nondestructive techniques can be integrated and compared effectively. Both the 3D-EBSD and HEDM datasets have been made publicly available, allowing further investigation of best practices for collection and analysis of HEDM data.
Material and Sample
The “low solvus, high refractory” (LSHR) alloy is a NASA-developed nickel-based superalloy [6], chosen for this study as a representative example of the superalloy family with a well-established procedure to produce a coarse-grained microstructure suitable for HEDM analysis. The recrystallized LSHR sample serves as a baseline for comparison of 3D-EBSD and HEDM and allows validation of the workflow in preparation for similar studies on deformed samples of other superalloys. For details of the thermomechanical processing steps used to produce a coarse-grained, fully recrystallized microstructure, see [7]; this sample was produced from the same batch of processed LSHR alloy used in that study. The test sample was cut from the raw material using wire electrical discharge machining (EDM) as a millimeter-scale tensile sample with a 1\(\times \)1 mm square cross section throughout its 8 mm gauge length. No further machining, surface processing, or heat treatment was done following EDM. Figure 1 shows the sample geometry. Note that despite the geometry being chosen to match that of an in situ tensile testing sample, this particular sample was never deliberately loaded.
Sample Marking
Similarly to previous AFRL studies [2, 8], a gold cube marker was attached to the sample surface prior to HEDM measurements to indicate the location of the VOI. In this study only a single such marker was used, as the height of the VOI along the scanning/sectioning axis was only 40 \(\upmu \)mFootnote 1 and we did not conduct any in situ loading that would elongate the gauge section, rendering additional such markers unnecessary. However, to aid in registering the HEDM and 3D-EBSD data, a focused ion beam (FIB) was used to deposit platinum marker lines on all four faces of the sample prior to HEDM scanning.Footnote 2 Fig. 2a and 2b shows scanning electron microscope (SEM) images of these markers on the sample surface. While only one set of lines was necessary to cover the full height of the VOI, an additional set was deposited on the face with the gold cube so that material removal could be monitored more precisely during serial sectioning when approaching the VOI. The high atomic number of platinum was found to give the deposited material sufficient contrast to be visible in X-ray computed tomography (XCT) reconstructions, as shown in Fig. 2c. Since the raw data for XCT can be captured immediately after the raw data for HEDM without dismounting the sample or significantly altering the experimental setup, the two data types are inherently very well registered with each other, and the reconstruction of the markers from XCT can be compared with the reconstruction from serial sectioning SEM images to assist with cross-modality alignment.
SEM and XCT images of the sample markers. The volume of interest for analysis is a 40 \(\upmu \)m thick segment of the 1\(\times \)1 mm sample cross section, located to the left of the gold cube in the SEM images. The sample surface roughness (visible in both SEM and XCT) is from the EDM recast layer
Overview of Characterization Techniques
HEDM and XCT Scanning
For HEDM and XCT, we used a multi-modal imaging instrument at the 1-ID-E endstation of the Advanced Photon Source (APS), Argonne National Laboratory [9,10,11,12]. Figure 3a shows a schematic of this instrument. It is capable of seamlessly switching between XCT, near-field HEDM (NF-HEDM), and far-field HEDM (FF-HEDM) modes by translating the XCT and NF-HEDM detectors along \(y_L\), moving them into or out of the beam path as needed. X-rays from a superconducting undulator [13] were monochromatized [14] to 71.68 keV, the rhenium k-edge. Throughout the measurement, peak-to-peak X-ray energy fluctuation was approximately 0.006 keV, with the X-ray energy remaining spatially uniform across the beam. The temperature inside the endstation was held at \(22.2^\circ \) C to within \(\pm 0.15^\circ \) C, providing the thermal stability necessary to characterize the VOI precisely and accurately. Figure 3b illustrates the four coordinate systems relevant to the HEDM and XCT measurements. Here, we list the important features of these coordinate systems; they are described further in [12].
-
The laboratory coordinate system (\(x_L\), \(y_L\), \(z_L\)) are defined by the beam propagation direction (\(x_L\)) and the tomographic rotation axis (\(z_L\)), with \(y_L\) defined as the cross-product between \(z_L\) and \(x_L\). In practice, the instrument is aligned such that the tomographic rotation axis is perpendicular to \(x_L\); the slits and focusing optics are aligned such that they are aligned with \(y_L\) and \(z_L\). The intersection between \(x_L\) and \(z_L\) is defined as \(O_L\).
-
The crystal coordinate system(s) (\(x_C\), \(y_C\), \(z_C\)) are associated with the crystal lattice orientations of individual grains identified using HEDM reconstruction.
-
The sample coordinate system (\(x_S\), \(y_S\), \(z_S\)) are defined as the significant directions associated with the sample. While often similar to the laboratory coordinate system, it is not identical, as it varies with sample alignment prior to scanning. The relationship between the sample and crystal frames is generally the desired result from HEDM reconstruction, but determining the relationship between the sample and laboratory frames is necessary to extract this information.
-
The detector coordinate system(s) (\(x_D\), \(y_D\), \(z_D\)) are associated with the measurement planes on the detectors used to collect each data type. The relationship between each detector coordinate system and the laboratory coordinate system needs to be established to correctly process the diffraction signals captured for each data type. Calibration is conducted using a combination of a \(\mathrm {CeO_{2}}\) or \(\mathrm {LaB_{6}}\) powder sample [15] and a sample with only a few constituent grains such as the attached gold cube marker [8] or ruby crystals [16].
XCT
XCT was used to obtain the overall shape of the sample cross section, locate the VOI with respect to the overall sample geometry, and establish the relationship between the sample coordinate system and the laboratory coordinate system. In this mode, an area detector [17] was placed approximately 140 mm from \(O_L\). Its nominal pixel size is 1.172 \(\upmu \)m. Its field of view (FOV) is approximately 2 mm along \(x_L\) and 1 mm along \(y_L\), defined by a set of slits placed approximately 1 m upstream of \(O_L\). The flyscan approach was used to continuously acquire XCT radiographs in 0.1\({}^{\circ }\) intervals from \(-\)180\({}^{\circ }\) to 180\({}^{\circ }\). The Microstructural Imaging using Diffraction Analysis Software (MIDAS) suite was used for XCT reconstruction [18]. Figure 2c shows some representative projections from this reconstruction. The LSHR sample, gold cube marker, and platinum lines are all clearly visible. These features were essential in accurately establishing the VOI position with respect to the incident X-ray beam and devising a measurement protocol to virtually section and characterize layers within the VOI.
HEDM
For the FF-HEDM and NF-HEDM measurements, we used a planar X-ray beam from a set of sawtooth refractive lenses [19] placed approximately 1.5 m upstream of \(O_L\). The planar beam’s nominal dimensions are approximately 1.5 \(\upmu \)m in \(z_L\) and 2 mm in \(y_L\), allowing it to illuminate the entire cross section of the sample. For FF-HEDM, we used a GE detector [20] with 200 \(\upmu \)m pixels; it was placed approximately 754 mm from \(O_L\). For NF-HEDM, we used a CCD detector coupled with scintillation optics resulting in 1.48 \(\upmu \)m pixels [21] placed at multiple distances from the sample in order to triangulate the source of the diffraction signals. Figure 4 illustrates the VOI characterized using HEDM. The first layer of the VOI was nominally aligned to the bottom of the Au cube using radiography, while the axial direction of the sample was aligned to \(z_L\). The subsequent layers were 2 \(\upmu \)m apart in \(z_L\) for 20 layers, resulting in a total VOI height of 40 \(\upmu \)m below the gold cube. At each layer, HEDM diffraction patterns were acquired in FF and NF modes. For the first layer, five sets of HEDM patterns were acquired to verify stability. The raw diffraction patterns and metadata associated with each layer are defined in a spreadsheet provided with the raw dataset, which is available via the Materials Data Facility (MDF) [22] at https://doi.org/10.18126/4y0p-v604.
HEDM Reconstruction Parameters
The MIDAS suite [18] was used for both FF-HEDM and NF-HEDM reconstruction. The underlying HEDM reconstruction framework used in MIDAS is described in [23,24,25]. Here, the most salient pieces of information necessary for reconstruction are presented. The full MIDAS parameter files used for reconstruction are available at the MDF (https://doi.org/10.18126/4y0p-v604).
HEDM reconstruction produces a 3D microstructure map of the VOI. For FF-HEDM, we obtain the centers of mass (COMs), crystal orientations, and elastic strain tensors associated with each grain. The COMs and elastic strain tensors are in the laboratory frame (\(x_L\), \(y_L\), \(z_L\)). For NF-HEDM, we obtain a space-filling map of crystal orientations, with the voxel positions similarly in the laboratory frame. In both FF- and NF-HEDM, crystal orientations are given as rotations that relate the laboratory frame to the appropriate crystal frame (\(x_C\), \(y_C\), \(z_C\)). It is worth noting that these quantities are commonly transformed from the laboratory frame to the sample frame (\(x_S\), \(y_S\), \(z_S\)) using XCT. For this work, these quantities are instead converted to a frame shared with the three-dimensional (3D) EBSD data; this procedure is described in §4.
FF-HEDM Table 1 summarizes the detector parameters that define the relationship between the laboratory and detector coordinate systems for FF-HEDM. A threshold of 80 counts was applied to the dark-current corrected diffraction patterns. The following \(\{h\, k\, l\}\) families of crystallographic planes were included in the analysis: \(\{2\, 2 \,0\}\), \(\{3 \,1 \,1\}\), \(\{2\, 2\, 2\}\), \(\{4 \,0 \,0\}\), \(\{3\, 3\, 1\}\), \(\{4 \,2 \,0\}\), and \(\{4\, 2\, 2\}\). The tolerances used for diffraction spot matching were 0.5\({}^{\circ }\) in \(\omega \) and 800 \(\upmu \)m in position on the detector. A confidence threshold of 70% was used in the reconstruction; this means that for each indexed grain, at least 171 of the 244 expected diffraction spots must be observed on the area detector [26]. For diffraction analysis, we assumed that LSHR has a face-centered cubic (FCC) structure. Its fitted lattice parameter was found to be 3.585 Å.
NF-HEDM An undeformed and isolated gold cube (the marker attached to the sample) was moved to \(O_L\), and a set of NF-HEDM diffraction patterns was collected using 5 detector distances. This dataset was used to determine the initial values of the NF-HEDM detector parameters. These values were further refined using the NF-HEDM patterns collected from the LSHR sample with the same detector placements; Table 2 presents these refined NF-HEDM detector parameters. As the detector is placed asymmetrically in NF-HEDM, only the diffraction spots with positive \(z_D\) were recorded and used in the analysis. A threshold of 5 counts was applied after removing the median from the diffraction patterns. The following \(\{h \,k \,l\}\) families of crystallographic planes were used in the analysis: \(\{1\, 1\, 1\}\), \(\{2\, 0\, 0\}\), \(\{2 \,2 \,0\}\), \(\{3\, 1\, 1\}\), \(\{2\, 2\, 2\}\), \(\{4\, 0 \,0\}\), \(\{3\, 3\, 1\}\), and \(\{4\, 2\, 0\}\), resulting in up to 112 expected diffraction spots per orientation.
The sample reconstruction space for NF-HEDM was divided into voxels, equilateral triangles of 1 \(\upmu \)m in length, placed 3 \(\upmu \)m apart in a hexagonal arrangement for each layer. The initial NF-HEDM reconstruction space was intentionally larger than the sample size; XCT reconstruction data were used as a mask to define the sample boundaries. The NF-HEDM reconstruction was seeded using FF-HEDM results for the corresponding layer, meaning that for each reconstruction voxel, MIDAS would first try the orientations found in FF-HEDM. If a satisfactory solution was not found for a particular voxel (\(\textrm{confidence} < 30\%\), i.e., less than 30% of the expected diffraction spots were observed for any of the seeded orientations), MIDAS would run an exhaustive orientation search for that voxel.
Serial Sectioning via Mechanical Polishing
Sample Mounting and Polishing
Following HEDM and XCT, the LSHR sample was carefully cut and prepared for three-dimensional serial sectioning via mechanical polishing (MPSS). We generally follow the procedure described in [2], with adjustments to the mounting and polishing procedures to preserve the gold marker and platinum lines, account for the material properties of LSHR, and accurately determine the material removal rate and surface flatness. §A provides a detailed description of the sample mounting process and polishing procedure.
Optical Imaging and Surface Profiling using Scanning White Light Interferometry
Automated optical imaging was used to determine the rate of material removal during sectioning and to provide baseline images for a registration workflow that removed distortions from the EBSD data. Both are critical to ensuring that the data obtained during MPSS can be accurately matched to the data obtained from HEDM. The full details of the optical microscope and imaging procedures used are available in §B. The mean material removal rate was found to be 2.16 \(\upmu \)m/section, slightly higher than the nominal value of 2 \(\upmu \)m/layer used during HEDM. This difference was accounted for as part of the data fusion procedure described in §4, including any variations from section to section.Footnote 3
Additionally, surface profiles of each section were collected using scanning white light interferometry (SWLI). This procedure was introduced to ensure that any deviations from flatness of the polished surface could be measured and included in the final MPSS reconstruction, rather than simply assuming that every section was perfectly flat. The full details of this procedure are available in §C.
SEM Imaging and EBSD Pattern Acquisition
A series of SEM images was acquired from each section, with the majority of these images used to automatically align the exposed sample cross section to the SEM FOV. A final high-quality backscattered electron (BSE) image was acquired at zero tilt prior to moving the sample to the EBSD acquisition position, with an example of this shown in Fig. 5. These high-quality BSE images were used as a bridge between the optical images (taken as the “ground truth” in terms of image distortion) and the EBSD maps of each section, as was done in an earlier study [2]. This registration procedure is critical to accurate MPSS reconstruction, as the EBSD maps are known to contain significant image distortions from tilt correction, dynamic focus, and other potential sources [27,28,29,30,31].
Custom Python scripts were developed to automate SEM imaging and EBSD scanning using manufacturer-provided APIs. These scripts ensured that the sample was rotationally aligned and centered in the SEM FOV, with a tolerance of \(\pm 1\upmu {\hbox {m}}\) for both horizontal and vertical centering in the FOV and a tolerance of \(\pm 0.1^{\circ }\) for rotational alignment to the FOV. The scripts also automatically moved the sample to the EBSD acquisition position and acquired EBSD patterns. §D presents further details on the SEM imaging and EBSD pattern acquisition procedures.
High-Precision EBSD Pattern Indexing
Spherical harmonic indexing (SHI) was chosen as the high-precision indexing technique used for this study. SHI was originally introduced by Lenthe et al. [32] as a computationally efficient method of reliably indexing noisy or otherwise low-quality EBSD patterns. Our earlier work [33] demonstrated that it could also provide high-precision orientation indexing given sufficiently high-quality EBSD patterns, with the signal-to-noise ratio (SNR) of the collected patterns being the most important parameter and the pattern pixel resolution having remarkably little effect on the precision. To further improve orientation accuracy, significant efforts were made to accurately determine the EBSD sample-detector geometry, also known as the pattern center (PC). Readers are referred to §E and §F for more details on PC determination and SHI, respectively.
Data Registration for Serial Sectioning Reconstruction
Multi-modal registration of the various data types collected during MPSS is needed for layer-to-layer alignment and removal of distortion effects inherent in SEM/EBSD data collection [27,28,29,30,31]. Many of the techniques used for MPSS data registration here were based off those developed during an earlier study [2]. Adjustments were made to the overall workflow to allow for as much of the dataset registration as possible to be completed with automated Python scripts, reducing the need for manual intervention or the use of other software packages for the registration sequence itself.
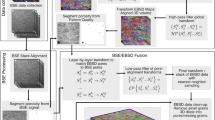
Following the same framework as in [2], the optical images were used as the ground truth for spatial position of the sample. First, the optical images were aligned against each other in series using a rigid transform. From there, the BSE images were registered to their corresponding optical images with a nonlinear transform, acting as an intermediate registration step to align the EBSD data to the optical data. Finally, the EBSD data were registered against the corrected BSE images with a nonlinear transform to remove distortions. The SWLI data were separately registered against the optical images so that it could be matched to the EBSD data for generation of a 3D point-cloud volume. Figure 6 summarizes the data registration workflow for each of the MPSS data modes, with detailed descriptions available in §G.
Alignment of 3D-EBSD and HEDM Data
The MPSS workflow described in §3.2 produces a registered, distortion-corrected 3D-EBSD map of the VOI. Even without further processing, a relatively close correspondence can be assigned between the HEDM data layers and a set of 3D-EBSD sections due to the careful control of the material removal rate and surface flatness during polishing. However, there are still several further steps required to optimize the alignment for data fusion, which are summarized in Fig. 7. For the full details of this alignment process, see §H.
First, it is necessary to bring the 3D-EBSD, NF-HEDM, and FF-HEDM data into a unified coordinate system. This includes compensating for the fact that the EBSD vendor data format uses different reference frames for recording position data and orientation data. The EBSD data are then rotated in 3D and nearest-neighbor resampled to compensate for any remaining misalignment between the physical sectioning planes for MPSS and the virtual sectioning planes for HEDM. Finally, a similarity transform is calculated to align the NF-HEDM data to the corresponding resampled 3D-EBSD data layers in 2D, and the NF-HEDM data points are themselves nearest-neighbor resampled from the hexagonal reconstruction grid onto the square 3D-EBSD position grid to allow for point-to-point quantitative comparisons. Here, we use the coordinate system employed by the 3D-EBSD position data as the unified coordinate system and bring the HEDM data into this coordinate system, as shown in Fig. 8. Subsequent maps use this coordinate system.
Results and Discussion
A detailed catalog of the data is tabulated and included in the dataset archived at the MDF. In this section, we present several comparison studies conducted using the 3D orientation map of the VOI generated from MPSS and NF-HEDM.
Layer-to-Layer Orientation Comparisons
Once the EBSD and NF-HEDM maps have been registered and resampled onto the unified coordinate system, they can be easily compared across corresponding points, whether those are from a different data type (i.e., comparing points with matching (\(x_U\), \(y_U\), \(z_U\)) values across data types) or a different layer in the same data type (i.e., comparing points with matching (\(x_U\), \(y_U\)) values but different \(z_U\) values within a single data type). In this work, a misorientation angle threshold of 3\({}^{\circ }\) is used to define whether a pair of points belong to the same grain in two compared datasets. Figure 9 shows that the observed misorientations between EBSD sections are generally either well above or well below this threshold, with the number of points above the threshold increasing as the distance in \(z_U\) between the compared sections is increased. Similar results were found for comparisons between layers in the NF-HEDM data. Note that specifically for comparisons of EBSD to EBSD in this work, the distortion-corrected data (processed as described in §3.2.5, but without the further alignments discussed in §4) are used to highlight differences between the individual sectioning runs.
Reference Frame Misalignments Between Different Layers of the Same Type
To estimate the rigid-body misalignment between two different layers of the same type, we first calculate the misorientations between the two layers at each (\(x_U\), \(y_U\)) point, similarly to Fig. 9. Any points with misorientation angles above the 3\({}^{\circ }\) threshold are discarded, and the remaining point-to-point misorientations are then averaged to produce a mean misorientation between the two different layers. These layer-to-layer misorientations can be averaged further to obtain a mean misorientation between one layer and every other layer, e.g., averaging the 19 results from layer 1 of the NF-HEDM data versus layers 2 through 20. Averaging is done in quaternion form to account for all misorientation parameters [34], although the discussion below focuses on the misorientation angles.
For the reconstructed NF-HEDM data, the misorientations are uniformly very small, with all layer-to-layer misorientation angles below 0.05\({}^{\circ }\). As shown in Fig. 10, higher misorientation angles are found between individual sections of EBSD data due to the sample needing to be removed from the SEM for mechanical polishing and the SEM stage needing to be moved between 0\({}^{\circ }\) and 70\({}^{\circ }\) tilt for every section. Despite the higher variation between sections for the EBSD data, the mean misorientation angle between one section and all other sections never exceeds 0.3\({}^{\circ }\), while the individual section-to-section misorientation angles never exceed 0.5\({}^{\circ }\), both low enough to be below the noise threshold of conventional EBSD. We attribute this high consistency to the combination of high-precision EBSD, pattern center fitting for every section, automated sample alignment routines to ensure that the sample is consistently positioned during SEM imaging and EBSD scanning, and an effective clamping solution to ensure that the sample mounting on the SEM stage is highly repeatable.
Misorientations Between Resampled EBSD and NF-HEDM Data Layers
Comparisons between matching layers of the resampled EBSD and NF-HEDM data were similar to layer-to-layer comparisons within a single data type, but a significantly higher fraction of noncorresponding pixels was observed relative to comparisons of adjacent layers within a single data type (see Fig. 11 versus Fig. 9). Across a subset of the VOI cropped to contain only the sample, the fraction of noncorresponding points (those with a misorientation angle of more than 3\({}^{\circ }\)) was found to be 46.2%. By comparing the IPF colormaps of both data types (as shown in Fig. 8), we found that these points typically involved regions where smaller grains or twins visible in the EBSD data were absent from the HEDM data, with the larger grains found in the HEDM reconstruction having expanded to cover those regions. This result is similar to the findings of previous EBSD/HEDM comparison studies [1,2,3,4], with improving the correspondence of reconstructed HEDM data to the ground truth being a critical research target.
In addition to the network of noncorresponding points near the outlines of larger grains, Fig. 11 also shows a relatively consistent misorientation axis and angle in the regions that do correspond. Further investigation showed that this was true for all resampled EBSD layers when compared with their matching HEDM layers. This prompted the idea of applying a per-layer uniform rotation to the EBSD orientation data to better align it with the HEDM orientation data, empirically determined from the average misorientation axis and angle for corresponding points in that pair of layers. This was highly successful at reducing the misorientation between the EBSD and HEDM data, with 95% of the corresponding points after such alignment having a misorientation angle below 0.46\({}^{\circ }\) across the entire volume of interest, with a mean value of only 0.24\({}^{\circ }\). While these alignment rotations were calculated per layer (allowing for the known variations between EBSD layers to be accounted for), the alignment rotations themselves were also remarkably consistent from layer to layer. The largest angle between any individual layer’s alignment rotation and the average alignment rotation across all layers was 0.21\({}^{\circ }\), similar in magnitude to the misorientation between EBSD sections shown in Fig. 10 (as might be expected given the high internal consistency of the HEDM data).
The source of this misorientation between the EBSD and HEDM orientation data has not been confirmed, but the high level of consistency across the entire dataset implies that it is a systematic error. For the EBSD analysis, plausible sources of systematic error include the true EBSD camera tilt angle differing from its nominal value of 10\({}^{\circ }\) or the true sample tilt in the EBSD position differing from its nominal value of 70\({}^{\circ }\). Either of these is known to produce a fictitious rotation of the crystal orientations about \(x_U\) in direct proportion to the difference between the nominal angle and the true angle. However, it is unlikely that the latter would produce such a consistent error across multiple sections, as the SEM stage is moved for every section. The SEM stage tilt axis could also be slightly misaligned relative to the EBSD camera, which is known to cause consistent errors in the measured orientations from EBSD [35]. Averaging the alignment rotations used to rotate the EBSD orientation data to match the HEDM orientation data across all layers produces a rotation of 1.67\({}^{\circ }\) about (\(x_U\), \(y_U\), \(z_U\)) \(=(0.954, -0.272, -0.124)\). Misalignment of the HEDM axes is unlikely to be a major cause of systematic error, as an error on the order of 1.67\({}^{\circ }\) would cause NF-HEDM reconstruction to fail completely. While it is very difficult to experimentally confirm the accuracy of orientation measurements (as even if a sample of known crystal orientation is available, it must also be positioned within the instrument with an accurately known physical orientation), it is convenient that the high precision of both analysis techniques used here allows these systematic errors to be empirically corrected with simple uniform rotations.
When averaged across all layers for each (\(x_U\), \(y_U\)) position (as shown in Fig. 12), the misorientation angles between the empirically aligned EBSD orientation data and the HEDM orientation data show a notable increase with distance from the center of the scan, which is consistent with the primary source of remaining misalignment being residual PC shift effects (see the end of §F). Given the same registration and alignment treatment as the spherical harmonic indexing data, the conventional Hough-indexed EBSD data show an even greater misorientation relative to the HEDM data, along with with an obvious trend versus position implying that there is insufficient PC shift correction across the large scan area. Interestingly, there is a far stronger trend in the Y-direction than the X-direction for the Hough data, so it is possible that the PC shift calibration in the Y-direction was not correctly adjusted for the large scan area used and might provide better performance if this was rectified. However, even in the central region where the PC shift effects are minimal, the Hough data retain a notably higher mean misorientation angle than the SHI data due to its lower precision.
Misorientation angle between corresponding points in the resampled EBSD and HEDM data (after per-layer empirical orientation alignment rotations) when using either spherical harmonic indexing or Hough indexing on the same raw EBSD patterns, averaged across \(z_U\) and plotted versus (\(x_U\), \(y_U\)) position. Note the significantly different color scales for the two plots
Grain-level Correspondence Between EBSD and NF-HEDM
DREAM.3D software (version 6.5.171) [36] was used for 3D grain segmentation. Prior to further processing, the individual “.ang” orientation data files for each layer of the aligned and resampled EBSD and HEDM data were combined into one “.h5ebsd” HDF5 data file per data type. Grains were segmented using a misorientation threshold of 3\({}^{\circ }\), followed by merging adjacent twinned regions with an axis tolerance of 3\({}^{\circ }\) and an angle tolerance of 2\({}^{\circ }\). The segmentation results were then saved in the HDF5-based “.dream3d” format. The .h5ebsd and .dream3d data files, as well as the DREAM.3D pipelines that were used for cleanup and segmentation, are available at the MDF.
Misorientations between corresponding voxels in the two datasets were calculated using the open-source MATLAB package MTEX [37], with matching voxels defined as those with orientations either within 3\({}^{\circ }\) of each other, or within 3\({}^{\circ }\) of the twinning misorientation for FCC crystals. Note that for the latter case, MTEX considers both the misorientation angle and the misorientation axis, so voxels that have a misorientation angle close to 60\({}^{\circ }\) around a nontwinning axis will not be defined as matching. As each voxel is also labeled with feature ID numbers indicating which grain it belongs to (both before and after merging twins),Footnote 4 identifying matching voxels allows determination of matched grains between the two data types, defined here as any grains that contain at least one matching voxel. The unmatched grains in either dataset are of particular interest, as they highlight regions where issues may exist in one or both techniques.
Initial examination of the segmented grains in the HEDM data showed that the majority are extremely small, even after merging twins. Out of the 13204 merged grains, 10127 (76.7%) have an equivalent diameter below 5\(\upmu \)m, containing at most 8 voxels per grain given the 2\(\times \)2\(\times \)2 \(\upmu \)m voxel size used here. In comparison, out of the 8610 merged grains in the EBSD data, only 2304 (26.8%) have an equivalent diameter below 5\(\upmu \)m. The volume fraction of the sample region occupied by these small grains was 0.3% in the HEDM data and 0.1% in the EBSD data. As they represent only a tiny fraction of the total sample volume and the low number of voxels per grain raises concerns about their validity, grains with an equivalent diameter below 5\(\upmu \)m are referred to as “trivial grains” and excluded from further grain-level analysis, while “nontrivial grains” are those above this size threshold.
Out of the 3077 nontrivial HEDM merged grains, only 71 have no matches in the EBSD data (2.3% of nontrivial HEDM merged grains, representing 0.04% of the total nontrivial HEDM merged grain volume). Furthermore, only 5 of those 71 unmatched grains have an equivalent diameter above 10\(\upmu \)m. To within a small margin of error, we can say that every significant grain observed in HEDM has at least some correspondence to the EBSD data; there are very few “false positives.” In contrast, out of the 6306 nontrivial EBSD merged grains, 3371 have no matches in the HEDM data (53.4% of nontrivial EBSD merged grains, representing 10.9% of the total nontrivial EBSD merged grain volume).Footnote 5 The likelihood of matching increases as the equivalent diameter of the EBSD grain increases, with the largest unmatched grain having an equivalent diameter of 25.1 \(\upmu \)m. The percentage of matched EBSD grains versus equivalent diameter (as calculated from the histogram counts for matched EBSD grains versus all EBSD grains) is impressively well fit by an error function curve, as shown in Fig. 13. Per the fitted parameters, a 5% probability of matching is reached at an equivalent diameter of 11.5 \(\upmu \)m, a 50% probability at 17.8 \(\upmu \)m, and 95% probability at 24.1 \(\upmu \)m.
While the volume fraction occupied by completely unmatched grains is relatively low, almost all grains in both data types contain a significant fraction of unmatched voxels. This fraction varies both between the two data types and with the size of the grains in question, as shown in Fig. 14. The majority of HEDM grains have a matched fraction close to 60%, with outliers both above and below this value becoming less common as the grain size increases. In contrast, the EBSD matched fraction per grain is heavily concentrated at values below 25%, albeit with a relatively long tail to the distribution and a positive correlation between grain size and matched fraction.
Examining the confidence index (CI) distributions of individual voxels within the sample region proves instructive. As shown in Fig. 15, for the EBSD data, there is a slight tendency for matched voxels to have a higher CI than unmatched voxels, but both matched and unmatched voxels generally cover the full CI range observed. However, for the HEDM data, we see a very significant difference in the CI distribution for matched voxels and unmatched voxels, with higher CI values being much more likely to match the EBSD data. At the other end of the scale, a HEDM CI below 0.25 generally indicates regions of the reconstruction volume that are outside of the sample, although the reconstruction may still provide a “most likely” orientation for those voxels, typically matching the orientation of nearby surface grains. The correlation between the HEDM CI and the probability of matching the EBSD data for a given voxel is both remarkably strong and very linear, as shown in Fig. 16.
The data from individual layers also support these findings. From Fig. 17, it is clear that the highest CI values for NF-HEDM are found in the largest grains, while lower CI values are found in the interstitial regions between those large grains. The EBSD data maintain a relatively consistent CI throughout the entire sample.
Differences in Grain Positions in FF-HEDM, NF-HEDM, and EBSD
FF-HEDM, NF-HEDM, and EBSD can all provide the centers of mass (COMs) of individual grains. We examine the differences in COMs provided by the three characterization techniques. For the purposes of this analysis, matching grains between different techniques are defined as those with COMs within 100 \(\upmu \)m of each other and with orientations within 1\({}^{\circ }\) of each other (after a uniform empirical alignment rotation is applied to the EBSD orientation data to reduce the systematic misorientation versus the HEDM data discussed in §5.1.2). Furthermore, we only consider the data from the first layer for simplicity, as merging FF-HEDM data from layer to layer is nontrivial; we anticipate that the trends observed in a single layer will not be significantly different from those in the full 3D volume.
Table 3 shows the number of grains identified by each technique for the first layer. The number of grains found by FF-HEDM is much closer to the number of grains found by EBSD than those found by NF-HEDM. This is likely because the FF-HEDM detector is more sensitive than the NF-HEDM detector.
For the FF-HEDM dataset, there were 753711 diffraction spots detected in the set of diffraction patterns acquired in this layer. Out of 753711 spots, 630115 spots were assigned to the 3775 grains through the indexing process. Table 4 shows how 630115 spots were assigned to the indexed grains. Out of 630115 spots, 398038 diffraction spots were uniquely associated with the indexed grains; 168391 spots were doubly associated, 48036 spots were triply associated, and so forth. This is because of the high number of twins in this material; any adjacent grains in a twin relationship share many diffraction spots. The remaining 123596 spots out of 753711 detected diffraction spots were not assigned to any indexed grains; they were “orphaned.”
Figure 18 shows a histogram of all the indexed vs nonindexed spots. In this figure, the equivalent grain size associated with a particular diffraction spot is computed using the spot’s measured intensity relative to the sum of the measured intensity for the particular family of planes and the formulation in [23, 24]. Figure 18 shows that the orphaned spots are typically less intense and therefore associated with the smaller grains. In fact, the orphaned spots account for only 3.705% of the total area illuminated by the planar beam. Notably, FF-HEDM diffraction spots retain a high probability of being associated with an indexed grain down to the equivalent grain size threshold of 5 \(\upmu \)m that was used to exclude “trivial grains” in EBSD or NF-HEDM grain segmentation. Furthermore, as FF-HEDM cannot resolve individual grains with similar orientations and small spatial separations, the FF-HEDM results would include merged twin orientations that are otherwise spatially resolved in EBSD and NF-HEDM.
For the grain pairs found across these techniques, Fig. 19 shows a histogram of the Euclidean distances between the COMs of the paired grains (with these distances referred to as the “Error”), and Table 5 summarizes the data presented in Fig. 19. As this figure illustrates, the majority of the matching grains found across the two variants of HEDM and EBSD have COM distances below 20 \(\upmu \)m. This figure also shows that the matching grains from NF-HEDM and EBSD have the least positional discrepancy out of the three comparison scenarios. Table 5 similarly shows that the NF-HEDM vs. EBSD comparison yields the smallest mean and median errors compared to the other cases. It is interesting to note that the FF-HEDM vs. EBSD comparison yields the largest error. It is reasonable to assume that the positional accuracy of FF-HEDM is relatively poor compared to the other two mapping techniques. This is likely because the detector position with respect to the sample and its pixel size are optimized to measure the angles associated with diffraction spots rather than their source positions. Both NF-HEDM and FF-HEDM can also fail to resolve adjacent twinned regions that were resolved by EBSD, which could shift the calculated grain COMs and increase the error. Despite this, the NF-HEDM data still match the EBSD data relatively well.
A histogram of Euclidean distances between matching grains found using different characterization techniques. “FF\(\leftrightarrow \)EBSD,” “FF\(\leftrightarrow \)NF,” and “NF\(\leftrightarrow \)EBSD” refer to distances between grains found in FF-HEDM vs. EBSD, FF-HEDM vs. NF-HEDM, and NF-HEDM vs. EBSD, respectively
In any case, this experimental finding implies that generating virtual microstructures for simulations relying solely on a grain map acquired through HEDM needs careful consideration. For instance, as [38] illustrates, the micromechanical stress state of a grain in a crystal plasticity simulation can vary significantly depending on the geometry of the virtual microstructure. It seems that understanding and quantifying that spread in stress state, solely arising due to the change in the virtual microstructure, is a critical step before we use FF-HEDM and associated simulation tools to draw conclusions on the mechanical behavior of materials at the grain length scale.
Resolving Smaller Grains in NF-HEDM
Figure 8 and Fig. 13 show that the NF-HEDM map is generally successful at capturing larger grains, but often fails to detect smaller grains. The robust data registration exercise detailed in §4 provides an excellent avenue to investigate why smaller grains can be missed and presents an opportunity to improve the HEDM reconstruction. Using the EBSD maps as the ground truth and conducting a forward simulation to the NF-HEDM detector planes, we can begin to examine whether the diffraction signals from the smaller grains are missing altogether, or if those signals exist but were neglected during reconstruction. Assessing these possibilities allows us to devise appropriately tailored solutions. While a full examination of these scenarios is a topic for a follow-up publication, we provide some preliminary results here.
EBSD and HEDM comparison of a twin pair. (a) EBSD map (red/blue twin pair) overlaid with the HEDM reconstruction of the parent grain (red spheres). (b) Confidence map for HEDM voxels for the parent grain. (c-g) HEDM raw data showing diffraction spots for the two grains. Arrows show the projection direction of the diffraction signal (black: signal from both grains, red/blue: signal from corresponding grain). (c) Diffraction spot shared between the two grains. (d,f) Diffraction spots with signal from parent grain only. (e,g) Diffraction spots with signal from twin grain only. The diffraction spots’ widths are between 40 \(\upmu \)m and 60 \(\upmu \)m depending on the projection angle
Figure 20 highlights a pair of grains (the parent grain in red and its twin in blue) that were detected by EBSD. The corresponding HEDM map of the same region (represented by colored spheres overlaid on the EBSD map) shows only the parent grain; the smaller twin region was not identified during HEDM reconstruction (Fig. 20(a)). The confidence map for the HEDM data shows relatively high values, particularly in the region where the twin should exist (Fig. 20(b)). However, when we conduct a forward simulation onto the NF-HEDM detector plane using the EBSD map data from the parent grain and its twin, we observe that the diffraction signal for the twinned region does exist as isolated diffraction peaks (Fig. 20(e) and (g)) or gaps in the parent grain’s diffraction signal (Fig. 20(d)). However, in many cases, the signal from the twin is lost due to being overwhelmed by the signal from the parent grain (Fig. 20(c)). These observations indicate that the diffraction conditions and the resulting projection angles have to be “just right” in order to detect the diffraction signals from a smaller grain, especially a narrow twin within a larger parent grain. Advanced signal processing that can capture minute intensity variations in a diffraction peak or fitting the intensity profile of the NF-HEDM diffraction peaks is a possible solution paths to consider in order to capture these smaller grains and features that reside next to a larger neighbor and share reflections with that neighbor. We can also start to consider a more flexible confidence threshold, moving away from a single rigid confidence threshold value that dictates whether a voxel should have a particular orientation or not. Newer detector technologies with reduced noise and point spread function can also be considered to enhance the raw measured diffraction signal.
Summary
A selection of raw and processed data types from this study is available at the MDF (https://doi.org/10.18126/4y0p-v604). Major results of this work are listed below:
-
The experimental procedures were described in detail for reference.
-
A workflow for registering BSE images to optical images and EBSD data to BSE images was described and implemented, allowing the removal of image distortions from the BSE and EBSD data. The same workflow was also used to register SWLI surface heightmaps to the optical images, allowing their application to the 3D positions of the EBSD data points.
-
A workflow to determine any 3D misalignment of the sectioning planes between the EBSD and NF-HEDM modalities using BSE and XCT image data was described and implemented, allowing the EBSD data to be 3D rotated and resampled into layers with a one-to-one correspondence to the HEDM layers.
-
An algorithm to determine matching grains between the EBSD and NF-HEDM modalities was described and implemented, allowing the calculation of the 2D similarity transform required to best align the HEDM data layers with the resampled EBSD data layers and the resampling of the HEDM data into a point-to-point correspondence with the EBSD data.
-
High-precision EBSD from spherical harmonic indexing was introduced to the serial sectioning workflow, with section-to-section consistency better than 0.3\({}^{\circ }\) based on the measured crystal orientations. The noise floor for this technique given the scan conditions used here was found to be 0.06\({}^{\circ }\) for adjacent points within a single scan.
-
Comparison of 3D-EBSD and NF-HEDM orientation data showed a systematic error of 1.67\({}^{\circ }\) about (\(x_U\), \(y_U\), \(z_U\)) \(=(0.954, -0.272, -0.124)\) after application of the alignment workflows, with either a difference between the nominal and true EBSD camera tilt angle or a misalignment of the SEM stage tilt axis relative to the EBSD camera proposed as the primary cause of this error.
-
Due to the high precision of both techniques, empirical correction of the systematic error in the orientation data using per-layer uniform rotations improved the mean misorientation angle between 3D-EBSD and NF-HEDM data to 0.24\({}^{\circ }\), with 95% of corresponding points having a misorientation angle below 0.46\({}^{\circ }\).
-
Comparisons between 3D-EBSD and NF-HEDM data showed that 46.2% of voxels in the sample did not match between the two data types, with a direct correlation between the HEDM confidence index and the likelihood of matching the EBSD data. 97.7% of grains identified in HEDM were present in EBSD, but 53.4% of grains identified in EBSD were not present in HEDM, with a strong correlation between their equivalent diameter and the likelihood of matching data being present in the HEDM reconstruction. Due to this correlation, wholly unmatched EBSD grains represent only 10.9% of the total sample volume, with the majority of the unmatched voxels coming from partially matched grains.
-
Overall, many grains were detected by both techniques. Out of 4496 grains detected by EBSD on a single layer, FF-HEDM detected 3409 grains, while NF-HEDM detected 1595 grains. The grain COMs identified by the three techniques were within 20 \(\upmu \)m of each other.
-
It was demonstrated that for a parent grain containing a narrow twin visible in EBSD but absent from the NF-HEDM reconstruction, the signal from the twinned region was present in the raw X-ray data, but was neglected during reconstruction due to certain reflections being overwhelmed by the signal from the larger parent grain.
Notes
The total height of the VOI was primarily dictated by the availability of beam time for HEDM.
A previous study used similar marker lines [2], but they were applied to the sample after HEDM scanning and only used to correlate the optical and SEM image modes.
The term “section” in this article is used to specifically refer to data from a single step of MPSS. The term “layer” is less specific, as it is also used for data from HEDM, or for 3D-EBSD data after rotation and resampling to match the HEDM layers (see §4), which combines data from multiple sections of MPSS into each resampled EBSD layer.
Note that a feature ID number of 0 is assigned to voxels that were not part of any grain after segmentation.
The number of matched grains (3006 for HEDM and 2935 for EBSD) is not identical for the two data types due to differences in the underlying data producing differences in grain segmentation. It is possible for one HEDM grain to overlap and match multiple EBSD grains, or vice versa.
Registration based on the sample itself is not recommended, as it has previously been shown to incorrectly remove any existing sample tilt within the mount by aligning the sample edges from section to section [2].
Note that the required rotation depends on the reference frame setting used in the EDAX software during data collection. This rotation is correct for .ang data collected with “Setting 2” and is used in the built-in MTEX import function for such data. Other EBSD data formats also generally require different rotations.
Version 2022.2 for Windows, Comet Technologies Canada Inc., Montreal, Canada; available at https://www.theobjects.com/dragonfly
References
Menasche DB, Shade PA, Suter RM (2020) Accuracy and precision of near-field high-energy diffraction microscopy forward-model-based microstructure reconstructions. J Appl Crystallogr 53(1):107–116
Chapman MG, Shah MN, Donegan SP, Scott JM, Shade PA, Menasche D, Uchic MD (2021) AFRL additive manufacturing modeling series: challenge 4, 3D reconstruction of an IN625 high-energy diffraction microscopy sample using multi-modal serial sectioning. Integrat Mater Manuf Innov 10:129–141
Louca K, Abdolvand H (2021) Accurate determination of grain properties using three-dimensional synchrotron X-ray diffraction: a comparison with EBSD. Mater Char 171:110753
Ball JA, Oddershede J, Davis C, Slater C, Said M, Vashishtha H, Michalik S, Collins DM (2023) Registration between DCT and EBSD datasets for multiphase microstructures. Mater Char 204:113228
Lenthe WC, Echlin MP, Trenkle A, Syha M, Gumbsch P, Pollock TM (2015) Quantitative voxel-to-voxel comparison of TriBeam and DCT strontium titanate three-dimensional data sets. J Appl Crystallogr 48(4):1034–1046
Gabb TP, Gayda J, Telesman J, Kantzos PT (2005) Thermal and mechanical property characterization of the advanced disk alloy LSHR, thermal and mechanical property characterization of the advanced disk alloy LSHR, tech. rep. NASA/TM-2005-213645, NASA
Musinski WD, Shade PA, Pagan DC, Bernier JV (2021) Statistical aspects of grain-level strain evolution and reorientation during the heating and elastic-plastic loading of a Ni-base superalloy at elevated temperature. Materialia 16:101063
Shade PA, Menasche DB, Bernier JV, Kenesei P, Park JS, Suter RM, Schuren JC, Turner TJ (2016) Fiducial marker application method for position alignment of in situ multimodal X-ray experiments and reconstructions. J Appl Crystallogr 49:700–704
Wielewski E, Menasche DB, Callahan PG, Suter RM (2015) Three-dimensional colony characterization and prior-\(\beta \) grain reconstruction of a lamellar Ti-6Al-4V specimen using near-field high-energy X-ray diffraction microscopy. J Appl Crystallogr 48:1165–1171
Bernier JV, Suter RM, Rollett AD, Almer JD (2020) High-energy x-ray diffraction microscopy in materials science. Ann Rev Mater Res 50:395–436
Menasche DB, Musinski WD, Obstalecki M, Shah MN, Donegan SP, Bernier JV, Kenesei P, Park J-S, Shade PA (2021) AFRL additive manufacturing modeling series: challenge 4, in situ mechanical test of an IN625 sample with concurrent high-energy diffraction microscopy characterization. Integrat Mater Manuf Innov 10:338–347
Park J-S, Sharma H, Kenesei P (2021) Repeatability and sensitivity characterization of the far-field high-energy diffraction microscopy instrument at the Advanced Photon Source. J Synchrot Radiat 28:1786–1800
Ivanyushenkov Y, Harkay K, Borland M, Dejus R, Dooling J, Doose C, Emery L, Fuerst J, Gagliano J, Hasse Q, Kasa M, Kenesei P, Sajaev V, Schroeder K, Sereno N, Shastri S, Shiroyanagi Y, Skiadopoulos D, Smith M, Sun X, Trakhtenberg E, Xiao A, Zholents A, Gluskin E (2017) Development and operating experience of a 1.1-m-long superconducting undulator at the Advanced Photon Source. Phys Rev Accelerat Beams 20:100701
Shastri SD, Fezzaa K, Mashayekhi A, Lee W-K, Fernandez PB, Lee PL (2002) Cryogenically cooled bent double-Laue monochromator for high-energy undulator X-rays (50–200 keV). J Synchrot Radiat 9:317–322
Kaiser DL, Watters JRL (2007) National Institute of Standards and Technology standard reference material 674b x-ray powder diffraction intensity set for quantitative analysis by x-ray powder diffraction.
Kaiser DL, Watters JRL (2007) National Institute of Standards and Technology Standard reference material676 alumina internal standard for quantitative analysis by x-ray powder diffraction.
Khounsary A, Kenesei P, Collins J, Navrotski G, Nudell J (2013) High energy x-ray micro-tomography for the characterization of thermally fatigued GlidCop specimen. J Phys Conf Ser 425:212015
MIDAS, https://www.github.com/marinerhemant/MIDAS
Shastri SD, Almer J, Ribbing C, Cederström B (2007) High-energy X-ray optics with silicon saw-tooth refractive lenses. J Synchrot Radiat D 14:204–211
Lee J, Almer J, Aydéner C, Bernier J, Chapman K, Chupas P, Haeffner D, Kump K, Lee P, Lienert U, Miceli A, Vera G (2007) Characterization and application of a GE amorphous silicon flat panel detector in a synchrotron light source. Nuclear Instrum Methods Phys Res Sect A Accelerat Spectromet Detect Associat Equip 582:182–184
Lienert U, Lind J, Hefferan CM, Pantleon W, Mills MJ, Miller MP, Suter RM, Li SF, Brandes MC, Bernier JV, Jakobsen B, Barton NR (2011) JOM 63:70–77
Blaiszik B, Chard K, Pruyne J, Ananthakrishnan R, Tuecke S, Foster I (2016) The materials data facility: data services to advance materials science research. JOM 68:2045–2052
Sharma H, Huizenga RM, Offerman SE (2012) A fast methodology to determine the characteristics of thousands of grains using three-dimensional X-ray diffraction. I. Overlapping diffraction peaks and parameters of the experimental setup. J Appl Crystallogr 45:693–704
Sharma H, Huizenga RM, Offerman SE (2012) A fast methodology to determine the characteristics of thousands of grains using three-dimensional X-ray diffraction. II. Volume, Centre-of-mass position, crystallographic orientation and strain state of grains. J Appl Crystallogr 45:705–718
Wozniak JM, Sharma H, Armstrong TG, Wilde M, Almer JD (2015) Big data staging with MPI-IO for interactive x-ray science. I. Foster in Institute of Electrical and Electronics Engineers, pp 26–34
Poulsen HF, Jensen DJ, Vaughan GB (2011) Three-dimensional x-ray diffraction microscopy using high-energy x-rays. MRS Bull 29:166–169
Nolze G (2007) Image distortions in SEM and their influences on EBSD measurements. Ultramicroscopy 107:172–183
Tong VS, Ben Britton T (2021) TrueEBSD: Correcting spatial distortions in electron backscatter diffraction maps. Ultramicroscopy 221:113130
Winiarski B, Gholinia A, Mingard K, Gee M, Thompson G, Withers P (2021) Correction of artefacts associated with large area EBSD. Ultramicroscopy 226:113315
Nguyen LT, Rowenhorst DJ (2021) The alignment and fusion of multimodal 3d serial sectioning datasets. JOM 73:3272–3284
Stinville JC, Hestroffer JM, Charpagne MA, Polonsky AT, Echlin MP, Torbet CJ, Valle V, Nygren KE, Miller MP, Klaas O, Loghin A, Beyerlein IJ, Pollock TM (2022) Multi-modal dataset of a polycrystalline metallic material: 3D microstructure and deformation fields. Sci Data 9:460
Lenthe W, Singh S, Graef MD (2019) A spherical harmonic transform approach to the indexing of electron back-scattered diffraction patterns. Ultramicroscopy 207:112841
Sparks G, Shade PA, Uchic MD, Niezgoda SR, Mills MJ, Obstalecki M (2021) High-precision orientation mapping from spherical harmonic transform indexing of electron backscatter diffraction patterns. Ultramicroscopy 222:113187
Markley FL, Cheng Y, Crassidis JL, Oshman Y (2007) Averaging quaternions. J Guid Control Dyn 30:1193–1197
Zaefferer S, Elhami NN (2014) Theory and application of electron channelling contrast imaging under controlled diffraction conditions. Acta Mater 75:20–50
Groeber MA, Jackson MA (2014) DREAM.3D: A digital representation environment for the analysis of microstructure in 3D. Integrat Mater Manuf Innov 3:56–72
Bachmann F, Hielscher R, Schaeben H (2011) Grain detection from 2d and 3d EBSD data-specification of the MTEX algorithm. Ultramicroscopy 111:1720–1733
Wong SL, Park JS, Miller MP, Dawson PR (2013) A framework for generating synthetic diffraction images from deforming polycrystals using crystal-based finite element formulations. Comput Mater Sci 77:456–466
Mingard K, Day A, Maurice C, Quested P (2011) Towards high accuracy calibration of electron backscatter diffraction systems. Ultramicroscopy 111:320–329
Zhu C, Kurniawan C, Ochsendorf M, An D, Zaefferer S, De Graef M (2022) Orientation, pattern center refinement and deformation state extraction through global optimization algorithms. Ultramicroscopy 233:113407
Pang EL, Larsen PM, Schuh CA (2020) Global optimization for accurate determination of EBSD pattern centers. Ultramicroscopy 209:112876
Shi Q, Zhong H, Loisnard D, Wang L, Chen Z, Wang H, Roux S (2023) Enhanced EBSD calibration accuracy based on gradients of diffraction patterns. Mater Character 202:113022
Lenthe W (2004) EMSphInx online documentation, EMSphInx online documentation. https://emsphinx.readthedocs.io/en/latest/index.html. Accessed: 2024-05-06
Chen YH, Park SU, Wei D, Newstadt G, Jackson MA, Simmons JP, De Graef M, Hero AO (2015) A dictionary approach to electron backscatter diffraction indexing. Microsc Microanal 21:739–752
Singh S, Ram F, De Graef M (2017) Application of forward models to crystal orientation refinement. J Appl Crystallogr 50:1664–1676
Gulsoy E, Simmons J, De Graef M (2009) Application of joint histogram and mutual information to registration and data fusion problems in serial sectioning microstructure studies. Scripta Mater 60:381–384
Johnstone DN, Martineau BH, Crout P, Midgley PA, Eggeman AS (2020) Density-based clustering of crystal (mis)orientations and the orix Python library. J Appl Crystallogr 53:1293–1298
Ånes HW, Martineau B, Harrison P, Crout P, Johnstone D, Cautaerts N, Gerlt A, Mathisen AC, Høgås S, Clausen A (2023) pyxem/orix: orix 0.11.1, version v0.11.1,
Rowenhorst D, Rollett AD, Rohrer GS, Groeber M, Jackson M, Konijnenberg PJ, De Graef M (2015) Consistent representations of and conversions between 3D rotations. Modell Simul Mater Sci Eng 23:083501
Acknowledgements
This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science user facility at Argonne National Laboratory, and is based on research supported by the US DOE Office of Science-Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. Electron microscopy was performed at the Materials Characterization Facility (MCF) at the Air Force Research Laboratory, supported under agreement number FA2394-23-C-B028. This research was supported by the AFRL under agreement number FA8650-18-2-5295 and by the University of Dayton Research Institute under agreement number FA8650-20-D-5211. The US Government is authorized to reproduce and distribute reprints for governmental purposes notwithstanding any copyright notation thereon. The views and conclusions contained herein are those of the authors and should not be interpreted as necessarily representing the official policies or endorsements, either expressed or implied, of the AFRL or the US Government. All introductions of product names in this article are only to describe the measurement and analysis setup and do not imply endorsement. M.D. Uchic, P.A. Shade, and M. Obstalecki acknowledge additional support from the Air Force Research Laboratory.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix A: MPSS Sample Mounting and Polishing
Following HEDM and XCT, a 3D-Micromac microPREP™ PRO picosecond laser ablation mill was used to cut the sample into two pieces. The cut was placed on the opposite side of the gold cube marker from the VOI, with the cut face at a distance of approximately 1 mm from the gold cube. Testing on a dummy sample of LSHR showed that the heat-affected zone from the laser cut was less than 100 \(\upmu \)m thick, so this distance was more than sufficient to ensure that the VOI was unaffected. It also provided a substantial amount of noncritical material that could be used to refine the serial sectioning workflow prior to sectioning the VOI, including both testing newly introduced workflow steps and adjusting the polishing parameters to produce the desired section thickness. Following this cut, a FIB was used to mill a diagonal trench into the face of the sample that was previously marked with a gold cube to simplify tracking sectioning depth while approaching the volume of interest. Figure 21a shows the cut face and FIB-milled trench, along with the previously attached gold cube and FIB-deposited platinum lines.
Figure 21b shows the sample mounted for serial sectioning, with several major features labeled. We followed the mounting method described in [2], with two notable changes. In this study, the support frame used to protect the sample during hot mounting was made from nickel-based superalloy rather than titanium to reduce any potential mismatch between the polishing behavior of the sample and the support frame. In addition, a slightly deeper step was milled around the outside of the Polyfast sample mount and a polished stainless steel ring was attached using vacuum-compatible conductive silver epoxy to serve as a height reference. This reference ring greatly improved the consistency of height measurements over the milled Polyfast surface, which had significant surface roughness. The optical microscope auto-focus algorithm could also locate the surface of the polished reference ring far more reliably than the milled Polyfast. This allowed precise tracking of material removal during sectioning, as discussed in §B.
Mechanical polishing for serial sectioning was accomplished in a similar manner to that described in [2], using the same RoboMet.3D™ automated polishing system. All polishing pads and polishing media used were identical to [2]. A constant force of \(\sim 20\)N was used during all polishing steps, and the duration of the material removal step using 1 \(\upmu \)m polycrystalline diamond abrasive was adjusted to produce a \(\sim 2\upmu {\hbox {m}}\) depth per section, which for this sample required a polishing time of approximately 2 min at a platen speed of 100 RPM. The colloidal silica polishing step was extended to 10 min at a lower platen speed of 60 RPM to help eliminate residual deformation from the material removal step, which was found to be visible under high-precision EBSD during preliminary testing. All sample cleaning steps were identical to those in [2].
Appendix B: MPSS Optical Imaging and Material Removal Measurement
Automated optical imaging was performed using a Zeiss Imager.Z2m optical microscope outfitted with an AxioCam 705 mono camera which collected 14-bit grayscale bright-field images. The serial sectioning region of interest (ROI) containing the sample was acquired as a 7\(\times \)7 tile montage using a 20\(\times \) objective lens (Epiplan, 0.5 NA) and 1\(\times \) tube lens, with auto-focusing performed at each tile position. The effective pixel size at this magnification was 0.345 \(\upmu \)m, which was verified against a calibration artifact. The ROI montage covered an area of approximately 2\(\times \)2 mm, ensuring that it included both the full 1\(\times \)1 mm cross section of the sample and a sufficiently large region of Polyfast mounting material on all sides of the sample to allow for image registration based solely on the mounting material.Footnote 6 An example optical image is shown in Fig. 22.
Cropped optical montage of the same section shown in Fig. 5. The raw optical images must be rotated 180\({}^{\circ }\) in-plane to bring them into the same reference frame as the BSE or EBSD data
The optical imager was also used to determine the rate of material removal during sectioning. Differential depth values were measured between four points on the reference ring and one point on the polished mounting material near the center of the mount. The latter point was used instead of the sample itself due to the ease of focusing on the mounting material versus the polished sample, which has relatively few visible features in optical microscopy. It should be noted that while there was a slight offset (\(< {0.5\,\mathrm{\upmu \text {m}}}\)) between the sample surface and the mounting material, as can be seen in Fig. 6, this offset was very consistent from section to section. All of the height-measurement points were collected as 3\(\times \)3 montages using a 50\(\times \) objective lens (Epiplan, 0.55 NA) and 1\(\times \) tube lens with auto-focusing performed for every tile, with the average of the auto-focus heights for each montage taken as the height of that point. The image data from these non-ROI montages were not used further, outside of acting as a cross-check that the auto-focus algorithm succeeded and that the correct locations on the reference ring were imaged. The four reference ring heights were averaged together and then subtracted from the measured surface height for each section, producing relative height values which are then referenced against a user-selected section to determine the material removal rate. As shown in Fig. 23, a linear fit to the removal rate in the volume of interest (sections 310-339) gave a value of 2.16 \(\upmu \)m/section, slightly higher than the nominal value of 2 \(\upmu \)m/layer used during HEDM. The section thickness statistics are summarized in Table 6. Note that the VOI collected during serial sectioning was thicker than the HEDM VOI to ensure that the latter was fully covered, as the precise correspondence between the two regions was difficult to determine during sectioning.
Appendix C: MPSS Surface Topography Mapping Using SWLI
Surface profiles of each polished layer were collected using scanning white light interferometry (SWLI). The instrument used was a Bruker ContourGT-X8 optical profiler with Vision64 control software. Data collection was automated using the manufacturer-provided control API and custom Python scripts. Before collecting the sample surface profile, the objective lens tip-tilt stage was used to align the lens with the sample by acquiring a single low-magnification heightmap from the center of the sample and using a 2D least-squares linear fit to determine the tip and tilt adjustments required for leveling. This leveling operation was always performed twice, as two runs were found to produce better leveling than a single run, but further iterations did not result in any significant improvement. Accurate lens-to-sample leveling was critical to eliminate sinusoidal artifacts in the surface profile data. After leveling, an autofocus function was used to position the scanner near the interference fringe envelope position, minimizing the scan length required to collect the entire fringe envelope and therefore reducing the total scan time. Individual montage regions were captured at the native camera resolution of 640\(\times \)480 pixels using a 0.55\(\times \) zoom lens and 50\(\times \) structured-light objective lens (0.55 NA). The effective pixel size at this magnification was 0.365 \(\upmu \)m. The ROI montage covered an area of 1.2\(\times \)1.2 mm, ensuring that it included both the full 1\(\times \)1 mm cross section of the sample and a small amount of the mounting material on all sides of the sample. Montage region overlap was specified as 20%, which required a 9\(\times \)7 tile array to cover the desired area. VSI acquisition mode was used with white-light illumination and 1\(\times \) scan speed, a backscan length of 2 \(\upmu \)m, and a scan length of 10 \(\upmu \)m.
Appendix D: MPSS SEM Image and EBSD Pattern Acquisition
The SEM used for imaging was a ThermoFisher Scientific Apreo C, set to an accelerating voltage of 20 kV and a beam current of approximately 20 nA. The working distance used for zero-tilt imaging was 5 mm, and the image pixel size was 0.5 \(\upmu \)m, with a total image resolution of 3072\(\times \)2048 pixels. The resulting 1536\(\times \)1024 \(\upmu \)m field of view allowed the entire sample cross section to be captured in a single image, with a cropped example shown in Fig. 5. Secondary electron (SE) images were also collected using the same parameters, although these images were primarily intended for clarification of any ambiguous features in the BSE images and were not used for registration.
Custom Python scripts were developed to automate SEM imaging and EBSD scanning using the manufacturer-provided control APIs. In addition to the centering and rotational alignment discussed in §3.2.3, the working distance was similarly refined to within \(\pm 5\upmu {\hbox {m}}\) of the desired 5 mm prior to zero-tilt imaging, and the contrast/brightness settings were automatically adjusted for optimal image contrast. In the case of the BSE images, this was specifically optimized only on the sample region in order to maximize grain-to-grain contrast, although SE images were adjusted to a lower contrast value where both the mounting material and the sample were visible. Following zero-tilt imaging, the sample was then automatically moved to the nominal EBSD working distance of 20 mm, tilted to 70\({}^{\circ }\), the working distance refined to within \(\pm 10\upmu {\hbox {m}}\) of the desired 20 mm after tilting, and the sample centered in the SEM FOV with a tolerance of \(\pm 1\upmu {\hbox {m}}\) for both horizontal and vertical centering. The sample rotation was left unchanged from the zero-tilt alignment step. Automated EBSD scans were then run according to pre-defined parameters, with the exception of the EBSD camera exposure time, which was automatically adjusted at the beginning of each scan.
The EBSD patterns were captured using an EDAX Velocity Super camera at a resolution of 120x120 pixels. In the APEX control software for the Velocity camera, the “Super Mode” parameter was active and the “High Range” parameter was inactive. The former option sets the camera into high-speed 8-bit acquisition mode, while the latter choice effectively decreases the camera gain, allowing for an improved SNR by increasing beam current or exposure time. The SEM used for EBSD was the same ThermoFisher Scientific Apreo C used for imaging, still set to an accelerating voltage of 20 kV and a beam current of approximately 20 nA. Exposure time was automatically adjusted to ensure that the brightest point on the EBSD camera for any pattern in the scan was just below the saturation limit of the camera. No frame averaging was used for the primary EBSD maps, but a custom image processing sequence was applied to the captured patterns in APEX. This sequence consisted of dynamic background subtraction with 5 passes and a balance value of 100%, followed by image histogram normalization with a brightness value of 55% and contrast value of 75%. Figure 24 shows an example EBSD pattern captured using these settings. For each layer, an 1100\(\times \)1100 \(\upmu \)m EBSD map was captured on a square grid with a 2 \(\upmu \)m scan point spacing, covering the entire 1\(\times \)1 mm sample cross section plus a small amount of the mounting material on all sides of the sample.
Appendix E: MPSS EBSD Pattern Center Determination
Accurate representation of the EBSD sample-detector geometry, also known as the pattern center (PC), is critical for orientation accuracy when using simulation-based indexing techniques [39]. To allow for empirical PC fitting, prior to the main EBSD scan for each layer, a secondary 800\(\times \)800 \(\upmu \)m scan was captured on a square grid with an 80 \(\upmu \)m scan point spacing (this produces an 11\(\times \)11 grid in APEX, for a total of 121 scan points). This scan was roughly centered on the middle of the sample using a known X/Y offset from the main scan to ensure that patterns were only collected from the sample and not from the mounting material. For this “PC scan,” 16\(\times \) frame averaging with no other image processing was used for pattern capture. The camera exposure time was automatically adjusted before this scan in the same manner as for the main EBSD scan. All other scan parameters (including the 120\(\times \)120 pixel pattern resolution) were left unchanged between the PC scan and the main scan. Static background correction was applied in post-processing by averaging all patterns in the PC scan, dividing each pattern by this average pattern, and then re-scaling the resulting floating-point values to the full [0, 255] integer range for 8-bit image intensities. Examples of such patterns before and after correction are shown in Fig. 25, along with an example averaged pattern background.
The background-corrected PC scan data were fed into the EMsoft program EMEBSDGlobalOpt for empirical fitting of orientation and pattern center at each scan point as described by Zhu et al. [40]. The program parameters were generally set according to the recommendations from that study, except that a larger search range of 10% of the pattern width in PC space and 10\({}^{\circ }\) in orientation space was used here. To ensure robustness against stochastic variance in the PC fitting search algorithm, biases from the initial crystal orientation [41, 42], or the occasional low-quality pattern collected near surface debris or porosity, a least-squares plane fit was generated for each PC component (x*, y*, and z*) versus X and Y position in the scan. These plane fits were evaluated at the position corresponding to the center of the main EBSD scan and the resulting values used for SHI of that scan.
Appendix F: MPSS Spherical Harmonic Indexing
SHI was performed using a bandwidth of 123, a circular mask radius of 50 pixels, no Gaussian background subtraction, no adaptive histogram equalization, un-normed cross-correlation, and Newton’s method refinement following initial indexing. These parameters specifically apply to the open-source implementation of SHI “EMSphInx”; for further documentation, see [43].
While SHI uses a single PC value for analysis of the entire scan to make the processing computationally tractable, this results in fictitious crystal rotations that increase with the distance from the point at which the PC was evaluated, as the PC should actually shift slightly as the beam scans across the sample [33]. The related technique of dictionary indexing [44] has an equivalent issue, and a method for correcting this fictitious rotation in post-processing was previously developed by Singh et al. [45]. It was found that the best corrective rotation to apply to the orientation at each scan point was the rotation which mapped the position of the initial (incorrect) PC on the EBSD detector screen to the position of the true PC on the EBSD detector screen for that scan point, with both positions defined as vectors in the sample coordinate frame. Here, rather than calculating the “true” PC based off the nominal geometry of the sample surface as was done in [45], it was calculated from the empirical fits versus X and Y position in the scan that were obtained earlier (see §E). The corrective rotations were calculated and applied to the EBSD data using the open-source MATLAB package MTEX [37]. However, for dictionary indexing this was followed with another local refinement step starting from the corrected orientation and pattern center at each point, as there was still a nontrivial level of residual misorientation following the initial correction (see Figure 8 in [45]). The SHI software used here cannot currently perform this secondary refinement step and therefore presumably still has some level of fictitious rotation in the corrected data. However, comparisons with HEDM orientation data show that the level of residual misorientation present in the corrected SHI data is as good or better than that present in the conventional Hough analysis (see Fig. 12), which has its own correction algorithm for the issue of PC shift versus position built into the vendor software.
Appendix G: MPSS Data Registration Procedure
The first step in the registration workflow was to self-align the optical image series with a rigid transform, creating a 3D volume to serve as the ground-truth spatial position for the MPSS data. An initial 180\({}^{\circ }\) in-plane rotation was needed in order to align the optical images to the same spatial reference frame as the BSE and EBSD datasets. This rotation was determined by examining the location of the gold cube with respect to the sample in the different data types (see Fig. 5 and Fig. 22). In order to reduce contrast/exposure differences in the optical images, they were histogram-matched using Scikit-Image’s “match_histograms” function, with the first optical image serving as the reference histogram for the later layers to match against. From there, the images were spatially registered in series using OpenCV’s implementation of the SIFT algorithm. The default parameters were used, and feature matching was done using OpenCV’s brute force matcher, keeping the top 30% of matched features for the calculation of the transformation matrix using “cv2.estimateAffinePartial2D.” This function calculates a rigid transform, only allowing for rotation and translation. After alignment, a binary mask of the sample was automatically found using a combination of Otsu thresholding, binary hole filling, and segmenting out the binary feature with the largest area. This binary mask is used to mask out the mounting medium (Polyfast) and the gold cube while registering the BSE data against the optical data, as well as determining the sample center for later alignment with the BSE and EBSD data.
The same general SITK multi-resolution image registration framework as outlined in [2] was used for the BSE to optical and EBSD to BSE registrations, with differences in resolution and hyperparameters as needed. This framework allows for intra- and inter-modal image registration using a variety of different similarity metrics and transformation types. The parameters used were as follows: B-Spline transform, mutual information as the similarity metric [46], L-BFGS-B as the optimizer, and nearest-neighbor interpolation for final image resampling. As SITK image objects are array-based images defined on a physical-space region, each data mode’s image is provided along with its physical grid spacing ( \(\upmu \)m/pixel). This allows for data modes at different resolutions to be registered, as well as the easy resampling of higher-resolution data modes down to the lowest resolution reference grid.
The BSE images were registered against the aligned optical data using an updated version of the Python script from [2]. One difference from the earlier study is that only the BSE images and not the optical images were contrast-enhanced, as contrast enhancement of the optical images from this study emphasized features such as surface scratches that were not shared with the BSE images and resulted in a poor nonlinear registration. The BSE images were treated with Scikit-Image’s “equalize_adapthist” function to perform adaptive histogram equalization on each layer, followed by a “match_histograms” treatment, matching each sequential layer with the first BSE layer to improve grayscale consistency across the image series. Binary masks for the BSE images were then calculated using the same method as the optical images.
For the BSE to optical registration, the B-Spline parameters were a mesh size of 1x1 and a transformation order of 2. The B-Spline transform was calculated using histogram-matched optical images and contrast-enhanced BSE images and subsequently applied to the raw images and binary mask data types. As the first layer of the optical data serves as the reference location of the sample’s spatial information, the optical data were centered in physical space around the center point of the first layer’s binary mask using SITK’s “CenteredTransformInitializer” function. The centering transform from the first optical layer was then applied to each subsequent layer to ensure that any global alignment shifts due to shear or sample rotation captured in the SIFT registration of the optical images were carried through to the BSE and EBSD registration steps.
The EBSD data were then registered against the updated BSE images using the same SITK framework, with the B-Spline mesh size being updated to 3x3 as determined in [2]. Grayscale conversions of EBSD inverse pole figure (IPF) colormaps were registered against the contrast-enhanced BSE images, allowing for the nonlinear registration of grain boundary information to remove the distortions from EBSD data collection. The IPF colormaps were generated as part of a Python script using ORIX, an open-source Python EBSD analysis suite [47, 48]. ORIX’s “orientation2color” function was used to generate the IPF colormaps for each layer, while binary masks for the EBSD data were calculated using the Otsu threshold method on the confidence index values from the EBSD scan data. The BSE images were downsampled from a 2500\(\times \)2500 grid with a spacing of 0.5 \(\upmu \)m/pixel to a 600\(\times \)600 grid with a spacing of 2 \(\upmu \)m/pixel using SITK’s “Resample” function to match the SITK grid used for processing the EBSD data. Note that both the BSE and EBSD grids in SITK were larger than the original images to allow for motion during registration without loss of data; added “padding” pixels near the boundaries of the grids were filled with zero values to match the dark background of both images. The B-Spline transform was then calculated on a per-layer basis and applied to the raw EBSD orientation data files with nearest-neighbor interpolation using a custom Python input–output function. Any orientations in the exported EBSD data corresponding to padding pixels were set to the identity rotation with a confidence index of zero.
Separately from the SEM data modes, the SWLI data were also registered against the self-aligned optical images. The raw heightmap data were corrected to remove any tilt present from data collection, which was done by calculating the surface plane of the sample using least squares estimation and subtracting the resulting plane from the raw data. The tilt-corrected heightmaps were then registered against the optical images using the same SITK framework and subsequently downsampled to a 600\(\times \)600 grid with 2 \(\upmu \)m spacing to match the grid used for the corrected EBSD data, ensuring that each height value corresponded to a single EBSD data point. Figure 6 (d) and (h) shows an example of the raw and tilt-corrected heightmaps.
Appendix G.1: Registration Challenges
Challenges with the B-Spline registration of EBSD to BSE data occurred when one corner of the sample was accidentally damaged within the mount partway through the serial sectioning series. As the B-Spline transform and mutual information similarity metric operate on a pixel-by-pixel basis, the SITK optimizer distorts the moving image as needed to maximize the metric score. For the damaged region, there is no longer a clear grain structure or much shared information between the EBSD and BSE data types, causing the optimizer to incorrectly distort the damaged region with respect to the rest of the sample. In some areas of the sample, the poor distortion correction from the damaged region causes neighboring grains to be pulled away from their true locations as well. Overall, the data in the damaged region are not usable, and attempts were made to minimize its impact on the nearby regions by thresholding out low-confidence EBSD data, or potentially masking out the entire damaged region from the registration optimization. Figure 26a and Fig. 26b show the BSE and EBSD scans on section 331, at the start of the damaged region, as well as the impact of the damaged region on the final registration in Fig. 26c. As shown in Fig. 26c, there is a slight “pull” effect in the area around the damaged region, moving neighboring grains closer to the top left corner than desired. Thankfully, this region represents a relatively small fraction of the total VOI and therefore does not significantly affect the overall results.
Appendix H: Aligning 3D-EBSD and HEDM Maps
Appendix H.1: Data Import and Aligning EBSD/NF-HEDM Reference Frames
EBSD and NF-HEDM orientation data formats both define position and orientation values for each data point. Positions are usually given as distances from the origin of a chosen reference frame, while crystal orientations are generally specified as rotations relative to an “origin” reference frame. It is possible for the reference frame used to define the position values (the spatial reference frame) to be different than the reference frame used to define the rotations representing orientations (the grain reference frame), so both must be considered separately. The recorded rotations may also differ in one or more choices required when specifying a rotation in numerical form [49], such as using either the active or passive rotation convention. As all of these choices frequently vary between different data formats, cross-format comparisons require significant attention to these details to ensure that any differences have been accounted for.
We employ MTEX [37] to manipulate the EBSD and NF-HEDM orientation maps, as it has native support for many existing orientation data formats and can be extended as necessary to accommodate new ones. The (\(x_U\), \(y_U\), \(z_U\)) unified coordinate system shown in Fig. 8 is also the one used in MTEX, with the origin in the upper left corner, \(x_U\) pointing to the right, \(y_U\) pointing down, and \(z_U\) pointing into the plane (which may require adjusting the MTEX display conventions). By default, data import into MTEX includes forcing the spatial and grain reference frames to coincide by applying format-dependent rotations to either the position data or the orientation data, as using the same reference frame for both greatly simplifies calculations and comparisons across data formats. For the “.ang” format EBSD data files used here, the rotation needed to align the spatial and grain reference frames is 180\({}^{\circ }\) about \((1, -1, 0)\)Footnote 7. This rotation was applied to the orientation data, altering the grain reference frame to align with the spatial reference frame. Orientations are numerically represented in the .ang files as active rotations in the Bunge Euler angle convention.
Appendix H.2: MTEX Import of NF-HEDM Orientation Data
MTEX did not have a pre-existing import function for the “.mic” orientation data format output by MIDAS [18] for NF-HEDM reconstruction, so the correct import settings and rotations required to align the HEDM spatial and grain reference frames with the imported EBSD data had to be determined manually. As each data layer must remain in the (\(x_U\), \(y_U\)) plane and the in-plane axes must be either horizontal or vertical, the possible choices for the rotation to align the HEDM spatial reference frame to the imported EBSD data were 180\({}^{\circ }\) about \(x_U\), 180\({}^{\circ }\) about \(y_U\), any multiple of 90\({}^{\circ }\) about \(z_U\), or any combination of these rotations. Analysis of the possible combinations showed that there were only eight distinct results, with four being either a 0\({}^{\circ }\), 90\({}^{\circ }\), 180\({}^{\circ }\), or 270\({}^{\circ }\) rotation about (0, 0, 1), and the remaining four being 180\({}^{\circ }\) rotations about either (1, 0, 0), (0, 1, 0), (1, 1, 0), or \((1, -1, 0)\). The HEDM orientation data were initially imported with no changes to either the position or orientation values, and IPF colormaps were then generated from a single layer of HEDM data with each of these possible rotations applied to the position data. This was done using both the active and passive rotation conventions for orientation data import, as the convention used by the data format was initially unknown. The correct choice of rotation convention, spatial rotation, and initial layer-to-layer correspondence between the EBSD and HEDM data were all determined by finding the maximum image similarity between these IPF colormaps and the IPF colormaps from every layer of the EBSD data.
IPF Z colormaps demonstrating the sequence for aligning the HEDM and EBSD spatial reference frames. Note that despite the visibly corresponding IPF maps, a rotation must also be applied to the HEDM orientation values to align the HEDM and EBSD grain reference frames. The residual spatial rotation about \(z_U\) from the initial position of the sample in the HEDM laboratory coordinate system is also visible. This will be corrected during final alignment of the HEDM data to the EBSD data
Figure 27 illustrates the process described above using data from the layers that were found to correspond to each other (HEDM layer 1 and MPSS section 320). While the IPF map of EBSD data in Fig. 27a does not seem to correspond to the IPF map of HEDM data imported using the passive rotation convention shown in Fig. 27b, a correspondence can be seen between the larger grains at the bottom of Fig. 27a and the top of the IPF map of HEDM data imported using the active rotation convention shown in Fig. 27c. Further examination shows that the top of the former and the bottom of the latter also have corresponding grains, leading to the choice of a 180\({}^{\circ }\) rotation about \(x_U\) to align the HEDM spatial reference frame with the imported EBSD data, as shown in Fig. 27d. Visual comparison of Fig. 27a and Fig. 27d clearly shows the general correspondence between the two datasets, outside of the slight remaining rotation about \(z_U\).
It should be noted that despite the visual correspondence of the IPF maps in Fig. 27a and Fig. 27d, the underlying orientations do not actually match until the rotation required to align the grain reference frames is also applied. The apparent visual match is due to the combination of the chosen IPF direction, the cubic symmetry of the crystal lattice, and the specific alignment rotation required, which together result in no changes to the IPF colormap shown in Fig. 27d before or after alignment of the grain reference frames. The orientation mismatch can be more reliably demonstrated by directly measuring misorientation angles between the two datasets or alternately by plotting the IPF colormaps along a nonstandard axis (e.g., an arbitrary nonsymmetric direction) rather than along \(z_U\) as is done in Fig. 27. In this case, the rotation required to align the grain reference frames was determined by loading both datasets into MTEX and examining the misorientations of the HEDM orientation data relative to the EBSD orientation data in corresponding grains. It was found to be 180\({}^{\circ }\) about \(x_U\), the same as for the spatial reference frames, implying that MIDAS uses the same reference frame for both position and orientation data (unlike several common EBSD data formats, including the .ang format used here). With the spatial and grain reference frames both aligned, the HEDM orientations were within 5\({}^{\circ }\) of corresponding EBSD orientations. Most of this residual misalignment is due to the nominal 0\({}^{\circ }\) position for the HEDM and XCT scans not being fully square with the sample faces (as can be seen in Fig. 27) and is easily corrected by a rigid rotation about \(z_U\) applied to both the position and orientation data.
It is noteworthy that the initial brute-force search for spatial alignment could be avoided by comparing fiducial markers visible in both the BSE data and the XCT data to determine the rotations required to align their spatial reference frames, as well as to identify the corresponding 2D slices of EBSD and NF-HEDM data. From there, the EBSD and HEDM grain reference frames could be aligned by testing both active and passive rotation conventions to find the best match and then comparing misorientations between corresponding grains as described above. However, if using this method, care must be taken to account for any differences between the XCT and NF-HEDM reconstruction reference frames. In this study, the XCT data were already spatially aligned with the BSE data and did not need the 180\({}^{\circ }\) rotation about \(x_U\) that was required to spatially align the NF-HEDM data with the EBSD data.
Appendix H.3: Alignment and Resampling of EBSD Sections to NF-HEDM Layers
While both 3D-EBSD and NF-HEDM collect data as a series of thin layers, there was almost certainly some level of misalignment between the sectioning planes for the two methods. Given the extremely high aspect ratio of the layers used here (over 1000 \(\upmu \)m in width/height versus \(\sim 2\upmu {\hbox {m}}\) in thickness), even a relatively small misalignment (e.g., 0.3\({}^{\circ }\)) between the sectioning planes can cause a single layer of one data type to intersect three or more layers of the other type. It is useful to align and resample one of the data types into new layers to produce a one-to-one correspondence between layers from the two data modes. Final alignment can then be handled in 2D, significantly simplifying data processing. Here, we chose to resample the EBSD data into layers corresponding to the HEDM data.
3D rotational alignment of the EBSD data versus the HEDM data was done using the BSE data as a proxy for the EBSD volume and the XCT data as a proxy for the HEDM volume. Dragonfly softwareFootnote 8 was used to find the 3D rotational misalignment. The X/Y/Z axes used in Dragonfly were confirmed to match the conventions of the unified coordinate system. However, the X/Y position of the sample within the BSE image volume is different than the \(x_U\)/\(y_U\) position of the sample within the EBSD data volume due to different amounts of padding data surrounding the sample in the two data types (i.e., the sample is effectively translated to a different position relative to the origin). Similarly, the X/Y position of the sample in the XCT image volume does not match the \(x_U\)/\(y_U\) position of the sample in the HEDM data volume. The Z-position of the first HEDM layer in the XCT image volume is also partway through said volume, aligned with the bottom of the gold cube, and the appropriate Z-position for the first BSE layer relative to the XCT data is not precisely known until we perform the 3D alignment. Due to these differences, we refer to coordinates of BSE or XCT data as X/Y/Z to distinguish them from \(x_U\)/\(y_U\)/\(z_U\) coordinates of EBSD or HEDM data.
Prior to further analysis, the XCT data were rotated \(-\)4.092\({}^{\circ }\) about Z so that the sample faces were as square as possible relative to the X/Y axes, allowing tighter cropping of irrelevant XCT data and therefore a reduction in overall data size, as well as better initial alignment with the BSE data. The rotated XCT reconstruction of the sample and the registered BSE data from serial sectioning were both saved as layered TIFF images and loaded into Dragonfly, generating two 3D image volumes. The pixel dimensions for the BSE data were 0.5 \(\upmu \)m in X and Y and 2.16 \(\upmu \)m in Z, while the pixel dimensions for the XCT data were 1.172 \(\upmu \)m on all axes. Manual translation was used to bring the two volumes roughly into alignment as a starting point (shown in Fig. 28); then, the 3D image registration algorithm built into Dragonfly was used to determine the 3D rotation, translation, and scaling that best aligned the two data volumes. For alignment, the BSE data were treated as the moving image and the XCT data as the static image, with mutual information used as the registration metric. A custom Python script was used to repeat the 3D alignment across a range of starting step-size values to ensure confidence in the final results, as the iterative alignment algorithm can fail to find a good match if the initial step size(s) are either too small or too large. The final 3D alignment rotation was found to be 0.3175\({}^{\circ }\) about \((x_U, y_U, z_U)=(-0.5447, 0.5830, -0.6029)\).
MTEX 6.0 (beta release, accessed September 2023) introduced pseudo-3D EBSD capabilities, most importantly allowing for EBSD data positions to be defined with arbitrary (\(x_U\), \(y_U\), \(z_U\)) positions for each point, with no requirement for any of the positions to be on a uniform grid. A 3D point-cloud representation of the EBSD data can thus be generated using the relative layer heights (as recorded during serial sectioning; see §B) along with the surface heightmaps from the SWLI profiler. The positive direction for the imager layer position data matches \(z_U\), while the positive direction for the SWLI heightmap data is the opposite of \(z_U\). As such, for each layer of distortion-corrected EBSD data, the \(z_U\) position for each data point was set to the nominal layer position from the optical imager minus the relative height values at each point from the plane-subtracted and registered SWLI data. Distortion-corrected EBSD data layers were loaded into MTEX, set to appropriate \(z_U\) values, and concatenated to create the full 3D-EBSD volume. The 3D sample rotation found using Dragonfly is then applied to the 3D-EBSD data volume in MTEX, altering both the (\(x_U\), \(y_U\), \(z_U\)) data point positions and the crystal orientations at those points, as it represents what is presumed to be a physical rotation of the sectioning plane between the two experiments. Note that only the 3D rotation found from Dragonfly is used; the translation or scaling values from Dragonfly are not included, as those alignments will be handled later.
Having defined the rotated 3D-EBSD volume, we then had to set up a 3D grid of position values for resampling the EBSD data into new layers. As the applied rotation was relatively small and the distortion-corrected EBSD maps already contained “padding” data around the region of interest in the \(x_U\) and \(y_U\) directions, the \(x_U\)/\(y_U\) resampling grid was simply defined to match the 600\(\times \)600 point grid with 2 \(\upmu \)m spacing used for the distortion-corrected EBSD data prior to this alignment rotation. The \(z_U\) dimensions of the resampling grid were defined to match the nominal dimensions of the HEDM data, in this case 20 layers with a spacing of 2 \(\upmu \)m, which resulted in a 600\(\times \)600\(\times \)20 point resampling grid with a uniform spacing of 2 \(\upmu \)m in all directions. However, defining the exact \(z_U\) position for the first layer of the resampling grid required more effort, as it needed to correspond to the position of the first HEDM data layer in the unified coordinate system, taking into account the 3D alignment from Dragonfly.
The XCT data were only rotated about Z, so the layers remain parallel to the HEDM layers. The first HEDM layer was collected at the bottom of the gold cube marker, which is clearly visible in the XCT image volume. We therefore identified the layer of XCT data corresponding to the first layer of HEDM data and thus obtained its Z position in the Dragonfly reference frame. We then needed to determine the effective Z position of the MTEX origin in the Dragonfly reference frame. While the \(x_U\)/\(y_U\) positions of the sample in the EBSD data do not match the X/Y positions of the sample in the BSE data, the \(z_U\) positions do match the Z positions prior to any further adjustments, since the two data types were collected from the same sections of the sample. Out of the translation, rotation, and scaling applied to align the BSE image volume with the XCT image volume, the position of the BSE origin is only affected by the translation component, as the other effects are applied about the origin. Therefore, the new Z position of the transformed BSE image volume origin is simply the Z-translation value extracted from the full transformation matrix applied to it for alignment in Dragonfly. The 3D rotation applied to the EBSD data in MTEX is similarly applied about the origin and does not affect its position, so the Z-translation value from the BSE transformation matrix directly represents the Z position of the MTEX origin in the Dragonfly reference frame.
As such, the value that should be used for the \(z_U\) position of the first resampling layer in the MTEX reference frame is the difference between the Z-translation extracted from the Dragonfly transformation matrix for the aligned BSE data and the Z-position of the first HEDM data layer as measured in Dragonfly. This was found to be 18.88 \(\upmu \)m. With the position of the first layer defined, the remaining layers of the resampling grid simply follow the specified 2 \(\upmu \)m spacing in \(z_U\) starting from that point. Having fully defined the resampling grid, the rotated 3D-EBSD data were resampled onto these positions using 3D nearest-neighbor interpolation and exported as individual data files per layer.
Appendix H.4: Alignment and Resampling of NF-HEDM Layers to EBSD Grid
After aligning the spatial and grain reference frames for the EBSD and HEDM data types and resampling the EBSD data into layers directly corresponding to the HEDM data layers, a final 2D transformation is required to bring the HEDM data into full alignment with the EBSD data. Direct overlap allows for resampling of the HEDM data onto the same uniform spatial grid as the EBSD data, making point-to-point comparisons very simple. The final spatial transformation was calculated by determining pairs of matching grains between HEDM and EBSD data and calculating the necessary similarity transformation (translation, rotation, and scaling) to bring the locations of grain centroids in HEDM to the locations of matching centroids in EBSD for the first pair of layers.
An algorithm was written to pair grains present in the HEDM data to those in the EBSD data based on a set of user-selectable grain-averaged properties calculated using MTEX. The grain properties used here were grain centroid location, grain area, and grain average orientation. This version of the grain matching algorithm does depend on a prior approximate alignment of datasets so that matching grains fall within the boundary of the local grain cutoff distance and thus cannot be used on datasets where the spatial reference frames have not yet been aligned. For these datasets, the 180\({}^{\circ }\) rotation of the HEDM data about \(x_U\) was enough to bring matching grains within the cutoff radius, even without correcting the residual misalignment about \(z_U\).
A summary of the grain-matching algorithm is shown in Fig. 29. For each grain in the HEDM data, a local neighborhood of EBSD grains was selected containing all grains with centroids within 100 \(\upmu \)m. From there, the misorientations between the chosen HEDM grain and all EBSD grains within the local region are calculated. The EBSD grain with the lowest misorientation is chosen as the potentially matching grain. If this grain is not already matched to another HEDM grain and its area is within ±75% of the area of the reference HEDM grain, a matched grain pair is defined. This process is then repeated for each grain in the HEDM data. Grain-pair selections are currently not optimized beyond a single pass through this algorithm.
While this method was not able to match every grain in the HEDM data to a corresponding grain in the EBSD data, it provided enough matched data points that MATLAB’s “fitgeotform2d” function could be used to find the similarity transformation needed to align the HEDM data to the EBSD data. The transformation calculated for aligning the matched grain centroids from the first layer of HEDM data versus the first layer of resampled EBSD data was a rotation of \(-\)3.9030\({}^{\circ }\) about \(z_U\), a translation of \(-\)32.8378 \(\upmu \)m in \(x_U\) and 44.7012 \(\upmu \)m in \(y_U\), and a scaling factor of 1.0053. MTEX was used to apply this transformation to the raw HEDM data, and the transformed HEDM data were then nearest-neighbor resampled onto the same uniform \(x_U\)/\(y_U\) grid as the EBSD data. The matching grains and aligned HEDM data are shown in Fig. 30. The transformed and resampled HEDM data were then exported in .ang format for ease of use with other orientation data processing software. To maintain the HEDM data as a self-consistent volume, the same 2D transformation was applied to every layer.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sparks, G., Mason, S.A., Chapman, M.G. et al. 3D Reconstruction of a High-Energy Diffraction Microscopy Sample Using Multi-modal Serial Sectioning with High-Precision EBSD and Surface Profilometry. Integr Mater Manuf Innov (2024). https://doi.org/10.1007/s40192-024-00370-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40192-024-00370-6