Abstract
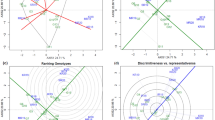
The main objective of this study was to compare the several statistical models, i.e., joint regression analysis (JRA), additive main effect and multiplicative interaction (AMMI) and genotype and genotype × environment (GE) interaction (GGE) biplot, for analyzing of GE interaction for grain yield of rainfed safflower multi-environment trials. The effectiveness of each model was compared for identifying the best performing genotypes across environments, identifying the best genotypes for mega-environment differentiation and evaluating the yield and stability performance. Grain yield data of 13 cold-tolerant safflower breeding lines along with a check cultivar grown in three rainfed research stations for two cropping seasons were used. Environment (E) main effect accounted for 57.1% of total variation, compared to 8.8 and 34.1% for G and GE interaction effects, respectively. Spearman’s rank correlation analysis indicated that the three methods (GGE biplot, AMMI analysis and JRA) were significantly correlated (P < 0.01) in ranking of genotypes for static (biological) stability, suggesting that they can be used interchangeably. All three methods identified genotypes G4 and G9 as the most stable genotypes with low-yielding performance, and the breeding line G3 as high-yielding stable genotype across environments. Based on the results, the Maragheh was an ideal test location with a demonstrated high efficiency in selecting new cultivars with a wide adaptability. The main conclusions were the similarity between the dominant genotypes in the three models. The GGE biplot was more versatile and flexible and provided a better understanding of GE interaction than the other methods. Positive increase in yield and yield stability is attributable predominately to genetic improvement in safflower breeding lines. The breeding line 415/338 could serve as a good genetic source for both high yielding and stability in safflower breeding programs for highland cold rainfed areas of Iran.


Similar content being viewed by others
References
Alwala S, Kwolek T, McPherson M, Pellow J, Meyer DA (2010) Comprehensive comparison between Eberhart and Russell joint regression and GGE biplot analyses to identify stable and high yielding maize hybrids. Field Crops Res 119:225–230
Annicchiarico P (1997) Joint regression vs AMMI analysis of genotype-environment interactions for cereals in Italy. Euphytica 94:53–62
Argiller O, Hebert Y, Barriere Y (1994) Statistical analysis and interpretation of line × environment interaction for biomass yield in maize. Agronomie 14:661–672
Becker HC, Leon J (1988) Stability analysis in plant breeding. Plant Breed 101:1–23
Blanche SB, Myers GO (2006) Identifying discriminating locations for cultivar selection in Louisiana. Crop Sci 46:946–949
Chaturvedi M, Srivastava N, Gudipati T, Bhadauria R, Khandelwal S, Das RR (2001) In vitro studies in the wild species of Carthamus—C. oxycantha L. In: Bergman J, Mündel HH (eds) Safflower: a multipurpose species with unexploited potential and world adaptability. Proceedings of the 5th international Safflower conference, Williston, ND, Sidney, MT, 23–27 July, pp 47–49
Crossa J (1990) Statistical analyses of multilocation trials. Adv Agron 44:55–85
Dimitrios B, Christos G, Jesus R, Eva B (2008) Separation of cotton cultivar testing sites based on representativeness and discriminating ability using GGE biplots. Agron J 100:1230–1236
Ebdon JS, Gauch HG (2002) Additive main effects and multiplicative interaction analysis of National Turfgrass performance trials: II. Genotype recommendation. Crop Sci 42:497–506
Eberhart SA, Russell WA (1966) Stability parameters for comparing varieties. Crop Sci 6:36–40
Fan XM, Kang MS, Chen H, Zhang Y, Tan J, Xu C (2007) Yield stability of maize hybrids evaluated in multi-environment trials in Yunnan, China. Agron J 99:220–228
Gabriel KR (1971) The bi-plot-graphical display of matrices with application to principal component analysis. Biometrika 58:453–467
Gauch HG (1992) AMMI analysis of yield trials. In: Kang MS, Gauch HG (eds) Genotype-by-environment interaction. CRC Press, Boca Raton, pp 1–40
Gauch HG (2006) Statistical analysis of yield trials by AMMI and GGE. Crop Sci 46:1488–1500
Gauch HG, Zobel RW (1997) Identifying mega-environment and targeting genotypes. Crop Sci 37:381–385
Gauch HG, Piepho HP, Annicchiarico P (2008) Statistical analysis of yield trials by AMMI and GGE: further considerations. Crop Sci 48:866–889
Goyal A, Beres BL, Randhawa HS, Navabi A, Salmon DF, Eudes F (2011) Yield stability analysis of broadly adaptive triticale germplasm in southern and central Alberta, Canada, for industrial end-use suitability. Can J Plant Sci 91:125–135
IRRI (International Rice Research Institute) (2005) IRRISTAT for Windows. Version 5, Los Baños, Philippines
Isik K, Kleinschmit J (2005) Similarities and effectiveness of test environments in selecting and deploying desirable genotypes. Theor Appl Genet 110:311–322
Kroonenberg PM (1995) Introduction to biplots for G × E tables. University of Queensland, Brisbane
Lacey DJ, Wellner N, Beaudoin F, Napier JA, Shewry PR (1998) Secondary structure of oleosins in oil bodies isolated from seeds of safflower (Carthamus tinctorius L.) and sunflower (Helianthus annuus L.). Biochem J 334:469–477
Ma BL, Yan W, Dwyer LM, Frégeau-Reid J, Voldeng HD, Dion Y, Nass H (2004) Graphic analysis of genotype, environment, Nitrogen fertilizer, and their interaction on Spring Wheat yield. Agron J 96:169–180
Mohammadi R, Pourdad SS (2009) Estimation, interrelationships and repeatability of genetic variability parameters in spring safflower using multi-environment trial data. Euphytica 165:313–324
Purchase JL (1997) Parametric analysis to describe G × E interaction and yield stability in winter wheat. Ph.D. thesis. Department of Agronomy, Faculty of Agriculture, University of the Orange Free State, Bloemfontein, South Africa
Ramaswamy NM (2001) Safflower biotechnology: progress and prospects. In: Bergman J, Mündel HH (eds) Safflower: a multipurpose species with unexploited potential and world adaptability. Proceedings of the 5th international Safflower conference, Williston, ND, Sidney MT, 23–27 July, p 59
Robins JG, Waldron BL, Vogel KP, Berdahl JD, Haferkamp MR, Jensen KB, Jones TA, Mitchell R, Kindiger BK (2007) Characterization of testing locations for developing cool-season grass species. Crop Sci 47:1004–1012
Romagosa I, Fox PN (1993) Genotype × environment interaction and adaptation. In: Hayward MD, Bosemark NO, Romagosa I (eds) Plant breeding: principles and prospects. Chapman & Hall, London, pp 373–390
Yan W (2001) GGEbiplot—a Windows application for graphical analysis of multi-environment trial data and other types of two-way data. Agron J 93:1111–1118
Yan W, Kang MS (2003) GGE biplot analysis: a graphical tool for breeders, geneticists, and agronomists. CRC Press, Boca Raton
Yan W, Tinker NA (2006) Biplot analysis of multi-environment trial data: principles and applications. Can J Plant Sci 86:623–645
Yan W, Hunt LA, Sheng Q, Szlavnics Z (2000) Cultivar evaluation and mega-environment investigation based on GGE biplot. Crop Sci 40:596–605
Yan W, Kang MS, Ma BL, Woods S, Cornelius PL (2007) GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci 47:643–653
Yau SK (1995) Regression and AMMI analysis of genotype × environment interactions: an empirical comparison. Agron J 87:121–126
Zhang Z, Lu C, Xiang ZH (1998) Stability analysis for varieties by AMMI model. Acta Agron Sin 24:304–309
Zobel RW, Wright MG, Gauch HG (1988) Statistical analysis of yield trial. Agron J 80:388–393
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alizadeh, K., Mohammadi, R., Shariati, A. et al. Comparative Analysis of Statistical Models for Evaluating Genotype × Environment Interaction in Rainfed Safflower. Agric Res 6, 455–465 (2017). https://doi.org/10.1007/s40003-017-0279-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40003-017-0279-1