Abstract
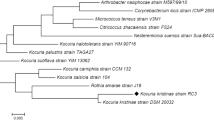
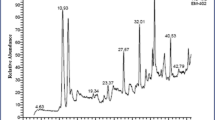
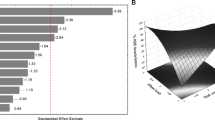
In this paper, the microbial degradation of methyl red (MR) was investigated by novel isolated bacteria S1 and K1 from wastewater effluent. Based on morphological, biochemical and molecular methods, strain S1 belonged to the Staphylococcus schleiferi subsp. coagulans GA 211 with 92.67% similarity (accession number MT573965) and strain K1 belonged to the Klebsiella michiganensis W14 with 99.85% similarity (accession number MT572941). To the best of our knowledge, this is the first report of dye degradation activity of these bacteria. The Taguchi Statistical Method (TSM) was used to optimize the experimental conditions. Taguchi’s L18 orthogonal array (OA) was performed, and the results were used to calculate signal-to-noise (S/N) ratios. According to S/N ratios, it was discovered that optimum experimental conditions are at pH 8, temperature 40°C, initial dye concentration 50 mg/l, carbon source glucose, rpm 75, inoculum 3% v/v and strain S1. Analyzing the results of the biodegradation assay by Minitab 19 statistical software, it was found that rpm and temperature are most and less effective in the MR biodegradation assay, respectively. Significant changes in the HPLC and UV-Vis spectrum of samples of strain S1 before and after biodegradation assay confirmed the complete biodegradation of MR by S1 strain.







Similar content being viewed by others
Abbreviations
- MR:
-
Methyl Red
- TSM:
-
Taguchi Statistical Method
- OA:
-
orthogonal array
- S/N:
-
signal-to-noise
- MSM:
-
mineral salt medium
- HPLC:
-
high-pressure liquid chromatography
- VP:
-
Voges Proskauer
- dB:
-
Decibel
- ANOVA:
-
Analysis of variance
- GLM:
-
General Linear Model
- Rt :
-
retention time
- ABA:
-
2-aminobenzoic acid
- DMPD:
-
N-N'-dimethyl-ρ-phenylenediamine
References
Abou-El-Souod GW, El-Sheekh MM (2016) Biodegradation of basic fuchsin and methyl red. Environ Eng Manag J 15:279–286
Adedayo O, Javadpour S, Taylor C, Anderson W, Moo-Young M (2004) Decolourization and detoxification of methyl red by aerobic bacteria from a wastewater treatment plant. World J Microb Biot 20:545–550. https://doi.org/10.1023/B:WIBI.0000043150.37318.5f
Angelova B, Avramova T, Stefanova L, Mutafov S (2008) Temperature effect on bacterial azo bond reduction kinetics: an Arrhenius plot analysis. Biodegradation 19:387–393. https://doi.org/10.1007/s10532-007-9144-4
Athreya S, Venkatesh Y (2012) Application of Taguchi method for optimization of process parameters in improving the surface roughness of lathe facing operation. IRJES 1:13–19
Ayed L, Zmantar T, Bayar S, Charef A, Achour S, Mansour HB, Mzoughi RE (2019) Potential use of probiotic consortium isolated from kefir for textile azo dye decolorization. J Microbiol Biotechnol 29:1629–1635. https://doi.org/10.4014/jmb.1906.06019
Berkane N, Meziane S, Aziri S (2020) Optimization of congo red removal from aqueous solution using Taguchi experimental design. Sep Sci Technol 55:278–288. https://doi.org/10.1080/01496395.2019.1577442
Bhattacharya A, Goyal N, Gupta A (2017) Degradation of azo dye methyl red by alkaliphilic, halotolerant Nesterenkonia lacusekhoensis EMLA3: application in alkaline and salt-rich dyeing effluent treatment. Extremophiles 21:479–90. https://doi.org/10.1007/s00792-017-0918-2
Ishrat SI, Khan ZA, Siddiquee AN, Badruddin IA, Algahtani A, Javaid S, Gupta R (2019) Optimising parameters for expanded polystyrene based pod production using Taguchi method. Mathematics 7:847. https://doi.org/10.3390/math7090847
Igimi S, Takahashi E, Mitsuoka T (1990) Staphylococcus schleiferi subsp. coagulans subsp. nov., isolated from the external auditory meatus of dogs with external ear otitis. Int J Syst Evol Microbiol 40:409–411. https://doi.org/10.1099/00207713-40-4-409
Jadhav J, Parshetti G, Kalme S, Govindwar SP (2007) Decolourization of azo dye methyl red by Saccharomyces cerevisiae MTCC 463. Chemosphere 68:394–400. https://doi.org/10.1016/j.chemosphere.2006.12.087
Jafari N, Soudi MR, Kasra-Kermanshahi R (2014) Biodegradation perspectives of azo dyes by yeasts. Microbiology 83:484–497. https://doi.org/10.1134/S0026261714050130
Kalyani D, Telke A, Dhanve R, Jadhav J (2009) Ecofriendly biodegradation and detoxification of Reactive Red 2 textile dye by newly isolated Pseudomonas sp. SUK1. J Hazard Mater 163:735–742. https://doi.org/10.1016/j.jhazmat.2008.07.020
Kaushik G, Thakur IS (2009) Isolation and characterization of distillery spent wash color reducing bacteria and process optimization by Taguchi approach. Int Biodeterior Biodegrad 63:420–426. https://doi.org/10.1016/j.ibiod.2008.11.007
Kodam K, Soojhawon I, Lokhande P, Gawai K (2005) Microbial decolorization of reactive azo dyes under aerobic conditions. World J Microb Biot 21:367–370. https://doi.org/10.1007/s11274-004-5957-z
Lin J, Zhang X, Li Z, Lei L (2010) Biodegradation of reactive blue 13 in a two-stage anaerobic/aerobic fluidized beds system with a Pseudomonas sp isolate. Bioresour Technol 101:34–40
Lewis, R. J. (Ed.). (2007). Hawley's condensed chemical dictionaryHoboken, NJ, USA: John Wiley & Sons, Inc. https://doi.org/10.1002/9780470114735
Mahmood S, Khalid A, Arshad M, Mahmood T, Crowley DE (2016) Detoxification of azo dyes by bacterial oxidoreductase enzymes. Crit Rev Biotechnol 36(4):2016. https://doi.org/10.3109/07388551.2015.1004518
Maniyam MN, Ibrahim AL, Cass AE (2020) Decolourization and biodegradation of azo dye methyl red by Rhodococcus strain UCC 0016. Environ Technol 41:71–85. https://doi.org/10.1080/09593330.2018.1491634
Misal SA, Gawai KR (2018) Azoreductase: a key player of xenobiotic metabolism. Bioresour Bioprocess. https://doi.org/10.1186/s40643-018-0206-8
Mishra S, Maiti A (2019) Optimization of process parameters to enhance the biodecolorization of Reactive Red 21 by Pseudomonas aeruginosa 23N1 Int J Environ Sci Techno 16: 6685-6698
Moutaouakkil A, Zeroual Y, Dzayri FZ, Talbi M, Lee K, Blaghen M (2003) Purification and partial characterization of azoreductase from Enterobacter agglomerans. Arch Biochem Biophys 413:139–146. https://doi.org/10.1016/S0003-9861(03)00096-1
Muthuraman G, Teng TT (2009) Extraction of methyl red from industrial wastewater using xylene as an extractant. Prog Nat Sci 19:1215–1220. https://doi.org/10.1016/j.pnsc.2009.04.002
Nachiyar CV, Rajakumar GS (2005) Purification and characterization of an oxygen insensitive azoreductase from Pseudomonas aeruginosa. Enzyme Microb Technol 36:503–509. https://doi.org/10.1016/j.enzmictec.2004.11.015
Nachiyar CV, Rajakumar GS (2003) Degradation of a tannery and textile dye, Navitan Fast Blue S5R by Pseudomonas aeruginosa. World J Microb Biot 19:609–614. https://doi.org/10.1023/A:1025159617260
Ooi T, Shibata T, Sato R, Ohno H, Kinoshita S, Thuoc TL, Taguchi S (2007) An azoreductase, aerobic NADH-dependent flavoprotein discovered from Bacillus sp.: functional expression and enzymatic characterization. Appl Microbiol Biotechnol 75:377–386. https://doi.org/10.1007/s00253-006-0836-1
Park C, Lee M, Lee B, Kim S-W, Chase HA, Lee J, Kim S (2007) Biodegradation and biosorption for decolorization of synthetic dyes by Funalia trogii. Biochem Eng J 36:59–65. https://doi.org/10.1016/j.bej.2006.06.007
Pokharia A, Ahluwalia SS (2013) Isolation and screening of dye decolorizing bacterial isolates from contaminated sites. TLIST 2:54–61
Pourbabaee A, Ramezani S, Javaheri Daneshmand H (2013) Biodegradation of malachite green by Klebsiella Terrigenaptcc 1650: the critical parameters were optimized using Taguchi optimization method. J Biorem Biodegrad 4:175. https://doi.org/10.4172/2155-6199.1000175
Ranga P, Saharan BS, Sharma D (2015) Bacterial degradation and decolorization of textile dyes by newly isolated Lysobacter sp. Afr J Microbiol Res 9:979–987. https://doi.org/10.5897/AJMR2014.7046
Saha R, Farrance CE, Verghese B, Hong S, Donofrio RS (2013) Klebsiella michiganensis sp. nov., a new bacterium isolated from a tooth brush holder. Curr Microbiol 66:72–78
Sahoo C, Gupta A, Pal A (2005) Photocatalytic degradation of methyl red dye in aqueous solutions under UV irradiation using Ag+ doped TiO2. Desalination 181:91–100. https://doi.org/10.1016/j.desal.2005.02.014
Saratale RG, Saratale GD, Chang J-S, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem E 42:138–157. https://doi.org/10.1016/j.jtice.2010.06.006
Saratale R, Saratale G, Chang J-S, Govindwar SP (2009) Decolorization and biodegradation of textile dye Navy blue HER by Trichosporon beigelii NCIM-3326. J Hazard Mater 166:1421–1428. https://doi.org/10.1016/j.jhazmat.2008.12.068
Sarayu K, Sandhya S (2010) Aerobic biodegradation pathway for remazol orange by pseudomonas aeruginosa. Appl Biochem Biotechnol 160:1241–1253. https://doi.org/10.1016/j.enzmictec.2004.11.015
Sari IP, Simarani K (2019) Decolorization of selected azo dye by Lysinibacillus fusiformis W1B6: biodegradation optimization, isotherm, and kinetic study biosorption mechanism adsorption. J Sci Technol 37:492–508
Schmidt C, Berghahn E, Ilha V, Granada C (2019) Biodegradation potential of citrobacter cultures for the removal of amaranth and congo red azo dyes. Int J Environ Sci Techno 16:6863–6872. https://doi.org/10.1007/s13762-019-02274-x
Srinivasan S, Murthy D (2009) Statistical optimization for decolorization of textile dyes using Trametes versicolor. J Hazard Mater 165:909–914. https://doi.org/10.1016/j.jhazmat.2008.10.072
Tan L, He M, Song L, Fu X, Shi S (2016) Aerobic decolorization, degradation and detoxification of azo dyes by a newly isolated salt-tolerant yeast scheffersomyces spartinae TLHS-SF1. Bioresour Technol 203:287–294. https://doi.org/10.1016/j.biortech.2015.12.058
Vijaya P, Sandhya S (2003) Decolorization and complete degradation of methyl red by a mixed culture. Environmentalist 23:145–149. https://doi.org/10.1023/A:1024839805387
Vinoda B, Vinuth M, Bodke Y, Manjanna J (2015) Photocatalytic degradation of toxic methyl red dye using silica nanoparticles synthesized from rice husk ash. J Environ Anal Toxicol 5(2161–0525):1000336. https://doi.org/10.4172/2161-0525.1000336
Wiklund P, Bergman J (2006) The chemistry of anthranilic acid. Curr Org Synth 3(3):379–402. https://doi.org/10.2174/157017906777934926
Wong P, Yuen P (1996) Decolorization and biodegradation of methyl red by Klebsiella pneumoniae RS-13. Water Res 30:1736–1744. https://doi.org/10.1016/0043-1354(96)00067-X
Zhao M, Sun PF, Du LN, Wang G, Jia XM, Zhao YH (2014) Biodegradation of methyl red by Bacillus sp. strain UN2: decolorization capacity, metabolites characterization, and enzyme analysis. Environ Sci Pollut Res 21:6136–6145. https://doi.org/10.1007/s11356-014-2579-3
Acknowledgments
The research leading to these results has received funding from the Ferdowsi University of Mashhad, Iran (grant number 52048). Authors express sincere gratitude to Dr. Marjan E. Shabestari (Department of Materials Science and Engineering & Chemical Engineering, Polytechnic School, Carlos III University of Madrid), Ehsan N. Kalali (IMDEA Materials Institute, Getafe, Madrid) for their scientific advice during the project. We also appreciate the helpful advisory of Dr. Mohammad Taghiabadi (Department of chemistry, Ferdowsi University of Mashhad, Iran.
Funding
The research leading to these results has received funding from the Ferdowsi University of Mashhad, Iran (Grant Number 52048).
Author information
Authors and Affiliations
Contributions
MB conceived and designed research. AM conducted experiments and HD helped him. AM, MG and MM analyzed data. MB, AM and MRS wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
I, as the corresponding author, declare, on behalf of all authors of the paper, that no financial conflict of interest exists in relation to the work described. Also, I, as the corresponding author, declare that the results/data/figures in this manuscript have not been published nor are they under consideration for publication elsewhere.
Availability of data and material
The authors confirm that the data supporting the findings of this study are available within the article.
Additional information
Editorial responsibility: A. Grobelak.
Rights and permissions
About this article
Cite this article
Marvi-Mashhadi, A., Sharifmoghadam, M.R., Goharimanesh, M. et al. Methyl red biodegradation based on Taguchi method by two novel bacteria. Int. J. Environ. Sci. Technol. 19, 1357–1368 (2022). https://doi.org/10.1007/s13762-021-03264-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-021-03264-8