Abstract
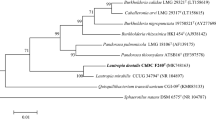
Isolate W14T recovered from a household tooth brush holder was found to be gram-negative, a facultative anaerobic, non-motile, capsulated, and a non-endospore-forming straight rod. Based on phylogenetic analysis with 16S rRNA gene sequence, isolate W14T was affiliated to the genus Klebsiella. The closest phylogenetic relative was K. oxytoca with 99 % similarity in the 16S rRNA gene sequence. The major whole-cell fatty acids were C16:0 (31.23 %), C18:1ω6c/C18:1ω7c (21.10 %), and C16:1ω7c/C16:1ω6c (19.05 %). The sequence similarities of isolate W14T based on rpoB, gyrA, and gyrB were 97, 98, and 98 % with K. oxytoca, and 97, 93, and 90 % with K. mobilis (=Enterobacter aerogenes), respectively. The ribotyping pattern showed a 0.46 similarity with K. oxytoca ATCC 13182T and 0.24 with K. mobilis ATCC 13048T. The DNA G+C content of isolate W14T was 54.6 mol%. The DNA–DNA relatedness was 55.7 % with K. oxytoca ATCC 13182T. Using the identification technology of MALDI-TOF mass spectrometry, the top matches for this isolate were K. oxytoca ATCC 13182T (Match Factor Score 1.998) and K. mobilis (Score 1.797). On the basis of phenotypic, biochemical, chemotaxonomic, and molecular studies, isolate W14T could be differentiated from other members of the genus Klebsiella including K. mobilis. Therefore, it is proposed that isolate W14T (=ATCC BAA-2403T=DSM 25444T) should be classified as the type strain of a novel species of the genus Klebsiella, K. michiganensis sp. nov.




Similar content being viewed by others
References
Barrow GL, Feltham RKA (2004) Cowan and steel’s manual for the identification of medical bacteria, 3rd edn. Cambridge University Press, Cambridge
Brisse S, Verhoef J (2001) Phylogenetic diversity of Klebsiella pneumoniae and Klebsiella oxytoca clinical isolates revealed by randomly amplified polymorphic DNA, gyrA and parC gene sequencing and automated ribotyping. Int J Syst Evol Microbiol 51:915–924
Brosius J, Palmer ML, Kennedy PJ et al (1978) Complete nucleotide sequence of a 16S ribosomal gene from Escherichia coli. Proc Natl Acad Sci 75:4801–4805
Cashion P, Holder-Franklin MA, McCully J et al (1977) A rapid method for base determination of bacterial DNA. Anal Biochem 81:461–466
Cole JR, Chai B, Marsh TL et al (2003) The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res 31:442–443
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Drancourt M, Bollet C, Carta A et al (2001) Phylogenetic analyses of Klebsiella species delineate Klebsiella and Raoultella gen. nov., with description of Raoultella ornithinolytica comb. nov., Raoultella terrigena comb. nov. and Raoultella planticola comb. nov. Int J Syst Evol Microbiol 51:925–932
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol 4:184–192
Kämpfer P, Ruppel S, Remus S (2005) Enterobacter radicincitans sp. nov., a plant growth promoting species of the family Enterobacteriaceae. Syst Appl Microbiol 28:213–221
Kimura M (1980) A simple method for estimating evolutionary rates of base substitution through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kovtunovych G, Lytvynenko T, Negrutska V et al (2003) Identification of Klebsiella oxytoca using a specific PCR assay targeting the polygalacturonase pehX gene. Res Microbiol 154:587–592
Li X, Zhang D, Chen F et al (2004) Klebsiella singaporensis sp. nov., a novel isomaltulose-producing bacterium. Int J Syst Evol Microbiol 54:2131–2136
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Mollet C, Drancourt M, Raoult D (1997) rpoB sequence analysis as a novel basis for bacterial identification. Mol Microbiol 26:1005–1011
Rosenblueth M, Martinez L, Silva J et al (2004) Klebsiella variicola, a novel species with clinical and plant associated isolates. Syst Appl Microbiol 27:27–35
Sistanich D, Sherriff M (2005) The RiboPrinter® characterization system: an overview of automated ribotyping. In: Miller MJ (ed) Encyclopedia of rapid microbiological methods, vol 3. DHI Publishing, PDA, Bethesda
Stephan R, Van Trappen S, Cleenwerck I et al (2007) Enterobacter turicensis, sp. nov. and Enterobacter helveticus, sp. nov., isolated from fruit powder. Int J Syst Evol Microbiol 57:820–826
Tamura K, Peterson D, Peterson N et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F et al (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Wayne LG, Brenner DJ, Colwell RR et al (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Yamamoto S, Kasai H, Arnold DL et al (2000) Phylogeny of the genus Pseudomonas: intrageneric structure reconstructed from the nucleotide sequences of gyrB and rpoD genes. Microbiol 146:2385–2394
Acknowledgments
The authors would like to extend their sincere thanks to NSF International and Accugenix Inc. for providing funding and facilities for the project and also to Jodie Lee of Bacteriology at American Type Culture Collection (ATCC), Virginia, USA for excellent technical support.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank accession numbers for the 16S rRNA gene, rpoB, gyrB, and the gyrA nucleotide sequences for strain W14T are JQ070300, JQ269337, JQ284304, and JQ990329.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saha, R., Farrance, C.E., Verghese, B. et al. Klebsiella michiganensis sp. nov., A New Bacterium Isolated from a Tooth Brush Holder. Curr Microbiol 66, 72–78 (2013). https://doi.org/10.1007/s00284-012-0245-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-012-0245-x