Abstract
Wheat stripe/yellow rust (WYR), caused by Puccinia striiformis f. sp. tritici (Pst), is a major constraint in global wheat production. A set of 766 hexaploid synthetic wheat lines, including primary crosses of Triticum turgidum x Aegilops tauschii and their derivatives, were screened in artificially rust inoculated field nurseries for three seasons. From this set, a core set of 94 non-lodging lines with unique pedigrees and resistance to Pst that was consistent across years was established. The core set was tested for adult plant field response under field conditions for three seasons in Australia and at least one crop season in Ethiopia, India, Kenya, Nepal and Pakistan. It was also challenged with an array of well-defined Pst pathotypes at seedling growth stages in the greenhouse, and genotyped with molecular markers linked to the adult plant resistance (APR) genes Yr18, Yr36 and Yr46. Combined analysis of field rust responses, multi-pathotype seedling phenotyping and marker genotyping resolved seven classes of Pst resistance: uncatalogued (new) APR (UAPR, 11%), uncatalogued seedling resistance (USR, 46%), known seedling resistance (KSR, 5%), KSR + USR (2%), Yr18 + UAPR (4%), Yr18 + USR (29%) and Yr18 + KSR (3%). A majority of the lines carrying UAPR and USR either singly or in combination showed high levels of field resistance across all field sites and years of testing, demonstrating that these lines represent a valuable resource for breeding wheat for resistance to Pst.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bread wheat (Triticum aestivum) and durum wheat (T. durum) are grown worldwide and provide important sources of food and nutrition for humans, providing about 19–20% of the total global dietary calories and 21% of protein (Braun et al. 2010; Tesfaye 2021). About 2.5 billion people eat wheat and its cultivation covers more than 218 million hectares of farmland worldwide under a wide range of climatic conditions (Listman and Ordóñez 2019). World wheat production in 2020 was estimated at 761.7 million tonnes, almost comparable to the world wheat production in 2019 (FAO 2020). World population is expected to reach 9.8 billion in the 2050s, approximately 2.2 billion more than in 2018 (United Nations 2017). To meet projected growing demand, wheat production must increase by at least 60% by 2050 (Wani et al. 2021). As per the FAO’s 2020 global report on crop prospects and global food supply, Pakistan was projected to import 1.7 million tonnes of wheat in 2020/21 due to tight domestic availability, despite it being traditionally a wheat exporting country. Ethiopia, India and Nepal were predicted to import 1940, 718.5 and 1260 thousand tonnes of cereals, respectively, during 2019/2020 or 2020 (FAO 2020).
The wheat stripe/yellow rust (WYR) causing pathogen Puccinia striiformis f. sp. tritici (Pst) is one of the major biotic limitations to global wheat production and has the potential to cause major yield losses wherever wheat is grown. Epidemics of WYR have resulted in yield losses as high as 100% (Wellings 2011; Chen 2020). Beddow et al. (2015) estimated that 88% of the world’s wheat crops were vulnerable to Pst infection, with annual loss estimates of 5.47 million tonnes of wheat worth US$979 million.
WYR can be successfully managed within and between cropping cycles by means of genetic resistance, reducing or even eliminating the need for fungicide intervention and resulting in increased productivity and a cleaner environment. To give some idea of the value of this approach, it was estimated in 2009 that genetic protection from WYR in Australia returns some $431 million annually (Murray and Brennan 2009). An example of the benefit of genetic resistance in controlling WYR in Australia is demonstrated by the much lower impact of Pst pathotype 198E16A-J+T+17+, a member of the PstS13 genetic lineage that caused widespread yield losses in Argentina and Europe (Park et al. 2020; Ding et al. 2021).
So far, 89 alleles at 83 loci governing resistance to Pst have been catalogued in wheat (KOMUGI 2023). Of these, 62 are regarded as major genes, conferring all stage resistance (ASR), and 27 as minor genes that confer adult plant resistance (APR). Of these, five ASR ((Yr5, Yr7, YrSP) (Marchal et al. 2018)), ((Yr10 (Liu et al. 2014)) and Yr15 (Klymiuk et al. 2018)), two APR ((Yr18 (Krattinger et al. 2009) and Yr46 (Moore et al. 2015)) and one HTP (Yr36 (Fu et al. 2009)) genes have been cloned so far.
Most of the ASR genes conferring resistance to Pst that have been catalogued so far have proven race specific (Chen 2005). Virulence for most of the commercially harnessed ASR genes (Yr1, Yr2, Yr3, Yr6, Yr7, Yr8, Yr9, Yr17, Yr19, Yr20, Yr21, Yr22, Yr23, Yr24, Yr25, Yr26, Yr27, Yr28, Yr31, Yr32, Yr33, Yr35, Yr43, Yr44, Yr76) has been reported from at least one or more wheat producing country (Calonnec and Johnson 1998; Chen 2005; Chen et al. 2010; Holtz et al. 2013; Kumar et al. 1988; Line and Qayoum 1992; Liu et al. 2016; McIntosh et al. 1995; Park et al. 2020; Sharma-Poudyal et al. 2013; Stubbs 1985; Wan and Chen 2012, 2014; Wan et al. 2004, 2016, 2017; Wellings and Burdon 1992; Wellings et al. 2009; Yang et al. 2013). However, virulence has been rarely recorded for Yr5 in Australia, India, Tajikistan and Turkey (Wellings et al. 2009) and for Yr10 in Chile, China, Ethiopia, Kenya, Hungary, Pakistan and Uzbekistan (Sharma-Poudyal et al. 2013; Wan et al. 2017) and Australia. Virulence for ASR gene Yr15 has been reported only from Afghanistan (Van Silfhout 1989).
In contrast to ASR genes, APR genes have proven to have a greater tendency to be non-race specific and durable, however they can be less effective at protecting against yield loss under high disease pressure (Bariana and McIntosh 1995; McIntosh et al. 1995; Chen and Line 1995; Johnson 1988; Chen 2013). High levels of durable rust resistance can be achieved by incorporating different minor genes. This strategy of combining APR genes with additive effect has been practiced successfully in Australia, to breed rust resistant cultivars (Singh et al. 2001).
The ability of Pst to evolve and overcome resistance has resulted in many catalogued ASR genes being rendered ineffective. Although APR genes have proven to be more durable, fewer of these are available to wheat breeders. In view of this, we conducted detailed tests of a large set of synthetic wheat lines (produced by crossing Aegilops tauschii Coss. (= Ae. squarrosa) with tetraploid wheat), which were previously reported to carry high and diverse levels of stripe rust resistance (Kishii et al. 2019).
Materials and methods
Germplasm
A set of 766 hexaploid synthetic wheat lines including primary crosses of T. turgidum x Ae. tauschii Coss. (= Ae. squarrosa L.) and their derivatives, developed at CIMMYT (Mujeeb-Kazi et al. 1996). were sourced from the “Australian Grains Genebank”. All lines were screened for rust response in artificially rust inoculated field nurseries for three consecutive seasons at the Plant Breeding Institute (PBI) Cobbitty. Of these, 591 lines were consistently resistant to Pst, from which a core set of 94 non-lodging lines with unique pedigrees was established (Table 1). The control genotypes AvocetS, and two lines near isogenic to AvocetS (NILs) carrying the APR genes Yr18 (AvocetS +Yr18) or Yr29 (AvocetS +Yr29) were sourced from the University of Sydney PBI germplasm collection. The 94 core set lines were increased by selecting a single representative plant from each, with the seed thus derived used for testing across environments, in greenhouse multi-pathotype testing, and for DNA used in genotyping for markers linked with the genes Yr18, Yr36 and Yr46.
Pathogen material
For seedling tests, nine Australian pathotypes of Pst (104E137A+, 110E143A+, 134E16A+, 134E16A+J+Yr27+, 134E16A+17+27+, 134E16A+J+T+, 150E16A+, 239E237A-17+33+ and 198E16A-J+T+17+) were used (Table 2). Field nurseries were inoculated with a mixture of two or more of these pathotypes (Table 2).
Greenhouse rust screening
For greenhouse tests, all lines along with differentials (Table 3) were planted in pots filled with a mixture of fine bark and coarse sand and fertilised using “Aquasol®” (100 gm per 10 L of water per 200 pots) prior to sowing. Seedlings of differentials and synthetic wheat lines were raised in 9 cm diameter pots by sowing four clumps (test lines) of each genotype at 8–10 seeds per clump. Following sowing, pots were kept in a growth room at 20 ± 2 °C for germination. Seven-day old seedlings were fertilised with granular urea using “Incitec Pivot” w/w 46% nitrogen (50 gm per 10 L of water per 200 pots). Seedlings at the 1.5–2 leaf growth stage (10–12 days old) were inoculated with a suspension of Pst urediniospores (2 mg urediniospores/1.0 ml of light mineral oil; Univar Solvent Naphtha L 100), using an airbrush attached to a motorized compressor. Following inoculation, plants were transferred to an incubation cabinet fitted with an ultrasonic humidifier that maintained greater than 95% relative humidity inside a dark room maintained at 10 °C darkness. After 24 h, plants were moved to a naturally lit microclimate room maintained at 18 ± 2 °C.
Field rust screening
All lines were field tested in Australia (Karalee; 2016 and Horse Unit; 2017 to 2020), Ethiopia (Kulumsa; 2020), India (Karnal; 2020), Kenya (Njoro; 2019 to 2020), Nepal (Khumaltar; 2020) and Pakistan (Islamabad and Nowshehra; 2020). One-meter row plots were sown during June (Australia), May (Ethiopia and Kenya) and in November (India, Nepal and Pakistan). A row of susceptible spreader (mixture of stripe rust-susceptible local genotypes) was sown after every fifth test line row to facilitate the build-up and uniform distribution of Pst inoculum. Four weeks after sowing, plots were fertilised using granular urea (w/w 46% nitrogen @ 100 kg/hectare) followed by irrigation. In Australia, plots were irrigated twice a week or as required, using fixed sprinklers. In other countries, plots were irrigated using flood irrigation.
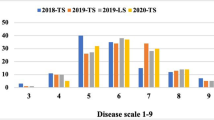
In Australia, field epidemics of stripe rust were created following the procedures described by McIntosh et al. (1995). Urediniospores (30–40 mg) were suspended in 1.5 L of light mineral oil (Shellsol®, Mobil Oil) and sprayed over buffer/spreader lines with an ultra-low-volume applicator (Microfit®, Micron Sprayer Ltd., UK). Four to five inoculations were performed during late evening on days that had a strong forecast of overnight dew. On the first and second inoculations, hot spots of disease were also established by placing susceptible, greenhouse inoculated Morocco seedlings raised in 9 cm diameter pots within the plots. In all other countries, epidemics of stripe rust were dependent on natural pathogen inoculum. Adult plant stripe rust response was scored at the flag leaf growth stage in all the crop seasons using 1–9 scale developed by Sandhu et al. (2021), as documented in Fig. 1 and Table 4.
Field stripe rust scoring scale (1–9). Source: Sandhu et al. (2021)
Molecular analysis
DNA extraction
Genomic DNA was extracted from leaf tissue using a CTAB (cetyltrimethylammonium bromide) protocol (Doyle and Doyle 1990). A 15 to 20 mm sample of leaf tissue was collected from 10–12 day old seedlings into 2 ml Eppendorf tubes from each test line and controls. The tubes were kept for 96 h above silica beads to dry the leaf tissue. Two small stainless steel ball bearings were added per tube and dried leaves were crushed to powder using a Retsch MM300 Mixer Mill (Retsch, Germany) for 3 min at 25 rpm. Pre-warmed (65 °C) 700 µl of CTAB extraction buffer was added per tube. Samples were incubated for 30 min at 65 °C and tubes were shaken vigorously after every 10 min of incubation. An equal amount of Chloroform: phenol (700 µl, 24:1 v/v) was added per tube and the contents were mixed properly by inverting each tube at least 100 times. Samples were centrifuged for 15 min at 12,000 rpm and 650 µl of supernatant was transferred to new 1.5 ml tubes. An equal volume (750 µl) of chilled (-20 °C) isopropanol was added per tube and the supernatant was mixed thoroughly by inverting tubes several times. The tubes were placed in a freezer (-20 °C) for 10 min to precipitate DNA and then centrifuged at 10,000 rpm for 10 min to pellet the DNA. The supernatant was discarded, and the pellet was washed with 500 µl of of wash buffer (76% v/v ethanol and 10 mM ammonium acetate). The DNA pellet was air dried and re-suspended in 200 µl of 10 mM Tris–HCl (pH 8.0). Rnase A @ 20 µl per 40 ml of 10 mM Tris–HCl was added before the re-suspension of the DNA pellet. Tubes were kept in an oven (37 °C) for 2 h to dissolve the DNA pellet properly. DNA was quantified using a Nanodrop ND-1000 spectrophotometer (Nanodrop® Technologies). Working dilutions (30 ng/µl) of all samples were prepared from these stocks, using doubled distilled autoclaved water (ddH2O).
Genotyping using molecular markers
Core set and control lines were genotyped with three gel-based markers and one Kompetitive allele specific PCR (KASP) marker, each linked to a stripe rust resistance gene. The codominant microsatellite marker csLV34 (Lagudah et al. 2006) linked with Yr18, gene specific WKS1 and WKS2 markers (Fu et al. 2009; Huang et al. 2016) linked with Yr36, and KASP marker TM4 linked with Yr46 (Moore et al. 2015) were used for genotyping. Details of all markers and PCR profiles used in this study are provided in Table 5.
For gel-based markers, PCR was performed using 10 µl of reaction volume containing 3.0 µl of genomic DNA (30 ng µl–1), 2.0 µl of 5 × MyFi™ (Bioline Australia Pty. Ltd. Alexandria NSW) reaction buffer formulation (containing dNTPs and MgCl2), 0.25 µl of each forward and reverse primer, 0.1 µl (5 U µl–1) of Taq (MyFi™ DNA Polymerase, Bioline Australia Pty. Ltd. Alexandria NSW) and 4.4 µl of ddH2O. Reactions were performed in a 96-well DNA thermocycler (Eppendorf AG 22331 Mastercycler, Hamburg, Germany). PCR products were resolved on 2% (w v-1) agarose (Agarose, Molecular Grade, Bioline Australia Pty. Ltd. Alexandria NSW) gels at 110 V electrophoresis for 1.5 h. For staining, 2.0 µl of GelRed™ (Biotium Inc., CA, USA) was added per 100 ml of gel solution. One hundred bp HyperLadder™ IV (Bioline Australia Pty. Ltd. Alexandria NSW) was used as molecular size marker. The separated fragments were visualized under an ultraviolet light unit fitted with a GelDoc-IT UVP camera.
KASP assays were performed using 8.0 μl of reaction volume containing 3.0 µl of genomic DNA (30 ng µl–1), 4.0 μl of 2 × KASP-TF Master Mix [(optimised buffer containing universal fluorescence resonance energy transfer assettes for FAM and HEX, ROX™ passive reference dye, Taq polymerase, nucleotides and MgCl2), (LGC Biosearch Technologies, USA)], 0.11 μl primer mix (mixture of 12 μM each allele-specific A1 and A2 primers and 30 μM of common reverse primer) and 0.89 μl of autoclaved ddH2O. PCR reactions were performed in T100™ thermal cycler (BioRad, USA) using a 96-well PCR plate. Amplification conditions included 15 min at 94 °C; 10 touchdown (TD) cycles of 20 s at 94 °C, 60 s at 65–57 °C (dropping annealing temperature by 0.8 °C per cycle); and 35–38 cycles of 20 s at 94 °C, 60 s at 57 °C. End product fluorescence readings were performed at 25 °C for 30 s using a CFX96 Touch™ real-time PCR detection system (BioRad, USA).
Results
None of the 94 lines tested positive for markers WKS1 and WKS2 linked to Yr36, and KASP marker TM4 linked with Yr46 (Fig. 2), indicating that they lacked these genes. Based on overall combined field rust screening tests, greenhouse multi-pathotype testing using nine Australian pathotypes of Pst (Table 3) and genotyping with APR genes linked markers (Table 5), the core set fell into seven categories (Fig. 3): uncatalogued APR (UAPR, 11%); uncatalogued seedling resistance (USR, 46%); known seedling resistance (KSR, 5%); KSR + USR (2%); Yr18 + UAPR (4%); Yr18 + USR (29%); and Yr18 + KSR (3%).
Multi-pathotype seedling tests and marker analysis
Multi-pathotype testing with nine Pst pathotypes revealed that 14 lines (AUS30288, AUS30332, AUS30509, AUS30515, AUS30577, AUS33403, AUS33406, AUS34091, AUS34097, AUS34148, AUS30619, AUS33376, AUS33381, AUS34169) did not carry any detectable seedling resistance to Pst in Australia (Table 3). All 14 lines were resistant to Pst across all environments, and were hence concluded to carry uncatalogued (new) APR (UAPR) to Pst that was effective under Australian conditions and possibly elsewhere (Table 6). Genotyping of these lines revealed that four (AUS30619, AUS33376, AUS33381, AUS34169) carried the Yr18 linked marker. Given the higher resistant response of these four lines as compared to the NIL AvocetS +Yr18, these lines were concluded to carry additional uncatalogued APR (UAPR). Lines AUS30288, AUS30515 and AUS34097 were more resistant than the other lines carrying UAPR (Table 6).
Seventy lines were seedling resistant to all nine test pathotypes. These lines could carry one or more ASR gene(s) that are effective to all pathotypes (eg. Yr15) or uncatalogued ASR (Table 3). The resistance in these lines was highly effective in the field in Australia, and although this masked the potential presence of APR, marker genotyping indicated the presence of Yr18 in all.
Line AUS30398 was susceptible to the Yr17-virulent pathotypes 134E16A+17+27+, 239E237A-17+33+, and 198E16A-J+T+17+, and resistant to all other six pathotypes used (Table 3), indicating that it likely carries Yr17. Similarly, the remaining nine lines (AUS30538, AUS30615, AUS30265, AUS30266, AUS30282, AUS30283, AUS30336, AUS30508, AUS30549) showed compatible infection types against at least one of the nine pathotypes and were postulated to carry known seedling resistance genes including Yr1 or Yr32 or Yr33 (AUS30538), Yr9 (AUS30615) and Yr10 (AUS30265, AUS30266, AUS30282, AUS30283, AUS30336, AUS30508, AUS30549). Of the 70 lines carrying USR, 27 carried the Yr18 linked marker (Table 7) and were also resistant in the field. Ten lines with no detectable ASR that lacked markers for Yr18 and Yr46 carried low to high levels of resistance in the field indicating that they likely carry APR distinct from Yr18. Another four lines (AUS30619, AUS33376, AUS33381and AUS34169) with no ASR in the background and carrying the Yr18 linked marker were more resistant as compared to Yr18 alone (Table 6), indicating that the UAPR carried by these lines is distinct from APR gene Yr18.
Multi-site field evaluation
The number of lines expressing high levels (VR–RMR) of resistance to Pst under field conditions varied across the different environments: Australia (92 lines), Ethiopia (30), India (63), Kenya (74), Nepal (48) and Pakistan (32)) (Fig. 4). The remaining lines expressed medium (MR-MS) to low levels (MS-S) of resistance to Pst that also varied between countries. The highest number of lines scored MR-MS occurred in Ethiopia (49 lines), followed by India (31), Nepal (17), Kenya (14), Pakistan (11) and Australia (2). A higher number of lines expressed low levels of Pst resistance under field conditions in Nepal and Pakistan, whereas only two and 14 lines were recorded in this category in Kenya and Ethiopia, respectively (Fig. 4). No line expressed low levels of resistance in Australia or in India (Karnal).
Eight lines (AUS30265, AUS30266, AUS30268, AUS30282, AUS30283, AUS30288, AUS30320 and AUS30505) were scored as VR–RMR (1–3) and 17 lines (AUS30269, AUS30273, AUS30279, AUS30336, AUS30341, AUS30353, AUS30538, AUS30552, AUS30615, AUS30617, AUS30635, AUS33388, AUS34097, AUS34116, AUS34142, AUS34198, AUS34219) as MR–MS (4–6) across all environments and all years tested (Table 8).
Discussion
Continued evolution in Pst globally has resulted in the emergence of virulence for many of the known and deployed ASR genes along with previously detected virulences for other ASR genes (eg Yr1, Yr2, Yr3, Yr6, Yr7, Yr8, Yr9, Yr17, Yr19, Yr20, Yr21, Yr22, Yr23, Yr24, Yr25, Yr26, Yr27, Yr28, Yr31, Yr32, Yr33, Yr35, Yr43, Yr44, Yr76; Calonnec and Johnson 1998; Chen 2005; Chen et al. 2010; Holtz et al. 2013; Kumar et al. 1988; Line and Qayoum 1992; Liu et al. 2016; McIntosh et al. 1995; Park et al. 2020; Sharma-Poudyal et al. 2013; Stubbs 1985; Wan and Chen 2012, 2014; Wan et al. 2004, 2016, 2017; Wellings and Burdon 1992; Wellings et al. 2009; Yang et al. 2013) and also combined virulence for important gene combinations (eg. Yr17 plus Yr33; Park et al. 2020). This highlights the importance of finding and deploying new durable sources of resistance to manage WYR. Synthetic hexaploid wheats developed by crossing A. tauschii with tetraploid wheat have been considered an important means of introducing new genetic diversity into the hexaploid wheat germplasm pool to combat abiotic and biotic stresses (Börner et al. 2015; Kishii et al. 2019). The present study targeted a collection of such synthetic hexaploid wheats to search for new sources of resistance to Pst.
We initially tested a set of 766 synthetic wheat lines for responses to Australian pathotypes of Pst. All lines were tested as seedlings under greenhouse conditions for ASR and for three years in Australia in field trials that were artificially inoculated with defined pathotypes of Pst. A core set was identified as carrying APR under Australian conditions based on field responses and seedling tests of ASR with nine Australian pathotypes of Pst. Characterising the ASR in the lines was important as it allowed the identification of lines carrying APR under Australian conditions. Comparative analyses of results from both greenhouse multi-pathotype testing and field-testing revealed the presence of APR in 14 lines AUS30288, AUS30332, AUS30509, AUS30515, AUS30577, AUS30619, AUS33376, AUS33381, AUS33403, AUS33406, AUS34091, AUS34097, AUS34148, and AUS34169, all of which did not carry any seedling resistance against any of the nine Australian pathotypes of Pst but were resistant in the field in all countries. While four of these lines were found to carry Yr18, it is likely that the remaining 10 carry uncatalogued new APR that is effective across the environments of eastern Africa, Oceania and south Asia. Exceptionally, three lines (AUS30288, AUS30515 and AUS34097) were found to be highly resistant across the field environments with a maximum of 1–3 field rust scores and it is predicted that these lines likely carry a combination of two or more resistance loci providing high levels of broadly effective APR. A combination of an APR gene Yr62 and a slow rusting QTL has been reported to provide high levels of APR to stripe rust at high temperature in spring wheat line PI 192252 (Lu et al. 2014). Similarly, Uauy et al. (2005) reported that wheat breeding lines developed from the backcrossing of Yr36 source ‘Glupro’ with ‘Anza’, and carrying both APR genes Yr18 and Yr36, expressed high levels of WYR resistance in comparison to genotypes carrying Yr18 alone. Lines AUS30288, AUS30515 and AUS34097 detected in this study represent useful sources of broadly effective APR for stripe rust for wheat breeding programs of countries in eastern Africa, Oceania and south Asia.
Four lines AUS30619, AUS33376, AUS33381 and AUS34169 carrying a combination of the Yr18 linked marker and UAPR expressed high to medium levels resistance across the environments, but lines AUS30619, AUS33376 and AUS33381 expressed low levels of resistance in Nepal and Pakistan. As Yr18 alone (NIL Avocet S+Yr18) provided low levels of resistance across all the environments, the expression of high to medium levels resistance in these four lines is most likely derived from the additive effect of Yr18 with the UAPR present in each. Previously, Singh et al. (2011) achieved high levels of stripe rust resistance by combining APR genes in different spring wheats. They advocated the pyramiding of 4–5 APR genes for accomplishing durable resistance against both stripe rust and leaf rust in wheat. In this context, synthetic wheat lines AUS30619, AUS33376, AUS33381, and AUS34169 carrying a combination of Yr18 and UAPR are useful sources for attaining durable stripe rust resistance.
So far, only one APR gene Yr18, which was derived from an Asian wheat landrace, has been used widely in modern wheat cultivars (McIntosh et al. 1995; Kolmer et al. 2008). Another APR gene Yr46, though widely distributed in wheat landraces in India and Pakistan (Moore et al. 2015) is not commonly used in the modern wheat cultivars grown in this region. Consequently, there is a dearth of APR genes for stripe rust resistance and the sources identified in this study are important contributors to fill this gap. In addition to the rust resistance, these synthetic wheat lines can also be used to improve much needed genetic diversity for other traits in bread wheat. Because the bulk of synthetic wheat lines characterised to carry UAPR or USR either singly or in combination with Yr18 showed high levels of resistance against Pst across the field conditions of Australia, Ethiopia, India, Nepal, and Pakistan, the lines represent an important source of new broadly effective stripe rust resistance for wheat improvement globally.
Data availability statement
The authors confirm that the research data and supporting information on this study are available within the article and in the form of its supplementary files. Seed of these lines can be provided on request and under SMTA.
References
Bariana HS, McIntosh RA (1995) Genetics of adult plant stripe rust resistance in four Australian wheats and the French cultivar ‘Hybride-Bersee.’ Plant Breed 14:485–491
Beddow JM, Pardey PG, Chai Y, Hurley TM, Kriticos DJ, Braun HJ, Park RF, Cuddy WS, Yonow T (2015) Research investment implications of shifts in the global geography of wheat stripe rust. Nature Plants 1:1–5
Börner A, Ogbonnaya FC, Röder MS, Rasheed A, Periyannan S, Lagudah ES (2015) Aegilops tauschii introgressions in wheat. In: Molnár-Láng M, Ceoloni C, Doležel J (eds) Alien introgression in wheat. Springer International, Switzerland
Braun HJ, Atlin G, Payne T (2010) Multi-location testing as a tool to identify plant response to global climate change. In: Reynolds MP (ed) Climate change and crop production. CABI Publishers, Wallingford (UK)
Calonnec A, Johnson R (1998) Chromosomal location of genes for resistance to Puccinia striiformis in the wheat line TP1295 selected from the cross of Soissonais-Desprez with Lemhi. Eur J Plant Pathol 104:835–847
Chen XM (2005) Epidemiology and control of stripe rust Puccinia striiformis f. sp. tritici on wheat. Can J Plant Pathol 27:314–337
Chen XM (2013) High-temperature adult-plant resistance, key for sustainable control of stripe rust. Am J Plant Sci 4:608–627
Chen XM (2020) Pathogens which threaten food security: Puccinia striiformis, the wheat stripe rust pathogen. Food Security 12:239–251
Chen XM, Line RF (1995) Gene-Action in wheat cultivars for durable, high-temperature, adult-plant resistance and interaction with race-specific, seedling resistance to Puccinia striiformis. Phytopathology 85:567–572
Chen XM, Penman L, Wan AM, Cheng P (2010) Virulence races of Puccinia striiformis f. sp. tritici in 2006 and 2007 and development of wheat stripe rust and distributions, dynamics, and evolutionary relationships of races from 2000 to 2007 in the United States. Can J Plant Pathol 32:315–333
Ding Y, Cuddy WS, Wellings CR, Zhang P, Thach T, Hovmøller MS, Qutob D, Brar GS, Kutcher HR, Park RF (2021) Incursions of divergent genotypes, evolution of virulence and host jumps shape a continental clonal population of the stripe rust pathogen Puccinia striiformis. Mol Ecol 30:6566–6584. https://doi.org/10.1111/mec.16182
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
FAO (2020) Crop Prospects and Food Situation. Quarterly Global Report No. 4, December 2020. Rome. https://doi.org/10.4060/cb2334en
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360
Holtz MD, Kumar K, Xi KQ (2013) Virulence phenotypes of Puccinia striiformis in Alberta from 2009–2011. Can J Plant Pathol 35:241–250
Huang L, Sela H, Feng LH, Chen QJ, Krugman T, Yan J, Dubcovsky J, Fahima T (2016) Distribution and haplotype diversity of WKS resistance genes in wild emmer wheat natural populations. Theor Appl Genet 129:921–934
Johnson R (1988) Durable resistance to yellow (stripe) rust in wheat and its implications in plant breeding. In: Simmonds NW, Rajaram S (eds) Breeding strategies for resistance to the rusts of wheat. CIMMYT, Mexico City, pp 63–75
Kishii M, Huerta J, Tsujimoto H et al (2019) Stripe rust resistance in wild wheat Aegilops tauschii Coss.: genetic structure and inheritance in synthetic allohexaploid Triticum wheat lines. Genet Resour Crop Evol 66:909. https://doi.org/10.1007/s10722-019-00758-w
Klymiuk V, Yaniv E, Huang L, Raats D, Fatiukha A, Chen S, Feng L, Frenkel Z, Krugman T, Lidzbarsky G, Chang W (2018) Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nat Commun 9(1):3735
Kolmer JA, Singh RP, Garvin DF, Viccars L, William HM, Huerta-Espino J, Ogbonnaya FC, Raman H, Orford S, Bariana HS, Lagudah ES (2008) Analysis of the Lr34/Yr18 rust resistance region in wheat germplasm. Crop Sci 48:1841–1852
KOMUGI (2023) Wheat Genetic Resources Database. https://shigen.nig.ac.jp/wheat/komugi/genes/symbolListPageAction.do?page=-1. Accessed 24 Mar 2023
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Kumar J, Nayar SK, Bahadur P, Nagarajan S, Bhardwaj SC, Prashar M, Singh SB (1988) Virulence survey of yellow rust of wheat (Puccinia striiformis f.sp. tritici) and barley (P. striiformis f.sp. hordei) during 1985–87. Cereal Rusts Powdery Mildews Bull 16:30–35
Lagudah ES, McFadden H, Singh RP, Huerta-Espino J, Bariana HS, Spielmeyer W (2006) Molecular genetic characterisation of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor Appl Genet 114:21–30
Line RF, Qayoum A (1992) Virulence aggressiveness, evolution, and distribution of race Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–87. USDA-ARS Technical Bulletin 1788. USDA-ARS, Washington, DC
Listman M, Ordóñez R (2019) Imagine a world without maize and wheat. CIMMYT Newsletter World Food Day 2019. https://www.cimmyt.org/news/ten-things-you-should-know-about-maize-and-wheat/. Accessed 24 Mar 2023
Liu W, Frick M, Huel R, Nykiforuk CL, Wang X, Gaudet DA, Eudes F, Conner RL, Kuzyk A, Chen Q, Kang Z, Laroche A (2014) The stripe rust resistance gene Yr10 encodes an evolutionary-conserved and unique CC-NBS-LRR sequence in wheat. Mol Plant 7:1740–1755
Liu TL, Wan AM, Chen XM (2016) Virulence characterization of Puccinia striiformis f. sp. tritici in the US for the past 48 years using the Yr single-gene differentials. Phytopathology S4:201
Lu Y, Wang MN, Chen XM, See D, Chao SM, Jing JX (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127:1449–1459
Marchal C, Zhang J, Zhang P, Fenwick P, Steuernagel B, Adamski NM, Boyd L, McIntosh R, Wulff BBH, Berry S, Lagudah E, Uauy C (2018) BED-domain-containing immune receptors confer diverse resistance spectra to yellow rust. Nature Plants 4(9):662–668. https://doi.org/10.1038/s41477-018-0236-4
McIntosh RA, Wellings CR, Park RF (1995) Wheat rust: an atlas of resistance genes. CSIRO Publishing, Melbourne, Australia, p 200
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong XY, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Singh R, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nature Genet 47:1494–1498
Mujeeb-Kazi A, Rosas V, Roldan S (1996) Conservation of the genetic variation of Triticum tauschii (Coss.) Schmalh. (Aegilops squarrosa auct. non L.) in synthetic hexaploid wheats (T. turgidum L. s.lat. x T. tauschii; 2n = 6x = 42, AABBDD) and its potential utilization for wheat improvement. Genet Resour Crop Evol 43:129–134
Murray GM, Brennan JP (2009) Estimating disease losses to the Australian wheat industry. Australas Plant Pathol 8:558–570
Park R, Ding Y, Singh D, Hovmøller M, Thach T, Justesen A, Cuddy W (2020) Confirmation of the exotic origins of two pathotypes of the wheat stripe rust pathogen detected in 2017 and 2018, and their impact. In: Cereal Rust Report, Cereal Rust Laboratory, The University of Sydney, Plant Breeding Institute, Cobbitty 17(4)
Sandhu KS, Singh D, Park RF (2021) A pictorial disease assessment scale for assessing wheat stripe rust at adult plant growth stage. Australas Plant Pathol. https://doi.org/10.1007/s13313-021-00827-8
Sharma-Poudyal D, Chen XM, Wan AM, Zhan GM, Kang ZS, Cao SQ, Jin SL, Morgounov A, Akin B, Mert Z, Shah SJA, Bux H, Ashraf M, Sharma RC, Madariaga R, Puri KD, Wellings CR, Xi KQ, Manninger K, Wanyera R, Ganzalez MI, Koyda M, Sanin S, Patzek LJ (2013) Virulence characterization of international collections of the wheat stripe rust pathogen, Puccinia striiformis f. sp. tritici. Plant Dis 97:379–386
Singh RP, Huerto-Espino J, William MH (2001) Slow rusting gene based resistance to leaf and yellow rusts in wheat: genetics and breeding at CIMMYT. Proceedings of the 10th assembly of the Wheat Breeding Society of Australia Inc., Mildura, Australia, 16–21 September. Wheat Breeding Society of Australia Inc., Australia, pp 103–108
Singh RP, Huerta-Espino J, Bhavani S, Herrera-Foessel SA, Singh D, Singh PK, Velu G, Mason RE, Jin Y, Njau P, Crossa J (2011) Race non-specific resistance to rust diseases in CIMMYT spring wheats. Euphytica 179:175–186. https://doi.org/10.1007/s10681-010-0322-9
Stubbs RW (1985) Stripe rust. In: Roelfs AP, Bushnell WR (eds) The cereal rusts II. Academic, Orlando, pp 61–101
Tesfaye K (2021) Climate change in the hottest wheat regions. In Population Division of United Nations, World Population Prospects: The 2017 Revision United Nations. Nature Food, vol 2 New York, NY, USA, pp 8–9
Uauy C, Brevis JC, Chen XM, Khan I, Jackson L, Chicaiza O, Distenfeld A, Fahima T, Dubcovsky J (2005) High-temperature adult-plant stripe rust resistance gene Yr36 from Triticum turgidum ssp. dicoccoides is closely linked to the grain protein content locus Gpc-B1. Theor Appl Genet 112:97–105
United Nations (2017) World population prospects revision, data booklet. https://www.un.org/development/desa/pd/sites/www.un.org.development.desa.pd/files/files/documents/2020/Jan/un_2017_world_population_prospects-2017_revision_databooklet.pdf. Accessed 24 Mar 2023
Van Silfhout CH (1989) Identification and characterization of resistance to yellow rust and powdery mildew in wild emmer wheat and their transfer to bread wheat. PhD thesis, Research Institute for Plant Protection, Wageningen, The Netherlands
Wan AM, Chen XM (2012) Virulence, frequency, and distribution of races of Puccinia striiformis f. sp. tritici and P. striiformis f. sp. hordei identified in the United States in 2008 and 2009. Plant Dis 96:67–74
Wan AM, Chen XM (2014) Virulence characterization of Puccinia striiformis f. sp. tritici using a new set of Yr single-gene line differentials in the United States in 2010. Plant Dis 98:1534–1542
Wan AM, Zhao ZH, Chen XM, He ZH, Jin SL, Jia QZ, Yao G, Yang JX, Wang BT, Li GB, Bi YQ, Yuan ZY (2004) Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici in China in 2002. Plant Dis 88:896–904
Wan AM, Chen XM, Yuen J (2016) Races of Puccinia striiformis f. sp. tritici in the United States in 2011 and 2012 and comparison with races in 2010. Plant Dis 100:966–975
Wan AM, Muleta KT, Zegeye H, Hundie B, Pumphrey MO, Chen XM (2017) Virulence characterization of wheat stripe rust fungus Puccinia striiformis f. sp. tritici in Ethiopia and evaluation of Ethiopian wheat germplasm for resistance to races of the pathogen from Ethiopia and the United States. Plant Dis 101:73–80
Wani SH, Mohan A, Singh GP (2021) Physiological, Molecular, and Genetic Perspectives of Wheat Improvement. Springer Nature Switzerland AG. https://doi.org/10.1007/978-3-030-59577-7 ISBN 978–3–030–59577–7 (eBook)
Wellings CR (2011) Global status of stripe rust: A review of historical and current threats. Euphytica 179:129–141
Wellings CR, Burdon JJ (1992) Variability in Puccinia striiformis f. sp. tritici in Australasia. Vorträge für Pflanzenzüchtung 24:114
Wellings CR, Singh RP, Yahyaoui A, Nazari K, McIntosh RA (2009) The development and application of near-isogenic lines for monitoring cereal rust pathogens. In: McIntosh RA (ed) Proc Borlaug Global Rust Initiative Technical Workshop. Mexico, BGRI Cd Obregon, pp 77–87
Yang EN, Rosewarne GM, Herrera-Foessel SA, Huerta-Espino J, Tang ZX, Sun CF, Ren ZL, Singh RP (2013) QTL analysis of the spring wheat “Chapio” identifies stable stripe rust resistance despite inter-continental genotype × environment interactions. Theor Appl Genet 126:1721–1732
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions This research was conducted under the project CIM/2014/081, funded by the Australian Centre for International Agricultural Research.
Author information
Authors and Affiliations
Contributions
Karanjeet S. Sandhu planned and conducted all the greenhouse, field, and lab experiments, and drafted this manuscript. Robert F. Park conceived the project and secured funding, and along with Davinder Singh provided valuable suggestions in writing this manuscript. Fikrte Y. Belayineh, Tamrat Negash, Hanif Khan, Subhash C. Bhardwaj, Suraj Baidya, Dhruba B. Thapa, Muhammad Fayyaz, Shahzad Asad and Mandeep S. Randhawa helped in the field rust screening of germplasm.
Corresponding authors
Ethics declarations
Conflict of interest
Authors declare that there is no conflict of interests exist in relation to this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sandhu, K.S., Singh, D., Belayineh, F.Y. et al. Identification of synthetic wheat lines with broadly effective stripe rust resistance. Australasian Plant Pathol. 53, 221–238 (2024). https://doi.org/10.1007/s13313-024-00971-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-024-00971-x