Abstract
Poststroke dementia (PSD) is associated with pathology in frontal brain regions, in particular dorsolateral prefrontal cortex (DLPFC) neurons and white matter, remote from the infarct. We hypothesised that PSD results from progressive DLPFC neuronal damage, associated with frontal white matter gliovascular unit (GVU) alterations. We investigated the transcriptomic profile of the neurons and white matter GVU cells previously implicated in pathology. Laser-capture microdissected neurons, astrocytes and endothelial cells were obtained from the Cognitive Function After Stroke cohort of control, PSD and poststroke non-dementia (PSND) human subjects. Gene expression was assessed using microarrays and pathway analysis to compare changes in PSD with controls and PSND. Neuronal findings were validated using NanoString technology and compared with those in the bilateral common carotid artery stenosis (BCAS) mouse model. Comparing changes in PSD compared to controls with changes in PSND compared to controls identified transcriptomic changes associated specifically with dementia. DLPFC neurons showed defects in energy production (tricarboxylic acid (TCA) cycle, adenosine triphosphate (ATP) binding and mitochondria), signalling and communication (MAPK signalling, Toll-like receptor signalling, endocytosis). Similar changes were identified in neurons isolated from BCAS mice. Neuronal findings accompanied by altered astrocyte communication and endothelium immune changes in the frontal white matter, suggesting GVU dysfunction. We propose a pathogenic model in PSD whereby neuronal changes are associated with frontal white matter GVU dysfunction leading to astrocyte failure in supporting neuronal circuits resulting in delayed cognitive decline associated with PSD. Therefore, targeting these processes could potentially ameliorate the dementia seen in PSD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Having a stroke over the age of 65 years old doubles the chance of developing dementia [1], while around 25% of stroke survivors develop progressive cognitive decline more than 3 months poststroke with the majority satisfying criteria for vascular dementia [2]. Previous pathological investigations in the Cognitive Function After Stroke (CogFAST) cohort, a longitudinal study of dementia incidence following stroke in the over 75’s [3], have shown that progressive cognitive decline, leading to poststroke dementia (PSD), is associated with pathology remote from the infarct, particularly in the frontal white matter. The infarcts themselves may be small so that direct infarct damage per se does not explain the progressive dementia. The association of PSD with pathology in frontal brain regions, specifically the dorsolateral prefrontal cortex (DLPFC) implicates damage to anterior cognitive circuits (ACC) involved in impaired executive function, which is a common characteristic of vascular dementia [4, 5].
Pathological changes have been identified in both the glutamatergic pyramidal neurons of the DLPFC and the astrocytes that form the gliovascular unit (GVU) of the frontal white matter in those stroke survivors who proceed to PSD compared to those that do not develop dementia [6, 7]. DLPFC neurons project to the dorsolateral head of the caudate, important for executive function incorporating anticipation, planning, goal setting and flexible thinking. Pyramidal neuronal cell volumes are reduced by 30–40% in layers III and V of the DLPFC of PSD subjects compared to poststroke non-dementia (PSND) and control (no dementia or stroke) subjects. This reduction in cellular volume directly correlates with an increase in neurofilament protein SMI-31 immunoreactivity in layer III of the DLPFC of PSD subjects. These neuronal changes are not associated with neurofibrillary tangle pathology suggesting a vascular origin of the observed neuronal atrophy [7] that differs between cortical layers. Additionally, an increased number of clasmatodendritic astrocytes, immunopositive for fibrinogen, have been identified within the frontal white matter of PSD subjects, indicative of blood brain barrier (BBB) disruption [6]. Astrocytic end-feet also show stripping of aquaporin-4, further evidence of pathology of the frontal WM GVU in PSD [6]. Gliovascular dysfunction is a key contributor to neuronal injury following ischaemic stroke and in other neurodegenerative diseases, with neuronal cell death, glial cell activation, BBB disruption and resulting serum protein accumulation and inflammatory response being associated with disease progression [8, 9]. In addition to gliovascular changes in PSD, there is a higher burden of white matter hyperintensities and diffuse myelin changes in PSD compared to PSND subjects [6].
The current study aimed to identify the cellular and molecular basis of PSD by investigating the transcriptomic profile of the cells that have been implicated by previous pathological studies. Our study focused particularly on the pyramidal neurons of the DLPFC, but we also compared this with changes in capillary endothelial cells and astrocytes of the frontal white matter. We hypothesised that PSD results from progressive neuronal damage in the DLPFC that is remote from the stroke and that this may be associated with alterations in the GVU of frontal white matter with BBB disruption, astrocyte dysregulation and a neuroinflammatory response.
Materials and Methods
Human Central Nervous System (CNS) Tissue
Human post-mortem brain tissue was obtained from the Newcastle CogFAST Study. Ethical approval was obtained from the Joint Ethics Committee of Newcastle and North Tyneside Health Authority, Newcastle University and the University of Northumbria at Newcastle, and participants gave written informed consent in accordance with the declaration of Helsinki. Cognitive function and dementia tests including Mini-Mental State Exam (MMSE) (PSD average: 13; PSND average: 24), CAMCOG (PSD average: 50; PSND average: 88), Consortium to Establish a Registry for Alzheimer’s Disease (CERAD) (PSD average: 0.7; PSND average: 1.0) and Braak staging (PSD average: 1.9; PSND average: 2.1) were considered when selecting participants as part of the current study (Supplementary Table 1). Full CogFAST study cohort characteristics have been described previously [3, 10]. Generally, most stroke survivors in the CogFAST study considered clinically demented by the time of their death showed a high burden of vascular pathology in the general absence of Alzheimer’s type pathology [11]. It has been acknowledged that stroke survivors may not be completely free of Alzheimer’s pathology of amyloid-β burden but most stroke survivors over the age of 75 years present with significant vascular dementia over Alzheimer’s [12]. Snap frozen DLPFC and frontal white matter samples (n = 10 for each group) were collected from PSD, PSND and control subjects (no dementia or stroke). All PSD and PSND samples were from brains with infarcts of < 5 mm remote from the DLPFC and frontal white matter investigated in this current study. Where possible, cases were selected equally based on age and sex (Table 1, Supplementary Table 2). The quantity of RNA from each sample was analysed using a NanoDrop 1000 spectrophotometer (Thermoscientific, UK) and the quality of RNA was assessed on a 2100 Bioanalyzer (Agilent, Palo Alto, CA, USA). The RNA integrity number (RIN) of each frozen tissue block was assessed prior to laser capture microdissection (LCM) using a previously reported method [13, 14]. The mean pre-LCM RIN of the full set of DLPFC samples was 2.4, standard deviation (SD) 1.02; range 1.0–7.1 comprising of the PSD group (2.3, SD 0.24; range 1.9–2.7), PSND group (2.0, SD 0.53; range 1.0–2.6) and control group (3.0, SD 1.62; range 1.6–7.1). The mean brain pH of the samples was 6.44, SD 0.40; range 5.77–7.19 comprising of the PSD group (6.39, SD 0.43; range 5.77–7.03), PSND group (6.45, SD 0.46; range 6.00–7.04) and control group (6.49, SD 0.37; range 5.87–7.09). The mean post-mortem interval (PMI) time of the samples was 41 h SD 25.24 h; range 12–98 h comprising of the PSD group (45 h, SD 28.07 h; range 12–88 h), PSND group (47 h, SD 22.99 h; range 19–81 h) and control group (38 h, SD 26.33; range 15–98 h). The mean age at death of the samples was 85 years, SD 8.3 years; range 72–99 years comprising of the PSD group (89 years, SD 6.9 years; range 75–97 years), PSND group (85 years, SD 6.0 years; range 78–99 years) and control group (80 years, SD 9.4 years; range 72–96 years).
Laser Capture Microdissection and RNA Extraction of Neurons, Astrocytes and Endothelial Cells
Toluidine blue histologically stained pyramidal neurons [15, 16], glial fibrillary acidic protein-positive (GFAP+) astrocytes and collagen IV-positive (Coll-IV+) endothelial cells [14, 17] were isolated from each case using an Arcturus PixCell II LCM system (Thermoscientific, UK). Total RNA (approximately 50 ng/cell type) was extracted from each isolated enriched group of cells using the Arcturus PicoPure RNA isolation kit, according to the manufacturer’s instructions (Thermoscientific, UK) (Supplementary Fig. 1). The quantity and quality of the extracted RNA were determined using a NanoDrop 1000 spectrophotometer (Thermoscientific, UK) and a 2100 Bioanalyzer, respectively (Agilent, USA). Only those samples with a RIN of > 2.0 post-LCM or in the absence of a RIN, samples not showing a peak shift towards a shorter fragment size representing a more degraded sample on a Bioanalyzer trace, were taken forward for microarray analysis.
RNA Amplification and Microarray Hybridisation
The target labelled mRNA was prepared following the GeneChip WT Pico amplification protocol (Applied Biosystems, ThermoFisher, UK). In brief, RNA amplification was achieved using low-cycle PCR followed by amplification using T7 in vitro transcription. The cRNA was converted to biotinylated sense-strand DNA hybridisation targets. Approximately 5.5 μg amplified cDNA was fragmented, labelled and hybridised to Clariom S Array chips for 16 h at 45 °C in a rotating oven at 60 rpm. Using the Fluidics Station 400 and GeneChip Operating System, the hybridisation cocktail was removed, and a series of stringency washing steps are followed to remove any unbound DNA, before each microarray was stained. Each microarray chip was scanned using the GeneChip 3000 scanner to determine the fluorescence intensity of hybridised transcripts.
Microarray Analysis
Transcription Analysis Console (TAC) was used for quality control and analysis of the data, the individual sample. CEL files were imported into TAC version 4.0.1 (Applied Biosystems, USA) and several quality control (QC) parameters were checked for each sample including Poly-A RNA labelling controls, hybridisation controls, positive vs. negative area under the curve (AUC) and signal intensity to identify any outlier samples. Samples that passed QC were normalised to the median of all genes, prior to statistical analysis, to identify differentially expressed genes (DEGs). Transcripts were defined as being significantly differentially expressed if they showed a p-value of < 0.05 (unpaired t-test). DEGs were classified according to their biological function using the Database for Annotation Visualisation and Integrated Discovery (DAVID) [18, 19] using Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis to identify specific pathways while Functional Annotation Clustering (FAC) was used to cluster together functionally similar genes. In the case of a significant result, the p value was adjusted for multiple testing using the Bonferroni correction method and considered significant if p < 0.05.
Neuronal Gene Expression Validation Using NanoString nCounter Analysis
Gene expression validation was performed on 14 samples consisting of 7 control and 7 PSD neuronal samples from the original cohort (Supplementary Table 3). Neuronal enriched RNA served as input into NanoString’s low RNA input amplification assay, designed to produce sufficient target for detection in an nCounter hybridisation assay. Multiplexed target enrichment (MTE) was achieved through the conversion of samples with low RNA concentration into cDNA, which was then amplified using target-specific primers. In brief, approximately 10 ng RNA was reverse transcribed using the following conditions: 10 min at 25 °C, 60 min at 42 °C and 5 min at 85 °C. For this work, NanoString human neuropathology gene expression panels were used for validation based on the initial microarray findings (NanoString Technologies, USA). Therefore, for MTE amplification 5 × dT Amp master mix with neuropathology panel gene specific primers were used. Following denaturation for 10 min at 95 °C, the products were amplified for 10 cycles under the following conditions, 15 s at 95 °C and 4 min at 60 °C.
Gene expression analysis was performed on the nCounter SPRINT Profiler system (NanoString Technologies, USA) according to manufacturer’s instructions and data analysed using the NanoString nSolver analysis v3.0 software (NanoString Technologies, USA). In brief hybridisation between target sample and reporter-capture probes pairs was performed for 22 h at 65 °C. Post hybridisation processing steps were carried out and excess probes removed. The probe/target complexes were aligned and immobilised on the nCounter cartridge. Image acquisition and data processing was carried out as per manufacturer’s protocol. Firstly, all data was background adjusted using internal negative controls followed by technical normalisation using internal positive controls. Data was then normalised to account for the variation in sample concentration and/or quality using the mean expression of the most stable housekeeping genes across samples and cartridges. Gene expression was calculated by counting the number of times each barcoded gene was detected above the defined background threshold and downstream differential gene expression between control and PSD cases was completed in the NanoString nSolver v3.0 software.
Neuronal Gene Expression in the BCAS Mouse Model
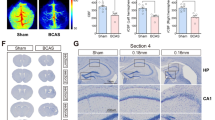
For additional validation and comparison, the bilateral carotid artery stenosis (BCAS) mouse model [20] that replicates features of vascular dementia was used alongside control sham animals to characterise the neuronal transcriptomic changes in the frontal cortex (full animal procedures, Supplementary Methods 1). Toluidine blue stained cryosections from 5 BCAS and 5 Sham animals were subjected to LCM to isolate neurons from the frontal cortex of each animal as previously described [15]. Target labelled mRNA was prepared following the GeneChip WT Pico amplification protocol (Applied Biosystems, ThermoFisher, UK) and mouse Clariom S Arrays were used to investigate the neuronal gene changes across the two groups. Within TAC, transcripts were considered significantly, differentially expressed if they had a minimum fold change (FC) of plus or minus 1.2 and p-value of < 0.05 (unpaired t-test). The annotated gene lists were exported into DAVID where KEGG pathway analysis and FAC was carried out. Analysis was completed on each unique combined gene list (up- and downregulated genes) alongside analysis from the separate distinct up- and downregulated gene lists.
Data Availability
The data that support the findings of this study are openly available in Gene Expression Omnibus at https://www.ncbi.nlm.nih.gov/geo/, reference number GSE186798.
Results
Gene Expression Changes in Neurons, Astrocytes and Endothelium
Quality control measures were applied to each microarray-generated dataset in TAC. Based on gene expression alone, a clear separation of the different patient groups was confirmed for all three cell types using principal components analysis (PCA) (Fig. 1A–C) and heatmaps (Fig. 1D–F). All significantly DEGs (p < 0.05) between patient group comparisons, i.e. PSD vs control, PSND vs control and PSD vs PSND for each cell type were tabulated (Fig. 1G and Supplementary Table 4). In total from the neuronal data generated, 784 genes (PSD vs control), 193 genes (PSND vs control) and 608 genes (PSD vs PSND) were specific and exclusive to each patient group comparison. From the astrocyte data, 817, 767 and 363 genes respectively were specific and exclusive to each patient group comparison. While from the endothelial cell data, 681, 1048 and 352 genes were specific and exclusive to each patient group comparison.
Gene expression analysis across cell types. Principal component analysis (PCA) reveals clear separation of control, poststroke dementia (PSD) and poststroke non-dementia (PSND) cases based on differential gene expression in neurons (A), astrocytes (B) and endothelial cells (C). Heatmap representation of differential gene expression in neurons (D), astrocytes (E) and endothelial cells (F). Transcription Analysis Console (TAC) analysis shows significantly expressed genes across patient comparison and cell types (G) (PSD; poststroke dementia, PSND; poststroke non-dementia)
These specific and exclusive gene lists from each patient comparison (PSD vs control, PSND vs control and PSD vs PSND) and cell type (neuron, astrocyte, and endothelial cells) were exported into DAVID where KEGG pathway analysis and FAC was carried out. Analysis was completed on each separate distinct up- and downregulated gene list alongside analysis from each unique combined gene list (up- and downregulated genes) (Supplementary Tables 5 and 6).
The Neuronal Gene Expression Profile Is Significantly Altered in PSD Compared to PSND
Our experiments and analysis focused particularly on changes in pyramidal neurons of the DLPFC. To identify gene expression and pathway changes that might specifically be related to the development of dementia poststroke, we compared gene changes that were identified in PSD cases versus controls with those seen in PSND cases vs controls. Only 193 DEGs occurred in PSND neurons compared to control neurons. In contrast, 784 DEGs were identified in PSD compared to control neurons, including KEGG pathways associated with neuronal metabolism/mitochondrial changes (ribosome [25 DEGs, p 3.39E-06], Parkinson’s disease [9 DEGs, p 4.47E-02], Alzheimer’s disease [11 DEGs, p 1.88E-02], tricarboxylic acid (TCA) cycle [5 DEGs, p 9.13E-03]) and signalling changes (MAPK signalling [25 DEGs, p 2.87E-02], endocytosis [13 DEGs, p 3.68E-02] and Toll-like receptor signalling [8 DEGs, p 2.83E-02]) (Fig. 2A). Additionally, FAC carried out in DAVID identified the mitochondrion cluster (enrichment score (ES); 1.68, 44 genes, p 6.90E-03) and neuronal metabolism-related clusters including nucleotide binding (ES; 1.90, 69 genes, p 9.60E-04), glycoprotein (ES; 1.73, 181 genes, p 2.20E-03) and adenosine triphosphate (ATP) binding (ES; 1.31, 61 genes, p 1.6E-02) (Fig. 2B).
Comparing neuronal KEGG pathway analysis and functional annotation clustering associated with having a stroke (PSND vs. control) and having dementia resulting from a stroke (PSD vs. control) (↓ down regulated gene list, ↑ red arrow, upregulated gene list, ↕ up- and downregulated gene lists). (PSD, poststroke dementia; PSND, poststroke non-dementia)
Poststroke Dementia Neuronal Changes Verified by NanoString nCounter Analysis
Validation candidate genes were selected from the initial microarray analysis based on the significant KEGG pathways, FAC and specific, exclusive gene changes associated with PSD neurons versus control neurons. In total, NanoString nCounter analysis showed 97/770 genes on the neuropathology panels to be significantly altered in PSD neurons compared to control neurons (Supplementary Table 7). Based on neuronal gene expression alone, a clear separation of the different patient groups was confirmed using PCA (Fig. 3A). Taking the 97 significantly altered genes (p < 0.05), KEGG pathway analysis and FAC showed the same altered pathways that were identified in the microarray study including MAPK signalling (ES; 2.89, 13 genes, p 3.07E-05), Toll-like receptor signalling pathway (ES; 2.89, 7 genes, p 1.39E-03) and Alzheimer’s disease (11 DEGs, p 2.18E-05) in addition to other signalling and synaptic changes (Fig. 3B and Supplementary Table 8).
NanoString nCounter gene expression validation of neuronal microarray data. Principal component analysis (PCA) reveals clear separation of poststroke dementia (PSD) and control neuronal cases based on differential gene expression changes analysed via the NanoString nCounter system (A). Functional annotation clustering (FAC) of altered genes analysed via the NanoString nCounter system in PSD vs control neurons (B). (ES; enrichment score)
Comparison with Neuronal Gene Changes in the BCAS Mouse Model
We also determined whether the changes associated with PSD in human autopsy tissue were found in a mouse model of chronic cerebral hypoperfusion (CCH) that simulates features of vascular cognitive impairment and vascular dementia. Quality control measures were applied to the mouse microarray dataset in TAC, and based on neuronal gene expression alone, a clear separation of the BCAS and sham animal groups was confirmed using PCA (Fig. 4A and Supplementary Table 9). A total of 1185 DEGs (FC ≥ 1.2, ≤ p0.05) were identified between the BCAS and sham group (575 upregulated, 610 downregulated). Following KEGG pathway analysis, several synaptic and signalling pathways were identified in the neuronal BCAS data that were identified in the human PSD neuronal NanoString validation including the downregulation of the dopaminergic synapse (9 DEGs, p 3.47E-02), altered glutamatergic synapse (13 DEGs, p 2.38E-02) and cholinergic synapse (12 DEGs, p 4.59E-02), alongside altered chemokine signalling (11 DEGs, p 4.91E-02), thyroid hormone signalling (12 DEGs, p 4.84E-02), oxytocin signalling (16 DEGs, p 1.77E-02) and retrograde endocannabinoid signalling (13 DEGs, p 1.06E-02) (Fig. 4B). Additional FAC displayed glycoprotein (ES: 5.81, 152 genes, p 2.30E-10) and transmembrane (ES: 5.43, 223 genes, p 9.11E-07) clusters of genes in the neurons isolated from the BCAS model similar to the initial human neuronal microarray findings that showed the same enriched clusters in PSD neurons: glycoprotein (ES; 1.73, 181 genes, p 2.20E-03) and cell membrane (ES: 1.45, 128 genes, p 7.90E-03) (Fig. 4C) (Supplementary Table 10).
BCAS model neuronal KEGG and FAC analysis. Principal component analysis (PCA) reveals clear separation of BCAS and sham neuronal cases based on differential gene expression changes analysed by microarray analysis (A). KEGG pathway analysis revealed the same synaptic and signalling changes in PSD neurons compared to BCAS neurons (B), while functional annotation clustering (FAC) revealed the same enriched gene clusters between the PSD and BCAS neuronal microarray studies (C) (BCAS, bilateral common carotid artery stenosis; ES, enrichment score)
Astrocytic and Endothelium Dysfunction in the White Matter Associated with Poststroke Dementia
Because changes in frontal white matter may also be important in the pathogenesis of dementia poststroke, and may affect neurons, we identified PSD-associated transcriptomic changes in astrocytes and capillary endothelial cells of this area.
As a result of the stroke, PSND astrocytes showed 767 DEGs compared to control astrocytes. Employing KEGG pathway analysis and FAC, DEGs were associated with changes in metabolism including altered proteolysis, RNA metabolism and phosphorylation alongside changes in Alzheimer’s disease and Parkinson’s disease pathways mostly associated with metabolic gene changes (Fig. 5A and B).
KEGG Pathway analysis and functional annotation clustering associated with having dementia resulting from a stroke (PSD vs. control) compared to changes associated with having a stroke (PSND vs. control) in astrocytes (A&B) and endothelial cells (C&D) (↓ downregulated gene list, ↑ red arrow, upregulated gene list, ↕ up- and downregulated gene lists). (PSD; poststroke dementia, PSND; poststroke non-dementia)
Overall PSD astrocytes showed the greatest number of gene changes compared to control astrocytes (817 DEGs). Astrocytes in the PSD group further showed the greatest number of altered KEGG pathways with most of these pathways downregulated (Fig. 5A). An overall loss of cellular communication was apparent in astrocytes as seen through impaired gap junctions (10 DEGs, p 2.56E-03) and axonal guidance (10 DEGs, p 2.60E-02), alongside both intra- and extracellular signalling changes with altered calcium signalling (18 DEGs p 9.25E-05), glucagon signalling (10 DEGs, p 5.66E-03) and Wnt signalling (10 DEGs, p 4.12E-02). Astrocytic neurotransmitter functions were also altered including changes in dopaminergic synapse (11 DEGs, p 1.04E-02), GABAergic synapse (13 DEGs, p 1.87E-02) and glutamatergic synapse (12 DEGs, p 1.42E-03). The altered KEGG pathways in PSD astrocytes showed downregulation of mitochondria-related genes which was also identified as the most enriched cluster of functionally similar genes in DAVID using FAC (mitochondrial inner membrane, ES: 6.13, 41 genes, p 4.00E-09) alongside changes in mRNA transport (mRNA export from nucleus, ES: 1.50, 10 genes, p 5.90E-03, mRNA transport, ES 1.39, 10 genes, p 5.10E-03) in addition to synapse and membrane clusters (membrane, ES 1.62, 274 genes, p 0.0051; synapse ES:2.44, 25 genes, p 2.70E-04) (Fig. 5B).
Endothelial cells showed the greatest number of DEGs in PSND compared to control cases (1048 DEGs). Through KEGG pathway analysis and FAC, changes in endothelial cell lipid and carbohydrate metabolism were most apparent as a result of a stroke (Fig. 5C and D). Overall, in PSD endothelial cells compared to control endothelial cells, 681 DEGs were identified; through KEGG pathway analysis, an upregulation of chemokine signalling (18 genes, p 5.40E-02) and changes in glucagon signalling (14 genes, p 4.06E-02) (Fig. 5C) were most apparent alongside altered mitochondria, again the most enriched cluster of functionally similar genes in DAVID being the mitochondrion (ES: 3.34, 41 genes, p 5.20E-05) (Fig. 5D).
Discussion
Progressive dementia after stroke is associated with pathological changes in DLPFC neurons and frontal white matter. As these changes can occur with small strokes not located in situ or in regions of the frontal lobe we assessed, it suggests that such strokes initiate a pathogenic process remote from site of ischemic injury. To elucidate this process further, we investigated transcriptomic changes in DLFPC neurons, and in astrocytes and endothelium of frontal white matter. To identify transcriptomic changes specific to those that develop dementia following a stroke rather than changes due to the stroke itself, we identified changes that were seen in the PSD vs control comparison but were not present in the PSND vs control comparison. In PSD DLPFC neurons, we identified changes in key pathways and genes associated with mitochondria, cellular communication, signalling and metabolism. The key neuronal pathway changes were validated using NanoString nCounter technology, and further experimental validation was completed on neurons isolated from the BCAS mouse model of chronic cerebral hypoperfusion. We also identified in PSD frontal white matter changes in astrocytes signalling and cellular communication, while frontal white matter endothelium showed changes in mitochondria and chemokine signalling pathways. Neuronal atrophy within the DLPFC, rather than neuronal loss, has been shown in PSD [7]. This study, which demonstrates significant dysregulation of numerous neuronal functions including cellular communication, signalling and metabolism, provides further evidence of DLPFC neuronal dysfunction as an important component of PSD pathogenesis, with pathway changes reminiscent of those in neurodegenerative states.
Neuronal endocytosis was dysregulated in the current study with neuronal physiology relying on the interplay between endocytosis and autophagy, as a method of communicating with the environment and maintaining cellular homeostasis [21]. Defects in endocytosis have been identified as a common theme in neurodegenerative diseases including Alzheimer’s disease [22], Parkinson’s disease [23], amyotrophic lateral sclerosis [24] and Huntington’s disease [25] often underlying the earliest events in neurodegeneration [21]. Endocytosis may prevent neurodegeneration by regulating protein turnover via autophagy. Proteins involved in endocytosis, including clathrin-mediated endocytosis (CME) adapters, assembly protein complex 2 (AP-2) and CALM2, control autophagy and regulate the degradation of components of amyloidosis in Alzheimer’s disease [26, 27]. The dysregulation of neuronal endocytosis in PSD as identified in the current study was not identified in those that did not progress to dementia (PSND) which may suggest a dysregulation in endocytosis could contribute to neuronal dysfunction and cognitive decline.
Mitogen-activated protein kinases (MAPKs) are typically activated through extracellular stresses including oxidative stress in addition to other stimuli such as hormones and cytokines and have been extensively studied in association with inflammation [28, 29]. Many of the neuronal genes in the MAPK signalling pathway in the current study were upregulated reinforcing this statement. Seen as a potential mechanism of Alzheimer’s disease pathology, neuronal apoptosis is induced partly by the activation of MAPK signalling pathways [30,31,32] and human post-mortem studies have identified increased MAPK phosphorylation associated with amyloid beta plaques and neurofibrillary tangles [33]. MAPK signalling is needed for inducing synaptic plasticity [34] alongside brain-derived neurotrophic factor (BDNF), which also plays an important role in neural plasticity and brain recovery following stroke [35]. Interleukin-1 (IL1) is a family of major proinflammatory cytokines, with IL1β previously connected to the neurodegenerative effect of neuroinflammation on cognition and synaptic plasticity in the brain [36]. Chronic elevation of IL1β suppresses BDNF and is a potential therapeutic target in the treatment and recovery from ischaemic stroke [37]. This concept is corroborated in the current study with the upregulation of another IL1 cytokine, IL1α and corresponding downregulation of BDNF in PSD neurons. It has been reported that IL1β functions in suppressing the effect of BDNF on synaptic plasticity through the induction of p38 MAPK on central nervous system (CNS) neurons [38,39,40]. The dysregulation of MAPK signalling in PSD neurons and what effect this may have on driving dementia following a stroke requires further investigation.
In the current study, neuronal Toll-like receptor (TLR) signalling was altered in PSD. During brain injury, such as in stroke, the BBB is breached leading to a dysregulation in communication and signalling between the immune system and the CNS [6, 41, 42]. TLRs play a role in the innate immune system by responding to pathogen-derived and tissue damage-related ligands. The interaction between neuronal TLRs and the immune system is well known with alterations in neuronal TLRs having been reported in several neurodegenerative diseases with varying effects [43]. For example, targeting the TLR4/NFκB pathway downregulates the proinflammatory response in both a mouse model of Alzheimer’s disease [44] and traumatic brain injury [45]. Despite not being directly studied in relation to neurodegenerative disease, TLR3, known to recognise double-stranded RNA associated with viral infection, was downregulated in the current study [43]. In vivo data collected from early life studies may suggest that neuronal TLR3 stimulation is detrimental in neurodegenerative disease, especially in the context of viral infections including Epstein-Barr virus and hepatitis C [46]. Therefore, identifying whether the dysregulation of neuronal TLR signalling in PSD mediates neuroinflammation and neurodegeneration or forms part of a neuroprotective preconditioning of the brain to respond and handle insult better needs to be established [47].
The current study also identified altered neuronal metabolism in PSD. The TCA cycle functions in the mitochondrial matrix and connects multiple catabolic and anabolic pathways, such as glycolysis and gluconeogenesis that result in the release of stored energy in cells [48]. The decrease in neuronal TCA cycle genes in PSD alongside altered neuronal mitochondrial function may infer a potential pathogenic avenue for neuronal dysfunction. CCH models, which include the BCAS mouse model, have shown energetic deficits reflected by changes in energy metabolites including ATP, adenosine diphosphate (ADP) and adenosine monophosphate (AMP) alongside alteration of ATP-related enzymes [49]. Additionally, these CCH models show mitochondria dysfunction because of decreased cerebral blood flow. Mitochondria also play a role in regulating calcium storage [50], innate and adaptive immunity [51], activating apoptosis [52] and signalling pathways [50]. Altered neuronal mitochondria could serve as a master regulator of the pathology seen in PSD and is worth further study.
To further explore the link between neuronal changes and vascular processes, we also investigated frontal neuronal transcriptome alterations in the BCAS mouse model. The BCAS model is a CCH injury presenting with glial responses and GVU disruption associated with long-term white matter damages accompanied by cognitive decline and is commonly used as an experimental model for vascular dementia [20, 53, 54]. Neuronal changes associated with cell membrane and glycoprotein were remarkably validated in the experimental model, as also key pathway changes including chemokine signalling, axon guidance alongside additional neuronal synaptic changes associated with neuronal dysfunction. In the BCAS mice, similar changes in neurons were identified as seen in PSD neurons, and this strengthens the notion that neuronal degenerative processes maybe linked to vascular processes. As the BCAS models show some of the pathway changes observed in PSD, it will be an important system to investigate aspects of poststroke pathophysiology relevant to the development of dementia.
Previous histological frontal white matter gliovascular alterations involving astrocytes and endothelial cells suggest they are associated with the neuronal changes observed in PSD [6]. We identified an overall dysregulation of astrocyte communication with altered astrocyte calcium signalling in PSD. Astrocyte calcium signalling has a functional role in astrocyte-neuron communication [55, 56], with dysregulated astrocyte calcium signalling occurring as Alzheimer’s type pathology develops [57]. During synaptic activity, the release of neuronal transmitters leads to changes in astrocyte function including calcium activity [58] and these astrocytic calcium signalling changes determine the release of other neuroactive molecules including gliotransmitters: glutamate, GABA and ATP [59]. As a result of a stroke (i.e. PSND vs controls), our study shows a general alteration in astrocytic synaptic effect, yet calcium signalling dysregulation is only apparent in PSD, corroborating the potential importance of modulating astrocytic calcium signalling in PSD that warrants further investigation.
The main endothelial change in PSD was altered chemokine signalling, suggesting immune activation that corroborates the gliovascular histological findings in PSD. The BBB not only protects and maintains homeostasis of the CNS, but also allows tightly controlled communication with the periphery. In multiple sclerosis, brain endothelial cells lose their protective function leading to increased BBB permeability, neuroinflammation and neurodegeneration [60]. Similarly, schizophrenia has been linked to endothelial dysfunction and inflammation [61]. Overall, within frontal white matter, PSD is associated with gliovascular dysfunction, altered astrocyte communication and immune changes in endothelium creating an overall disrupted immune environment. Although it is not possible to pinpoint the sequence of events that lead to PSD from the current study, it may be hypothesised that gliovascular changes in the white matter cause a lack of axonal process support or that frontal white matter neuroinflammation could directly damage axons. Either scenario could result in an impaired ACC and overtime a deterioration in neuronal processing leading to PSD.
These transcriptomic findings, and the previous histological studies, propose a possible pathogenic model (Fig. 6) that indicates that in PSD, neuronal changes in the DLPFC consist of a dysregulation of cellular communication, signalling and metabolic processes. These neuronal changes may result from gliovascular changes within the frontal white matter that arise through pathogenic changes resulting from a remote stroke. Thus, in PSD, the GVU in the frontal white matter becomes dysfunctional resulting in altered vascular permeability, neuroinflammation and an overall astrocyte failure in supporting axonal processes that pass through the frontal white matter that form the ACC.
PSD model of pathology in poststroke dementia (PSD)-specific neuronal changes in the dorsal lateral prefrontal cortex (DLPFC) consisted of dysregulation of metabolic processes (TCA cycle, glycoprotein, ATP binding), signalling (MAPK signalling, Toll-like receptor signalling, chemokine signalling, thyroid hormone signalling, oxytocin signalling and retrograde endocannabinoid signalling) and cellular communication (endocytosis, nucleotide binding, cadherin binding, rRNA binding and DNA binding) (A). These PSD neuronal changes within the DLPFC may result from gliovascular changes within the frontal white matter that arise through pathogenic changes resulting from a remote stroke. Astrocytes in PSD showed the greatest number of altered pathways related to cellular communication (gap junction, membrane), signalling (calcium signalling, oestrogen signalling, GnRH Signalling) and neurotransmitter function (dopaminergic synapse, ATP binding) (B). These astrocytic changes were also corroborated by the changes observed in the endothelial cells of blood vessels with altered mitochondria and inflammatory response underpinning an overall loss of frontal white matter gliovascular unit function (C). Ultimately, these changes we propose result in astrocyte failure to support axonal processes that form the cognitive network responsible for executive function (D). Consequently, over time, this would lead to a disconnection of the frontal network resulting in DLFPC neuronal changes apparent in PSD
Transcriptomic work using LCM and microarray analysis has limitations. Firstly, transcriptomic analysis is a powerful hypothesis generating tool, but our study has not linked this to proteome changes, while any temporal sequences or functional outcomes may be inferred require further experimental investigation in model systems. Secondly, the current study investigated DLPFC neurons and frontal white matter astrocytes and endothelial cells, yet other cell types including other glial cells such as microglia and oligodendrocytes as well as other cells involved in the GVU including pericytes will undoubtedly play a role in the pathology and must be considered. Thirdly, LCM does not isolate specific cell populations; rather the method allows for an enriched population of cells to be isolated [14]. Therefore, the population of endothelial cells isolated is likely to include pericytes and astrocytic end-feet, yet still gives an assessment of GVU changes.
In conclusion, this study identified key cellular processes that are affected in atrophic DLPFC neurons in PSD. These neuronal pathways share common links to neurodegeneration suggesting a neurodegenerative process in those neurons that underlie the dementia seen in PSD. This may in turn appear related to the changes seen in the frontal white matter. Understanding these changes may allow better prediction of individuals who will develop dementia poststroke, while targeting these neurodegenerative cellular processes could potentially ameliorate this progressive dementia.
References
Savva GM, Stephan BC, Alzheimer’s Society Vascular Dementia Systematic Review G. Epidemiological studies of the effect of stroke on incident dementia: a systematic review. Stroke. 2010;41:e41-46. https://doi.org/10.1161/STROKEAHA.109.559880.
Allan LM, Rowan EN, Firbank MJ, Thomas AJ, Parry SW, Polvikoski TM, O’Brien JT, Kalaria RN. Long term incidence of dementia, predictors of mortality and pathological diagnosis in older stroke survivors. Brain : a journal of neurology. 2011;134:3716–27. https://doi.org/10.1093/brain/awr273.
Ballard C, Rowan E, Stephens S, Kalaria R, Kenny RA. Prospective follow-up study between 3 and 15 months after stroke: improvements and decline in cognitive function among dementia-free stroke survivors >75 years of age. Stroke. 2003;34:2440–4. https://doi.org/10.1161/01.STR.0000089923.29724.CE.
Bonelli RM, Cummings JL. Frontal-subcortical circuitry and behavior. Dialogues Clin Neurosci. 2007;9:141–51.
Pohjasvaara T, Leskela M, Vataja R, Kalska H, Ylikoski R, Hietanen M, Leppavuori A, Kaste M, Erkinjuntti T. Post-stroke depression, executive dysfunction and functional outcome. Eur J Neurol. 2002;9:269–75. https://doi.org/10.1046/j.1468-1331.2002.00396.x.
Chen A, Akinyemi RO, Hase Y, Firbank MJ, Ndung’u MN, Foster V, Craggs LJ, Washida K, Okamoto Y, Thomas AJ, et al. Frontal white matter hyperintensities, clasmatodendrosis and gliovascular abnormalities in ageing and post-stroke dementia. Brain : a journal of neurology. 2016;139:242–58. https://doi.org/10.1093/brain/awv328.
Foster V, Oakley AE, Slade JY, Hall R, Polvikoski TM, Burke M, Thomas AJ, Khundakar A, Allan LM, Kalaria RN. Pyramidal neurons of the prefrontal cortex in post-stroke, vascular and other ageing-related dementias. Brain : a journal of neurology. 2014;137:2509–21. https://doi.org/10.1093/brain/awu172.
Cai W, Zhang K, Li P, Zhu L, Xu J, Yang B, Hu X, Lu Z, Chen J. Dysfunction of the neurovascular unit in ischemic stroke and neurodegenerative diseases: an aging effect. Ageing Res Rev. 2017;34:77–87. https://doi.org/10.1016/j.arr.2016.09.006.
Lo EH, Rosenberg GA. The neurovascular unit in health and disease: introduction. Stroke. 2009;40:S2-3. https://doi.org/10.1161/STROKEAHA.108.534404.
Ballard C, Stephens S, Kenny R, Kalaria R, Tovee M, O’Brien J. Profile of neuropsychological deficits in older stroke survivors without dementia. Dement Geriatr Cogn Disord. 2003;16:52–6. https://doi.org/10.1159/000069994.
Kalaria RN, Kenny RA, Ballard CG, Perry R, Ince P, Polvikoski T. Towards defining the neuropathological substrates of vascular dementia. J Neurol Sci. 2004;226:75–80. https://doi.org/10.1016/j.jns.2004.09.019.
Altieri M, Di Piero V, Pasquini M, Gasparini M, Vanacore N, Vicenzini E, Lenzi GL. Delayed poststroke dementia: a 4-year follow-up study. Neurology. 2004;62:2193–7. https://doi.org/10.1212/01.wnl.0000130501.79012.1a.
Bahn S, Augood SJ, Ryan M, Standaert DG, Starkey M, Emson PC. Gene expression profiling in the post-mortem human brain–no cause for dismay. J Chem Neuroanat. 2001;22:79–94. https://doi.org/10.1016/s0891-0618(01)00099-0.
Waller R, Woodroofe MN, Francese S, Heath PR, Wharton SB, Ince PG, Sharrack B, Simpson JE. Isolation of enriched glial populations from post-mortem human CNS material by immuno-laser capture microdissection. J Neurosci Methods. 2012;208:108–13. https://doi.org/10.1016/j.jneumeth.2012.04.014.
Ferraiuolo L, Heath PR, Holden H, Kasher P, Kirby J, Shaw PJ. Microarray analysis of the cellular pathways involved in the adaptation to and progression of motor neuron injury in the SOD1 G93A mouse model of familial ALS. J Neurosci. 2007;27:9201–19. https://doi.org/10.1523/JNEUROSCI.1470-07.2007.
Simpson JE, Ince PG, Minett T, Matthews FE, Heath PR, Shaw PJ, Goodall E, Garwood CJ, Ratcliffe LE, Brayne C, et al. Neuronal DNA damage response-associated dysregulation of signalling pathways and cholesterol metabolism at the earliest stages of Alzheimer-type pathology. Neuropathol Appl Neurobiol. 2016;42:167–79. https://doi.org/10.1111/nan.12252.
Goodall EF, Leach V, Wang C, Cooper-Knock J, Heath PR, Baker D, Drew DR, Saffrey MJ, Simpson JE, Romero IAet al (2019) Age-associated mRNA and miRNA expression changes in the blood-brain barrier. Int J Mol Sci 20: https://doi.org/10.3390/ijms20123097
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57. https://doi.org/10.1038/nprot.2008.211.
Huang da W, Sherman BT, Zheng X, Yang J, Imamichi T, Stephens R, Lempicki RA (2009) Extracting biological meaning from large gene lists with DAVID. Curr Protoc Bioinformatics Chapter 13: Unit 13 11 https://doi.org/10.1002/0471250953.bi1311s27
Shibata M, Ohtani R, Ihara M, Tomimoto H. White matter lesions and glial activation in a novel mouse model of chronic cerebral hypoperfusion. Stroke. 2004;35:2598–603. https://doi.org/10.1161/01.STR.0000143725.19053.60.
Overhoff M, De Bruyckere E, Kononenko NL. Mechanisms of neuronal survival safeguarded by endocytosis and autophagy. J Neurochem. 2021;157:263–96. https://doi.org/10.1111/jnc.15194.
Merthan L, Haller A, Thal DR, von Einem B, von Arnim CAF. The role of PTB domain containing adaptor proteins on PICALM-mediated APP endocytosis and localization. Biochem J. 2019;476:2093–109. https://doi.org/10.1042/BCJ20180840.
Gomez-Suaga P, Rivero-Rios P, Fdez E, Blanca Ramirez M, Ferrer I, Aiastui A, Lopez De Munain A, Hilfiker S. LRRK2 delays degradative receptor trafficking by impeding late endosomal budding through decreasing Rab7 activity. Hum Mol Genet. 2014;23:6779–96. https://doi.org/10.1093/hmg/ddu395.
Liu G, Byrd A, Warner AN, Pei F, Basha E, Buchanan A, Buchan JR (2020) Cdc48/VCP and endocytosis regulate TDP-43 and FUS toxicity and turnover. Mol Cell Biol 40: https://doi.org/10.1128/MCB.00256-19
Borgonovo JE, Troncoso M, Lucas JJ, Sosa MA. Mutant huntingtin affects endocytosis in striatal cells by altering the binding of AP-2 to membranes. Exp Neurol. 2013;241:75–83. https://doi.org/10.1016/j.expneurol.2012.11.025.
Bera S, Camblor-Perujo S, Calleja Barca E, Negrete-Hurtado A, Racho J, De Bruyckere E, Wittich C, Ellrich N, Martins S, Adjaye J, et al. AP-2 reduces amyloidogenesis by promoting BACE1 trafficking and degradation in neurons. EMBO Rep. 2020;21: e47954. https://doi.org/10.15252/embr.201947954.
Tian Y, Chang JC, Fan EY, Flajolet M, Greengard P. Adaptor complex AP2/PICALM, through interaction with LC3, targets Alzheimer’s APP-CTF for terminal degradation via autophagy. Proc Natl Acad Sci U S A. 2013;110:17071–6. https://doi.org/10.1073/pnas.1315110110.
Correa SA, Eales KL. The role of p38 MAPK and its substrates in neuronal plasticity and neurodegenerative disease. J Signal Transduct. 2012;2012: 649079. https://doi.org/10.1155/2012/649079.
Kim EK, Choi EJ. Pathological roles of MAPK signaling pathways in human diseases. Biochim Biophys Acta. 2010;1802:396–405. https://doi.org/10.1016/j.bbadis.2009.12.009.
Chiarini A, Dal Pra I, Marconi M, Chakravarthy B, Whitfield JF, Armato U. Calcium-sensing receptor (CaSR) in human brain’s pathophysiology: roles in late-onset Alzheimer’s disease (LOAD). Curr Pharm Biotechnol. 2009;10:317–26. https://doi.org/10.2174/138920109787847501.
Guo Y, Liu C, Zhang J, Tian BB, Wang L, Liu GK, Liu Y. A relationship between MAPK/ERK pathway expression and neuronal apoptosis in rats with white matter lesions. Eur Rev Med Pharmacol Sci. 2020;24:4412–9. https://doi.org/10.26355/eurrev_202004_21023.
Hashimoto Y, Tsuji O, Niikura T, Yamagishi Y, Ishizaka M, Kawasumi M, Chiba T, Kanekura K, Yamada M, Tsukamoto E, et al. Involvement of c-Jun N-terminal kinase in amyloid precursor protein-mediated neuronal cell death. J Neurochem. 2003;84:864–77. https://doi.org/10.1046/j.1471-4159.2003.01585.x.
Hensley K, Floyd RA, Zheng NY, Nael R, Robinson KA, Nguyen X, Pye QN, Stewart CA, Geddes J, Markesbery WR, et al. p38 kinase is activated in the Alzheimer’s disease brain. J Neurochem. 1999;72:2053–8. https://doi.org/10.1046/j.1471-4159.1999.0722053.x.
Falcicchia C, Tozzi F, Arancio O, Watterson DM, Origlia N (2020) Involvement of p38 MAPK in synaptic function and dysfunction. Int J Mol Sci 21: https://doi.org/10.3390/ijms21165624
Berretta A, Tzeng YC, Clarkson AN. Post-stroke recovery: the role of activity-dependent release of brain-derived neurotrophic factor. Expert Rev Neurother. 2014;14:1335–44. https://doi.org/10.1586/14737175.2014.969242.
Song C, Zhang Y, Dong Y. Acute and subacute IL-1beta administrations differentially modulate neuroimmune and neurotrophic systems: possible implications for neuroprotection and neurodegeneration. J Neuroinflammation. 2013;10:59. https://doi.org/10.1186/1742-2094-10-59.
Alam JJ, Krakovsky M, Germann U, Levy A. Continuous administration of a p38alpha inhibitor during the subacute phase after transient ischemia-induced stroke in the rat promotes dose-dependent functional recovery accompanied by increase in brain BDNF protein level. PLoS ONE. 2020;15: e0233073. https://doi.org/10.1371/journal.pone.0233073.
Pozzi D, Menna E, Canzi A, Desiato G, Mantovani C, Matteoli M. The communication between the immune and nervous systems: the role of IL-1beta in synaptopathies. Front Mol Neurosci. 2018;11:111. https://doi.org/10.3389/fnmol.2018.00111.
Stefanoska K, Bertz J, Volkerling AM, van der Hoven J, Ittner LM, Ittner A. Neuronal MAP kinase p38alpha inhibits c-Jun N-terminal kinase to modulate anxiety-related behaviour. Sci Rep. 2018;8:14296. https://doi.org/10.1038/s41598-018-32592-y.
Tong L, Prieto GA, Kramar EA, Smith ED, Cribbs DH, Lynch G, Cotman CW. Brain-derived neurotrophic factor-dependent synaptic plasticity is suppressed by interleukin-1beta via p38 mitogen-activated protein kinase. J Neurosci. 2012;32:17714–24. https://doi.org/10.1523/JNEUROSCI.1253-12.2012.
Ding R, Hase Y, Ameen-Ali KE, Ndung’u M, Stevenson W, Barsby J, Gourlay R, Akinyemi T, Akinyemi R, Uemura MT, et al. Loss of capillary pericytes and the blood-brain barrier in white matter in poststroke and vascular dementias and Alzheimer’s disease. Brain Pathol. 2020;30:1087–101. https://doi.org/10.1111/bpa.12888.
Hase Y, Horsburgh K, Ihara M, Kalaria RN. White matter degeneration in vascular and other ageing-related dementias. J Neurochem. 2018;144:617–33. https://doi.org/10.1111/jnc.14271.
Rietdijk CD, Wezel RJA, Garssen J, Kraneveld AD (2016) Neuronal toll-like receptors and neuro-immunity in Parkinson’s disease, Alzheimer’s disease and stroke. Neuroimmunology and Neuroinflammation: 27-37 https://doi.org/10.20517/2347-8659.2015.28
Li L, Jin X, Zhang H, Yin J. Protective effect of picroliv against lipopolysaccharide-induced cognitive dysfunction and neuroinflammation by attenuating TLR4/NFkappaB pathway. Folia Neuropathol. 2018;56:337–45. https://doi.org/10.5114/fn.2018.80867.
Yu N, Hu S, Hao Z. Benificial effect of stachydrine on the traumatic brain injury induced neurodegeneration by attenuating the expressions of Akt/mTOR/PI3K and TLR4/NFkappa-B pathway. Transl Neurosci. 2018;9:175–82. https://doi.org/10.1515/tnsci-2018-0026.
Zhou L, Miranda-Saksena M, Saksena NK. Viruses and neurodegeneration. Virol J. 2013;10:172. https://doi.org/10.1186/1743-422X-10-172.
Goulopoulou S, McCarthy CG, Webb RC. Toll-like receptors in the vascular system: sensing the dangers within. Pharmacol Rev. 2016;68:142–67. https://doi.org/10.1124/pr.114.010090.
Ryan DG, Murphy MP, Frezza C, Prag HA, Chouchani ET, O’Neill LA, Mills EL. Coupling Krebs cycle metabolites to signalling in immunity and cancer. Nat Metab. 2019;1:16–33. https://doi.org/10.1038/s42255-018-0014-7.
Du SQ, Wang XR, Xiao LY, Tu JF, Zhu W, He T, Liu CZ. Molecular mechanisms of vascular dementia: what can be learned from animal models of chronic cerebral hypoperfusion? Mol Neurobiol. 2017;54:3670–82. https://doi.org/10.1007/s12035-016-9915-1.
van Vliet AR, Verfaillie T, Agostinis P. New functions of mitochondria associated membranes in cellular signaling. Biochim Biophys Acta. 2014;1843:2253–62. https://doi.org/10.1016/j.bbamcr.2014.03.009.
Weinberg SE, Sena LA, Chandel NS. Mitochondria in the regulation of innate and adaptive immunity. Immunity. 2015;42:406–17. https://doi.org/10.1016/j.immuni.2015.02.002.
Jian H, Yi-Fang W, Qi L, Xiao-Song H, Gui-Yun Z. Cerebral blood flow and metabolic changes in hippocampal regions of a modified rat model with chronic cerebral hypoperfusion. Acta Neurol Belg. 2013;113:313–7. https://doi.org/10.1007/s13760-012-0154-6.
Hase Y, Craggs L, Hase M, Stevenson W, Slade J, Chen A, Liang D, Ennaceur A, Oakley A, Ihara M, et al. The effects of environmental enrichment on white matter pathology in a mouse model of chronic cerebral hypoperfusion. J Cereb Blood Flow Metab. 2018;38:151–65. https://doi.org/10.1177/0271678X17694904.
Hase Y, Craggs L, Hase M, Stevenson W, Slade J, Lopez D, Mehta R, Chen A, Liang D, Oakley A, et al. Effects of environmental enrichment on white matter glial responses in a mouse model of chronic cerebral hypoperfusion. J Neuroinflammation. 2017;14:81. https://doi.org/10.1186/s12974-017-0850-5.
Bazargani N, Attwell D. Astrocyte calcium signaling: the third wave. Nat Neurosci. 2016;19:182–9. https://doi.org/10.1038/nn.4201.
Chai H, Diaz-Castro B, Shigetomi E, Monte E, Octeau JC, Yu X, Cohn W, Rajendran PS, Vondriska TM, Whitelegge JP, et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron. 2017;95(531–549): e539. https://doi.org/10.1016/j.neuron.2017.06.029.
Garwood C, Faizullabhoy A, Wharton SB, Ince PG, Heath PR, Shaw PJ, Baxter L, Gelsthorpe C, Forster G, Matthews FE, et al. Calcium dysregulation in relation to Alzheimer-type pathology in the ageing brain. Neuropathol Appl Neurobiol. 2013;39:788–99. https://doi.org/10.1111/nan.12033.
Nanclares C, Baraibar AM, Araque A, Kofuji P (2021) Dysregulation of astrocyte-neuronal communication in Alzheimer's disease. Int J Mol Sci 22: https://doi.org/10.3390/ijms22157887
Harada K, Kamiya T, Tsuboi T. Gliotransmitter release from astrocytes: functional, developmental, and pathological implications in the brain. Front Neurosci. 2015;9:499. https://doi.org/10.3389/fnins.2015.00499.
Derada Troletti C, de Goede P, Kamermans A, de Vries HE. Molecular alterations of the blood-brain barrier under inflammatory conditions: the role of endothelial to mesenchymal transition. Biochim Biophys Acta. 2016;1862:452–60. https://doi.org/10.1016/j.bbadis.2015.10.010.
Cai HQ, Catts VS, Webster MJ, Galletly C, Liu D, O’Donnell M, Weickert TW, Weickert CS. Increased macrophages and changed brain endothelial cell gene expression in the frontal cortex of people with schizophrenia displaying inflammation. Mol Psychiatry. 2020;25:761–75. https://doi.org/10.1038/s41380-018-0235-x.
Acknowledgements
This project was funded by the Alzheimer’s Society (AS-PG-17-007). CogFAST tissue for this study was provided by the Newcastle Brain Tissue Resource which is funded in part by a grant from the UK Medical Research Council (G0400074), by NIHR Newcastle Biomedical Research Centre and Unit awarded to the Newcastle upon Tyne NHS Foundation Trust and Newcastle university, and as part of the Brains for Dementia Research Programme jointly funded by Alzheimer’s Research UK and Alzheimer’s Society. BCAS tissue for this study was provided by Professor Raj Kalaria and Dr Yoshiki Hase at the Translational and Clinical Research Institute, Newcastle University, Newcastle upon Tyne. We would like to thank the Alzheimer’s Society Research Network Monitors for their support and advice during the project. We are grateful to our respondents and their families for their generous gift to medical research, which has made this study possible.
Author information
Authors and Affiliations
Contributions
SBW, RK: conception of the project. RW, JES, PRH, MW, SBW: study design. RW, MW: experimental work and data collection. RW, YH: data analysis. RW, JES, SBW: writing of first draft. JES, PRH, SBW: supervision of lab work. RW: collation of author contributions into final draft. All authors: contribution to interpretation and to the final manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12975_2022_1038_MOESM12_ESM.jpg
Supplementary file12 (JPG 1002 kb) Supplementary Figure 1: Laser capture microdissection cell isolation enrichment. Endothelial cells of vessels were identified by rapid Coll-V immunostaining. The laser is fired causing the film to melt and adhere to the underlying cell (A). The cap is lifted off with the adhered captured cells (B) leaving behind the remaining tissue (C). For LCM of neurons and astrocyte see references 52 and 58. RT-PCR analysis of cells isolated by immuno-LCM (D). Toluidine blue positive neurons are associated with high levels of NFL transcripts; GFAP positive astrocytes are associated with high levels of GFAP transcripts. Coll-IV positive vessels are associated with high levels of vWF transcripts. Key: Astro, astrocytes; GFAP, glial fibrillary acidic protein; LCM: laser capture microdissection; NFL, neurofilament light; vWF, von willibrand factor.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Waller, R., Hase, Y., Simpson, J.E. et al. Transcriptomic Profiling Reveals Discrete Poststroke Dementia Neuronal and Gliovascular Signatures. Transl. Stroke Res. 14, 383–396 (2023). https://doi.org/10.1007/s12975-022-01038-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12975-022-01038-z