Abstract
The timber of the species Lophira alata (azobe) is very popular for outdoor constructions, which favours its overexploitation and illegal logging. We sampled individuals from Liberia, Ivory Coast, Ghana, Nigeria, Cameroon, Gabon, Congo Brazzaville and Republic Democratic of Congo to discover new nuclear and plastidial SNP and INDEL loci through restriction associated DNA sequencing (RADSeq) and low coverage MiSeq genome sequencing. From an initial set of 397 loci, a final set of 126 loci was selected for timber tracking purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Lophira alata is a tropical pioneer timber species typical from secondary humid forests. Its distribution ranges from Sierra Leone in Western Africa to Congo in Central Africa. The timber is highly valuable because of its resistance against insects and moulding, which enables it’s use in outdoor constructions. Despite a minimum cutting diameter of at least 60 cm in all producer countries, this species is heavily exploited and classified as vulnerable in the IUCN red list.
The genus Lophira has been poorly studied until now and only consist of two species: L. alata typical from humid forest habitats while the species L. lanceolata is mostly found in savannahs. However, when both species are growing in sympatry, species identification is difficult (Biwole et al. 2012). Genetic studies are limited to the development and use of SSRs markers (Ewédjè et al. 2020; Pineiro et al. 2015), which aimed at studying the genetic structure and hybridisation between L. alata and L. lanceolata. With 10 SSRs, the presence of a cryptic species in L. alata was suggested in West Gabon (Ewédjè et al 2020). Thus, little information is available on the diversity and differentiation within both species.
After the entry into force of timber regulations (among others European Timber Regulation and US Lacey Act) requiring that the species identity and the country of origin are declared and correct, tracking tools to control the claims on species and origin received increasing attention (Dormontt et al. 2015). Although many genetic markers have been successfully developed on tropical and temperate species, the low DNA quality obtained from timber strongly hinders the use of SSRs markers in timber tracking (Blanc-Jolivet and Liesebach 2015). By contrast, SNP loci usually provide good results and sets of SNP loci for origin identification have already been developed for other African species (Jardine et al. 2016; Blanc-Jolivet et al. 2017, 2018).
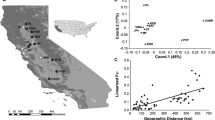
Cambium or leaf samples from 103 individual trees stemming from West and Central Africa (Table 1) were dried in silicagel in the field. DNA was extracted according to Dumolin et al. (1995). Three and nine samples were selected for respectively, restriction site associated DNA sequencing [RADSeq, (Miller et al. 2007)] and low coverage Illumina MiSeq genome sequencing (Straub et al. 2012). The use of both NGS methods allowed the SNP and INDEL discovery in the nuclear and in the plastidial genomes. A MassARRAY® iPLEX™ design [Assay Design Suite v2.0 (Agena Bioscience™, San Diego, USA)] included loci which showed polymorphism among sequenced individuals and for which the sequences did not show the presence of another SNP locus in a 50 bp neighbourhood. 397 loci were run in 12 multiplexes on a MassARRAY® iPLEX™ platform (Agena Bioscience™, San Diego, USA) using the iPLEX™ GOLD chemistry (Supplementary material S1) on 94 samples (Table 1). Genotyping was done with Typer Viewer v.4.0.24.71 (Agena Bioscience™, San Diego, USA).
In order to select loci showing a differentiation over the species distribution range, we grouped all individuals according to their country of origin. We estimated, when applicable, average diversity, Gregorius’ differentiation (Gregorius 1987) and correlation among genetic and geographic distance for each locus (Supplementary material S2). Because most African species show a strong genetic split between Western and Central African populations, we also estimated differentiation parameters within Western African and Central African countries to catch loci showing local genetic differentiation. On the three datasets, we selected the loci with the highest genetic differentiation, the highest genetic-geographic distance correlation, while loci with very low calling rates were excluded (GDA-NT, unpublished). 126 selected loci were included in four MassARRAY® iPLEX™ multiplexes (Supplementary material S3), from which 75, 20 and 28 SNP loci were located in the nuclear, chloroplastic and mitochondrial genome respectively. Two chloroplastic and one mitochondrial INDELs were also included. Theoretical performance for the verification of the country of origin reached 86% (self-assignment test with leave-one-Out procedure using the Bayesian criteria of Rannala and Mountain 1997) when we splited the Nigeria and Cameroon individuals according to a STRUCTURE analysis with two putative genetic groups (Pritchard et al. 2000). Indeed, the genetic boarder between West and Central Africa seems to be located between Nigeria and Cameroon where both groups are present in sympatry.
Our new set of 126 loci including both nuclear and plastidial loci will be of great importance for further population genetics and biogeographic studies as well as timber origin tracking in Lophira alata.
References
Biwole AB, Bourland N, Dainou K, Doucet JL (2012) Definition of the ecological profile of Lophira alata (ekki), a major important African timber species: literature review and perspectives for future studies. Biotechnol Agron Soc Environ 16:217–228
Blanc-Jolivet C, Liesebach M (2015) Tracing the origin and species identity of Quercus robur and Quercus petraea in Europe: a review. Silvae Genet 64:182–193
Blanc-Jolivet C, Kersten B, Dainou K, Hardy O, Guichoux E, Delcamp A, Degen B (2017) Development of nuclear SNP markers for genetic tracking of Iroko, Milicia excelsa and Milicia regia. Conserv Genet Resour 9:531–533
Blanc-Jolivet C, Kersten B, Bourland N, Guichoux E, Delcamp A, Doucet JL, Degen B (2018) Development of nuclear SNP markers for the timber tracking of the African tree species Sapelli, Entandrophragma cylindricum. Conserv Genet Resour 10:539–541
Dormontt EE, Boner M, Braun B, Breulmann G, Degen B, Espinoza E, Gardner S, Guillery P, Hermanson JC, Koch G, Lee SL, Kanashiro M, Rimbawanto A, Thomas D, Wiedenhoeft AC, Yin Y, Zahnen J, Lowe AJ (2015) Forensic timber identification: it's time to integrate disciplines to combat illegal logging. Biol Conserv 191:790–798
Dumolin S, Demesure B, Petit RJ (1995) Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor Appl Genet 91:1253–1256
Ewédjè EBK, Jansen S, Koffi GK et al (2020) Species delimitation in the African tree genus Lophira (Ochnaceae) reveals cryptic genetic variation. Conserv Genet 21:501–514
Gregorius HR (1987) The relationship between the concepts of genetic diversity and differentiation. Theor Appl Genet 74:397–401
Jardine DI, Blanc-Jolivet C, Dixon RRM, Dormontt EE, Dunker B, Gerlach J, Kersten B, van Dijk KJ, Degen B, Lowe AJ (2016) Development of SNP markers for Ayous (Triplochiton scleroxylon K. Schum) an economically important tree species from tropical West and Central Africa. Conserv Genet Resour 8:129–139
Miller MR, Dunham JP, Amores A, Cresko WA, Johnson EA (2007) Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Res 17:240–248
Pineiro R, Staquet A, Hardy OJ (2015) Isolation of nuclear microsatellites in the african timber tree Lophira alata (Ochnaceae) and cross-amplification in L-Lanceolata. Appl Plant Sci. https://doi.org/10.3732/apps.1500056
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Straub SCK, Parks M, Weitemier K, Fishbein M, Cronn RC, Liston A (2012) Navigating the tip of the genomic iceberg: next-generation sequencing for plant systematics. Am J Bot 99:349–364
Acknowledgements
This research was supported by the German Federal Ministry of Food and Agriculture in the frame of the “Large scale project on genetic timber verification” (Project No. 28I-001-01). Genotyping was performed at the Genome Transcriptome Facility of Bordeaux (Grants from ANR-10-EQPX-16), with the help of Erwan Guichoux and Adline Delcamp. The CNIAF of republic of Congo, the MINFOF of Cameroon, the Tshopo provincial ministry in charge of environment, the Institut Supérieur des Etudes Agronomiques de Bengamisa (ISEA), the Institut Faculté des Sciences Agronomiques de Yangambi (IFA/Yangambi) of DRC, the Forestry Commission of Ghana, the SODEFOR of Cote d'Ivoire, the Liberia Forest Authority and the Nigeria Forest Service assisted the sampling activities. Laboratory work was conducted by Susanne Bein and Maike Paulini, Lennart Meyer-Sand and Birte Pakull organised sampling, sample preparation and documentation.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Blanc-Jolivet, C., Mader, M., Bouda, HN. et al. Development of new SNP and INDEL loci for the valuable African timber species Lophira alata. Conservation Genet Resour 13, 85–87 (2021). https://doi.org/10.1007/s12686-020-01173-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-020-01173-5