Abstract
Long noncoding RNAs (lncRNAs), which are more than 200 nucleotides in length and do not encode proteins, play crucial roles in governing gene expression at both the transcriptional and posttranscriptional levels. These molecules demonstrate specific expression patterns in various tissues and developmental stages, suggesting their involvement in numerous developmental processes and diseases, notably cancer. Despite their widespread acknowledgment and the growing enthusiasm surrounding their potential as diagnostic and prognostic biomarkers, the precise mechanisms through which lncRNAs function remain inadequately understood. A few lncRNAs have been studied in depth, providing valuable insights into their biological activities and suggesting emerging functional themes and mechanistic models. However, the extent to which the mammalian genome is transcribed into functional noncoding transcripts is still a matter of debate. This review synthesizes our current understanding of lncRNA biogenesis, their genomic contexts, and their multifaceted roles in tumorigenesis, highlighting their potential in cancer-targeted therapy. By exploring historical perspectives alongside recent breakthroughs, we aim to illuminate the diverse roles of lncRNA and reflect on the broader implications of their study for understanding genome evolution and function, as well as for advancing clinical applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Background: the emergence of long noncoding RNAs
Long noncoding RNAs (lncRNA) have become integral to our understanding of genome evolution, size, and complexity, a process that began in the 1950s. The investigation of the C-value, which measures the DNA content of a haploid genome, revealed intriguing paradoxes, notably the limited correlation between DNA quantity and organismal complexity [1, 2]. This “C-value paradox” sparked widespread discussion, particularly with discoveries that some organisms, considered evolutionarily “simpler,” possess genomes significantly larger than those of “higher” organisms, including humans [3, 4]. A pivotal moment in resolving this paradox came with the realization that a substantial fraction of the genome is noncoding and not dedicated to protein synthesis [5, 6]. Initially, termed “junk DNA,” these noncoding regions, which make up 50–70% of the human genome, were later found to play crucial roles beyond their initial dismissal as genomic filler [6,7,8,9,10,11]. The 1970s marked the beginning of a shift in perspective with the observation of “pervasive transcription,” indicating that a vast portion of the genome, beyond known coding regions, is transcribed. These RNAs include not only coding genes but also heterogeneous nuclear RNAs (hnRNAs) and other RNA types, such as rRNA and tRNA [6, 12, 13]. The late 1990s and early 2000s brought technological advancements that further illuminated the genome’s transcriptional landscape, such as the recognition of ribozymes in 1989, which highlighted the catalytic capabilities of RNA, and the discovery of XIST in 1991, which provided a new understanding of X chromosome inactivation. The subsequent discovery of lin-4 in 1993 pioneered the exploration of microRNAs (miRNAs), a class of small noncoding RNAs that, similar to lncRNAs, are involved in the posttranscriptional regulation of gene expression [14,15,16,17,18]. Early hypotheses by Jacob and Monod and later by Britten and Davidson proposed regulatory roles for noncoding RNAs, suggesting their involvement in gene expression modulation and signal transmission [19, 20]. The discovery of lncRNAs such as H19 and Xist in the early 1990s underscored their significance in epigenetic regulation [21,22,23,24]. Challenging the notion of “transcriptional noise” and sparking debates about the functional relevance of these transcripts [25,26,27,28]. The ongoing exploration of lncRNA is progressively unveiling the complex mechanisms of gene expression regulation, revealing a complex landscape of genomic functionality that extends significantly beyond the coding sequences traditionally emphasized in molecular biology. This burgeoning field of research, as illustrated in Fig. 1, promises to substantially deepen our understanding of genome architecture and its regulation, implicating the expression of lncRNAs in pivotal developmental stages, physiological conditions, and a spectrum of pathologies, notably cancer.
2 Complexity and diverse functional implications in genomic contexts
2.1 General features of lncRNA
LncRNAs, which are characterized by a length surpassing 200 nucleotides and lack protein-coding capability, are predominantly transcribed by RNA polymerase II. This mirrors the intricate and adaptable nature of mRNA in terms of structure, composition, and function [29, 30]. These molecules play pivotal roles in the intricate dance of genomic regulation, engaging with proteins, DNA, and RNA to orchestrate the organization of the genome, cellular architecture, and nuanced layers of gene expression regulation [29, 31,32,33]. Their influence extends from the nucleus to the cytoplasm, where they oversee critical processes such as translation, metabolism, and signal transduction, demonstrating a remarkable specificity to tissue types and developmental stages [34,35,36,37]. Among the diverse classes of lncRNAs, stand-alone lncRNA, or "lincRNA," represent unique transcription units that operate independently of protein-coding genes. These lncRNAs, including notable examples such as Xist, H19, HOTAIR, and MALAT1, are characterized by their transcription by RNA Pol II, subsequent polyadenylation, and splicing, typically spanning an average length of 1 kb [38,39,40,41]. This independence from coding sequences allows them to serve as crucial regulators of gene expression and chromatin architecture.
Natural antisense transcripts (NATs) add another layer of complexity, often forming sense‒antisense pairs with coding transcripts, such as Xist/Tsix and Kcnq1/Kcnq1ot1. Despite their abundance, the full extent of the biological functions of NATs remains an area ripe for exploration [42,43,44,45,46]. Similarly, pseudogenes, the genomic remnants once considered mere evolutionary artifacts, have revealed their capacity to influence gene expression, challenging our understanding of genomic ‘junk’ [47,48,49]. Intriguingly, long intronic ncRNAs, found within the introns of annotated genes, exhibit diverse expression patterns and have been implicated in specific biological processes, such as the regulation of plant vernalization by COLDAIR [50, 51]. Moreover, the genomic landscape was further enriched by divergent, promoter-linked, and enhancer-associated RNAs, including TSSa-RNAs, uaRNAs, and PROMPTs. Although these short transcripts are often rapidly degraded, their production at transcription start sites and enhancers hints at potential regulatory roles yet to be fully understood [52,53,54,55,56,57,58].
2.2 Functions and mechanisms of lncRNA
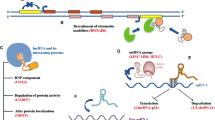
Insights into the multifaceted mechanisms of lncRNA are pivotal for elucidating their roles in diseases, particularly cancer. Studies have highlighted the integration of lncRNAs into cellular molecular networks, underscoring their importance in developing cancer therapies [59,60,61,62]. Table 1 categorizes a spectrum of lncRNAs associated with various cancers, delineating their unique roles and actions. These lncRNAs are intricately regulated, integral to the complex network of cellular regulation, and instrumental to both disease pathogenesis and normal physiological processes. They interact with DNA, RNA, and proteins, significantly influencing gene expression (Fig. 2). As signals, certain lncRNAs, such as Xist, which is transcribed from the inactive X chromosome, indicate cellular states and trigger X chromosome inactivation [63, 64]. Scaffold lncRNA, such as HOTAIR, form structural bases for regulatory complexes, recruiting enzymes for chromatin modification and consequently altering gene expression [65]. Decoy lncRNA, such as PANDA, bind transcription factors, thereby preventing them from activating proapoptotic genes [66]. Competing with endogenous RNA (ceRNA), lncRNA can sponge miRNAs, protecting target mRNAs from degradation—a mechanism exhibited by LINC00680, which binds to miR-423-5p to modulate PAK6 expression in esophageal squamous cell carcinoma [67]. Guide lncRNA, exemplified by MEG3, direct transcriptional machinery to specific genomic regions, modulating gene expression patterns [68].
Functional mechanisms of lncRNA A In response to diverse stimuli, lncRNA regulates signaling pathways by binding to transcription factors. B Acting as miRNA sponges, lncRNA inhibits mRNA degradation. C Serving as scaffolds, lncRNA aids in forming protein complexes, thereby regulating target gene transcription. D LncRNA also guides ribonucleoprotein complexes to specific DNA sequences, influencing gene expression
3 lncRNA biogenesis during pathological processes
3.1 Transcriptional mechanisms
During pathological processes such as cancer, the landscape of lncRNAs undergoes significant changes, with specific classes of lncRNA emerging as key players in disease progression (Fig. 3). The abnormal regulation of transcription processes can lead to the production of lncRNAs closely associated with tumorigenesis and metastasis, known as oncogenic lncRNAs. These lncRNAs are instrumental in driving tumor growth and conferring resistance to therapeutic interventions [69,70,71,72]. This complex interplay between lncRNAs and the cellular machinery adds layers of regulation and functionality, underscoring the versatility of lncRNAs in gene expression and cellular dynamics. Some lncRNAs may not play direct functional roles but instead contribute to cellular processes as structural scaffolds or decoys. This lncRNA can influence gene expression indirectly by modulating the transcriptional landscape or affecting the stability of other RNA molecules.
3.2 Posttranscriptional modifications
Furthermore, lncRNAs interact with various RNA modifications, notably N6-methyladenosine (m6A), which influences their stability, localization, and overall function within the cell. m6A modification, which occurs at specific genomic loci, plays a crucial role in the posttranscriptional regulation of lncRNAs, affecting their role in cancer and other diseases [73]. Recent studies have shed light on how the modulation of enzymes responsible for m6A methylation can significantly alter the biogenesis and functionality of lncRNAs, highlighting a subtle yet impactful mechanism of gene regulation with minimal effects on the coding transcriptome [74]. In some instances, lncRNAs are produced as a result of transcriptional noise, serving no clear benefit to the cell, yet their presence underscores the complexity of genomic transcription [33]. Despite the intricate nature of lncRNA biology and the challenges it presents, the critical role of lncRNAs in a wide array of diseases, particularly cancer, cannot be overstated. The involvement of lncRNAs in key cellular processes and disease mechanisms positions lncRNAs as promising targets for therapeutic intervention, offering new avenues for treatment strategies aimed at modulating their expression or function.
LncRNAs are categorized into five primary types based on their genomic location: antisense, bidirectional, intronic, enhancer-associated, and intergenic. Intergenic and enhancer-associated lncRNAs have distinct promoters and are independent of protein-coding genes. Conversely, bidirectional lncRNAs share a common promoter but are transcribed from the opposite strand of a protein-coding gene. Intronic lncRNAs originate within the introns of genes that encode proteins (Fig. 4).
Regulation and classification of lncRNA. A The figure’s lower left section illustrates the transcriptional regulation mechanisms of lncRNA. It shows basal transcriptional activation ‘Part 1’, and its enhancement during inflammatory responses ‘Part 2’ triggered by the activation of pattern recognition receptors (PRR). It presents three lncRNA gene examples, labeled A, B, and C. ‘Part 3’ demonstrates the co-transcriptional regulation of lncRNA through differential isoform expression, which can involve alternative splicing or the employment of novel transcription start sites in response to inflammatory stimuli like lipopolysaccharides (LPS). The representation of post-transcriptional regulation of lncRNAs is categorized into three segments: 4, 5, and 6. Following transcription, lncRNAs undergo diverse processes. In ‘Part 4,’ RNA modifications are depicted, influencing the structural configuration of the lncRNA molecule. These modifications can be reversible, contingent upon the cellular inflammatory condition. ‘Part 5’ delineates the transformation of lncRNA into mature miRNA during miRNA biogenesis. ‘Part 6’ signifies the potential translation of lncRNAs containing small open reading frames (smORFs). B The lower right section of the figure illustrates the regulatory roles of lncRNAs within the nucleus and cytoplasm. It indicates that during transcription, both basal ‘Part 1’ and inflammatory ‘Part 2’ lncRNAs can either suppress (mRNA genes A and C) or enhance (mRNA genes A and B) gene expression
3.3 LncRNAs as regulators of transcription
By playing a crucial role in the extensive landscape of the human genome, lncRNAs function as transcriptional regulators, coordinating gene expression at the transcriptional level [75]. Their ability to modulate chromatin architecture and interact with transcription factors, DNA, and other molecular entities, such as RNA‒DNA hybrids (R-loops), positions them as key players in gene regulation [75,76,77,78]. This regulatory capacity is particularly evident in pathological states such as cancer, where aberrant lncRNA expression correlates with altered gene expression profiles, presenting new avenues for therapeutic intervention [79]. In the context of posttranscriptional regulation, lncRNAs act as miRNA sponges, influencing gene expression by sequestering miRNAs and preventing them from degrading target mRNAs. This mechanism is crucial in various cancers, underscoring the significance of lncRNA‒miRNA interactions in disease progression [80].
3.4 LncRNAs as miRNA sponges
Recent studies have highlighted the crucial roles played by lncRNAs in cellular processes, with a particular emphasis on their role as miRNA sponges in cancer biology. The posttranscriptional modulation of gene activity by lncRNAs through miRNA sponging is a significant factor in various cancer types, underscoring the importance of the interplay between lncRNAs and miRNAs in tumorigenesis. Notably, the lncRNA UCA1, characterized by its structural stability attributed to specific genetic variations, has been linked to an elevated risk of endometriosis, a condition exhibiting notable parallels to cancer [81]. By functioning as a competing endogenous RNA (ceRNA), UCA1 hinders the activity of miRNAs, resulting in the dysregulation of downstream genes. Gene network analyses revealed alterations in UCA1 expression associated with pivotal pathways, such as fatty acid metabolism and mitochondrial beta-oxidation, in the context of endometriosis. In a parallel manner, LINC00662, which is upregulated in prostate cancer, serves as a miR-34a sponge, exerting influence on the progression of the disease [82, 83]. In hepatocellular carcinoma (HCC), ZFPM2-AS1 functions as a competing endogenous RNA (ceRNA), competitively binding with miR-139 and consequently influencing the expression of GDF10, a gene associated with the progression of HCC [84]. Research in breast cancer has focused on the involvement of miRNA sponges in the regulation of protein‒protein interactions (PPIs), emphasizing the essential role of interactions between lncRNAs and miRNAs in cancer [85].
3.5 LncRNA as modulators of RNA stability
LncRNAs, including mRNAs and other noncoding RNAs, are instrumental in regulating the stability of RNA molecules. This regulatory function influences gene expression and impacts diverse cellular processes [86]. For instance, CiRS-7/CDR1as stabilizes its target mRNA through RNA duplex formation [87], and HNF1A-AS1 engages the ELAV1 protein to augment the stability and translation of particular mRNAs, such as CKAP5, involved in microtubule function and cancer cell cycle progression [88]. By forming an RNA‒protein complex with IGF2BP2, the lncRNA PCAT6 increases the stability of IGF1R mRNA, affecting IGF1R expression [89]. Additionally, the m6A modification of lncRNAs, as observed for HNF1A-AS1, alters their stability, revealing the nuanced regulation of gene expression via posttranscriptional modifications [88].
4 LncRNA and RBP interactions
In the regulatory landscape, lncRNAs serve as crucial partners for RNA binding proteins (RBPs), guiding their localization and function within cells [90]. This interaction allows lncRNAs to influence cellular processes significantly, from modulating the nuclear translocation of proteins to acting as scaffolds for complex molecular structures [91, 92]. Notably, HuR is upregulated in cancer and plays a pivotal role in disease progression by post-transcriptionally regulating cancer-related mRNAs [93, 94]. It enhances the stability of NEAT1 in ovarian cancer, influencing crucial cancer pathways as a ceRNA [95], and stabilizes lncRNA-HGBC in gallbladder carcinoma to promote cell proliferation and invasion [96]. Conversely, HuR can degrade lncRNAs such as p53-regulated lncRNA-p21, demonstrating its context-dependent dual role in RNA stability [97]. Similarly, serine/arginine-rich splicing Factor 1 (SRSF1) is implicated in glioma, where its knockdown alters mRNA and lncRNA expression, notably downregulating NEAT1, confirming its role in maintaining NEAT1 levels [98]. This finding indicates the nuanced roles of RBPs such as SRSF1 and HuR in modulating lncRNA stability, which impacts cancer progression [95, 98]. The AUF1 complex, which arises from alternative splicing of the HNRNPD gene, also plays a significant role in posttranscriptional lncRNA regulation [99]. It competes with HuR to bind U-rich sequences of AREs, influencing the decay or stabilization of lncRNAs such as NEAT1, and the effects of MALAT1 AUF1 vary, destabilizing NEAT1, which disperses nuclear paraspeckles and affects mRNA export; however, it does not impact MALAT1 stability, underscoring the complexity of its lncRNA interactions [100]. PABPN1, another nuclear RBP, binds the 3ʹ poly(A) tail of RNA, influencing the stability of lncRNAs such as NEAT1 and TUG1. The depletion of PABPN1 in HeLa cells impacts specific polyadenylated lncRNAs, indicating its role in sophisticated regulatory networks involving the nuclear exosome [101, 102]. IGF2BP1, which is frequently reactivated in cancers such as HCC, interacts with the lncRNA HULC to uniquely promote its degradation, a departure from its usual stabilizing role with mRNAs. This interaction involving the CCR4-NOT complex highlights the impact of structural motifs and protein partners on RBP–lncRNA interactions [103]. Tristetraprolin (TTP), unlike HuR and AUF1, generally acts as a tumor suppressor by destabilizing oncogenic RNAs such as HOTAIR. Its downregulation in cancer cells leads to reduced RNA degradation, underscoring the importance of RBPs such as TTP in cancer progression and their potential as therapeutic targets [104, 105]. The binding of lncRNAs to RBPs can alter the function of these proteins, influencing mRNA stability and translation, RNA splicing, and other critical regulatory processes. In cancer, these interactions often lead to changes in cell proliferation, apoptosis, and metastasis, making them potential targets for therapeutic intervention.
5 LncRNAs as translated peptides
Challenging the traditional view of lncRNAs as merely noncoding RNAs, recent studies have revealed their potential to engage with ribosomes and translate into functional peptides [106, 107]. This novel understanding, supported by comprehensive studies such as those by L. Minati et al. and ribosome profiling efforts by Bernardo Bonilauri, opens up exciting possibilities for lncRNA functionality beyond mere transcriptional regulation. The identification of lncRNAs with small open reading frames (smORFs) suggests a hidden layer of proteomic complexity and functional diversity within the realm of noncoding RNAs [108, 109]. Among these, several lncRNAs were found to contain small open reading frames (smORFs) that potentially encode functional microproteins. In colorectal cancer research, the lncRNA EVADR, influenced by Fusobacterium nucleatum infection, has been shown to play a role in cancer metastasis by acting as a scaffold for YBX1 and promoting EMT-related translation [110]. The development of tools such as LncDC by Minghua Li and Chun Liang further enhanced our ability to uncover lncRNAs with translational capabilities, suggesting that a subset of these molecules may indeed encode functional peptides, challenging their conventional classification and highlighting their potential in novel therapeutic strategies [107, 111, 112].
6 Challenges and approaches in identifying and characterizing lncRNAs
The integration of lncRNAs in clinical practice for cancer treatment necessitates accurate RNA analysis to identify and quantify novel RNA species. The unique sequences of lncRNAs and their potential overlap with other RNA types present significant challenges. For example, the HOTTIP lncRNA has been identified as a key regulator in gastrointestinal cancers, presenting new opportunities for diagnosis and treatment. Furthermore, research into plant stress responses and colorectal cancer has underscored the importance of lncRNAs in elucidating disease mechanisms and their utility as biomarkers in exosomes for early detection and intervention [113, 114].
6.1 Techniques for identifying lncRNAs
The landscape of transcriptomics has been transformed by RNA sequencing (RNA-seq) and other high-throughput sequencing technologies. These advancements allow for comprehensive annotation and quantification of a diverse range of RNAs, encompassing both coding and noncoding transcripts [115]. The selective extraction of polyadenylated RNA, excluding ribosomal RNA (rRNA), has been instrumental in isolating lncRNAs [116]. Continued progress, including the creation of libraries depleted of rRNA and the implementation of random priming during cDNA synthesis, has enhanced the precision of characterizing lncRNAs [117]. Traditional methods, including RT‒qPCR and Northern blotting, are fundamental for the validation and quantification of specific lncRNAs [118]. Given the vast diversity of lncRNAs, achieving specificity in their study is crucial. Techniques employing RNAe H to target RNA‒DNA hybrids have been developed to identify particular lncRNAs within complex RNA landscapes [119]. Microarray analysis, using probes designed for specific lncRNA sequences, is an initial screening tool, although its results require rigorous validation due to potential inconsistencies [120]. The visualization and quantification of lncRNAs within cells are crucial for comprehending their biological functions. Techniques such as RNA fluorescence in situ hybridization (RNA-FISH) are employed to evaluate the spatial distribution of lncRNAs, revealing their interactions with proteins and miRNAs. This approach offers valuable insights into the functional mechanisms of lncRNAs [121]. Additionally, RNA-FISH has played a crucial role in revealing the interactions between lncRNAs and other cellular components, including proteins and miRNAs [122]. Recent progress has resulted in the creation of assays characterized by improved sensitivity and specificity. Notably, reverse transcription-droplet digital polymerase chain reaction (RT-ddPCR) is a robust technique for accurately quantifying even low-abundance lncRNAs. Additionally, the rolling circle amplification (RCA) technique represents another innovative method that facilitates the amplification of specific lncRNA sequences to achieve clear and precise detection [123].
6.2 Approaches for the functional assessment of lncRNAs
Beyond detection, understanding the function of lncRNAs requires a comprehensive approach. lncRNAs interact with a broad spectrum of molecular entities, including DNA, RNA, microRNA (miRNA), and proteins, thereby influencing gene expression and cellular signaling pathways [124]. For instance, the lncRNA GAS5 impacts hepatic lipid metabolism, with its knockdown resulting in a reduction in lipid accumulation, suggesting its potential as a target for treating conditions such as NAFLD [125]. Similarly, the lncRNA MG828507, located upstream of the FLT1 gene, has been associated with preeclampsia, highlighting its significance in disease pathogenesis [126]. Computational tools have become invaluable in elucidating lncRNA functions, especially in mapping lncRNA interactions with proteins and RNA. These tools utilize a range of models, from ensemble-based and machine-learning approaches to molecular docking and network analyses, facilitating a deeper understanding of lncRNA roles [127]. Cutting-edge sequencing technologies, such as Oxford Nanopore, provide thorough insights into the sequences of lncRNAs, while NanoString platforms allow precise detection and quantification without the necessity for amplification or reverse transcription [128, 129].
7 LncRNA as potential biomarkers
The roles of lncRNAs in numerous physiological processes, such as cell differentiation and proliferation, are gaining increasing recognition. This finding positions them as significant contributors to the development and progression of cancer [130]. Their association with specific clinical features, such as tumor grade, size, metastasis stage, and overall aggressiveness, underscores their potential as biomarkers for cancer [131]. The distinct expression profiles, stability, specificity, and distribution of lncRNAs enable their identification and measurement in bodily fluids through liquid biopsy. This method provides a noninvasive approach for early cancer detection, diagnosis, prognosis, and monitoring of therapeutic responses (Fig. 5). Ongoing research has revealed the potential of lncRNAs as liquid-based diagnostic biomarkers, particularly for head and neck cancer (HNC) and other malignancies [132]. Cutting-edge sequencing technologies, such as Oxford Nanopore, offer an in-depth understanding of lncRNA sequences. Moreover, NanoString platforms enable the accurate detection and quantification of these sequences without the necessity for amplification or reverse transcription [133].
Certain lncRNA have shown significant prognostic value across various cancers. For instance, in gastric cancer, early detection linked to specific lncRNA profiles correlates with improved 5-year survival rates [134]. In colorectal cancer (CRC), a 3-lncRNA signature has been identified as a potential prognostic marker, emphasizing the role of lncRNAs in predicting disease progression and outcomes. Research at the Oncology Institute of Porto into lncRNA/miRNA ceRNA networks highlights their importance in cancer prognosis [135, 136]. Pancancer analyses have revealed the prognostic significance of lncRNAs, positioning them as crucial elements in cancer research and potential therapeutic targets [137]. In breast cancer, the analysis of lncRNA-TF-associated ceRNA networks has led to the identification of novel prognostic biomarkers, illustrating the intricate interplay between lncRNAs and other cellular components in cancer progression [138].
8 lncRNA as novel cancer biomarkers
8.1 Biomarkers for diagnosis
LncRNA has emerged as a significant biomarker in cancer, offering insights into diagnosis, prognosis, and the prediction of therapeutic responses. The increasing mortality rate from cancer has intensified the search for reliable diagnostic tools, with circulating lncRNAs showing considerable promise. These molecules are being investigated for their potential to detect various cancers early, including hepatocellular carcinoma, colorectal cancer, gastric cancer, renal cell carcinoma, and prostate cancer, through liquid biopsies [132, 139,140,141,142]. Prostate cancer antigen 3 (PCA3) has become an instrumental biomarker for prostate cancer, notably enhancing early detection and screening accuracy beyond traditional prostate-specific antigen (PSA) testing [143]. The higher specificity of PCA3 helps mitigate the frequency of unnecessary biopsies, presenting a significant advantage over PSA tests, whose levels may vary with prostate size. [144, 145]. Notably, PCA3 is the first long noncoding RNA biomarker to gain FDA approval for cancer screening, establishing a groundbreaking precedent in cancer diagnostics [144, 146]. In practice, PCA3 is quantified from urine samples obtained post-digital rectal exam, offering a noninvasive assessment tool for men with elevated PSA levels [145]. The integration of PCA3 into existing diagnostic frameworks is continually advancing, with research efforts focused on improving diagnostic accuracy by combining it with other markers and imaging techniques. Additionally, its potential in personalized medicine is being explored to tailor treatment strategies based on individual biomarker profiles.
8.2 Biomarkers for prognosis
Specifically, the lncRNA HOTAIR has been identified as a crucial marker in glioma, where its elevated expression correlates with higher disease grades and poorer patient outcomes, highlighting its role in both diagnosis and prognosis [147]. In addition to their diagnostic utility, lncRNAs serve as valuable prognostic indicators, with HOTAIR standing out for its independent prognostic value in glioma. This finding not only suggests the potential for targeted therapeutic strategies but also underscores the broader applicability of lncRNAs in predicting disease progression. Furthermore, a deep learning study demonstrated the effectiveness of specific lncRNAs in predicting significant disease associations, reinforcing their role in prognosis [147, 148].
9 LncRNAs as novel therapeutic targets
9.1 Inhibiting oncogenic lncRNAs
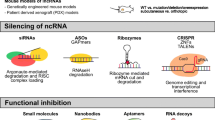
The modulation of lncRNA expression through RNA interference (RNAi) techniques, such as the use of small interfering RNA (siRNA) or short hairpin RNA (shRNA), offers a targeted approach to silence oncogenic lncRNAs in vivo. For instance, targeting HOTAIR in breast cancer and MALAT1 across various cancers has demonstrated potential in reducing tumor growth and improving prognosis [149, 150]. Antisense oligonucleotides (AONs) further enable the selective inhibition of lncRNAs such as GAS5, showing efficacy in tumor reduction [151].
9.2 Enhancing tumor-suppressive lncRNAs
Conversely, strategies to upregulate lncRNAs, particularly those that act as miRNA sponges or competing endogenous RNAs (ceRNAs), have been explored to counteract oncogenic miRNAs. Techniques employing lentivirus or adeno-associated virus vectors and nanoparticle encapsulation have facilitated the overexpression of lncRNAs such as GAS5 and UCA1, sensitizing cancer cells to radiation or chemotherapy and offering new avenues for overcoming treatment resistance [151,152,153].
9.3 Combining RNA therapy with conventional therapies
Advances in delivery methods, including the use of nanoparticles such as gold nanoparticles (AuNPs), have enhanced the efficiency of RNAi and AONs in targeting lncRNAs such as NEAT1 and XIST, contributing to tumor growth inhibition and increased survival in preclinical models [154, 155]. Additionally, drug inhibitors targeting lncRNAs such as ANRIL have shown promise in diminishing tumorigenicity, highlighting the therapeutic potential of modulating lncRNA activity [156]. In gastric cancer, the lncRNA CRNDE has been recognized as a crucial modulator of autophagy-related chemoresistance. Elevated CRNDE levels sensitize gastric cancer cells to chemotherapy by suppressing autophagy. Targeting the E2F6-CRNDE axis has emerged as a potential therapeutic strategy against chemoresistance in gastric cancer [157].
10 Overcoming treatment resistance and sensitivity through lncRNA modulation
The expression of lncRNAs is a key factor in cancer treatment resistance. LncRNAs such as H19 and UCA1 have been implicated in chemoresistance mechanisms in ovarian and gastric cancers, respectively, through their interaction with miRNAs and signaling pathways [158, 159]. Targeting these lncRNAs offers a strategy to enhance the efficacy of conventional therapies, such as cisplatin and tamoxifen, and improve patient outcomes. Moreover, the role of lncRNAs in promoting drug resistance through exosomal transfer, as observed with TP73-AS1 in glioblastoma and its contribution to temozolomide resistance, emphasizes the complexity of cancer biology and the potential of lncRNAs as therapeutic targets [160,161,162]. In cancer therapy, lncRNAs are pivotal for understanding treatment resistance. The expression profiles of these genes can change in response to chemotherapy, radiotherapy, and immunotherapy, affecting treatment outcomes [163]. For example, the lncRNA SNHG6, which acts as a miR-101 sponge, promotes epithelial–mesenchymal transition (EMT), a process linked to metastasis and resistance, particularly by influencing the responsiveness of breast cancer to tamoxifen [164]. Similarly, in colorectal cancer, UCA1 has been shown to confer resistance to cetuximab by modulating the miR-495 and HGF/c-MET signaling pathways, indicating its role in therapeutic resistance [165]. Prostate cancer cells resistant to androgen receptor inhibitors such as enzalutamide exhibit altered lncRNA expression patterns, suggesting a mechanism for the development of resistance [166]. In chemotherapy-sensitive lung cancer tissues, the lncRNA MEG3 is expressed at lower levels, and its overexpression reduces autophagy levels, thereby enhancing the effectiveness of vincristine in lung cancer chemotherapy [167]. Reducing the expression of the lncRNA HOXD-AS1 in glioma cells decreased their proliferation, migration, and invasion while increasing their sensitivity to cisplatin (DDP). This effect is mediated through the sequestration of miR-204, suggesting that HOXD-AS1 could serve as a viable therapeutic target for glioma treatment [168].
11 Conclusion and future prospects for lncRNA research
Advances in sequencing technologies have significantly refined our understanding of lncRNAs, increasingly distinguishing functional molecules from mere transcriptional noise. This progress, however, underscores the need for rigorous validation to confirm the biological roles of lncRNAs, with current guidelines aiming to streamline these efforts. Emerging at the forefront of cancer research, lncRNAs offer promising avenues for diagnostic and therapeutic innovations due to their involvement in key physiological processes. However, their full impact on cancer progression and utility in clinical settings warrants careful scrutiny, especially considering the potential unintended effects in healthy tissues and the complexity of their molecular pathways. The evolution of lncRNA research necessitates rigorous biological validation using patient samples alongside comprehensive functional studies to harness their potential effectively. Enhancing the sensitivity, specificity, and accuracy of lncRNA biomarkers by employing advanced molecular tools such as RNA-seq, RNA-FISH, ic-SHAPE, and quantitative real-time PCR is crucial. These techniques are essential for accurately measuring lncRNA levels in biological samples, providing insights into their roles in health and disease. Moreover, the safety, toxicity, and side effects associated with lncRNA-targeted therapies require thorough evaluation. Innovations in delivery methods, such as nanoparticle-based systems, and the integration of RNA therapy with other treatments are critical to advancing therapeutic efficacy and improving patient outcomes. Additionally, employing computational methods to understand how lncRNAs predict disease associations is pivotal, utilizing machine learning and other data-driven approaches to model lncRNA interactions and their functional impacts. Although lncRNAs hold significant promise for revolutionizing cancer diagnosis and therapy, realizing this potential involves overcoming numerous challenges. By focusing on these strategic areas, we aim to inspire ongoing research and innovation within the field, advancing toward effective clinical applications of lncRNAs.
Data availability
Not applicable.
References
Swift H. The constancy of desoxyribose nucleic acid in plant nuclei. Proc Natl Acad Sci U S A. 1950;36(11):643–54.
Thomas CA Jr. The genetic organization of chromosomes. Annu Rev Genet. 1971;5:237–56.
Nowoshilow S, et al. The axolotl genome and the evolution of key tissue formation regulators. Nature. 2018;554(7690):50–5.
Shao C, et al. The enormous repetitive Antarctic krill genome reveals environmental adaptations and population insights. Cell. 2023;186(6):1279-1294 e19.
Ohno S. So much “junk” DNA in our genome. Brookhaven Symp Biol. 1972;23:366–70.
Lewin B. Gene expression 2: eucaryotic chromosomes. New York: John Wiley & Sons; 1980.
Britten RJ, Davidson EH. Repetitive and non-repetitive DNA sequences and a speculation on the origins of evolutionary novelty. Q Rev Biol. 1971;46(2):111–38.
Yunis JJ, Yasmineh WG. Heterochromatin, satellite DNA, and cell function. Structural DNA of eucaryotes may support and protect genes and aid in speciation. Science. 1971;174(4015):1200–9.
Comings DE. The structure and function of chromatin. Adv Hum Genet. 1972;3:237–431.
John B, Miklos GL. Functional aspects of satellite DNA and heterochromatin. Int Rev Cytol. 1979;58:1–114.
Orgel LE, Crick FH. Selfish DNA: the ultimate parasite. Nature. 1980;284(5757):604–7.
Holmes DS, et al. Chromosomal RNA: its properties. Science. 1972;177(4043):72–4.
Pierpont ME, Yunis JJ. Localization of chromosomal RNA in human G-banded metaphase chromosomes. Exp Cell Res. 1977;106(2):303–8.
Okazaki Y, et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs: the FANTOM Consortium and the RIKEN Genome Exploration Research Group Phase I & II team. Nature. 2002;420:563–73.
Rinn JL, et al. The transcriptional activity of human Chromosome 22. Genes Dev. 2003;17(4):529–40.
Bertone P, et al. Global identification of human transcribed sequences with genome tiling arrays. Science. 2004;306(5705):2242–6.
Ota T, et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat Genet. 2004;36(1):40–5.
Carninci P, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309(5740):1559–63.
Jacob F, Monod J. Genetic regulatory mechanisms in the synthesis of proteins. J Mol Biol. 1961;3:318–56.
Britten RJ, Davidson EH. Gene regulation for higher cells: a theory. Science. 1969;165(3891):349–57.
Brannan CI, et al. The product of the H19 gene may function as an RNA. Mol Cell Biol. 1990;10(1):28–36.
Bartolomei MS, Zemel S, Tilghman SM. Parental imprinting of the mouse H19 gene. Nature. 1991;351(6322):153–5.
Brockdorff N, et al. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell. 1992;71(3):515–26.
Brown CJ, et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell. 1992;71(3):527–42.
Huttenhofer A, Schattner P, Polacek N. Non-coding RNAs: hope or hype? Trends Genet. 2005;21(5):289–97.
Struhl K. Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat Struct Mol Biol. 2007;14(2):103–5.
Ebisuya M, et al. Ripples from neighbouring transcription. Nat Cell Biol. 2008;10(9):1106–13.
Wunderlich Z, Mirny LA. Different gene regulation strategies revealed by analysis of binding motifs. Trends Genet. 2009;25(10):434–40.
Mattick JS, et al. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat Rev Mol Cell Biol. 2023;24(6):430–47.
Graf J, Kretz M. From structure to function: Route to understanding lncRNA mechanism. BioEssays. 2020;42(12): e2000027.
Ferre F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Brief Bioinform. 2016;17(1):106–16.
Marchese FP, Raimondi I, Huarte M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 2017;18(1):206.
Statello L, et al. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22(2):96–118.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9.
Moran VA, Perera RJ, Khalil AM. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res. 2012;40(14):6391–400.
Li X, et al. lncRNAs: insights into their function and mechanics in underlying disorders. Mutat Res Rev Mutat Res. 2014;762:1–21.
Fernandes JCR, et al. Long non-coding RNAs in the regulation of gene expression: physiology and disease. Noncoding RNA. 2019;5(1):17.
Bhan A, Mandal SS. LncRNA HOTAIR: a master regulator of chromatin dynamics and cancer. Biochim Biophys Acta. 2015;1856(1):151–64.
Brockdorff N. Localized accumulation of Xist RNA in X chromosome inactivation. Open Biol. 2019;9(12):190213.
Lecerf C, Le Bourhis X, Adriaenssens E. The long non-coding RNA H19: an active player with multiple facets to sustain the hallmarks of cancer. Cell Mol Life Sci. 2019;76(23):4673–87.
Arun G, Aggarwal D, Spector DL. MALAT1 long non-coding RNA: functional implications. Noncoding RNA. 2020;6(2):22.
Kanduri C, Thakur N, Pandey RR. The length of the transcript encoded from the Kcnq1ot1 antisense promoter determines the degree of silencing. EMBO J. 2006;25(10):2096–106.
Faghihi MA, Wahlestedt C. Regulatory roles of natural antisense transcripts. Nat Rev Mol Cell Biol. 2009;10(9):637–43.
Hoki Y, et al. A proximal conserved repeat in the Xist gene is essential as a genomic element for X-inactivation in mouse. Development. 2009;136(1):139–46.
Sun Y, et al. Strategies to identify natural antisense transcripts. Biochimie. 2017;132:131–51.
Santos F, et al. Non-coding antisense transcripts: fine regulation of gene expression in cancer. Comput Struct Biotechnol J. 2022;20:5652–60.
Zhang Z, et al. Millions of years of evolution preserved: a comprehensive catalog of the processed pseudogenes in the human genome. Genome Res. 2003;13(12):2541–58.
Hu X, Yang L, Mo YY. Role of pseudogenes in tumorigenesis. Cancers. 2018;10(8):256.
Salmena L. Pseudogenes: four decades of discovery. Methods Mol Biol. 2021;2324:3–18.
Duret L, et al. The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science. 2006;312(5780):1653–5.
Elisaphenko EA, et al. A dual origin of the Xist gene from a protein-coding gene and a set of transposable elements. PLoS ONE. 2008;3(6): e2521.
Louro R, Smirnova AS, Verjovski-Almeida S. Long intronic noncoding RNA transcription: expression noise or expression choice? Genomics. 2009;93(4):291–8.
Heo JB, Sung S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science. 2011;331(6013):76–9.
Guil S, et al. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat Struct Mol Biol. 2012;19(7):664–70.
Seila AC, et al. Divergent transcription from active promoters. Science. 2008;322(5909):1849–51.
Yu D, et al. Classification of transcription boundary-associated RNAs (TBARs) in animals and plants. Front Genet. 2018;9:168.
Ye R, Cao C, Xue Y. Enhancer RNA: biogenesis, function, and regulation. Essays Biochem. 2020;64(6):883–94.
Wen X, Zhong S. Alu transposable elements rewire enhancer-promoter network through RNA pairing. Mol Cell. 2023;83(18):3234–5.
Chan JJ, Tay Y. Noncoding RNA:RNA regulatory networks in cancer. Int J Mol Sci. 2018;19(5):1310.
Panni S, et al. Non-coding RNA regulatory networks. Biochim Biophys Acta Gene Regul Mech. 2020;1863(6):194417.
Su K, et al. The role of a ceRNA regulatory network based on lncRNA MALAT1 site in cancer progression. Biomed Pharmacother. 2021;137:111389.
Moldogazieva NT, et al. Differentially expressed non-coding RNAs and their regulatory networks in liver cancer. Heliyon. 2023;9(9): e19223.
Wang W, et al. Biological function of long non-coding RNA (LncRNA) xist. Front Cell Dev Biol. 2021;9:645647.
Brockdorff N, Bowness JS, Wei G. Progress toward understanding chromosome silencing by Xist RNA. Genes Dev. 2020;34(11–12):733–44.
Tsai MC, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329(5992):689–93.
Hung T, et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet. 2011;43(7):621–9.
Xue ST, et al. Long non-coding RNA LINC00680 functions as a ceRNA to promote esophageal squamous cell carcinoma progression through the miR-423-5p/PAK6 axis. Mol Cancer. 2022;21(1):69.
Yuan C, Ning Y, Pan Y. Emerging roles of HOTAIR in human cancer. J Cell Biochem. 2020;121(5–6):3235–47.
Li J, et al. Regulation of lncRNA and its role in cancer metastasis. Oncol Res. 2016;23(5):205–17.
Ghafouri-Fard S, Esmaeili M, Taheri M. H19 lncRNA: roles in tumorigenesis. Biomed Pharmacother. 2020;123:109774.
Liu SJ, et al. Long noncoding RNAs in cancer metastasis. Nat Rev Cancer. 2021;21(7):446–60.
Hashemi M, et al. Long non-coding RNA (lncRNA) H19 in human cancer: from proliferation and metastasis to therapy. Pharmacol Res. 2022;184:106418.
Zhou KI, et al. N(6)-methyladenosine modification in a long noncoding RNA hairpin predisposes its conformation to protein binding. J Mol Biol. 2016;428(5 Pt A):822–33.
Lence T, et al. Mechanistic insights into m(6)A RNA enzymes. Biochim Biophys Acta Gene Regul Mech. 2019;1862(3):222–9.
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–41.
Yang C, Chen K. Long non-coding RNA in esophageal cancer: a review of research progress. Pathol Oncol Res. 2022;28:1610140.
Yang Z, et al. Insights into the role of long non-coding RNAs in DNA methylation mediated transcriptional regulation. Front Mol Biosci. 2022;9:1067406.
Chedin F. Nascent connections: R-loops and chromatin patterning. Trends Genet. 2016;32(12):828–38.
Shi M, et al. Therapeutic potential of POU3F3, a novel long non-coding RNA, alleviates the pathogenesis of osteoarthritis by regulating the miR-29a- 3p/FOXO3 axis. Curr Gene Ther. 2022;22(5):427–38.
Chen LL. Linking long noncoding RNA localization and function. Trends Biochem Sci. 2016;41(9):761–72.
Chang CY, et al. Genetic variations in UCA1, a lncRNA functioning as a miRNA sponge, determine endometriosis development and the potential associated infertility via regulating lipogenesis. PLoS ONE. 2022;17(7): e0271616.
Li N, et al. Long noncoding RNA LINC00662 functions as miRNA sponge to promote the prostate cancer tumorigenesis through targeting miR-34a. Eur Rev Med Pharmacol Sci. 2019;23(9):3688–98.
Dang SC, et al. MicroRNA-505 suppresses gastric cancer cell proliferation and invasion by directly targeting Polo-like kinase-1. Onco Targets Ther. 2019;12:795–803.
He H, et al. Long noncoding RNA ZFPM2-AS1 acts as a miRNA sponge and promotes cell invasion through regulation of miR-139/GDF10 in hepatocellular carcinoma. J Exp Clin Cancer Res. 2020;39(1):159.
Zhang J, et al. Inferring miRNA sponge co-regulation of protein-protein interactions in human breast cancer. BMC Bioinform. 2017;18(1):243.
Sebastian-delaCruz M, et al. The role of lncRNAs in gene expression regulation through mRNA stabilization. Noncoding RNA. 2021;7(1):3.
Rahmati Y, et al. CiRS-7/CDR1as; An oncogenic circular RNA as a potential cancer biomarker. Pathol Res Pract. 2021;227:153639.
Bian Y, et al. m(6)A modification of long non-coding RNA HNF1A-AS1 facilitates cell cycle progression in colorectal cancer via IGF2BP2-mediated CCND1 mRNA stabilization. Cells. 2022;11(19):3008.
Lang C, et al. m(6) A modification of lncRNA PCAT6 promotes bone metastasis in prostate cancer through IGF2BP2-mediated IGF1R mRNA stabilization. Clin Transl Med. 2021;11(6): e426.
Reis EM, Verjovski-Almeida S. Perspectives of long non-coding RNAs in cancer diagnostics. Front Genet. 2012;3:32.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43(6):904–14.
Fang Y, Fullwood MJ. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genom Proteom Bioinform. 2016;14(1):42–54.
Schultz CW, et al. Understanding and targeting the disease-related RNA binding protein human antigen R (HuR). Wiley Interdiscip Rev RNA. 2020;11(3): e1581.
Wang J, et al. Multiple functions of the RNA-binding protein HuR in cancer progression, treatment responses and prognosis. Int J Mol Sci. 2013;14(5):10015–41.
Chai Y, et al. HuR-regulated lnc RNA NEAT 1 stability in tumorigenesis and progression of ovarian cancer. Cancer Med. 2016;5(7):1588–98.
Hu Y-P, et al. LncRNA-HGBC stabilized by HuR promotes gallbladder cancer progression by regulating miR-502-3p/SET/AKT axis. Mol Cancer. 2019;18:1–18.
Yoon J-H, et al. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012;47(4):648–55.
Zhou X, et al. The RNA-binding protein SRSF1 is a key cell cycle regulator via stabilizing NEAT1 in glioma. Int J Biochem Cell Biol. 2019;113:75–86.
White EJ, Matsangos AE, Wilson GM. AUF1 regulation of coding and noncoding RNA. Wiley Interdiscip Rev RNA. 2017;8(2): e1393.
Yoon J-H, et al. PAR-CLIP analysis uncovers AUF1 impact on target RNA fate and genome integrity. Nat Commun. 2014;5(1):5248.
Beaulieu YB, et al. Polyadenylation-dependent control of long noncoding RNA expression by the poly (A)-binding protein nuclear 1. PLoS Genet. 2012;8(11): e1003078.
Meola N, et al. Identification of a nuclear exosome decay pathway for processed transcripts. Mol Cell. 2016;64(3):520–33.
Hämmerle M, et al. Posttranscriptional destabilization of the liver-specific long noncoding RNA HULC by the IGF2 mRNA-binding protein 1 (IGF2BP1). Hepatology. 2013;58(5):1703–12.
Park J-M, Lee T-H, Kang T-H. Roles of tristetraprolin in tumorigenesis. Int J Mol Sci. 2018;19(11):3384.
Tian F-J, et al. RETRACTED: elevated tristetraprolin impairs trophoblast invasion in women with recurrent miscarriage by destabilization of HOTAIR. Mol Ther-Nucl Acids. 2018;12:600–9.
Xing J, et al. LncRNA-encoded peptide: functions and predicting methods. Front Oncol. 2020;10:622294.
Choi SW, Kim HW, Nam JW. The small peptide world in long noncoding RNAs. Brief Bioinform. 2019;20(5):1853–64.
Minati L, et al. One-shot analysis of translated mammalian lncRNAs with AHARIBO. Elife. 2021. https://doi.org/10.7554/eLife.59303.
Bonilauri B, Holetz FB, Dallagiovanna B. Long non-coding RNAs associated with ribosomes in human adipose-derived stem cells: from RNAs to microproteins. Biomolecules. 2021;11(11):1673.
Lu X, et al. Long non-coding RNA EVADR induced by Fusobacterium nucleatum infection promotes colorectal cancer metastasis. Cell Rep. 2022;40(3):111127.
Li M, Liang C. LncDC: a machine learning-based tool for long non-coding RNA detection from RNA-Seq data. Sci Rep. 2022;12(1):19083.
Wu P, et al. Emerging role of tumor-related functional peptides encoded by lncRNA and circRNA. Mol Cancer. 2020;19(1):22.
Budak H, Kaya SB, Cagirici HB. Long non-coding RNA in plants in the era of reference sequences. Front Plant Sci. 2020;11:276.
Ivkovic TC, et al. Abstract 2802: Molecular and functional characterization of colorectal cancer derived-exosomes and exosomal coding and long non-coding RNA. Cancer Res. 2022;82(12_Supplement):2802–2802.
D’Agostino N, Li W, Wang D. High-throughput transcriptomics. Sci Rep. 2022;12(1):20313.
Jathar S, et al. Technological developments in lncRNA biology. Adv Exp Med Biol. 2017;1008:283–323.
O’Neil D, Glowatz H, Schlumpberger M. Ribosomal RNA depletion for efficient use of RNA-seq capacity. Curr Protoc Mol Biol. 2013. https://doi.org/10.1002/0471142727.mb0419s103.
Tellez-Gabriel M, Heymann D. Exosomal lncRNAs: the newest promising liquid biopsy. Cancer Drug Resist. 2019;2(4):1002–17.
Bostick M, et al. Strand-specific transcriptome sequencing using SMART technology. Curr Protoc Mol Biol. 2016;116:4 27 1-4 27 18.
Chowdhary A, Satagopam V, Schneider R. Long non-coding RNAs: mechanisms, experimental, and computational approaches in identification, characterization, and their biomarker potential in cancer. Front Genet. 2021;12:649619.
Dunagin M, et al. Visualization of lncRNA by single-molecule fluorescence in situ hybridization. Methods Mol Biol. 2015;1262:3–19.
Zhu J, et al. Function of lncRNAs and approaches to lncRNA-protein interactions. Sci China Life Sci. 2013;56(10):876–85.
Yao Y, et al. Long non-coding RNA detection based on multi-probe-induced rolling circle amplification for hepatocellular carcinoma early diagnosis. Anal Chem. 2023;95(2):1549–55.
Zhang X, et al. Mechanisms and functions of long non-coding RNAs at multiple regulatory levels. Int J Mol Sci. 2019;20(22):5573.
Xu S, et al. LncRNA GAS5 knockdown mitigates hepatic lipid accumulation via regulating MiR-26a-5p/PDE4B to activate cAMP/CREB pathway. Front Endocrinol. 2022;13:889858.
Yoshizawa H, et al. Characterization of the MG828507 lncRNA located upstream of the FLT1 gene as an Etiology for pre-eclampsia. J Clin Med. 2022;11(15):4603.
Jha P, et al. Computational methods for functional characterization of lncRNAs in human diseases: a focus on co-expression networks. Curr Bioinform. 2023;19:21–38.
Krusnauskas R, et al. Identification and comparison of m6A modifications in glioblastoma non-coding RNAs with MeRIP-seq and Nanopore dRNA-seq. Epigenetics. 2023;18(1):2163365.
Pinskaya M, et al. Reference-free transcriptome exploration reveals novel RNAs for prostate cancer diagnosis. Life Sci Alliance. 2019;2(6): e201900449.
Kabzinski J, Kucharska-Lusina A, Majsterek I. RNA-based liquid biopsy in head and neck cancer. Cells. 2023;12(14):1916.
Toden S, Goel A. Non-coding RNAs as liquid biopsy biomarkers in cancer. Br J Cancer. 2022;126(3):351–60.
Barth DA, et al. Circulating non-coding RNAs in Renal cell carcinoma-pathogenesis and potential implications as clinical biomarkers. Front Cell Dev Biol. 2020;8:828.
Kamel LM, et al. Circulating long non-coding RNA GAS5 and SOX2OT as potential biomarkers for diagnosis and prognosis of non-small cell lung cancer. Biotechnol Appl Biochem. 2019;66(4):634–42.
Fattahi S, et al. LncRNAs as potential diagnostic and prognostic biomarkers in gastric cancer: a novel approach to personalized medicine. J Cell Physiol. 2020;235(4):3189–206.
Liu S, et al. Identification of the 3-lncRNA signature as a prognostic biomarker for colorectal cancer. Int J Mol Sci. 2020;21(24):9359.
Fernandes M, et al. ceRNA network of lncRNA/miRNA as circulating prognostic biomarkers in non-hodgkin lymphomas: bioinformatic analysis and assessment of their prognostic value in an NHL cohort. Int J Mol Sci. 2021;23(1):201.
Isaev K, et al. Pan-cancer analysis of non-coding transcripts reveals the prognostic onco-lncRNA HOXA10-AS in gliomas. Cell Rep. 2021;37(3):109873.
Li X, Zhu J, Qiu J. Identification of potential prognostic biomarkers for breast cancer based on lncRNA-TF-associated ceRNA network and functional module. Biomed Res Int. 2020;2020:5257896.
Di Meo A, et al. Liquid biopsy: a step forward towards precision medicine in urologic malignancies. Mol Cancer. 2017;16(1):80.
Necula L, et al. Recent advances in gastric cancer early diagnosis. World J Gastroenterol. 2019;25(17):2029–44.
Yuan S, et al. Circulating long noncoding RNAs act as diagnostic biomarkers in non-small cell lung cancer. Front Oncol. 2020;10:537120.
Sukowati CHC, et al. Circulating long and circular noncoding RNA as non-invasive diagnostic tools of hepatocellular carcinoma. Biomedicines. 2021;9(1):90.
Roobol MJ, et al. Performance of the prostate cancer antigen 3 (PCA3) gene and prostate-specific antigen in prescreened men: exploring the value of PCA3 for a first-line diagnostic test. Eur Urol. 2010;58(4):475–81.
Groskopf J, et al. APTIMA PCA3 molecular urine test: development of a method to aid in the diagnosis of prostate cancer. Clin Chem. 2006;52(6):1089–95.
Marks LS, et al. PCA3 molecular urine assay for prostate cancer in men undergoing repeat biopsy. Urology. 2007;69(3):532–5.
Rönnau C, et al. Noncoding RNAs as novel biomarkers in prostate cancer. BioMed Res Int. 2014;2014:1.
Xavier-Magalhaes A, et al. The long non-coding RNA HOTAIR is transcriptionally activated by HOXA9 and is an independent prognostic marker in patients with malignant glioma. Oncotarget. 2018;9(21):15740–56.
Lin X, et al. LncRNADisease v3.0: an updated database of long non-coding RNA-associated diseases. Nucleic Acids Res. 2023. https://doi.org/10.1093/nar/gkad828.
Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015;12(1):1–9.
Xu C, et al. MALAT-1: a long non-coding RNA and its important 3’ end functional motif in colorectal cancer metastasis. Int J Oncol. 2011;39(1):169–75.
Hu G, Lou Z, Gupta M. The long non-coding RNA GAS5 cooperates with the eukaryotic translation initiation factor 4E to regulate c-Myc translation. PLoS ONE. 2014;9(9): e107016.
Escors D, Breckpot K. Lentiviral vectors in gene therapy: their current status and future potential. Arch Immunol Ther Exp. 2010;58(2):107–19.
Li JL, et al. Long noncoding RNA UCA1 regulates proliferation and apoptosis in multiple myeloma by targeting miR-331-3p/IL6R axis for the activation of JAK2/STAT3 pathway. Eur Rev Med Pharmacol Sci. 2019;23(21):9238–50.
Jiang L, et al. NEAT1 scaffolds RNA-binding proteins and the Microprocessor to globally enhance pri-miRNA processing. Nat Struct Mol Biol. 2017;24(10):816–24.
Hall LL, Lawrence JB. XIST RNA and architecture of the inactive X chromosome: implications for the repeat genome. Cold Spring Harb Symp Quant Biol. 2010;75:345–56.
Sanchez A, et al. The long non-coding RNA ANRIL in cancers. Cancers. 2023;15(16):4160.
Zhang F, et al. The lncRNA CRNDE is regulated by E2F6 and sensitizes gastric cancer cells to chemotherapy by inhibiting autophagy. J Cancer. 2022;13(10):3061–72.
Tian X, et al. LncRNA-H19 regulates chemoresistance to carboplatin in epithelial ovarian cancer through microRNA-29b-3p and STAT3. J Cancer. 2021;12(19):5712–22.
Cheng H, et al. LncRNA UCA1 enhances cisplatin resistance by regulating CYP1B1-mediated apoptosis via miR-513a-3p in human gastric cancer. Cancer Manag Res. 2021;13:367–77.
Wang J, et al. The long noncoding RNA H19 promotes tamoxifen resistance in breast cancer via autophagy. J Hematol Oncol. 2019;12(1):81.
Mazor G, et al. The lncRNA TP73-AS1 is linked to aggressiveness in glioblastoma and promotes temozolomide resistance in glioblastoma cancer stem cells. Cell Death Dis. 2019;10(3):246.
Shao Z, et al. Non-coding RNAs that regulate the Wnt/beta-catenin signaling pathway in gastric cancer: Good cop, bad cop? (Review). Oncol Rep. 2020;44(4):1314–21.
Chen B, et al. Targeting non-coding RNAs to overcome cancer therapy resistance. Signal Transduct Target Ther. 2022;7(1):121.
Khan MI, Ahmad A. LncRNA SNHG6 sponges miR-101 and induces tamoxifen resistance in breast cancer cells through induction of EMT. Front Oncol. 2022;12:1015428.
Yuan HH, et al. LncRNA UCA1 mediates cetuximab resistance in colorectal cancer via the MiR-495 and HGF/c-MET pathways. J Cancer. 2022;13(1):253–67.
Zhang B, et al. LncRNA PCBP1-AS1-mediated AR/AR-V7 deubiquitination enhances prostate cancer enzalutamide resistance. Cell Death Dis. 2021;12(10):856.
Xia H, et al. LncRNA MEG3 promotes the sensitivity of vincristine by inhibiting autophagy in lung cancer chemotherapy. Eur Rev Med Pharmacol Sci. 2018;22(4):4185.
Zhou H, et al. Knockdown of lncRNA HOXD-AS1 suppresses proliferation, migration and invasion and enhances cisplatin sensitivity of glioma cells by sponging miR-204. Biomed Pharmacother. 2019;112:108633.
Tan J, et al. Double-negative feedback loop between long non-coding RNA TUG1 and miR-145 promotes epithelial to mesenchymal transition and radioresistance in human bladder cancer cells. FEBS Lett. 2015;589(20 Pt B):3175–81.
Jiang H, et al. Down-regulation of LncRNA TUG1 enhances radiosensitivity in bladder cancer via suppressing HMGB1 expression. Radiat Oncol. 2017;12(1):65.
Zhang S, et al. LncRNA HOTAIR enhances breast cancer radioresistance through facilitating HSPA1A expression via sequestering miR-449b-5p. Thorac Cancer. 2020;11(7):1801–16.
Liu L, et al. Long noncoding RNA LINC00511 involves in breast cancer recurrence and radioresistance by regulating STXBP4 expression via miR-185. Eur Rev Med Pharmacol Sci. 2019;23(17):7457–68.
Wang B, et al. Long noncoding RNA LINC02582 acts downstream of miR-200c to promote radioresistance through CHK1 in breast cancer cells. Cell Death Dis. 2019;10(10):764.
Zhang N, et al. LncRNA LINC00963 promotes tumorigenesis and radioresistance in breast cancer by sponging miR-324-3p and Inducing ACK1 expression. Mol Ther Nucleic Acids. 2019;18:871–81.
Qian L, et al. lncRNA HOTAIR promotes DNA repair and radioresistance of breast cancer via EZH2. DNA Cell Biol. 2020. https://doi.org/10.1089/dna.2020.5771.
Zhong G, et al. H19 Knockdown suppresses proliferation and induces apoptosis by regulating miR-130a-3p/SATB1 in breast cancer cells. Onco Targets Ther. 2020;13:12501–13.
Xiao J, et al. Long noncoding RNA TRPM2-AS acts as a microRNA sponge of miR-612 to promote gastric cancer progression and radioresistance. Oncogenesis. 2020;9(3):29.
Jiang Y, et al. Long noncoding RNA NEAT1 regulates radio-sensitivity via microRNA-27b-3p in gastric cancer. Cancer Cell Int. 2020;20(1):581.
Ge X, et al. Knockdown of lncRNA PCAT1 enhances radiosensitivity of cervical cancer by regulating miR-128/GOLM1 axis. Onco Targets Ther. 2020;13:10373–85.
Zhao H, et al. Long noncoding RNA LINC00958 regulates cell sensitivity to radiotherapy through RRM2 by binding to microRNA-5095 in cervical cancer. J Cell Physiol. 2019;234(12):23349–59.
Gao J, et al. LncRNA GAS5 confers the radio sensitivity of cervical cancer cells via regulating miR-106b/IER3 axis. Int J Biol Macromol. 2019;126:994–1001.
Li N, et al. Overexpression of HOTAIR leads to radioresistance of human cervical cancer via promoting HIF-1α expression. Radiat Oncol. 2018;13(1):210.
Jing L, et al. HOTAIR enhanced aggressive biological behaviors and induced radio-resistance via inhibiting p21 in cervical cancer. Tumour Biol. 2015;36(5):3611–9.
Liu Y, et al. Long non-coding RNA HOTAIR knockdown enhances radiosensitivity through regulating microRNA-93/ATG12 axis in colorectal cancer. Cell Death Dis. 2020;11(3):175.
Liu R, et al. Long noncoding RNA lnc-RI regulates DNA damage repair and radiation sensitivity of CRC cells through NHEJ pathway. Cell Biol Toxicol. 2020;36(5):493–507.
Liang H, et al. Long noncoding RNA LINC00958 suppresses apoptosis and radiosensitivity of colorectal cancer through targeting miR-422a. Cancer Cell Int. 2021;21(1):477.
Li C, et al. LncRNA EGOT/miR-211-5p affected radiosensitivity of rectal cancer by competitively regulating ErbB4. Onco Targets Ther. 2021;14:2867–78.
Cheng W, et al. LncRNA MAGI2-AS3 overexpression sensitizes esophageal cancer cells to irradiation through down-regulation of HOXB7 via EZH2. Front Cell Dev Biol. 2020;8:552822.
Zhang H, et al. Cancer-associated fibroblast-promoted LncRNA DNM3OS confers radioresistance by regulating DNA damage response in esophageal squamous cell carcinoma. Clin Cancer Res. 2019;25(6):1989–2000.
Chen M, et al. Long noncoding RNA FAM201A mediates the radiosensitivity of esophageal squamous cell cancer by regulating ATM and mTOR expression via miR-101. Front Genet. 2018;9:611.
Sun Y, et al. Radiation induces NORAD expression to promote ESCC radiotherapy resistance via EEPD1/ATR/Chk1 signalling and by inhibiting pri-miR-199a1 processing and the exosomal transfer of miR-199a-5p. J Exp Clin Cancer Res. 2021;40(1):306.
Wang P, et al. lncTUG1/miR-144-3p affect the radiosensitivity of esophageal squamous cell carcinoma by competitively regulating c-MET. J Exp Clin Cancer Res. 2020;39(1):7.
Li Z, et al. Long noncoding RNA MALAT1 affects the efficacy of radiotherapy for esophageal squamous cell carcinoma by regulating Cks1 expression. J Oral Pathol Med. 2017;46(8):583–90.
Ahmadov U, et al. The long non-coding RNA HOTAIRM1 promotes tumor aggressiveness and radiotherapy resistance in glioblastoma. Cell Death Dis. 2021;12(10):885.
Li J, Ji X, Wang H. Targeting long noncoding RNA HMMR-AS1 suppresses and radiosensitizes glioblastoma. Neoplasia. 2018;20(5):456–66.
Li W, et al. LncRNA RBPMS-AS1 promotes NRGN transcription to enhance the radiosensitivity of glioblastoma through the microRNA-301a-3p/CAMTA1 axis. Transl Oncol. 2022;15(1):101282.
Tang T, et al. lncRNA TPTEP1 inhibits stemness and radioresistance of glioma through miR-106a-5p-mediated P38 MAPK signaling. Mol Med Rep. 2020;22(6):4857–67.
Zheng J, et al. Linc-RA1 inhibits autophagy and promotes radioresistance by preventing H2Bub1/USP44 combination in glioma cells. Cell Death Dis. 2020;11(9):758.
Gao W, Qiao M, Luo K. Long noncoding RNA TP53TG1 contributes to radioresistance of glioma cells via miR-524-5p/RAB5A axis. Cancer Biother Radiopharm. 2021;36(7):600–12.
Zheng R, et al. Upregulation of long noncoding RNA small Nucleolar RNA host gene 18 promotes radioresistance of glioma by repressing semaphorin 5A. Int J Radiat Oncol Biol Phys. 2016;96(4):877–87.
Wang B, et al. NCK1-AS1 enhances glioma cell proliferation, radioresistance and chemoresistance via miR-22-3p/IGF1R ceRNA pathway. Biomed Pharmacother. 2020;129:110395.
Yang W, et al. MiR-146b-5p overexpression attenuates stemness and radioresistance of glioma stem cells by targeting HuR/lincRNA-p21/β-catenin pathway. Oncotarget. 2016;7(27):41505–26.
Tian W, et al. LINC01123 potentially correlates with radioresistance in glioma through the miR-151a/CENPB axis. Neuropathology. 2022;42(1):3–15.
Wang YP, et al. Overexpression of XIST facilitates cell proliferation, invasion and suppresses cell apoptosis by reducing radio-sensitivity of glioma cells via miR-329-3p/CREB1 axis. Eur Rev Med Pharmacol Sci. 2020;24(6):3190–203.
Li J, et al. Silencing of long non-coding RNA LINC00520 promotes radiosensitivity of head and neck squamous cell carcinoma cells. Free Radic Res. 2020;54(4):254–70.
Cui X, et al. Exosomes-derived long non-coding RNA HOTAIR reduces laryngeal cancer radiosensitivity by regulating microRNA-454-3p/E2F2 axis. Onco Targets Ther. 2019;12:10827–39.
Tang T, Shan G. DGCR5 promotes cancer stem cell-like properties of radioresistant laryngeal carcinoma cells by sponging miR-506 via Wnt pathway. J Cell Physiol. 2019;234(10):18423–31.
Tang T, Shan G, Zeng F. Knockdown of DGCR5 enhances the radiosensitivity of human laryngeal carcinoma cells via inducing miR-195. J Cell Physiol. 2019;234(8):12918–25.
Wang Y, et al. Long non-coding RNA NEAT1 regulates epithelial membrane protein 2 expression to repress nasopharyngeal carcinoma migration and irradiation-resistance through miR-101-3p as a competing endogenous RNA mechanism. Oncotarget. 2017;8(41):70156–71.
Wang Y, et al. The lncRNA PVT1 regulates nasopharyngeal carcinoma cell proliferation via activating the KAT2A acetyltransferase and stabilizing HIF-1α. Cell Death Differ. 2020;27(2):695–710.
Guo Z, et al. LncRNA linc00312 suppresses radiotherapy resistance by targeting DNA-PKcs and impairing DNA damage repair in nasopharyngeal carcinoma. Cell Death Dis. 2021;12(1):69.
Han YY, et al. LINC00114 promoted nasopharyngeal carcinoma progression and radioresistance in vitro and in vivo through regulating ERK/JNK signaling pathway via targeting miR-203. Eur Rev Med Pharmacol Sci. 2020;24(5):2491–504.
Han Y, et al. PVT1 mediates cell proliferation, apoptosis and radioresistance in nasopharyngeal carcinoma through regulating miR-515-5p/PIK3CA Axis. Cancer Manag Res. 2020;12:10077–90.
Liu H, et al. lncRNA CASC19 contributes to radioresistance of nasopharyngeal carcinoma by promoting autophagy via AMPK-mTOR pathway. Int J Mol Sci. 2021;22(3):1407.
Ma X, et al. LncRNA ANCR promotes proliferation and radiation resistance of nasopharyngeal carcinoma by inhibiting PTEN expression. Onco Targets Ther. 2018;11:8399–408.
Wang YH, et al. LINC-PINT impedes DNA repair and enhances radiotherapeutic response by targeting DNA-PKcs in nasopharyngeal cancer. Cell Death Dis. 2021;12(5):454.
Yi L, et al. Long noncoding RNA PTPRG-AS1 acts as a microRNA-194-3p sponge to regulate radiosensitivity and metastasis of nasopharyngeal carcinoma cells via PRC1. J Cell Physiol. 2019;234(10):19088–102.
Jin C, et al. The role of MALAT1/miR-1/slug axis on radioresistance in nasopharyngeal carcinoma. Tumour Biol. 2016;37(3):4025–33.
Chen Y, et al. Long non-coding RNA ROR promotes radioresistance in hepatocelluar carcinoma cells by acting as a ceRNA for microRNA-145 to regulate RAD18 expression. Arch Biochem Biophys. 2018;645:117–25.
Yu C, et al. LncRNA GAS5 enhances radiosensitivity of hepatocellular carcinoma and restricts tumor growth and metastasis by miR-144-5p/ATF2. Am J Transl Res. 2021;13(9):10896–907.
Jin Q, et al. lncRNA MIR22HG-derived miR-22-5p enhances the radiosensitivity of hepatocellular carcinoma by increasing histone acetylation through the inhibition of HDAC2 activity. Front Oncol. 2021;11:572585.
Yang QS, et al. Long noncoding RNA LINC00483/microRNA-144 regulates radiosensitivity and epithelial-mesenchymal transition in lung adenocarcinoma by interacting with HOXA10. J Cell Physiol. 2019;234(7):11805–21.
Chen J, et al. Radiotherapy induced Lewis lung cancer cell apoptosis via inactivating β-catenin mediated by upregulated HOTAIR. Int J Clin Exp Pathol. 2015;8(7):7878–86.
Zhang F, et al. M2 macrophage-derived exosomal long non-coding RNA AGAP2-AS1 enhances radiotherapy immunity in lung cancer by reducing microRNA-296 and elevating NOTCH2. Cell Death Dis. 2021;12(5):467.
Hou J, et al. Silencing of LINC00461 enhances radiosensitivity of lung adenocarcinoma cells by down-regulating HOXA10 via microRNA-195. J Cell Mol Med. 2020;24(5):2879–90.
He H, et al. Upregulation of KCNQ1OT1 promotes resistance to stereotactic body radiotherapy in lung adenocarcinoma by inducing ATG5/ATG12-mediated autophagy via miR-372-3p. Cell Death Dis. 2020;11(10):883.
Yu Z, et al. LncRNA SBF2-AS1 affects the radiosensitivity of non-small cell lung cancer via modulating microRNA-302a/MBNL3 axis. Cell Cycle. 2020;19(3):300–16.
Liu AM, et al. Long noncoding RNA FAM201A involves in radioresistance of non-small-cell lung cancer by enhancing EGFR expression via miR-370. Eur Rev Med Pharmacol Sci. 2019;23(13):5802–14.
Wu D, et al. Knockdown of Lncrna PVT1 enhances radiosensitivity in non-small cell lung cancer by sponging Mir-195. Cell Physiol Biochem. 2017;42(6):2453–66.
Jiang G, et al. lncRNA cytoskeleton regulator reduces non-small cell lung cancer radiosensitivity by downregulating miRNA-206 and activating prothymosin α. Int J Oncol. 2021. https://doi.org/10.3892/ijo.2021.5268.
Liu S, et al. Long noncoding RNA CBR3-AS1 mediates tumorigenesis and radiosensitivity of non-small cell lung cancer through redox and DNA repair by CBR3-AS1 /miR-409-3p/SOD1 axis. Cancer Lett. 2022;526:1–11.
Wang Z, et al. The HNF1A-AS1/miR-92a-3p axis affects the radiosensitivity of non-small cell lung cancer by competitively regulating the JNK pathway. Cell Biol Toxicol. 2021;37(5):715–29.
Brownmiller T, et al. Y Chromosome LncRNA are involved in radiation response of male non-small cell lung cancer cells. Cancer Res. 2020;80(19):4046–57.
Xue Y, et al. Long noncoding RNA GAS5 inhibits tumorigenesis and enhances radiosensitivity by suppressing miR-135b expression in non-small cell lung cancer. Oncol Res. 2017;25(8):1305–16.
Zhu C, et al. RBM5-AS1 promotes radioresistance in medulloblastoma through stabilization of SIRT6 protein. Acta Neuropathol Commun. 2021;9(1):123.
Liu Y, et al. Long noncoding RNA LINC00518 induces radioresistance by regulating glycolysis through an miR-33a-3p/HIF-1α negative feedback loop in melanoma. Cell Death Dis. 2021;12(3):245.
Cui Y, et al. LINC01224 facilitates the proliferation and inhibits the radiosensitivity of melanoma cells through the miR-193a-5p/NR1D2 axis. Kaohsiung J Med Sci. 2022;38(3):196–206.
Yang H, et al. Downregulation of lncRNA XIST represses tumor growth and boosts radiosensitivity of neuroblastoma via modulation of the miR-375/L1CAM Axis. Neurochem Res. 2020;45(11):2679–90.
Mou L, et al. Long noncoding RNA LINC01410 suppresses tumorigenesis and enhances radiosensitivity in neuroblastoma cells through regulating miR-545-3p/HK2 Axis. Onco Targets Ther. 2021;14:3225–38.
Chen C, et al. LncRNA HULC mediates radioresistance via autophagy in prostate cancer cells. Braz J Med Biol Res. 2018;51(6): e7080.
Xiu D, et al. Knockdown of lncRNA TUG1 enhances radiosensitivity of prostate cancer via the TUG1/miR-139-5p/SMC1A Axis. Onco Targets Ther. 2020;13:2319–31.
Ma X, et al. Long non-coding RNA GAS5 suppresses tumor progression and enhances the radiosensitivity of prostate cancer through the miR-320a/RAB21 Axis. Cancer Manag Res. 2020;12:8833–45.
Zhou X, et al. LINC02532 contributes to radiosensitivity in clear cell renal cell carcinoma through the miR-654–5p/YY1 Axis. Molecules. 2021;26(22):7040.
Chen W, et al. Roles of the SNHG7/microRNA-9-5p/DPP4 ceRNA network in the growth and (131)I resistance of thyroid carcinoma cells through PI3K/Akt activation. Oncol Rep. 2021. https://doi.org/10.3892/or.2021.7954.
Li L, et al. LncRNA GAS5 sponges miR-362-5p to promote sensitivity of thyroid cancer cells to (131) I by upregulating SMG1. IUBMB Life. 2020;72(11):2420–31.
Funding
This research was funded by the Basic Platform Project of the Ministry of Science and Technology of China (No. TDRC-2019-194-30).
Author information
Authors and Affiliations
Contributions
Qais Ahmad Nasser did the conception and design of the review, the acquisition of data, and the drafting of the manuscript. Fengyuan Zhang and Abdul Malik provided critical insights into the content, focusing on the analysis and interpretation of the data related to lncRNAs in cancer biology. Shengxia Chen*, as the corresponding author, played a pivotal role in revising the manuscript critically for important intellectual content, ensuring the accuracy and integrity of the work. All authors have read and approved the final manuscript for publication.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable. All the data/samples were collected from the articles in online databases and used anonymously.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Naseer, Q.A., Malik, A., Zhang, F. et al. Exploring the enigma: history, present, and future of long non-coding RNAs in cancer. Discov Onc 15, 214 (2024). https://doi.org/10.1007/s12672-024-01077-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01077-y