Abstract
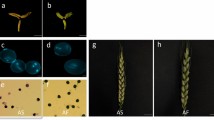
Seed shattering is one of the major domestication traits of crops. In wheat, except for the Q gene whose mutation renders free threshing and changing of rachis fragility, not much is known about the molecular mechanism for this process. We report here the cloning and characterization of TaqSH1, the ortholog of the rice seed shattering gene qSH1. TaqSH1 encodes a BEL1-like protein that is conserved between monocots and eudicots. TaqSH1 was located on the homoeologous group 3, a potential new genetic locus for seed threshability in wheat. Over expression of TaqSH1 in Arabidopsis resulted in dwarfed plants. The inflorescences of transgenic plants were more compact with larger pedicel angles. Scanning Electron Microscopy (SEM) showed that the transgenic siliques had a narrower replum where the dehiscence zone was altered. In addition, abscission of petals was significantly delayed due to delayed abscission zone development. Real-time PCR assays showed that over expression of TaqSH1 down regulated known Arabidopsis abscission related genes IDA, HAESA, KNAT1/6 and SHP1/2 in the transgenic plants. Taken together, our data suggest that TaqSH1 may represent another example of conserved mechanisms across monocots and eudicots for fruit/grain abscission and should have potential application in genetic manipulation of wheat seed shattering.
Similar content being viewed by others
References
Alvarez J, Smyth DR (2002) CRABS CLAW and SPATULA genes regulate growth and pattern formation during gynoecium development in Arabidopsis thaliana. Int J Plant Sci 163:17–41
Arnaud N, Lawrenson T, Østergaard L, Sablowski R (2011) The same regulatory point mutation changed seed-dispersal structures in evolution and domestication. Curr Biol 21:1215–1219
Butenko MA, Patterson SE, Grini PE, Stenvik GE, Amundsen SS, Mandal A, Aalen RB (2003) Inflorescence deficient in abscission controls floral organ abscission in Arabidopsis and identifies a novel family of putative ligands in plants. Plant Cell 15:2296–2307
Byrne ME, Simorowski J, Martienssen RA (2002) ASYMMETRIC LEAVES1 reveals knox gene redundancy in Arabidopsis. Development 129:1957–1965
Chalupska D, Lee H, Faris J, Evrard A, Chalhoub B, Haselkorn R, Gornicki P (2008) Acc homoeoloci and the evolution of wheat genomes. Proc Natl Acad Sci USA 105:9691–9696
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316: 1862–1866
Faris JD, Gill BS (2002) Genomic targeting and high-resolution mapping of the domestication gene Q in wheat. Genome 45:706–718
Faris JD, Zhang Z, Fellers JP, Gill BS (2008) Micro-colinearity between rice, Brachypodium, and Triticum monococcum at the wheat domestication locus Q. Funct Integr Genomics 8:149–164
Ferrándiz C, Pelaz S, Yanofsky MF (1999) Control of carpel and fruit development in Arabidopsis. Annu Rev Biochem 68:321–354
Ferrandiz C, Liljegren SJ, Yanofsky MF (2000) Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis fruit development. Science 289:436–438
Jinn TL, Stone JM, Walker JC (2000) HAESA, an Arabidopsis leucinerich repeat receptor kinase, controls floral organ abscission. Genes Dev 14:108–117
Koncz C, Schell J (1986) The promoter of T L-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gen Genet 204:383–396
Konishi S, Izawa T, Lin SY, Ebana K, Fukuta Y, Sasaki T, Yano M (2006) An SNP caused loss of seed shattering during rice domestication. Science 312:1392–1396
Lewis MW, Leslie ME, Liljegren SJ (2006) Plant separation: 50 ways to leave your mother. Curr Opin Plant Biol 9:59–65
Li C, Zhou A, Sang T (2006) Rice domestication by reducing shattering. Science 311:1936–1939
Li W, Gill BS (2006) Multiple genetic pathways for seed shattering in the grasses. Funct Integr Genomics 6:300–309
Liljegren SJ, Ditta GS, Eshed Y, Savidge B, Bowman JL, Yanofsky MF (2000) SHATTERPROOF MADS-box genes control seed dispersal in Arabidopsis. Nature 404:766–770
Liljegren SJ, Roeder AHK, Kempin SA, Gremski K, Østergaard L, Guimil S, Reyes DK, Yanofsky MF (2004) Control of Fruit Patterning in Arabidopsis by INDEHISCENT. Cell 116:843–853
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2CT method. Methods 25:402–408
Martinez-Trujillo M, Limones-Briones V, Cabrera-Ponce JL, Herrera-Estrella L (2004) Improving transformation efficiency of Arabidopsis thaliana by modifying the floral dip method. Plant Mol Biol Rep 22:63–70
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Nalam VJ, Vales MI, Watson CJW, Johnson EB, Riera-Lizarazu O (2007) Map-based analysis of genetic loci on chromosome 2D that affect glume tenacity and threshability, components of the free-threshing habit in common wheat (Triticum aestivum L.). Theor Appl Genet 116:135–145
Ragni L, Belles-Boix E, Günl M, Pautot V (2008) Interaction of KNAT6 and KNAT2 with BREVIPEDICELLUS and PENNYWISE in Arabidopsis inflorescences. Plant Cell 20:888–900
Rajani S, Sundaresan V (2001) The Arabidopsis myc/bHLH gene ALCATRAZ enables cell separation in fruit dehiscence. Curr Biol 11:1914–1922
Roeder AHK, Ferrándiz C, Yanofsky MF (2003) The Role of the REPLUMLESS Homeodomain Protein in Patterning the Arabidopsis Fruit Curr Biol 13:1630–1635
Rutjens B, Bao D, Eck-Stouten V, Brand M, Smeekens S, Proveniers M (2009) Shoot apical meristem function in Arabidopsis requires the combined activities of three BEL1-like homeodomain proteins. Plant J 58:641–654
Shi CL, Stenvik GE, Vie AK, Bones AM, Pautot V, Proveniers M, Aalen RB, Butenko MA (2011) Arabidopsis class I KNOTTED-like homeobox proteins act downstream in the IDA-HAE/HSL2 floral abscission signaling pathway. Plant Cell 23:2553–2567
Sood S, Kuraparthy V, Bai G, Gill BS (2009) The major threshability genes soft glume (sog) and tenacious glume (Tg), of diploid and polyploid wheat, trace their origin to independent mutations at non-orthologous loci. Theor Appl Genet 119:341–351
Stenvik GE, Butenko MA, Urbanowicz BR, Rose JKC, Aalen RB (2006) Overexpression of INFLORESCENCE DEFICIENT IN ABSCISSION activates cell separation in vestigial abscission zones in Arabidopsis. Plant Cell 18:1467–1476
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Venglat S, Dumonceaux T, Rozwadowski K, Parnell L, Babic V, Keller W, Martienssen R, Selvaraj G, Datla R (2002) The homeobox gene BREVIPEDICELLUS is a key regulator of inflorescence architecture in Arabidopsis. Proc Natl Acad Sci USA 99:4730–4735
Wang XQ, XU WH, Ma LG, Fu ZM, Deng XW, Li JY, Wang YH (2006) Requirement of KNAT1/BP for the development of abscission zones in Arabidopsis thaliana. J Integr Plant Biol 48:15–26
Zhang Z, Belcram H, Gornicki P, Charles M, Just J, Huneau C, Magdelenat G, Couloux A, Samain S, Gill BS (2011) Duplication and partitioning in evolution and function of homoeologous Q loci governing domestication characters in polyploid wheat. Proc Natl Acad Sci USA 108:18737–18742
Zhou Y, Lu D, Li C, Luo J, Zhu BF, Zhu J, Shangguan Y, Wang Z, Sang T, Zhou B, Han B (2012) Genetic control of seed shattering in rice by the APETALA2 transcription factor SHATTERING ABORTION1. Plant Cell 24:1034–1048
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, L., Liu, D., Wang, D. et al. Over expression of the wheat BEL1-like gene TaqSH1 affects floral organ abscission in Arabidopsis thaliana . J. Plant Biol. 56, 98–105 (2013). https://doi.org/10.1007/s12374-012-0438-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-012-0438-7