Abstract
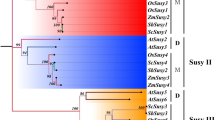
Sugarcane, the world’s largest source of sugar and biofuel, still carries an unassembled ~ 10 GB polyaneuploid genome, which is a major bottleneck in studying this crop at molecular level. Sugar is mainly derived from sugarcane; hence, the sucrose metabolism pathway is one of the most intriguing hotspot for sugar crop researchers. Four main gene families have been reported to be involved in sucrose metabolism including the SuSy (sucrose synthase) gene family. Sucrose synthase in plants degrades sucrose to UDP-glucose and fructose to ensure their availability for growth and developmental purposes. Here, we attempted to get deep insights into sugarcane SuSy gene family and carried out bioinformatics and expression analyses to get a high-resolution holistic snapshot. Multiple stress-responsive cis motifs, abscisic acid-responsive element (ABRE), anaerobic induction-responsive element (ARE), auxin-responsive element (AuxRE), low-temperature-responsive (LTR), wounding response element (WUN), WRKY transcription factors binding site (W-motif), predicted in the regulatory region showed their involvement in the stress signaling pathways, in addition to sucrose metabolism. Similarly, the protein interaction network analysis predicted an array of proteins of diverse range of functions. Moreover, the expression pattern of SuSy gene family in two varieties CPF251 (higher sugar level) and CPF252 (lower sugar level) was compared in leaf and internodes (top and bottom). qRT-PCR indicated the differential expression pattern of the SuSy genes in these two varieties. The expression of SoSuSy2 was high in leaf and top internodes, while low in bottom internodes indicating its activity in this tissue. In CPF251, both SoSuSy2 and SoSuSy4 displayed higher expression level; however, only SuSy2 had higher expression in CPF252. In contrast, SoSuSy6 and SoSuSy7 were the least expressed genes followed by SoSuSy1. This study highlights the candidate genes for gene manipulation and consequent metabolic engineering of sugarcane for enhanced sucrose contents.






Similar content being viewed by others
References
Abdullah, M., Y. Cao, X. Cheng, D. Meng, Y Chen, A. Shakoor, J. Gao, and Y. Cai. 2018. The sucrose synthase gene family in Chinese pear (Pyrus bretschneideri Rehd.): structure, expression, and evolution. Molecules 23: 1144
Ahmed, M., A. Iqbal, A. Latif, S.U. Din, M.B. Sarwar, X. Wang, A.Q. Rao, T. Husnain, and A.A. Shahid. 2020. Overexpression of a sucrose synthase gene indirectly improves cotton fiber quality through sucrose cleavage. Frontiers in Plant Science 11: 1639.
An, X., Z. Chen, J. Wang, M. Ye, L. Ji, J. Wang, W. Liao, and H. Ma. 2014. Identification and characterization of the Populus sucrose synthase gene family. Gene 539: 58–67.
Anur, R.M.N., W.D. Mufithah, H. Sakakibara. Sawitri, and B. Sugiharto. 2020. Overexpression of sucrose phosphate synthase enhanced sucrose content and biomass production in transgenic sugarcane. Plants 9: 200.
Barratt, D.H.P., P. Derbyshire, K. Findlay, M. Pike, N. Wellner, J. Lunn, R. Feil, C. Simpson, A.J. Maule, and A.M. Smith. 2009. Normal growth of Arabidopsis requires cytosolic invertase but not sucrose synthase. Proceedings of the National Academy of Sciences 106: 13124–13129.
Baud, S., M. Vaultier, and C. Rochat. 2004. Structure and expression profile of the sucrose synthase multigene family in Arabidopsis. Journal of Experimental Botany 55: 397–409.
Bieniawska, Z., D.H.P. Barratt, A.P. Garlick, V. Thole, N.J. Kruger, C. Martin, R. Zrenner, and A.M. Smith. 2007. Analysis of the sucrose synthase gene family in Arabidopsis. The Plant Journal: For Cell and Molecular Biology 49: 810–828.
Chen, L.Q., B.H. Hou, S. Lalonde, H. Takanaga, M.L. Hartung, X.Q. Qu, W.J. Guo, J.G. Kim, W. Underwood, and B. Chaudhuri. 2010. Sugar transporters for intercellular exchange and nutrition of pathogens. Nature 468: 527–532.
Chen, A., S. He, F. Li, Z. Li, M.Q. Ding, Q.P. Liu, and J.K. Rong. 2012a. Analyses of the sucrose synthase gene family in cotton: Structure, phylogeny and expression patterns. BMC Plant Biology 12: 1–17.
Chen, L.Q., X.Q. Qu, B.H. Hou, D. Sosso, S. Osorio, A.R. Fernie, and W.B. Frommer. 2012b. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335: 207–211.
Chen, Z.L., Y.Y. Gui, C.X. Qin, M. Wang, Q. Liao, D.L. Huang, and Y.R. Li. 2016. Isolation and expression analysis of sucrose synthase gene (ScSuSy4) from sugarcane. Sugar Tech 18: 134–140.
Cheng, J.T., S.Y. Wen, S. Xiao, B.Y. Lu, M.R. Ma, and Z.L. Bie. 2018. Overexpression of the tonoplast sugar transporter CmTST2 in melon fruit increases sugar accumulation. Journal of Experimental Botany 69: 511–523.
Coleman, H.D., J. Yan, and S.D. Mansfield. 2009. Sucrose synthase affects carbon partitioning to increase cellulose production and altered cell wall ultrastructure. Proceedings of the National Academy of Sciences 106: 13118–13123.
Cursi, D.E., H.P. Hoffmann, G.V.S. Barbosa, J.A. Bressiani, R. Gazaffi, R.G. Chapola, A.R.F. Junior, et al. 2021. History and current status of sugarcane breeding, germplasm development and molecular genetics in Brazil. Sugar Tech 24: 112–133.
Dejardin, A., L.N. Sokolov, and L.A. Kleczkowski. 1999. Sugar/osmoticum levels modulate differential abscisic acid-independent expression of two stress-responsive sucrose synthase genes in Arabidopsis. Biochemical Journal 344: 503–509.
Delmer, D.P., and Y. Amor. 1995. Cellulose biosynthesis. The Plant Cell 7: 987–1000.
Eid, A., C. Mohan, S. Sanchez, D. Wang, and F. Altpeter. 2021. Multiallelic, targeted mutagenesis of Magnesium Chelatase with CRISPR/Cas9 provides a rapidly scorable phenotype in highly polyploid sugarcane. Frontiers in Genome Editing 3: 654996.
FAOSTAT. 2020. Sugarcane production in 2019, Crops/Regions/World list/Production Quantity (pick lists).
Fu, H.Y., and W.D. Park. 1995. Sink-and vascular-associated sucrose synthase functions are encoded by different gene classes in potato. The Plant Cell 7: 1369–1385.
Gaudin, V., P.A. Lunness, P.R. Fobert, M. Towers, C.R. Khamlichi, J.A.H. Murray, E. Coen, and J.H. Doonan. 2000. The expression of D-cyclin genes defines distinct developmental zones in snapdragon apical meristems and is locally regulated by the Cycloidea gene. Plant Physiology 122: 1137–1148.
Gerber, L., B. Zhang, M. Roach, U. Rende, A. Gorzsás, M. Kumar, I. Burgert, T. Niittylä, and B. Sundberg. 2014. Deficient sucrose synthase activity in developing wood does not specifically affect cellulose biosynthesis, but causes an overall decrease in cell wall polymers. The New Phytologist 203: 1220–1230.
Goldner, William, Margaret Thom, and Andrew Maretzki. 1991. Sucrose metabolism in sugarcane cell suspension cultures. Plant Science 73: 143–147.
Haigler, C.H., M.I. Datcheva, P.S. Hogan, V.V. Salnikov, S. Hwang, K. Martin, and D.P. Delmer. 2001. Carbon partitioning to cellulose synthesis. Plant Cell Walls 47: 29–51.
Hänggi, E., and A.J. Fleming. 2001. Sucrose synthase expression pattern in young maize leaves: Implications for phloem transport. Planta 214: 326–329.
Hesse, H., and L. Willmitzer. 1996. Expression analysis of a sucrose synthase gene from sugar beet (Beta vulgaris L.). Plant Molecular Biology 30: 863–872.
Hirose, T., G.N. Scofield, and T. Terao. 2008. An expression analysis profile for the entire sucrose synthase gene family in rice. Plant Science 174: 534–543.
Islam, M.D.Z., X.M. Hu, L.F. Jin, Y.Z. Liu, and S.A. Peng. 2014. Genome-wide identification and expression profile analysis of citrus sucrose synthase genes: Investigation of possible roles in the regulation of sugar accumulation. PLoS One 9: e113623.
Jiang, N., L.F. Jin, J.A.T. Silva, M.D.Z. Islam, H.W. Gao, Y.Z. Liu, and S.A. Peng. 2014. Activities of enzymes directly related with sucrose and citric acid metabolism in citrus fruit in response to soil plastic film mulch. Scientia Horticulturae 168: 73–80.
Julius, B.T., K.A. Leach, T.M. Tran, R.A. Mertz, and D.M. Braun. 2017. Sugar transporters in plants: New insights and discoveries. Plant and Cell Physiology 58: 1442–1460.
Kandel, R., X.P. Yang, J. Song, and J.P. Wang. 2018. Potentials, challenges, and genetic and genomic gesources for sugarcane biomass improvement. Frontiers in Plant Science 9: 151.
Khan, Q., J.Y. Chen, X.P. Zeng, Y. Qin, D.J. Guo, A. Mahmood, L.T. Yang, et al. 2021. Transcriptomic exploration of a high sucrose mutant in comparison with the low sucrose mother genotype in sugarcane during sugar accumulating stage. GCB Bioenergy 13: 1448–1465.
Koch, K. 2004. Sucrose metabolism: Regulatory mechanisms and pivotal roles in sugar sensing and plant development. Current Opinion in Plant Biology 7: 235–246.
Leggewie, G., A. Kolbe, R. Lemoine, U. Roessner, A. Lytovchenko, E. Zuther, J. Kehr, W.B. Frommer, J.W. Riesmeier, and L. Willmitzer. 2003. Overexpression of the sucrose transporter So SUT1 in potato results in alterations in leaf carbon partitioning and in tuber metabolism but has little impact on tuber morphology. Planta 217: 158–167.
Lei, M.H., B.Y. Ye, B.M. Wang, Z.X. Huang, H. Zhang, L.P. Xu, Y.Q. Chen, and R.K. Chen. 2008. Cloning of sucrose synthase gene from sugarcane (Saccharum officinarum L.). Chinese Journal of Applied and Environmental Biology 14: 177–179.
Li, J., E.B. Fernández, A. Bahaji, F.J. Muñoz, M. Ovecka, M. Montero, M.T. Sesma, et al. 2013a. Enhancing sucrose synthase activity results in increased levels of starch and ADP-glucose in maize (Zea mays L.) seed endosperms. Plant and Cell Physiology 54: 282–294.
Li, H.P. P.S. Ming, L.H. Ming. 2013b. Cloning and sequence analysis of a chewing cane sucrose synthase gene (SoSuS1) and investigation of expression profiling. Biotechnology Bulletin 0(11): 58-62.
Li, J.M., M.F. Qin, X. Qiao, Y.S. Cheng, X.L. Li, H.P. Zhang, and J. Wu. 2017. A new insight into the evolution and functional divergence of SWEET transporters in Chinese white pear (Pyrus bretschneideri). Plant and Cell Physiology 58: 839–850.
Lin, I.W., D. Sosso, L.Q. Chen, K. Gase, S.G. Kim, D. Kessler, P.M. Klinkenberg, M.K. Gorder, B.H. Hou, and X.Q. Qu. 2014. Nectar secretion requires sucrose phosphate synthases and the sugar transporter SWEET9. Nature 508: 546–549.
Lingle, S.E., and J.M. Dyer. 2001. Cloning and expression of sucrose synthase-1 cDNA from sugarcane. Journal of Plant Physiology 158: 129–131.
Mizuno, H., S. Kasuga, and H. Kawahigashi. 2016. The sorghum SWEET gene family: Stem sucrose accumulation as revealed through transcriptome profiling. Biotechnology for Biofuels 9: 1–12.
Moore, P.H., A.H. Paterson, and T. Tew. 2013. Sugarcane: The crop, the plant, and domestication. Sugarcane: Physiology, Biochemistry, and Functional Biology.
Ohto, M., K. Onai, Y. Furukawa, E. Aoki, T. Araki, and K. Nakamura. 2001. Effects of sugar on vegetative development and floral transition in Arabidopsis. Plant Physiology 127: 252–261.
Pan, Y.Q., H.L. Luo, and Y.R. Li. 2009. Soluble acid invertase and sucrose phosphate synthase: Key enzymes in regulating sucrose accumulation in sugarcane stalk. Sugar Tech 11: 28–33.
Paterson, A.H., P.H. Moore, and T.L. Tew. 2013. The gene pool of Saccharum species and their improvement. In Genomics of the Saccharinae, ed. A.H. Paterson, 43–71. USA: Springer, New York.
Poovaiah, C.R., M. Mazarei, S.R. Decker, G.B. Turner, R.W. Sykes, M.F. Davis, and C.N.J. Stewart. 2015. Transgenic switchgrass (Panicum virgatum L.) biomass is increased by overexpression of switchgrass sucrose synthase (PvSUS1). Biotechnology Journal 10: 552–563.
Rolland, F., B. Moore, and J. Sheen. 2002. Sugar sensing and signaling in plants. The Plant Cell 14: S185–S205.
Rosche, E., D. Blackmore, M. Tegeder, T. Richardson, H. Schroeder, T.J.V. Higgins, W.B. Frommer, C.E. Offler, and J.W. Patrick. 2002. Seed-specific overexpression of a potato sucrose transporter increases sucrose uptake and growth rates of developing pea cotyledons. The Plant Journal 30: 165–175.
Ruan, Y.L., D.J. Llewellyn, and R.T. Furbank. 2003. Suppression of sucrose synthase gene expression represses cotton fiber cell initiation, elongation, and seed development. The Plant Cell 15: 952–964.
Ruan, Y.L., D.J. Llewellyn, Q. Liu, S.M. Xu, L.M. Wu, L. Wang, and R.T. Furbank. 2008. Expression of sucrose synthase in the developing endosperm is essential for early seed development in cotton. Functional Plant Biology 35: 382–393.
Stein, O., and D. Granot. 2019. An overview of sucrose synthases in plants. Frontiers in Plant Science 10: 95.
Sturm, A., S. Lienhard, S. Schatt, and M. Hardegger. 1999. Tissue-specific expression of two genes for sucrose synthase in carrot (Daucus carota L.). Plant Molecular Biology 39: 349–360.
Sugiyama, Akifumi, Yuka Saida, Mayuko Yoshimizu, Kojiro Takanashi, Davide Sosso, Wolf B. Frommer, and Kazufumi Yazaki. 2017. Molecular characterization of LjSWEET3, a sugar transporter in nodules of Lotus japonicus. Plant and Cell Physiology 58: 298–306.
Thirugnanasambandam, P.P., P.J. Mason, N.V. Hoang, A. Furtado, F.C. Botha, and R.J. Henry. 2019. Analysis of the diversity and tissue specificity of sucrose synthase genes in the long read transcriptome of sugarcane. BMC Plant Biology 19: 160.
Van, M., J. Margaretha, J.H. Groenewald, M. Stitt, J. Kossmann, and F.C. Botha. 2010. Downregulation of pyrophosphate: d-fructose-6-phosphate 1-phosphotransferase activity in sugarcane culms enhances sucrose accumulation due to elevated hexose-phosphate levels. Planta 231: 595–608.
Verma, Anjali K., S.K. Upadhyay, P. Cetal Verma, Sushil Solomon, and Shyam B. Singh. 2011. Functional analysis of sucrose phosphate synthase (SPS) and sucrose synthase (SS) in sugarcane (Saccharum) cultivars. Plant Biology 13: 325–332.
Wang, Z., P. Wei, M.Z. Wu, Y.L. Xu, F. Li, Z.P. Luo, J.F. Zhang, et al. 2015. Analysis of the sucrose synthase gene family in tobacco: Structure, phylogeny, and expression patterns. Planta 242: 153–166.
Wei, Z.G., Z.S. Qu, L.J. Zhang, S.J. Zhao, Z.H. Bi, X.H. Ji, X.W. Wang, and H.R. Wei. 2015. Overexpression of poplar xylem sucrose synthase in tobacco leads to a thickened cell wall and increased height. PLoS ONE 10: e0120669–e0120669.
Wu, J., W. Liu, L. Yuan, W.Q. Guan, C.S. Brennan, Y.Y. Zhang, J. Zhang, and Z.D. Wang. 2017. The influence of postharvest UV-C treatment on anthocyanin biosynthesis in fresh-cut red cabbage. Scientific Reports 7: 1–11.
Xiao, X.H., C.R. Tang, Y.J. Fang, M. Yang, B.H. Zhou, J.Y. Qi, and Y. Zhang. 2014. Structure and expression profile of the sucrose synthase gene family in the rubber tree: Indicative of roles in stress response and sucrose utilization in the laticifers. The FEBS Journal 281: 291–305.
Zhang, D.Q., B.H. Xu, X.H. Yang, Z.Y. Zhang, and B.L. Li. 2011. The sucrose synthase gene family in Populus: Structure, expression, and evolution. Tree Genetics & Genomes 7: 443–456.
Zhu, Y.J., E. Komor, and P.H. Moore. 1997. Sucrose accumulation in the sugarcane stem is regulated by the difference between the activities of soluble acid invertase and sucrose phosphate synthase. Plant Physiology 115: 609–616.
Zrenner, R., M. Salanoubat, L. Willmitzer, and U. Sonnewald. 1995. Evidence of the crucial role of sucrose synthase for sink strength using transgenic potato plants (Solanum tuberosum L.). The Plant Journal 7: 97–107.
Funding
This research was funded by Pakistan Agricultural Research Council (PARC), grant number PSDP No. 719. The APC was funded by PARC.
Author information
Authors and Affiliations
Contributions
Conceptualization was done by MN and MRK; methodology was done by MN; software was done by MN; validation was done by BS, IS and SI; formal analysis was carried out by SA; investigation was done by MN; resources were done by MRK; data curation was done by IS; writing—original draft preparation were done by MN; writing—review and editing were done by MRK and SA; visualization was done by IS; supervision was done by MN; project administration was done by MRK; funding acquisition was done by MRK.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Consent for publication
All authors have read and agreed to the current version of the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Noman, M., Siddique, I., Saleem, B. et al. In Silico Dissection and Expression Analysis of Sucrose Synthase Gene Family in Sugarcane. Sugar Tech 24, 1766–1777 (2022). https://doi.org/10.1007/s12355-022-01151-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-022-01151-1