Abstract
Over the past two decades, genetic dissection of complex phenotypes of economic and biological interest has revealed the chromosomal locations of many quantitative trait loci (QTLs) in rice and their contributions to phenotypic variation. Mapping resolution has varied considerably among QTL studies owing to differences in population size and number of DNA markers used. Additionally, the same QTLs have often been reported with different locus designations. This situation has made it difficult to determine allelic relationships among QTLs and to compare their positions. To facilitate reliable comparisons of rice QTLs, we extracted QTL information from published research papers and constructed a database of 1,051 representative QTLs, which we classified into 21 trait categories. This database (QTL Annotation Rice Online database; Q-TARO, http://qtaro.abr.affrc.go.jp/) consists of two web interfaces. One interface is a table containing information on the mapping of each QTL and its genetic parameters. The other interface is a genome viewer for viewing genomic locations of the QTLs. Q-TARO clearly displays the co-localization of QTLs and distribution of QTL clusters on the rice genome.
Similar content being viewed by others
Introduction
During the last two decades, quantitative trait locus (QTL) analysis has been extensively used to identify the chromosomal locations and phenotypic contributions of QTLs in rice. In addition, QTL data from rice have been useful in genetic studies of other grass species, which can be compared using information from syntenic regions in their genomes (Armstead et al. 2008; Jo et al. 2008). The availability of the rice genome sequence (IRGSP 2005) has accelerated the development of large numbers of molecular markers useful for fine mapping of target QTLs and cloning of QTLs for morphological and physiological traits (reviewed by Yamamoto et al. 2009).
Rice QTL information has recently been extracted from reports published between 1994 and 2006 and summarized in the Gramene-QTL database (Ni et al. 2009). These reports have identified many QTLs associated with a wide range of traits, but that information might be redundant for several reasons. One reason is that the same primary mapping populations have been used for QTL analysis in many different studies, mainly to avoid the time required to develop new mapping populations. QTL analyses performed under multiple environmental conditions and using different genetic interaction analyses might also generate redundant QTL information associated with a given trait. Although redundant QTL information is useful for verification, it makes comparison of QTLs among different studies more complex, and it produces noise in the determination of the accurate physical position of QTLs.
To compare QTLs among different studies, the method of meta-QTL analysis has been developed (Goffinet and Gerber 2000) and can be implemented using interactive software (Veyrieras et al. 2007). However, if there are false-positive QTLs in the studies being analyzed, these might produce false-positive intervals for the meta-QTLs. We believe that effective comparison of QTLs across studies requires (1) the organization of informative QTLs without redundancy and (2) the definition of the physical positions of markers linked to these QTLs. An analysis tool based on these characteristics might help researchers see and compare the genomic positions of QTLs categorized into different trait categories and inform them of QTL clusters or co-localized regions on the rice genome map. Such “cluster” regions are responsible for genetic features such as linkage drag and pleiotropy, and they are important for their implications in rice breeding.
To reduce the effort necessary to discriminate informative QTLs, we have developed a representative QTL database, the QTL Annotation Rice Online Database (Q-TARO). In this database, we have collected rice QTL information from published papers and summarized informative QTL information by removing redundant QTLs. Q-TARO consists of two web-based interfaces: a table of summary information and a genome browser integrated with information on anchor markers, genes and representative QTLs. Using this database, we have identified QTL clusters on several chromosomes.
Results
Extraction of QTL information
To obtain information about the physical positions of anchor restriction fragment length polymorphism (RFLP) markers for the mapping of QTLs, the program BlastN was used to compare the nucleotide sequences of RFLP probes against rice genome sequences. As a result, the physical positions of 3,704 markers were specified. Additionally, the physical positions of 16,582 simple sequence repeat (SSR) markers were estimated via the program ePCR (Schuler 1997). The map positions of 3,455 bacterial artificial chromosome (BAC)/plant artificial chromosome (PAC) and 29,389 RAP2 (Rice Annotation Project et al. 2008) clones were also used as landmarks for determining the physical position of QTLs. All information on the anchor markers, genomic clones, and gene loci is contained in the Q-TARO database (http://qtaro.abr.affrc.go.jp/).
To extract the non-redundant QTL information, we first focused on the datasets for 5,096 QTLs reported in 1,214 articles and examined 29 categories of related information (listed in Table 1). Eventually, 1,051 QTLs were selected from 463 articles and listed as representative QTLs. Figure 1 summarizes the papers used in this study, categorized by year of publication and type of marker used for QTL mapping. The earliest papers were published in 1990, and the number of publications peaked in 2002. RFLP markers were the most commonly used markers for QTL localization until 2005 when SSR markers became the most frequently used type.
Summary of references containing representative rice QTL map information (1988–2008). Within each publication year, each article was classified into one of four categories by the type of anchor marker used: SSR simple sequence repeat, RFLP restriction fragment length polymorphism, Mixture two or more types of markers were used, Other markers other than RFLP and SSR were used.
Cross combinations and population structures used for QTL analysis
In the early 1990s, QTL analysis was usually performed using F2 or F3 populations. Recombinant inbred lines (RILs) and backcross inbred lines (BILs) have been used frequently since the late 1990s. More than 75% of the populations used for QTL analysis were primary mapping populations such as RILs, F2, doubled haploid lines (DHLs), BILs, and backcross populations of the F1 to the recurrent parent (BC1F1; Fig. 2a). For the accurate evaluation of genetic effects of a donor allele in a uniform genetic background, advanced backcross progeny were widely used. In our dataset, 11% of the populations were repeated backcross progeny (Fig. 2a). Cloning of QTLs has usually been carried out using large-scale analyses of F2 populations with many individuals. In our dataset, 7% of the populations were F2 populations with a large number of plants (1,000 or more) used for positional cloning of QTLs.
Categories of mapping populations and cross combinations used for the detection of representative rice QTLs. a Percentage of total articles using each type of mapping population. RIL recombinant inbred line, DHL doubled haploid line, BIL backcross inbred line, BC 1 F 1 backcross population of F1 × recurrent parent. The F2 population category is further divided according to the number of individuals comprising each population (<1,000 individuals vs ≥1,000 individuals) because these two categories are often used for different objectives. b Cross combinations used in more than 10 articles.
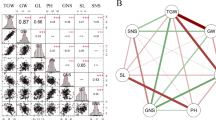
In total, QTL information has been generated mainly using 13 genetic combinations (22% of the 2,054 total populations; Fig. 2b), as reported in 454 papers. In particular, the top two combinations, “Azucena” × “IR64” (Guiderdoni et al. 1992) and “Minghui 63” × “Zhenshan 97” (Yu et al. 1997) comprised 8% of the total combinations reported.
Positioning of representative QTLs on the rice genome map
Physical positions for half of the 1,051 representative QTLs were determined within intervals of 1 to 10 Mb (Table 2). Candidate genomic regions of 63 QTLs were narrowed down to <100 kb. Among these 63 QTLs, positional candidate genes or BAC/PAC clones were identified in 50 QTLs by high-resolution mapping. Among traits affecting flowering time or heading date, 12 of the 49 QTLs detected were cloned or mapped to a known locus. Chromosomal positions could not be precisely determined for more than 20% of all QTLs owing to the lack of two flanking markers.
The largest number of QTLs were mapped on chromosome 1 (190 QTLs), followed by 149 QTLs on chromosome 6 and 142 QTLs on chromosome 3. QTLs for five morphological traits and tolerance to drought and soil stresses (e.g., salinity and heavy metals) were detected on chromosomes 1 and 3. Many QTLs for physiological traits such as eating quality and heading date were mapped to chromosome 6.
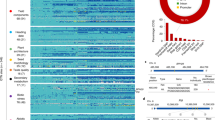
Figure 3 shows the frequency distribution of representative QTLs within 2-Mb intervals on each of the 12 rice chromosomes. The number of QTLs detected varied both among the chromosomes and among regions within each chromosome. Few QTLs were located in the pericentromeric regions of most chromosomes. In several chromosomal regions, relatively higher numbers of QTL (15 or more QTLs per 2-Mb interval) were mapped; these were designated as “QTL clusters.” QTL clusters were found in two regions (4–6 and 38–44 Mbp) on chromosome 1, 0–2 Mbp on chromosome 3, 2–8 Mbp on chromosome 6, 32–34 Mbp on chromosome 4, and 20–22 Mbp on chromosome 9.
Distribution of representative QTLs on the 12 rice chromosomes. The position of each QTL was assigned as the intermediate point between two flanking markers or between the start and end position of a single marker. The total number of QTLs and the number within each trait category were calculated for every 2-Mb interval.
QTL information table and QTL genome viewer
The Q-TARO database can be queried via two web interfaces. QTL information such as trait and trait category, population (parent), mapping method, accuracy (LOD value), map position, and reference can be obtained from the QTL Information table (http://qtaro.abr.affrc.go.jp/qtab/table/). Figure 4a shows a representative QTL Information table listing of mapped QTLs and their genetic parameters. Information is displayed by selecting QTL categories and chromosomes from the drop-down boxes and by specifying search text. Information on any QTL can be exported in text format. A graphical image of QTL positions can be obtained by using the QTL Genome Viewer (http://qtaro.abr.affrc.go.jp/cgi-bin/gbrowse/; Fig. 4b), which allows the user to select a given QTL by its physical position on the chromosome. The QTLs are displayed by trait categories using different colors, allowing the locations of QTLs within each trait category to be easily compared. Users can change the order of information bars (called “tracks”) by dragging and dropping bars next to each other, allowing them to compare the positions of interesting QTLs.
Screenshots of Q-TARO. a QTL information table. This table shows details of the representative QTLs. All displayed information can be exported as text format data. b QTL Genome Viewer. The viewer shows co-localized QTLs on the 12 rice chromosomes. QTLs are categorized into three major trait categories, as described in Table 2.
Discussion
Anchors for QTL positions on the rice genome map
To determine the physical position of a QTL, it is necessary to know the physical position of one or more linked markers. Sequence information for RFLP and SSR markers used in QTL mapping allowed us to determine their physical positions through the use of BLASTN and e-PCR searches. The positions of SSR markers were determined on the basis of the alignment between the primers and the genomic sequence of Nipponbare; a maximum of one mismatch or indel was allowed in e-PCR. For markers designed from genomic clones of other grass species, it was sometimes difficult to determine the map position using sequence alignment. Our experience with this database leads us to recommend that anchor markers with defined chromosomal positions be used to localize QTLs.
The size of the confidence interval for a QTL map position depends both on mapping population size and on marker density. The number of RFLP markers is limited to a few thousand, but over 20,000 SSR markers are currently available (IRGSP 2005). Additionally, detection of RFLPs requires Southern hybridization, whereas SSR polymorphisms are easily detected by PCR. For these reasons, mapping studies using SSR markers usually include many more markers than those using RFLP markers. The use of SSR markers is probably the reason that mapping resolution seems to be improved in more recent studies. Recently, single nucleotide polymorphisms (SNPs) have been found at 160,000 non-redundant positions in the rice genome (McNally et al. 2009). Markers derived from these SNPs will allow us to determine QTL positions more precisely than ever before.
Distribution of QTL clusters in the rice genome map
Co-localizations of 15 or more QTLs within the same 2-Mb chromosomal region were classified as “QTL clusters.” In this study, we identified six QTL clusters. It is unclear whether QTL clusters associated with multiple traits represent the pleiotropic effects of a single gene or close linkage of different genes affecting different traits. Through our comprehensive search of the rice QTL mapping literature, we have identified a possible explanation for QTL clusters of QTLs for the same trait: we suggest that co-localized QTLs identified in different studies might be caused by the same gene, with slight differences in localization caused by different measurement methods.
A cluster of 19 QTLs collected from 16 articles was localized in region 4–6 Mb of chromosome 1. Of these, ten QTLs controlling seed number or seed weight were detected using the same experimental materials and methods (i.e., mapping population, population size, and marker type) but reported in different articles. This suggests that these ten QTLs represent a single QTL (i.e., that they are redundant QTLs). All 10 QTLs were mapped in the range of 4.6–5.8 Mb, which also includes Gn1a, a gene at position 5.3 Mb controlling grain number per panicle (Ashikari et al. 2005). We also identified a second QTL cluster on chromosome 1. The largest number of QTLs (79) was detected in region 38–44 Mb on chromosome 1. Out of these 79 QTLs, 25 were detected in analyses for dwarfism, drought resistance, or nitrogen response. However, these seemingly different traits are all likely to be associated with plant height because height has been often used as the criterion for response to biotic or abiotic stress. Within the region of this QTL cluster, the semidwarfing gene sd1 has been identified at the 40.1-Mb position (Sasaki et al. 2002). It is very likely that several of the QTLs detected in this region were generated by the action of allelic differences at the Sd1 locus.
A second large cluster consisting of 68 QTLs was identified in region 2–8 Mb of chromosome 6. In this region, we observed multiple genes associated with eating quality (e.g., starch synthase IIa; SSIIa; Jiang et al. 2004) and heading date [e.g., Hd3a (Kojima et al. 2002) and Hd3b (Monna et al. 2002)]. Moreover, other genes controlling these traits were found near this cluster. For example, waxy has been mapped to 1.8 Mb (Okagaki 1992), and Hd1, which controls photosensitivity, was mapped at 9.3 Mb (Yano et al. 2000). The interval including these QTLs is likely to have been detected across multiple studies. Heading date and eating quality are easy characteristics to measure, and because they differ significantly between japonica and indica populations, they are relatively easy to map. Moreover, it is predictable that a gene or QTL for heading date could have a pleiotropic effect on eating quality, or vice versa.
A cluster of 19 QTLs was detected at region 20–22 Mb on chromosome 9. Many of the QTLs in this cluster are associated with tillering and leaf angle traits. Recently, TAC1, a gene responsible for controlling tiller angle, was cloned at 23.9 Mb (Yu et al. 2007). Although clear evidence for the association between TAC1 and QTLs related to grain yield and drought tolerance has not been obtained (one QTL for yield has been mapped near but not at the TAC1 locus; Xie et al. 2008), TAC1 might be detected as a QTL for those traits because tillering angle can affect photosynthetic ability and thus be associated with yield potential and drought tolerance.
In the clusters we have detected, several QTLs with major effects could be the primary cause of the cluster. When a new QTL is detected at the position of a QTL cluster presented here, it should be considered whether the QTL might represent a pleiotropic effect of a previously identified major QTL rather than being an entirely new one.
Comparison with QTL clusters identified in other studies
In this study, we found many interesting regions showing a relatively high density of QTLs and genes that did not meet our definition of a “QTL cluster” (i.e., they did not have more than 15 QTLs in 2 Mb). Some of these regions were found in similar locations to QTL clusters identified in other studies. In their reports on disease resistance genes, Zhou et al. (2004) and Wisser et al. (2005) describe clusters composed of nucleotide-binding site/leucine-rich repeat genes on chromosomes 11 and 12. In our study, these regions were detected as relatively dense regions associated with disease resistance, but they did not contain enough QTLs to be classified as QTL clusters, primarily because potentially redundant QTLs were removed.
In a study of rice domestication, QTLs for domestication traits were found to be clustered within certain chromosome regions (Sweeney and McCouch 2007). The authors identified the pericentromeric region of chromosome 7 as a QTL cluster for seed color, panicle structure, dormancy, shattering, and other traits. This site was not detected as a QTL cluster in our study. However, we detected a QTL cluster in the 32–34 Mb region of chromosome 4 containing 24 QTLs including domestication traits such as grain shattering. The Sh4 gene, responsible for grain shattering, has been cloned near this cluster at 34.6 Mb (Li et al. 2006a). A major cluster related to rice domestication seems to be located on the different position from ours (Li et al. 2006b). However, we cannot determine whether there has been crossing over between our cluster and the cluster of domestication reported on the long arm of chromosome 4 (Li et al. 2006b) because there is no precise information for the physical position of that cluster. Similarly, it is difficult to compare the positions of clusters reported on rice chromosomes 3, 6, and 9 with the positions of the QTL clusters we observed on the same chromosomes.
Rice breeders have long known that certain desirable traits are tightly linked with unfavorable characters, but until recently, it has been difficult to determine whether these associations are caused by linked genes or pleiotropy. For example, the reason for the relationship between the field blast resistance gene pi21 and poor eating quality was unknown for 80 years. Recently, this tight linkage was broken by positional cloning and marker-assisted selection, and the new cultivar with blast resistance and good eating quality has been developed (Fukuoka et al. 2009).
If linkage drag among QTLs appears to be caused by a QTL cluster, the Q-TARO database could be used to illustrate the position of the cluster. It might be also a useful tool for determining the genetic position at which crossing over could break the linkage drag.
New genetic combinations and tools will provide novel QTL information for Q-TARO
In QTL studies over the past 20 years, major QTLs controlling a wide range of phenotypes have been identified. However, many of these QTLs have been detected within limited cross populations made from genetically distant lines—indica and japonica—because of the considerable level of phenotypic variation between these two subspecies. The high level of sequence polymorphism between indica and japonica also encouraged researchers to use these wide crosses as mapping populations, leading to highly redundant results in QTL analyses.
More recently, the increased availability of SSR markers has enabled QTL analysis using populations between closely related lines, such as between temperate japonica lines. These analyses have identified several unique QTLs for heading date (Matsubara et al. 2008), culm length (Hori et al. 2009), pre-harvest sprouting resistance (Hori et al. 2010), and eating quality (Takeuchi et al. 2008; Kobayashi and Tomita 2008; Wada et al. 2008). The success of this approach implies that the use of a wide range of cross combinations might increase the opportunity to detect unique QTLs. Genome-wide SNP discovery can be performed using closely related cultivars (Yamamoto et al. 2010) as well as diverse germplasm accessions (McNally et al. 2009). This situation should allow us to perform QTL mapping using almost any cross combination. The availability of diverse populations and high-density SNP markers will make it possible to discover novel QTLs and accumulate this information in Q-TARO.
Methods
Extraction of representative QTL/gene information from articles
To extract QTL information from the literature, we searched databases for QTL/gene mapping papers published during the 20 years from 1988 to 2008. English-language articles were retrieved from both PubMed (http://www.ncbi.nlm.nih.gov/sites/entrez) and HighWire (http://highwire.stanford.edu/) databases using the search formula “Rice” AND (“quantitative trait” OR “QTL” OR “mapping”). Journal articles published by Japanese academic societies were retrieved from JStage (http://www.jstage.jst.go.jp/browse/-char/ja) with the same search formula. Journals that were not registered in any of these three databases were manually searched using journal web sites and hard copies. In addition, articles related to QTL mapping research were extracted from the Rice Genetics Newsletter (http://www.gramene.org/newsletters/rice_genetics/).
QTL/gene information was extracted from 1,214 selected articles in two steps. First, only highly reliable QTL information was selected; reports involving epistasis or genetic × environmental interaction effects were excluded. Next, QTLs retrieved from multiple papers but describing the same trait within the same genetic interval were screened for redundancy; if the QTLs appeared to be redundant, a single representative QTL was selected. Relevant background information in 29 categories (e.g., mapping method, type of marker, year of publication) was extracted for each representative QTL by careful reading of each article and entered into the database. The QTLs themselves were placed into three major categories: morphological traits, physiological traits, and resistance or tolerance. Each of these main categories was subdivided into six or eight groups (Table 2). Traits not fitting into any of these categories were classified as “other.”
Determination of physical positions of QTLs
Sequences of both RFLP probes and SSR primers were mapped to rice genomic sequence locations using IRGSP Pseudomolecules build 4.0 (http://rgp.dna.affrc.go.jp/J/IRGSP/Build4/build4.html).
The sequences of RFLP probes were compared with RAP2 transcript sequences using BLASTN searches (Altschul et al. 1997). When the RFLP and RAP2 sequences overlapped by >70% in the first alignment and the two sequences showed >70% identity, the physical position of the RAP2 transcript on the rice genome map was identified as the position of the RFLP marker. RFLP probes with low similarity to RAP2 transcripts were positioned using their sequences as queries in BLASTN searches against rice genomic sequences.
Physical positions of SSR markers were determined by one of two methods depending on which genome had been used for SSR marker design. If the Nipponbare genome had been used, flanking primer sequences were used as queries in a BLASTN search, and the locations of both primers were determined on the basis of perfect match alignment between the primers and Nipponbare genome sequences. The positions of SSR markers designed from the sequences of genotypes other than Nipponbare were estimated using e-PCR to search Nipponbare genomic sequences (IRGSP Pseudomolecules build 4.0). e-PCR was run with a five-word size and default parameters for a 50-bp margin on the product size, with one mismatch and one indel allowed in the SSR primers. Locations were identified by selecting the position with the smallest absolute values of gap and indel and then subtracting the observed size of the hit from the expected size of the PCR product. The files of BAC/PAC and RAP2 information were downloaded from the RAP ftp site (http://rapdb.dna.affrc.go.jp/archive/build4.html) and used as landmarks for determining the physical position of the QTL/gene of interest.
For QTLs not associated with RFLP or SSR markers, other methods were used to determine their physical positions. If the map interval for a QTL had been determined using two flanking markers as anchors, the minimum and maximum physical positions of the two markers were defined as the start and end of the target interval, respectively. If the map location of a QTL had been based on co-segregation with a single marker, the position of the QTL was placed at the terminal position of the marker. If a QTL position had been narrowed down to the position of a certain BAC/PAC or gene locus, the physical position of the QTL was placed between the start and end of the corresponding BAC/PAC or locus.
System construction
The Q-TARO database consists of a relational database and two web applications, “QTL Information Table” and “QTL Genome Viewer.” The web applications were implemented as Perl scripts and CGI modules. The database was constructed using MySQL, a relational database management system. We use a GBrowse viewer (http://gmod.org/wiki/Main_Page) configured to access the Q-TARO genome viewer.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402.
Armstead IP, Turner LB, Marshall AH, Humphreys MO, King IP, Thorogood D. Identifying genetic components controlling fertility in the outcrossing grass species perennial ryegrass (Lolium perenne) by quantitative trait loci analysis and comparative genetics. New Phytol. 2008;178:559–71.
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, et al. Cytokinin oxidase regulates rice grain production. Science. 2005;309:741–5.
Fukuoka S, Saka N, Koga H, Ono K, Shimizu T, Ebana K, et al. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science. 2009;325:998–1001.
Goffinet B, Gerber S. Quantitative trait loci: a meta-analysis. Genetics. 2000;155:463–73.
Guiderdoni E, Galinato E, Luistro J, Vergara G. Anther culture of tropical japonica × indica hybrids of rice (Oryza sativa L.). Euphytica. 1992;62:219–24.
Hori K, Yamamoto T, Ebana K, Takeuchi Y, Yano M. A novel quantitative trait locus, qCL1, involved in semi-dwarfism derived from Japanese rice cultivar Nipponbare. Breed Sci. 2009;59:285–95.
Hori K, Sugimoto K, Nonoue Y, Ono N, Matsubara K, Yamanouchi U, et al. Detection of quantitative trait loci controlling pre-harvest sprouting resistance by using backcrossed populations of japonica rice cultivars. Theor Appl Genet. 2010;120:1547–57.
International Rice Genome Sequencing Project (IRGSP). The map-based sequence of the rice genome. Nature. 2005;436:793–800.
Jiang H, Dian W, Liu F, Wu P. Molecular cloning and expression analysis of three genes encoding starch synthase II in rice. Planta. 2004;218:1062–70.
Jo YK, Barker R, Pfender W, Warnke S, Sim SC, Jung G. Comparative analysis of multiple disease resistance in ryegrass and cereal crops. Theor Appl Genet. 2008;117:531–43.
Kobayashi A, Tomita K. QTL detection for stickiness of cooked rice using recombinant inbred lines derived from crosses between japonica rice cultivars. Breed Sci. 2008;58:419–26.
Kojima S, Takahashi Y, Kobayashi Y, Monna L, Sasaki T, Araki T, et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol. 2002;43:1096–105.
Li C, Zhou A, Sang T. Rice domestication by reducing shattering. Science. 2006a;311:1936–9.
Li C, Zhou A, Sang T. Genetic analysis of rice domestication syndrome with the wild annual species, Oryza nivara. New Phytol. 2006b;170:185–93.
Matsubara K, Kono I, Hori K, Nonoue Y, Ono N, Shomura A, et al. Novel QTLs for photoperiodic flowering revealed by using reciprocal backcross inbred lines from crosses between japonica rice cultivars. Theor Appl Genet. 2008;117:935–45.
McNally KL, Childs KL, Bohnert R, Davidson RM, Zhao K, Ulat VJ, et al. Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci USA. 2009;106:12273–8.
Monna L, Lin X, Kojima S, Sasaki T, Yano M. Genetic dissection of a genomic region for a quantitative trait locus, Hd3, into two loci, Hd3a and Hd3b, controlling heading date in rice. Theor Appl Genet. 2002;104:772–8.
Ni J, Pujar A, Youens-Clark K, Yap I, Jaiswal P, Tecle I, et al (2009) Gramene QTL database: development, content and applications. Database bap005
Okagaki RJ. Nucleotide sequence of a long cDNA from the rice waxy gene. Plant Mol Biol. 1992;19:513–6.
Rice Annotation Project (RAP), Tanaka T, Antonio BA, Kikuchi S, Matsumoto T, Nagamura Y, et al. The Rice Annotation Project Database (RAP-DB): 2008 update. Nucleic Acids Res. 2008;36:D1028–33.
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, et al. A mutant gibberellin-synthesis gene in rice. Nature. 2002;416:701–2.
Schuler GD. Sequence mapping by electronic PCR. Genome Res. 1997;7:541–50.
Sweeney M, McCouch S. The complex history of the domestication of rice. Ann Bot. 2007;100:951–7.
Takeuchi Y, Hori K, Suzuki K, Nonoue Y, Takemoto-Kuno Y, Maeda H, et al. Major QTLs for eating quality of an elite Japanese rice cultivar, Koshihikari, on the short arm of chromosome 3. Breed Sci. 2008;58:437–45.
Veyrieras JB, Goffinet B, Charcosset A. MetaQTL: a package of new computational methods for the meta-analysis of QTL mapping experiments. BMC Bioinformatics. 2007;8:49.
Wada T, Ogata T, Tsubone M, Uchimura Y, Matsue Y. Mapping of QTLs for eating quality and physicochemical properties of the japonica rice “Koshihikari”. Breed Sci. 2008;58:427–35.
Wisser RJ, Sun Q, Hulbert SH, Kresovich S, Nelson RJ. Identification and characterization of regions of the rice genome associated with broad-spectrum, quantitative disease resistance. Genetics. 2005;169:2277–93.
Xie X, Jin F, Song MH, Suh JP, Hwang HG, Kim YG, et al. Fine mapping of a yield-enhancing QTL cluster associated with transgressive variation in an Oryza sativa × O. rufipogon cross. Theor Appl Genet. 2008;116:613–22.
Yamamoto T, Yonemaru J-i, Yano M. Towards the understanding of complex traits in rice: substantially or superficially? DNA Res. 2009;16:141–54.
Yamamoto T, Nagasaki H, J-i Y, Ebana K, Nakajima M, Shibaya T, et al. Fine definition of the pedigree haplotypes of closely related rice cultivars by means of genome-wide discovery of single-nucleotide polymorphisms. BMC Genomics. 2010;11:267.
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell. 2000;12:2473–84.
Yu SB, Li JX, Xu CG, Tan YF, Gao YJ, Li XH, et al. Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA. 1997;94:9226–31.
Yu B, Lin Z, Li H, Li X, Li J, Wang Y, et al. TAC1, a major quantitative trait locus controlling tiller angle in rice. Plant J. 2007;52:891–8.
Zhou T, Wang Y, Chen JQ, Araki H, Jing Z, Jiang K, et al. Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR NBS-LRR genes. Mol Genet Genomics. 2004;271:402–15.
Acknowledgments
We thank N. Namiki for his assistance with the determination of physical positions of both RFLP and SSR markers. We also thank E. Kibukawa for construction of the web-based interfaces for the database. This work was supported by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Integrated Research Project for Plant, Insect and Animal using Genome Technology, QT-1006, and Genomics for Agricultural Innovation, GIR-1003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Yonemaru, Ji., Yamamoto, T., Fukuoka, S. et al. Q-TARO: QTL Annotation Rice Online Database. Rice 3, 194–203 (2010). https://doi.org/10.1007/s12284-010-9041-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12284-010-9041-z