Abstract
Molecular markers that enable monitoring of fungi in their natural environment or assist in the identification of specific strains would facilitate Trichoderma utilization, particularly as an agricultural biocontrol agent (BCA). In this study, sequence analysis of internal transcribed spacer regions 1 and 2 (ITS1 and ITS2) of the ribosomal RNA (rRNA) gene cluster, a fragment of the translation elongation factor 1-alpha (tef1) gene, and random amplified polymorphic DNA (RAPD) markers were applied to determine the genetic diversity of Trichoderma atroviride strains collected in Poland, and also in order to identify loci and PCR-based molecular markers useful in genetic variation assessment of that fungus. Although tef1 and RAPD analysis showed limited genetic diversity among T. atroviride strains collected in Poland, it was possible to distinguish major groups that clustered most of the analyzed strains. Polymorphic RAPD amplicons were cloned and sequenced, yielding sequences representing 13 T. atroviride loci. Based on these sequences, a set of PCR-based markers specific to T. atroviride was developed and examined. Three cleaved amplified polymorphic sequence (CAPS) markers could assist in distinguishing T. atroviride strains. The genomic regions identified may be useful for further exploration and development of more precise markers suitable for T. atroviride identification and monitoring, especially in environmental samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Trichoderma is a fungal genus that includes more than 200 species that occur all over the world in different geographical regions and climatic zones (Atanasova et al. 2013; Hoyos-Carvajal et al. 2009; Kredics et al. 2014). They are remarkable for their rapid growth under various environmental conditions, as well as their ability to use different substrates as carbon sources (Kubicek et al. 2003; Atanasova and Druzhinina 2010); thus, they are recognized as efficient decomposers of woody and herbal materials and other organic matter (Atanasova et al. 2013; Schuster and Schmoll 2010). Trichoderma interacts with other microorganisms and plants in diverse ways (Harman et al. 2004; Lorito et al. 2010; Druzhinina et al. 2011; Woo et al. 2006). Trichoderma strains, including Trichoderma atroviride strains, that are promoting efficient plant growth and stress resistance are used as biological control agents (BCAs) in sustainable farming systems (Chet and Inbar 1994; Benítez et al. 2004; Stewart and Hill 2014).
Various molecular techniques have been used in Trichoderma research, in order to investigate genetic diversity within the genus. These include restriction fragment length polymorphism (RFLP) analysis (Dodd et al. 2004a), random amplified polymorphic DNA (RAPD) analysis (Hermosa et al. 2001), amplified fragment length polymorphism (AFLP) analysis (Buhariwalla et al. 2005), sequence-characterized amplified region (SCAR) analysis (Hermosa et al. 2001; Dodd et al. 2004b; Cordier et al. 2007), microsatellite markers (Naef et al. 2006), and sequence analysis (Kindermann et al. 1998; Dodd et al. 2003; Druzhinina et al. 2005; Błaszczyk et al. 2011; Atanasova et al. 2013). One of the simplest and cheapest techniques is RAPD, which is characterized by low repeatability and transferability between different laboratories. Nevertheless, it delivers solid data at minimal expense when used strictly in a single laboratory. Moreover, RAPD markers can be easily transformed into PCR-based SCAR markers, as demonstrated in several studies for T. atroviride (Hermosa et al. 2001; Cordier et al. 2007; Feng et al. 2011).
Application of molecular techniques in Trichoderma taxonomy solved inaccuracies related to phenotypical T. atroviride species identification, as well as demonstrated interspecies genetic variability. In order to reevaluate biocontrol strains reported as Trichoderma harzianum and Trichoderma viride, Hermosa et al. (2000) utilized sequences of internal transcribed spacer regions 1 and 2 (ITS1 and ITS2) of the ribosomal RNA (rRNA) gene cluster and analyzed the hybridization patterns of mitochondrial DNA. Authors identified four groups of Trichoderma including T. atroviride. Identification of mycoparasitic strains performed with assistance of ITS1, ITS2, mtSSUrDNA, and partial chi18-5 gene sequences revealed that the strains belonged to T. harzianum, T. atroviride, and Trichoderma asperellum (Kullnig et al. 2001). Based on the sequences of ITS1, ITS2, and translation elongation factor 1-alpha (tef1) gene fragments, four major clades (A, B, C, and D) of T. atroviride were distinguished, and the fifth clade E was suggested (Dodd et al. 2003; Jaklitsch et al. 2006; Samuels et al. 2006; Mulaw et al. 2010; Gal-Hemed et al. 2011).
Application of Trichoderma BCA strains raised questions about fungal survival and spread in the environment (Dodd et al. 2004a, b). The molecular strategies were applied to develop markers useful for Trichoderma monitoring at different levels: genus-specific (Hagn et al. 2007; Devi et al. 2011; Chakraborty et al. 2011; Friedl and Druzhinina 2012), species-specific (Chen et al. 1999; Kredics et al. 2009; Miyazaki et al. 2009; Friedl and Druzhinina 2012), and strain-specific (Hermosa et al. 2001; Rubio et al. 2005; Cordier et al. 2007; Savazzini et al. 2008). Strain-specific molecular markers proposed for Trichoderma BCA strain monitoring included RAPD, RFLP (Zimand et al. 1994; Bowen et al. 1996; Abbasi et al. 1999), and SCAR markers (Hermosa et al. 2001; Dodd et al. 2004b; Rubio et al. 2005; Cordier et al. 2007; Savazzini et al. 2008; Longa et al. 2009; Feng et al. 2011; Naeimi et al. 2011). Due to the complex taxonomy of Trichoderma, development of strain-specific markers is a long and laborious process (Druzhinina and Kubicek 2005). The most beneficial is development of species-specific markers, which can also be used to simplify and assist in taxonomic identification, based on sequence differences at the interspecies level, useful for elaboration of such markers (Chen et al. 1999; Friedl and Druzhinina 2012; Kredics et al. 2009; Miyazaki et al. 2009).
The aim of this study was to utilize sequences of ITS1 and ITS2 region, a fragment of the tef1 gene, and RAPD markers to analyze the genetic diversity of T. atroviride strains collected in Poland, develop PCR-based markers, and identify genomic regions to enable genetic diversity studies and further development of reliable methods of T. atroviride identification. In this study, RAPD markers were converted into PCR-based markers suitable for detection of T. atroviride. Additionally, application of CAPS markers developed in this study is expected to improve the accuracy of T. atroviride classification.
Materials and methods
Fungal material and strain identification
All Trichoderma strains used in this study were obtained from a stock culture collection maintained by the Microbiology Laboratory at the Research Institute of Horticulture in Skierniewice, Poland. Stock cultures were deep-frozen in glycerol and stored at −80 °C. Forty strains of T. atroviride were used in this study; these included 38 strains of T. atroviride collected at different locations in Poland, and the two reference strains CBS 693.94 (Dodd et al. 2003) and IMI206040 (Kubicek et al. 2011) (Table 1). Strain identification was based on the internal transcribed spacer regions 1 and 2 (ITS1 and ITS2) of the rRNA gene cluster sequences and TrichOKey v. 2.0 (Druzhinina et al. 2005) and sequences of the fragment of the gene encoding translation elongation factor 1-alpha (tef1) and TrichoBLAST v. 1.0 analysis (Kopchinskiy et al. 2005). All sequences were obtained by PCR and direct amplicon sequencing from both directions. ITS1 and ITS2 regions were PCR-amplified with ITS6 and ITS4 primers (Cooke and Duncan 1997; White et al. 1990). For the tef1 sequencing, primers EF1-728 F and TEFLLErev were used (Carbone and Kohn 1999; Jaklitsch et al. 2005). ITS and tef1 sequences of the strains are available in the NCBI GenBank (Accession numbers are provided in Table 1). The following strains were used for the PCR-based marker species specificity evaluation: Trichoderma aggressivum f. aggressivum CBS 100528, T. aggressivum f. europaeum CBS 100526, T. asperellum TRS705, Trichoderma citrinoviride TRS119, Trichoderma cerinum 38.24.06.2, Trichoderma crassum TRS113, Trichoderma gamsii TRS123, Trichoderma hamatum TRS127, T. harzianum sensu lato II subclade CBS 466.94, T. harzianum s. l. III subclade TRS60, T. harzianum s. l. X subclade CBS 115901, T. harzianum s. l. Lixii subclade TRS66, T. harzianum sensu stricto TRS71, Trichoderma longibrachiatum TRS708, Trichoderma pleuroticola TRS120, Trichoderma spirale TRS111, Trichoderma tomentosum TRS82, Trichoderma virens Gv29-8, Trichoderma viride TRS575, Trichoderma viridescens TRS35, and Trichoderma velutinum 29.24.06.1. For all above mentioned strains, species/subclade affiliation was confirmed by ITS and tef1 sequencing as described above for T. atroviride. Mycelia taken from stock cultures were placed on potato dextrose agar (PDA) plates (Sigma-Aldrich, St. Louis, MO, USA) and cultured in the dark at 25 °C for 4–7 days. Mycelia grown on PDA were then used to inoculate 250-mL flasks containing 50 mL potato dextrose broth (Sigma-Aldrich), and cultures were incubated in the dark on a rotary shaker at 200 rpm for 7 days at room temperature (21 ± 2 °C).
DNA extraction
Total fungal DNA was isolated using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany), according to the manufacturer’s instructions with slight modifications. Fungal cells of tested Trichoderma strains were harvested by centrifugation of liquid cultures and dried on filter paper at room temperature (21 ± 2 °C), prior to grinding in liquid nitrogen with quartz sand (Sigma-Aldrich). Proteinase K and RNase A were added to the lysis buffer. The purity and quantity of isolated DNA were evaluated by agarose gel electrophoresis followed by ethidium bromide staining, and by determination of A260/A280 using a NanoDrop 8000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). Finally, the DNA concentration was adjusted to 1 ng/μL by diluting the samples with 10 mmol/L Tris-HCl, pH 8.0.
RAPD analysis
A set of 520 decamer primers (primer kits OPA to OPZ from Operon Technology, Alameda, CA, USA) was used for initial RAPD screening of eight strains of T. atroviride: 35.24.06.4, TRS7, TRS12, TRS14, TRS20, TRS25, TRS40, and TRS43. Then, 68 primers that had given the best results were further tested on 39 strains to assess the genetic diversity of T. atroviride strains collected in Poland. RAPD-PCR was performed as described by Olczak-Woltman et al. (2009). Reactions contained 1× Green PCR Buffer containing 2 mmol/L MgCl2, 0.2 mmol/L each dNTP, 1 μg bovine serum albumin, 0.5 U Taq polymerase (Fermentas, Vilnius, Lithuania), 1 μmol/L arbitrary primer, and 2 ng DNA template. PCR was carried out under conditions of initial incubation at 95 °C for 1 min, followed by 10 cycles of 95 °C for 5 s, 37 °C for 30 s, and 72 °C for 30 s; 35 cycles of 95 °C for 5 s, 37 °C for 30 s, and 72 °C for 60 s; and a final extension at 72 °C for 7 min. RAPD-PCR was carried out in a PTC-200 Thermal Cycler (Bio-Rad, Hercules, CA, USA). Amplified DNA fragments were separated by electrophoresis with 2 % agarose–0.5× Tris-borate-EDTA (TBE) gels containing ethidium bromide and visualized under UV light. Gel images were edited in CorelDRAW version 12 (Corel Corporation, Ottawa, Ontario, Canada). The binary matrices were constructed based on evaluation of clear RAPD amplicons. Amplicons were scored for presence (1) or absence (0) for each strain. Cluster analysis of the binary data was performed using the NTSYS-pc v.2.1 program (Exeter Software, Setauket, NY, USA). Similarity matrices were generated using Jaccard’s coefficients, and an unweighted pair-group method using arithmetic averages (UPGMA) was chosen to generate the dendrogram from RAPD similarity matrices. To determine how faithfully the dendrogram preserves pairwise distances between original unmolded data, the cophenetic correlation coefficient parameter (r) was calculated.
Cloning and sequencing
Polymorphic RAPD amplicons were excised from the gel, and DNA was purified using the QIAquick Gel Extraction Kit (Qiagen) according to the manufacturer’s protocol. Next, purified amplicons were reamplified under the conditions described for RAPD-PCR, resolved using a 1 % agarose–0.5× TBE gel, excised from the gel, and purified. Efficiently reamplified RAPD amplicons were cloned using a TOPO TA Cloning Kit (Invitrogen, Carlsbad, CA, USA). Recombinant plasmids were isolated, and two independent plasmids were used for sequencing of each amplicon (with the exception of Q01-420, for which only one clone was available) using universal M13 forward and reverse primers. The Sequencher 5.1 program (Gene Codes Corporation, Ann Arbor, MI, USA) was used to assemble single reads to obtain consensus sequences in FASTA format. The consensus sequences were deposited at the National Center for Biotechnology Information (NCBI), and GenBank accession numbers are summarized in Table 2 (RAPD-PCR amplicons) and Table 3 (CAPS amplicons).
Bioinformatics analysis
For phylogenetic reconstruction, sequences of the fragment of tef1 gene were assembled with ClustalX (Thompson et al. 1997) and manually edited in CLC Genomic Workbench 7.5 (CLCBio, Aarhus, Denmark), and final block alignment was prepared with a less stringent selection option using GBlocksServer (http://molevol.cmima.csic.es/castresana/Gblocks_server.html). The unconstrained GTR + I + G substitution model, estimated nucleotide frequencies, and substitution values were selected according to Samuels et al. (2006). Metropolis-coupled Markov chain Monte Carlo (MCMCMC) analysis was performed with two runs for one million generations with four chains, with the heating coefficient λ = 0.1 with MrBayes 3.2.2 × 64 (Ronquist et al. 2012).
RAPD amplicon sequences were compared using basic local alignment search tool (BLAST) (Altschul et al. 1997) with publicly available sequences deposited at both NCBI and the Genome Portal of the Joint Genome Institute (JGI, Grigoriev et al. 2012). At NCBI, blastn and blastx were used to explore the non-redundant protein sequence, nucleotide collection, and high-throughput genomic sequence databases. At JGI, sequences were aligned using blastn to the T. atroviride IMI206040 genome version 2 unmasked assemblies. Proteins containing amino acid sequences similar to those encoded by RAPD amplicons were collected and further analyzed using CLC Genomics Workbench 7.5 (CLCBio), and Pfam database search (Punta et al. 2012). RAPD sequences were scanned for sequence repeats using RepeatMasker (http://www.repeatmasker.org/) and mfold (Zuker 2003).
Cleaved amplified polymorphic sequence marker development
The consensus RAPD sequences were mapped on the T. atroviride IMI 206040 v. 2 genome sequence deposited at JGI (http://genome.jgi-psf.org/Triat2/Triat2.home.html). Several sets of primers were designed to amplify RAPD regions or extended RAPD regions based on genome sequence. A standard PCR was performed using Taq polymerase (Fermentas) according to the manufacturer’s instructions. PCR was carried out under conditions of initial incubation at 94 °C for 3 min, followed by 35 cycles of 94 °C for 30 s, optimized annealing temperature or gradient temperature for 30 s, and 72 °C for 2 min, followed by a final extension at 72 °C for 10 min. The amplified products for selected T. atroviride strains were analyzed on agarose gels, purified, and sequenced. Derived sequences were compared and searched for polymorphic restriction sites using Sequencher 5.1 (Gene Codes). PCR amplification was performed using selected primer pairs. PCR products were digested with corresponding restriction enzyme (BslI, DraI, and TaqI; all from Fermentas) according to the manufacturer’s instructions, and digestion products were separated on 1.5 % agarose–0.5× TBE gel. The respective primer pairs with optimized PCR conditions and corresponding restriction enzymes are shown in Table 3. To confirm specificity of the CAPS PCR to T. atroviride species, multiplex PCR was performed with DNA of 19 Trichoderma species mentioned above, CAPS primers, and additional primers 5.8S-R and LR6 (Vilgalys and Hester 1990). The pair of primers 5.8S-R and LR6 amplified a 1.45-kb fragment of the ITS region and served as a positive control for the PCR.
Results and discussion
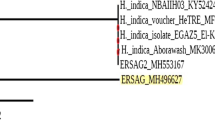
Molecular identification of strains based on sequences of ITS1, ITS2, and the fragment of tef1 gene confirmed that all of the strains used in this study belong to T. atroviride. Based on tef1 sequences, all strains were classified to clade A described by Dodd et al. (2003), except for TRS18 that did not fit any clade. TrichOKEY 2 analysis revealed one new ITS1 sequence variant that possesses a single nucleotide change in comparison to the IMI206040 sequence. Six strains, TRS15, TRS24, TRS28, TRS30, TRS34, and TRS45, were characterized by the new ITS1 variant. Seven variants of tef1 sequence which differed at least by a single nucleotide were found within the studied T. atroviride strains. Bayesian phylogeny analysis of tef1 sequences allowed strain clustering (Fig. 1). TRS18 strain was found to be the most distant one. Four groups of strains were distinguished, with the two biggest groups differing within by a single nucleotide change only. Strains TRS43 and CBS 693.94 did not fall into any groups (Fig. 1).
In addition, RAPD markers were used for further evaluation of genetic diversity of T. atroviride strains. A total of 520 primers were tested on eight T. atroviride strains, including 35.24.06.4, TRS7, TRS12, TRS14, TRS20, TRS25, TRS40, and TRS43, in order to select RAPD primers useful for analysis. Amplicons were obtained for 482 primers, with an average of eight amplicons of different sizes identified for each primer. Based on electrophoretic evaluation of amplification pattern quality, 68 primers that produced polymorphic amplicons were chosen for the assessment of genetic diversity of the T. atroviride strains. RAPD PCR was performed with selected primers on 38 strains of T. atroviride collected at different locations in Poland (Table 1). As a control, T. atroviride reference strains CBS 693.94 (Dodd et al. 2003) and IMI 206040 (Kubicek et al. 2011) were used. High-quality amplification products were obtained for 55 of the 68 RAPD primers. Only two primers (OPK11 and OPX09) did not show any polymorphisms among tested strains. The greatest number of polymorphic amplicons distinguishing groups of strains was obtained with primer OPA10. Amplification profiles of three primers, OPI18, OPQ01, and OPZ04, showed that the T. atroviride strains, with the exception of strain TRS18, can be classified into two main groups. The amplification profile generated using OPZ04 is shown in Fig. 2.
RAPD amplicons were scored for presence and absence, and 391 amplicons differentiating tested strains were used to construct a binary matrix, and then a dendrogram illustrating genetic diversity (Fig. 3). The cophenetic correlation coefficient (r) for the dendrogram was 0.979, indicating a good fit between the unmolded data and the dendrogram. Tested T. atroviride strains, of which all except TRS18 belonged to clade A, were classified into two main groups. The first group was represented by the reference strain IMI206040, as well as 28 of the tested strains, and corresponds to the two biggest groups of strains including TRS43, distinguished based on tef1 sequence. The second group contained reference strain CBS 693.94 and the following eight tested strains: TRS2, TRS6, TRS10, TRS16, TRS26, TRS40, TRS42, and TRS44. This group corresponds to CBS 693.94 and remaining two small groups of strains distinguished based on tef1. The observed variation among strains within the two major groups was rather limited; however, some subgroups could be distinguished (Fig. 3). T. atroviride strain TRS18 was clearly distinct from both groups of strains, based on both tef1 and RAPD analysis, and also did not fit any previously described clades. TRS18 tef1 intron 4 sequence BLAST similarity search with non-redundant nucleotide sequence database at NCBI revealed 95 % identity with any sequence deposited in GenBank. This suggests that TRS18 strain is characterized by a unique and distinct genotype that could represent new clade of T. atroviride.
Dendrogram generated using Jaccard’s coefficient and the UPGMA clustering method. Binary matrices for T. atroviride strains were constructed based on evaluation of RAPD amplicons generated using 55 primers. Cluster analysis of the binary data was performed using NTSYS-pc 2.1 software. Similarity matrices were generated using Jaccard’s coefficient and an unweighted pair-group method using arithmetic averages (UPGMA) was used to generate the dendrogram
Twenty polymorphic RAPD markers were chosen for cloning, and 13 RAPD-PCR products were successfully cloned and sequenced (Table 2). Resulting sequences were aligned with the T. atroviride IMI206040 genome at the JGI portal. Twelve RAPD sequences were identical to corresponding T. atroviride IMI206040 sequences (e-value = 0). Sequence R08-1283 showed some variation, but the e-value was very low, indicating strong similarity to the T. atroviride genome (Table 2). Seven sequences showed strong similarity to hypothetical proteins of T. atroviride deposited in the NCBI database, and Pfam protein domains were clearly identified for three of these (Table 2). The H08-917 sequence showed a high degree of similarity to tetratricopeptide repeat (TPR)-containing proteins. TPR motifs are short amino acid repeats that occur in many proteins and not only are responsible for interactions between proteins but also facilitate protein complex formation and protein transport within cells. TPR-containing proteins play a role in protein assembly and therefore have a significant impact on the cell cycle and a plethora of cellular processes (D’Andrea and Regan 2003). Interestingly, the H08-917 sequence also possessed short and degenerate DNA repeats that were identified by RepeatMasker and mfold. The R08-1283 and T07-429 sequences were recognized as proteins with ankyrin (ANK) repeats, which play a role in protein-protein interactions (Rubtsov and Lopina 2000). Moreover, the X16-1030 sequence was similar to the NIPA membrane protein, which functions as a magnesium transporter (Goytain et al. 2007, 2008).
Using the identified RAPD marker sequences and the T. atroviride genome sequence, pairs of specific primers were designed, followed by optimization of PCR conditions and testing of the primers on a set of Trichoderma strains belonging to different species, including T. atroviride (Table 3). Unique PCR amplification products for three pairs of primers, Q01-4, X18-35, and Z04-2, were obtained for T. atroviride strains and not for 16 other Trichoderma species/clades (Fig. 4). In the case of X18-35, a very faint PCR amplicon was obtained for T. gamsii. These suggest that the amplicons Q01-4, X18-35, and Z04-2 are specific to T. atroviride. We propose that pairs of primers Q01-4, Z18-35, and Z04-2 could be used for T. atroviride detection.
For selected T. atroviride strains representing two main groups of strains, amplicons Q01-4, X18-35, and Z04-2 were sequenced, and sequences were deposited in GenBank (Table 3). These sequences were compared, and polymorphic restriction sites were identified. Based on identified sequence differences (single nucleotide polymorphisms and insertions/deletions), restriction enzymes were selected to digest PCR amplicons and develop CAPS markers useful for T. atroviride identification (Table 3). In the case of CAPS marker Q01-4 (Q01-4FR amplification product digested with BslI), a two-band pattern was obtained for 29 strains of T. atroviride belonging to group 1, and a single band corresponding to the non-restricted Q01-4 F-4R amplicon was obtained for group 2 strains and TRS18 (Fig. 5). This CAPS marker allows the distinction of strains belonging to T. atroviride group 1. For Z04-2 amplification products, DraI digestion resulted in banding patterns reflecting the RAPD-based grouping. However, the products for strain TRS6 and TRS18 were digested and these strains were not classified to the group 1 as in the RAPD study (Fig. 5). We propose that analysis of both CAPS markers can be used to extend ITS/tef1-based T. atroviride identification and can assist in classification of strains within clade A into one of the two main groups of T. atroviride: group 1 (IMI206040-like), characterized by both Q01-4 and Z04-2 amplicons digested, and group 2 (CBS 693.60-like), characterized by both amplicons non-digested. In the case where Q01-4 amplicon is not digested and Z04-2 is digested, additional markers have to be developed to distinguish TRS6-like from TRS18-like strains. Developed CAPS markers have to be further tested on a larger number of strains that will confirm their usefulness in T. atroviride genetic studies. The variable RAPD loci found in this study and characterized at the sequence level may be useful in the development of more complex assays, such as multiplex PCR, real-time PCR, or next-generation sequencing (NGS)-based assays, that will be valuable in identification and monitoring of T. atroviride strains, as well as in genetic diversity studies including metagenomics approaches and further T. atroviride pan-genome understanding.
Q01-4 and Z04-2 CAPS marker profiles generated by PCR amplification with primers Q01_4F and Q01_4R and digestion of PCR products with BslI (upper part) and primers Z04_2F and Z04_2R and digestion of PCR products with DraI. IMI206040 and CBS 693.94 are the reference strains of T. atroviride. CAPS, cleaved amplified polymorphic sequences. M, 100-bp Plus DNA ladder
References
Abbasi PA, Miller SA, Meulia T, Hoitink HAJ, Kim J-M (1999) Precise detection and tracing of Trichoderma hamatum 382 in compost-amended potting mixes by using molecular markers. Appl Environ Microbiol 65:5421–5426
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Atanasova L, Druzhinina IS (2010) Global nutrient profiling by Phenotype MicroArrays: a tool complementing genomic and proteomic studies in conidial fungi. J Zhejiang Univ Sci B (Biomed and Biotechnol) 11:151–168
Atanasova L, Druzhinina IS, Jaklitsch WM (2013) Two hundred Trichoderma species recognized on the basis of molecular phylogeny. In: Mukherjee PK, Horwitz BA, Singh US, Mukherjee M, Schmoll M (eds) Trichoderma: biology and applications. CABI, Wallingford, pp 10–42
Benítez T, Rincón AM, Limón MC, Codón AC (2004) Biocontrol mechanisms of Trichoderma strains. Int Microbiol 7:249–260
Błaszczyk L, Popiel D, Chełkowski J, Koczyk G, Samuels GJ, Sobieralski K, Siwulski M (2011) Species diversity of Trichoderma in Poland. J Appl Genet 52:233–243
Bowen K, Franicevic SC, Crowhurst RN, Templeton MD, Stewart A (1996) Differentiation of a specific Trichoderma biological control agent by restriction fragment length polymorphism (RFLP) analysis. N Z J Crop Hortic 24:207–217
Buhariwalla HK, Srilakshmi P, Kannan S, Kanchi RS, Chandra S, Satyaprasad K, Waliyar F, Thakur RP, Crouch JH (2005) AFLP analysis of Trichoderma spp. from India compared with sequence and morphological–based diagnostics. J Phytopathol 153:389–400
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous Ascomycetes. Mycologia 91:553–556
Chakraborty BN, Chakraborty U, Sunar K, Dey PL (2011) RAPD profile and rDNA sequence analysis of Talaromyces flavus and Trichoderma species. Indian J Biotechnol 10:487–495
Chen X, Romaine CP, Tan Q, Schlagnhaufer B, Ospina-Giraldo MD, Royse DJ, Huff DR (1999) PCR-based genotyping of epidemic and preepidemic Trichoderma isolates associated with green mold of Agaricus bisporus. Appl Environ Microbiol 65:2674–2678
Chet I, Inbar J (1994) Biological control of fungal pathogens. Appl Biochem Biotechnol 48:37–43
Cooke DEL, Duncan JM (1997) Phylogenetic analysis of Phytophthora species based on ITS1 and ITS2 sequences of the ribosomal RNA gene repeat. Mycol Res 101:667–677
Cordier C, Edel-Hermann V, Martin-Laurent F, Blal B, Steinberg C, Alabouvette C (2007) SCAR-based real time PCR to identify a biocontrol strain (T1) of Trichoderma atroviride and study its population dynamics in soils. J Microbiol Methods 68:60–68
D’Andrea LD, Regan L (2003) TPR proteins: the versatile helix. Trends Biochem Sci 28:655–662
Devi TP, Kamil D, Prabhakaran N (2011) Development of genus specific rDNA based marker for detection of Trichoderma species. J Mycol Plant Pathol 41:600–604
Dodd SL, Lieckfeldt E, Samuels GJ (2003) Hypocrea atroviridis sp. nov., the teleomorph of Trichoderma atroviride. Mycologia 95:27–40
Dodd SL, Hill RA, Stewart A (2004a) Monitoring the survival and spread of the biocontrol fungus Trichoderma atroviride (C65) on kiwifruit using a molecular marker. Australas Plant Pathol 33:189–196
Dodd SL, Hill RA, Stewart A (2004b) A duplex-PCR bioassay to detect a Trichoderma virens biocontrol isolate in non-sterile soil. Soil Biol Biochem 36:1955–1965
Druzhinina I, Kubicek CP (2005) Species concepts and biodiversity in Trichoderma and Hypocrea: from aggregate species to species clusters? J Zhejiang Univ Sci B 6:100–112
Druzhinina IS, Kopchinskiy AG, Komoń M, Bissett J, Szakacs G, Kubicek CP (2005) An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet Biol 42:813–828
Druzhinina IS, Seidl-Seiboth V, Herrera-Estrella A, Horwitz BA, Kenerley CM, Monte E, Mukherjee PK, Zeilinger S, Grigoriev IV, Kubicek CP (2011) Trichoderma: the genomics of opportunistic success. Nat Rev Microbiol 9:749–759
Feng XM, Holmberg A-IJ, Sundh I, Ricard T, Melin P (2011) Specific SCAR markers and multiplex real-time PCR for quantification of two Trichoderma biocontrol strains in environmental samples. BioControl 56:903–913
Friedl MA, Druzhinina IS (2012) Taxon-specific metagenomics of Trichoderma reveals a narrow community of opportunistic species that regulate each other's development. Microbiology 158:69–83
Gal-Hemed I, Atanasova L, Komon-Zelazowska M, Druzhinina IS, Viterbo A, Yarden O (2011) Marine isolates of Trichoderma spp. as potential halotolerant agents of biological control for arid-zone agriculture. Appl Environ Microbiol 77:5100–5109
Goytain A, Hines RM, El-Husseini A, Quamme GA (2007) NIPA1(SPG6), the basis for autosomal dominant form of hereditary spastic paraplegia, encodes a functional Mg2+ transporter. J Biol Chem 282:8060–8068
Goytain A, Hines RM, Quamme GA (2008) Functional characterization of NIPA2, a selective Mg2+ transporter. Am J Physiol Cell Physiol 295:C944–C953
Grigoriev IV, Nordberg H, Shabalov I, Aerts A, Cantor M, Goodstein D, Kuo A, Minovitsky S, Nikitin R, Ohm RA, Otillar R, Poliakov A, Ratnere I, Riley R, Smirnova T, Rokhsar D, Dubchak I (2012) The genome portal of the department of energy joint genome institute. Nucleic Acids Res 40:D26–D32
Hagn A, Wallisch S, Radl V, Charles Munch J, Schloter M (2007) A new cultivation independent approach to detect and monitor common Trichoderma species in soils. J Microbiol Methods 69:86–92
Harman GE, Howell CR, Viterbo A, Chet I, Lorito M (2004) Trichoderma species — opportunistic, avirulent plant symbionts. Nat Rev Microbiol 2:43–56
Hermosa MR, Grondona I, Iturriaga EA, Diaz-Minguez JM, Castro C, Monte E, Garcia-Acha I (2000) Molecular characterization and identification of biocontrol isolates of Trichoderma spp. Appl Environ Microbiol 66:1890–1898
Hermosa MR, Grondona I, Díaz-Mínguez JM, Iturriaga EA, Monte E (2001) Development of a strain-specific SCAR marker for the detection of Trichoderma atroviride 11, a biological control agent against soilborne fungal plant pathogens. Curr Genet 38:343–350
Hoyos-Carvajal L, Orduz S, Bissett J (2009) Genetic and metabolic biodiversity of Trichoderma from Colombia and adjacent neotropic regions. Fungal Genet Biol 46:615–631
Jaklitsch WM, Komon M, Kubicek CP, Druzhinina IS (2005) Hypocrea voglmayrii sp. nov. from the Austrian Alps represents a new phylogenetic clade in Hypocrea/Trichoderma. Mycologia 97:1365–1378
Jaklitsch WM, Samuels GJ, Dodd SL, Lu B-S, Druzhinina IS (2006) Hypocrea rufa/Trichoderma viride: a reassessment, and description of five closely related species with and without warted conidia. Stud Mycol 56:135–177
Kindermann J, El-Ayouti Y, Samuels GJ, Kubicek CP (1998) Phylogeny of the genus Trichoderma based on sequence analysis of the internal transcribed spacer region 1 of the rDNA cluster. Fungal Genet Biol 24:298–309
Kopchinskiy A, Komoń M, Kubicek CP, Druzhinina IS (2005) TrichoBLAST: a multilocus database for Trichoderma and Hypocrea identifications. Mycol Res 109:658–660
Kredics L, Kocsubé S, Nagy L, Komoń-Żelazowska M, Manczinger L, Sajben E, Nagy A, Vágvölgyi C, Kubicek CP, Druzhinina IS, Hatvani L (2009) Molecular identification of Trichoderma species associated with Pleurotus ostreatus and natural substrates of the oyster mushroom. FEMS Microbiol Lett 300:58–67
Kredics L, Hatvani L, Naeimi S, Körmöczi P, Manczinger L, Vágvölgyi C, Druzhinina I (2014) Biodiversity of the genus Hypocrea/Trichoderma in different habitats. In: Gupta VK, Schmoll M, Herrera-Estrella A, Upadhyay RS, Druzhinina I, Tuohy MG (eds) Biotechnology and biology of Trichoderma. Elsevier, London, pp 3–24
Kubicek CP, Bissett J, Druzhinina IS, Kullnig-Gradinger C, Szakacs G (2003) Genetic and metabolic diversity of Trichoderma: a case study on South-East Asian isolates. Fungal Genet Biol 38:310–319
Kubicek CP, Herrera-Estrella A, Seidl-Seiboth V, Martinez DA, Druzhinina IS, Thon M, Zeilinger S, Casas-Flores S, Horwitz BA, Mukherjee PK, Mukherjee M, Kredics L, Alcaraz LD, Aerts A, Antal Z, Atanasova L, Cervantes-Badillo MG, Challacombe J, Chertkov O, McCluskey K, Coulpier F, Deshpande N, von Döhren H, Ebbole DJ, Esquivel-Naranjo EU, Fekete E, Flipphi M, Glaser F, Gómez-Rodríguez EY, Gruber S, Han C, Henrissat B, Hermosa R, Hernández-Oñate M, Karaffa L, Kosti I, Le Crom S, Lindquist E, Lucas S, Lübeck M, Lübeck PS, Margeot A, Metz B, Misra M, Nevalainen H, Omann M, Packer N, Perrone G, Uresti-Rivera EE, Salamov A, Schmoll M, Seiboth B, Shapiro H, Sukno S, Tamayo-Ramos JA, Tisch D, Wiest A, Wilkinson HH, Zhang M, Coutinho PM, Kenerley CM, Monte E, Baker SE, Grigoriev IV (2011) Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. Genome Biol 12:R40
Kullnig CM, Krupica T, Woo SL, Mach RL, Rey M, Benítez T, Lorito M, Kubicek CP (2001) Confusion abounds over identities of Trichoderma biocontrol isolates. Mycol Res 105:769–772
Longa CMO, Savazzini F, Tosi S, Elad Y, Pertot I (2009) Evaluating the survival and environmental fate of the biocontrol agent Trichoderma atroviride SC1 in vineyards in northern Italy. J Appl Microbiol 106:1549–1557
Lorito M, Woo SL, Harman GE, Monte E (2010) Translational research on Trichoderma: from ‘omics to the field. Annu Rev Phytopathol 48:1–23
Miyazaki K, Tsuchiya Y, Okuda T (2009) Specific PCR assays for the detection of Trichoderma harzianum causing green mold disease during mushroom cultivation. Mycoscience 50:94–99
Mulaw TB, Kubicek CP, Druzhinina IS (2010) The rhizosphere of Coffea arabica in its native highland forests of Ethiopia provides a niche for a distinguished diversity of Trichoderma. Diversity 2:527–549
Naef A, Senatore M, Défago G (2006) A microsatellite based method for quantification of fungi in decomposing plant material elucidates the role of Fusarium graminearum DON production in the saprophytic competition with Trichoderma atroviride in maize tissue microcosms. FEMS Microbiol Ecol 55:211–220
Naeimi S, Kocsubé S, Antal Z, Okhovvat SM, Javan-Nikkhah M, Vágvölgyi C, Kredics L (2011) Strain-specific SCAR markers for the detection of Trichoderma harzianum AS12-2, a biological control agent against Rhizoctonia solani, the causal agent of rice sheath blight. Acta Biol Hung 62:73–84
Olczak-Woltman H, Bartoszewski G, Madry W, Niemirowicz-Szczytt K (2009) Inheritance of resistance to angular leaf spot (Pseudomonas syringae pv. lachrymans) in cucumber and identification of molecular markers linked to resistance. Plant Pathol 58:145–151
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer ELL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40:D290–D301
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Rubio MB, Hermosa MR, Keck E, Monte E (2005) Specific PCR assays for the detection and quantification of DNA from the biocontrol strain Trichoderma harzianum 2413 in soil. Microb Ecol 49:25–33
Rubtsov AM, Lopina OD (2000) Ankyrins. FEBS Lett 482:1–5
Samuels GJ, Dodd SL, Lu BS, Petrini O, Schroers HJ, Druzhinina IS (2006) The Trichoderma koningii aggregate species. Stud Mycol 56:67–133
Savazzini F, Longa CMO, Pertot I, Gessler C (2008) Real-time PCR for detection and quantification of the biocontrol agent Trichoderma atroviride strain SC1 in soil. J Microbiol Methods 73:185–194
Schuster A, Schmoll M (2010) Biology and biotechnology of Trichoderma. Appl Microbiol Biotechnol 87:787–799
Stewart A, Hill R (2014) Applications of Trichoderma in plant growth promotion. In: Gupta VK, Schmoll M, Herrera-Estrella A, Upadhyay RS, Druzhinina I, Tuohy MG (eds) Biotechnology and biology of Trichoderma. Elsevier, London, pp 415–428
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Woo SL, Scala F, Ruocco M, Lorito M (2006) The molecular biology of the interactions between Trichoderma spp., phytopathogenic fungi, and plants. Phytopathology 96:181–185
Zimand G, Valinsky L, Elad Y, Chet I, Manulis S (1994) Use of the RAPD procedure for the identification of Trichoderma strains. Mycol Res 98:531–534
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Acknowledgments
This study was conducted as a part of the project “Polish Trichoderma strains in plant protection and organic waste management” under Priority 1.3.1, subject area “Bio,” co-financed by The European Union through the European Regional Development Fund within the Innovative Economy Operational Program, 2007-2013, project No UDA-POIG.01.03.01-00-129/09-08. The authors declare that there is no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Dominik Skoneczny and Michał Oskiera contributed equally to this work.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Skoneczny, D., Oskiera, M., Szczech, M. et al. Genetic diversity of Trichoderma atroviride strains collected in Poland and identification of loci useful in detection of within-species diversity. Folia Microbiol 60, 297–307 (2015). https://doi.org/10.1007/s12223-015-0385-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-015-0385-z