Abstract
Herpesvirus infections, such as those induced by human cytomegalovirus (HCMV), induce specific DNA damages. DNA damages can lead to cell mutation, death, apoptosis and immune system activation. Various types of DNA damage are repaired through multiple repair pathways, such as base excision, nucleotide excision, homologous recombination and nonhomologous end joining. Changes in the activity of DNA repair proteins during viral infection can cause disturbances in the DNA repair system and change its mechanisms. This report reviews results from studies, assaying a DNA repair system in HCMV infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA damage normally triggers activities of the cellular DNA repair machinery. There are six known systems of DNA repair: pathway of direct reversion of damage, base excision repair (BER), nucleotide-excision repair (NER), mismatch repair (MMR), homologous recombination (HR) and nonhomologous DNA end joining (NHEJ). Unrepaired DNA damage can lead to mutation, development of various diseases or cell death (Bartek et al. 2004; Jackson and Bartek 2009; Hoeijmakers 2009; Polo and Jackson 2011).
The viruses that induce cellular DNA damage include members of the herpesvirus group, adenovirus, mumps-virus, measles-virus, rubella-virus, poliovirus and papilloma-virus (Fortunato et al. 2000).
Human cytomegalovirus (HCMV) is a member of the betaherpesvirus subfamily and encodes more than 200 viral proteins (Chee et al. 1990; Salsman et al. 2008; Stern-Ginossar et al. 2012).
The viral infection process is very complex and driven by many mechanisms. It is known that HCMV can act as mutagen inducing various DNA damage (AbuBakar et al. 1988; Shen et al. 1997).
Herpesvirus infection is associated with apoptosis processes, immune activation, cell cycle control and DNA damage response (Chaurushiya and Weitzman 2009; Miller-Kittler and Sparer 2009; Weitzman et al. 2010).
It appears from literature that human cytomegalovirus enhances DNA repair capacity in host cells, without producing detectable lesions in cellular DNA or inhibiting DNA synthesis (Nishiyama and Rapp 1981).
Many studies have analysed cell repair capabilities, following viral infection. Those experimental studies focused on repairs of exogenously introduced damage in cellular DNA in the context of single viral protein expression (Becker et al. 1998; Prost et al. 1998; Groisman et al. 1999; Jia et al. 1999; Chipitsyna et al. 2004; Mathonnet et al. 2004; Liang et al. 2006; Sun et al. 2006; Trojanek et al. 2006; Durkin et al. 2008; Gruhne et al. 2009; Baydoun et al. 2011) and on effects of complete infection (Deng et al. 1992a; Philpott and Buehring 1999; Ranneberg-Nilsen et al. 2006; Duong et al. 2010; reviewed in Lilley et al. 2010; Pal et al. 2010; Kulkarni and Fortunato 2011).
Only several of those studies report some evidence of increased cell repair capacity after infection or viral protein overexpression (Chipitsyna et al. 2004; Kulkarni and Fortunato 2011; Ranneberg-Nilsen et al. 2006; Baydoun et al. 2011).
The control aspects of the five DNA repair mechanisms in virus-infected cells have not been well characterized. In this review, considerations and results of the experiments are presented, supporting the thesis on the important role of DNA repair systems in HCMV- infected cell.
HCMV-related changes in base excision repair
The base excision repair (BER) pathway corrects most endogenous base lesions, including alkylation, oxidation and deamination, apurinic/apyrimidinic (AP) sites as well as single-strand breaks.
A damaged base is recognized by a specific DNA glycosylase, which cleaves the bond between the base and sugar, creating an abasic site (AP), which is a mutagenic and cytotoxic intermediate. The resulting AP site is further processed by the AP lyase activity associated with bifunctional DNA glycosylases and AP endonucleases cleaving 3′or 5′at the AP site, respectively. The processed AP sites are further repaired by two sub pathways of BER: short-patch BER, a mechanism, where only 1 nucleotide is replaced, or long-patch BER, where 2–13 nucleotides are replaced (Fig. 1) (Wilson and Bohr 2007; Almeida and Sobol 2007).
The base excision repair (BER) system. BER is initiated by a specific DNA glycosylases, which recognize and remove damaged or incorrect bases, creating AP sites. The AP endonucleases cleave an AP site to yield a 3′ hydroxyl adjacent to a 5′ deoxyribosephosphate (dRP). During short-patch BER, 5′dRP is displaced by DNA polymerase β (Polβ), which inserts a single nucleotide. Polβ is also involved in long-patch BER inserting the first nucleotide at reduced AP sites. The ligation step is performed by DNA ligases I and III. Ligase I interacts with Polβ and participates mainly in long-patch BER. DNA ligase III interacts with X-ray repair cross-complementing protein 1 (XRCC1) and Polβ and is involved only in short-patch BER
If not repaired by AP endonucleases in dividing cells, AP site may have dramatic consequences, leading to single and, subsequently, double DNA strand breaks, which are lethal to the cell (Evans et al. 2000).
It is known that HCMV infection is genotoxic to host cells (AbuBakar et al. 1988; Fortunato and Spector 2003; Nystad et al. 2008). Viral infection to cell is associated with production of reactive oxygen species (ROS).
Speir et al. (1998) reported that reactive oxygen metabolites were created soon after uptake of HCMV-infected smooth muscle cells, indicating a potentially genotoxic effect of viral infection.
Literature data demonstrate that modulation of BER activities may play some role in HCMV infection. Ranneberg-Nilsen et al. (2006) analysed the capability of HCMV-infected human embryonic lung fibroblasts to carry out base excision repair system. These researchers have demonstrated that some of the initial steps in the base excision repair machinery are downregulated during human cytomegalovirus replication in human fibroblasts.
Ranneberg-Nilsen et al. (2006) have investigated viral infection effects on the initial steps of BER, which are possessed by DNA glycosylases, cleaving the N-glycosylic bond and subsequent incision of the phosphodiester bond at the abasic site by an AP endonuclease activity. It was shown that the DNA glycosylase activity for removal of oxidized and alkylated bases declined in HCMV-infected cells.
Moreover, the AP endonuclease activity was enhanced in HCMV-infected cells. However, the expression level of the major human AP endonuclease (APE1) was not altered, indicating that either HCMV had its own AP endonuclease activity or that APE1 could have been subjected to post-translation modifications that could modulate its activity.
Furthermore, they speculate that reduced BER system activity in HCMV-infected cells may contribute to increased genomic variation of the virus.
An experimental analysis has provided evidence that increased excision rates of uracil and enhanced AP endonuclease activity in infected cells may form gaps in viral DNA, required for efficient viral DNA synthesis at late stages of viral replication (Ranneberg-Nilsen et al. 2006).
In conclusion, Ranneberg-Nilsen et al. (2006) demonstrated an upregulation of base excision repair system activities that could be involved in HCMV replication, whereas other BER activities, such as glycosylase activities, initiating repairs of alkylated and oxidized bases, were downregulated in viral-infected cells.
Future studies will be aimed at clarifying the functions of BER during HCMV infection.
HCMV-related changes in nucleotide excision repair
The nucleotide excision repair system removes short DNA, damaged base-containing oligonucleotides (Hanawalt 2002). NER recognizes bulky lesions, caused by carcinogenic compounds, and covalent linkages between adjacent pyrimidines, resulting from exposure to UV. NER is a multistep process, involving numerous proteins, and is classified into global genome repair (GG–NER) that occurs in the genome, and transcription-coupled repair (TCR), which removes lesions in the transcribed strand of active genes (Fig. 2).
The nucleotide excision repair. a Global genomic repair (GGR), xeroderma pigmentosum, complementation group C (XPC), UV excision repair protein RAD23 homolog B (HR23B), replication protein A (RPA), xeroderma pigmentosum group A (XPA) or DNA damage-binding protein 1 and 2 (DDB1–DDB2) complex recognition DNA damage. DNA repair is performed by the transcription factor TFIIH, and excision of the lesion by xeroderma pigmentosum, complementation group G and F (XPG and XPF), excision repair cross-complementing 1 (ERCC1). Finally, resynthesis occurs by polymerase (Polδ or Polϵ) and ligation by DNA ligase I. b Transcription-coupled repair (TCR). First step, RNA polII (RNAPII) is blocked. This leads to assembly of Cockayne syndrome group A/B proteins (CSA, CSB) and/or the transcription elongation factor TFIIS at the site of the lesion, by which RNAPII is removed from the DNA or displaced from the lesion, making it accessible to the exonucleases XPF. Second step, XPG cleaving the lesion-containing DNA strand. Third step, resynthesis again occurs by Polδ or Polϵ and ligation by DNA ligase I
Because NER is involved in removing a substantial number of DNA damages, which can contribute to the genome instability, it is reasonable to check whether variability in repair efficiency may be associated with HCMV infection.
What is important, many reports introduce the role of NER systems in the development of viral infection. Those analyses targeted the expression of single viral proteins in evaluation of UV-induced damage. The results suggest that the expression of the hepatitis B X protein (HBX) in different cell types (Becker et al. 1998; Groisman et al. 1999; Jia et al. 1999; Mathonnet et al. 2004) or the expression of Epstein Barr virus proteins EBNA3C or LMP1 in transfected cells (Gruhne et al. 2009), decreased repair efficiency of UV-induced damage in transfected cells.
In the literature, recent studies are described, examining interactions with the NER machinery and/or with HCMV infection’s influence on cellular damage repair. These reports demonstrate that NER may play an important role during HCMV entry to cells (O’Dowd et al. 2012). It appears from the reports that differential association of certain cellular repair proteins, e.g. XP proteins with HCMV, may have far-reaching implications in the pathogenesis of viral infection (O’Dowd et al. 2012).
An in vitro analysis of infected fibroblasts revealed selective removal of UV lesions from the viral, and not cellular, DNA within infected cells (O’Dowd et al. 2012).
O’Dowd et al. exposed infected fibroblasts to ultraviolet (UV) radiation to inflict DNA damage. The ultraviolet radiation phenomenon is known to fuse adjacent thymine bases together into dimers, which are then cut out and replaced through NER system. Experimental analysis by comet assays technique showed that repair was initiated but not completed in the infected cells. The cyclobutane pyrimidine dimers (CPDs) were localized by immunofluorescence. CPDs were significantly reduced in viral DNA but were unchanged in the infected host DNA after a day of repair. These CPDs were efficiently repaired from viral DNA but not from the host cellular DNA suggesting that host genomes are unable to repair DNA damage (O’Dowd et al. 2012).
Recent studies demonstrate that a nucleotide excision repair-associated factor, DNA damage binding protein 2 (DDB2), is required for efficient HCMV DNA replication (Xiaofei et al. 2014). DDB2 is a component of a Cul4A-Ub ligase complex that participates in nucleotide excision repair. The DDB1-CUL4ADDB2 complex is a cullin-RING (i.e. E3) Ub-ligase that targets histone H2A at UV-damaged DNA sites (Xiaofei and Kowalik 2014).
In conclusion, little is provided in the literature about the role of nucleotide excision repair mechanisms during human cytomegalovirus infection. Future HCMV research should reveal more interesting, NER-modulating viral effectors.
HCMV-related changes in mismatch repair
Mismatch repair removes mispaired bases, resulting from replication errors, recombination between imperfectly matched sequences and deamination of 5-methyl-cytosine. Mismatch repair have escaped the proofreading function of the DNA polymerase and insertion/deletion (IDLs). The main MMR pathway is initiated by mismatch recognition by the heterodimer, consisting of MSH2 and MSH6 proteins (also called MutSα) (Fig. 3). MutSα is responsible for the recognition of base mismatches and insertion/deletion (IDLs) in mono to tetranucleotide repeats. This complex, i.e. MutSα, is able to recognize the majority of base–base mismatches and short IDLs (Hsieh and Yamane 2008). MMR is a highly conserved repair pathway that functions in improving replication fidelity by correcting replication-associated base–base and insertion/deletion mispairs. The MMR mechanism plays an important role in repair of oxidative damage by some mechanisms that are not well understood (Skinner and Turker 2005). MMR is essential for the maintenance of genome stability (Karran 1996; Ben Yehuda et al. 2000; Neri et al. 2005; Krichevsky et al. 2004).
The mismatch repair (MMR). Recognition of DNA lesions occurs by heterodimer hMSH2–hMSH6 (MutSα). Upon binding to the mismatch, MutSα associates with heterodimeric complex hMLH1-hPMS2 (MutLα). After formation of a complex composed of MutSα and MutLα, excision is performed by ExoI and repair synthesis by Polβ
To date, no studies have addressed the association between alterations in the MMR system during human cytomegalovirus infection. Changes in the MMR system during HCMV infection are not evident and further investigations are necessary, concerning its efficiency in infected organisms.
HCMV-related changes in double-strand break repair
Out of all the DNA damages, double-strand breaks (DSBs) are most mortal to cell. If not repaired, they cause losses of chromosomes and cell death. DSB accumulation destabilizes the genome and rearranges it, leading to transcription downregulation and development of various diseases (Jackson 2002).
HCMV can inflict site-specific chromosomal damage in a form of DSBs. Disruption of DSB repair system often leads to more intensive viral infection (Fortunato et al. 2000; Fortunato and Spector 2003).
The DSB repair system functions via two mechanisms: homologous recombination (HR) and nonhomologous DNA end joining (NHEJ) (Figs. 4, and 5) (Jackson 2002; Helleday 2003).
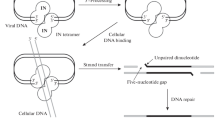
The homologous recombination (HR) system. HR starts with nucleolytic resection of the double strand break in the 5′ → 3′ direction by the MRE11–Rad50–NBS1 (MRN) complex, forming a 3′ single-stranded DNA fragment to which RAD52 homolog (S. cerevisiae) (Rad52) binds. Rad52 interacts with Rad51 homolog (RecA homolog, E. coli) (S. cerevisiae) (Rad51), provoking a DNA strand exchange with the undamaged, homologous DNA molecule. After DNA synthesis, ligation and branch migration, the resulting structure is resolved
The non-homologous end joining (NHEJ). The first step in NHEJ is the binding of a heterodimeric complex consisting of the proteins Ku70 and Ku80. Ku heterodimer associates with the catalytic subunit of DNA-dependent protein kinase (DNA–PKcs), forming the DNA–PK holoenzyme. One of the targets of DNA–PKcs is X-ray repair cross-complementing protein 4 (XRCC4). XRCC4 forms a stable complex with DNA ligase IV. DNA–PK activates XRCC4–ligase IV, which links the broken DNA ends together. Before re-ligation by XRCC4–ligase IV, the DNA ends are processed by the MRE11–Rad50–NBS1 complex, presumably involving Artemis
The nonhomologous end-joining pathway simply fuses two broken ends with little or no regard for sequence homology. It is known that NHEJ is involved in the immunological system maturation, cell–cell and cell–matrix adhesion (Tonegawa 1983).
The activities of NHEJ factors, such as Ku70/80, Artemis and Cernunnos protein, are very important for the immune system development (Weterings and van Gent 2004; Moshous et al. 2001; Buck et al. 2006)
Defects of NHEJ, observed during B and T lymphocyte receptors maturation, may be associated with immunodeficiency. The defective immune system is correlated with easy viral infection and HCMV diseases (Gamadia et al. 2003).
It appears from literature that a single viral protein can modulate homologous recombination repair (Nakai-Murakami et al. 2007; Trojanek et al. 2006; Kulkarni and Fortunato 2011).
Rad51 homolog (RecA homolog, Escherichia coli) (Saccharomyces cerevisiae) is involved in the homologous recombination and repair of double-strand breaks in DNA and DNA cross-links and in the maintenance of chromosome stability.
Experimental studies revealed Rad51 levels to be significantly increased in HCMV-infected human foreskin fibroblasts (HFFs) (Luo et al. 2007).
Literature data suggest that Rad51 levels do not generally increase in normal cells (Haaf et al. 1995). Raderschall et al. (2002) showed that increased RAD51 levels in tumour cells were found to be associated with unplanned HR and genetic instability. Therefore, elevated RAD51 levels may be signalling extensive DNA damage.
Human cytomegalovirus infection and expression may be specifically associated with human cancers, including malignant glioma, colorectal and prostate cancer (Cobbs et al. 2002; Harkins et al. 2002; Samanta et al. 2003, reviewed in Michaelis et al. 2009; Dziurzynski et al. 2012). HCMV can deregulate the signalling pathways, involved in initiation and promotion of various malignancy, including tumour suppressor, mitogenic signalling, inflammatory, immune regulation, angiogenesis and invasion. Human cytomegalovirus might play an important role in modulating tumour microenvironment as well as in the initiation and promotion of tumour cell development (Soroceanu and Cobbs 2011). The significance of HCMV infection in oncogenesis is an active area of scientific research (Cinatl et al. 2004). Although still controversial, there is a growing body of evidence that links viral infection to a various malignancy.
It is supposed that HCMV propensity to cause cancer may be associated with deregulation of homologous recombination of DNA repair machinery.
During HCMV infection, DSB repairing functions occur, leading to increased cancer development in infected cells because of genome rearrangement (Dziurzynski et al. 2011; Michaelis et al. 2011).
The oncogenic potential of human adenoviruses and HCMV are associated with site-specific chromosomal damage. HCMV induces specific damage in chromosome 1 at two loci 1q23 and 1q42 (Fortunato et al. 2000; Nystad et al. 2008).
These HCMV effects on chromosomal integrity and the synergistic activity of HCMV infection in combination with other cytotoxic agents contribute significantly to global genetic instability, which is a major cancer promoter (Albrecht et al. 2004; Siew et al. 2009). It has been shown that the HCMV protein UL76 can induce chromosomal aberrations (e.g. production of micronuclei, misaligned chromosomes, chromosomal lagging and bridging) (Siew et al. 2009).
Cellular responses to DNA damage are mediated by a number of protein kinases, including ATM (ataxia telangiectasia-mutated) and ATR (ATM and Rad3-related). HCMV has been shown to disrupt DNA repair pathways, including the activity of ATM and ATR (Luo et al. 2007).
HCMV interacts with your host DNA damage response (DDR) signalling molecules and repair machinery. Interactions between HCMV-encoded proteins pUL35 and pUL27 and DNA-damage DNA-binding protein (DDB1) contribute to DDR activation (Salsman et al. 2012; Reitsma et al. 2011; Costa et al. 2013; reviewed in Xiaofei and Kowalik 2014).
pUL35 can active DDR, causing γH2AX and 53BP1 foci formation and induce a cell cycle arrest which likely supports viral replication (Salsman et al. 2012). HCMV, in particular, encodes IE1, IE2, pp71, pUL97, pUL69 proteins that both modulate cell cycle controls and the host DDR to promote viral replication. The inactivation of RB family members by these four proteins and subsequent deregulation of E2F1 appears to result in double-strand breaks in human fibroblasts (Xiaofie et al. 2011). The mechanism by which E2F1 stimulates host DDR is not well understood.
It is known that viruses utilize initial DDR response to optimize infection (Weitzman et al. 2010; Weller 2010). HCMV genome replication requires host DDRs and raises the possibility that DNA repair pathways may influence viral replication. A recent study shows that a DNA repair factor, DDB2, can contribute to HCMV replication (Xiaofei et al. 2014).
Literature data have shown that in HCMV-infected human foreskin fibroblasts, DDR is activated at the time of viral deposition and during late-phase replication (Gaspar and Shenk 2006; Luo et al. 2007).
The progression of the HCMV life cycle also depends on the capability of the virus to modulate host cell processes such as cellular gene expression and apoptosis process to its advantage. Recently, US27 was identified as a modulator of cellular gene expression (Arnolds et al. 2013). The UL28-UL29, UL38, UL79 and RNA2.7 were found to control apoptosis (Terhune et al. 2007; Moorman et al. 2008; Siew et al. 2009; Qian et al. 2011; Fliss and Brune 2012; Costa et al. 2013; Savaryn et al. 2013).
In conclusion, several experimental studies have shown that double-strand break (DSB) repair undergoes viral-related changes, which are likely to contribute to the accumulation of DNA lesions.
DNA repair gene polymorphism and HCMV infection
As mentioned above, the human cytomegalovirus can inflict site-specific chromosomal damage in form of double-strand breaks (DSBs). The cellular reaction to the DNA damaging agents can modulate susceptibility to viral infection. This reaction is mainly determined by the efficacy of DNA repair which, in turn, may be influenced by the variability of DNA repair genes, expressed by their polymorphism.
DNA repair genes are highly polymorphic in nature. However, little is known about HCMV infection-related DNA repair gene polymorphisms.
Human cytomegalovirus infection and expression may be specifically associated with human cancers, including colon carcinoma, prostate, adenocarcinoma, cervical carcinoma, neuroblastoma and glioblastoma (Harkins et al. 2002; Samanta et al. 2003; Cobbs et al. 2002; Cinatl et al. 2004; reviewed in Michaelis et al. 2009; Dziurzynski et al. 2012).
Polymorphisms in DNA repair genes may alter the activity of the proteins and, thus, modulate susceptibility to cancer (Karahalil et al. 2012; Samulak et al. 2011; Smolarz et al. 2013).
It is supposed that HCMV propensity to cause cancer may be associated with DNA repair genes, previously implicated in tumour disease. Therefore, an effective analysis of DNA repair gene polymorphism during viral infection may represent a true advance in the study of viral pathogenesis.
References
AbuBakar S, Au WW, Legator MS, Albrecht T (1988) Induction of chromosome aberrations and mitotic arrest by cytomegalovirus in human cells. Environ Mol Mutagen 12:409–420
Albrecht T, Deng CZ, Abdel-Rahman SZ, Fons M, Cinciripini P, El-Zein RA (2004) Differential mutagen sensitivity of peripheral blood lymphocytes from smokers and nonsmokers: effect of human cytomegalovirus infection. Environ Mol Mutagen 43:169–178
Almeida KH, Sobol RW (2007) A unified view of base excision repair: lesion-dependent protein complexes regulated by post-translational modification. DNA Repair (Amst) 6:695–711
Arnolds KL, Lares AP, Spencer JV (2013) The US27 gene product of human cytomegalovirus enhances signaling of host chemokine receptor CXCR4. Virology 439:122–131
Bartek J, Lukas C, Lukas J (2004) Checking on DNA damage in S phase. Nat Rev Mol Cell Biol 5:792–804
Baydoun HH, Pancewicz J, Nicot C (2011) Human T-lymphotropic type 1 virus p30 inhibits homologous recombination and favors unfaithful DNA repair. Blood 117:5897–5906
Becker SA, Lee TH, Butel JS, Slagle BL (1998) Hepatitis B virus X protein interferes with cellular DNA repair. J Virol 72:266–272
Ben Yehuda A, Globerson A, Krichevsky S, Bar On H, Kidron M, Friedlander Y, Friedman G, Ben Yehuda D (2000) Ageing and the mismatch repair system. Mech Ageing Dev 121:173–179
Buck D, Malivert L, de Chasseval R, Barraud A, Fondanèche MC, Sanal O, Plebani A, Stéphan JL, Hufnagel M, le Deist F, Fischer A, Durandy A, de Villartay JP, Revy P (2006) Cernunnos, a novel nonhomologous endjoining factor, is mutated in human immunodeficiency with microcephaly. Cell 124:287–299
Chaurushiya MS, Weitzman MD (2009) Viral manipulation of DNA repair and cell cycle checkpoints. DNA Repair (Amst) 8:1166–1176
Chee MS, Satchwell SC, Preddie E, Weston KM, Barrell BG (1990) Human cytomegalovirus encodes three G protein-coupled receptor homologues. Nature 344:774–777
Chipitsyna G, Slonina D, Siddiqui K, Peruzzi F, Skorski T, Reiss K, Sawaya BE, Khalili K, Amini S (2004) HIV-1 Tat increases cell survival in response to cisplatin by stimulating Rad51 gene expression. Oncogene 23:2664–2671
Cinatl J, Vogel JU, Kotchetkov R, Doerr HW (2004) Oncomodulatory signals by regulatory proteins encoded by human cytomegalovirus: a novel role for viral infection in tumour progression. FEMS Microbiol Rev 28:59–77
Cobbs CS, Harkins L, Samanta M, Gillespie GY, Bharara S, King PH, Nabors LB, Cobbs CG, Britt WJ (2002) Human cytomegalovirus infection and expression in human malignant glioma. Cancer Res 62:3347–3350
Costa H, Nascimento R, Sinclair J, Parkhouse RM (2013) Human cytomegalovirus gene UL76 induces IL-8 expression through activation of the DNA damage response. PLoS Pathog 9:e1003609
Deng CZ, AbuBakar S, Fons MP, Boldogh I, Albrecht T (1992a) Modulation of the frequency of human cytomegalovirus-induced chromosome aberrations by camptothecin. Virology 189:397–401
Duong FH, Christen V, Lin S, Heim MH (2010) Hepatitis C virus-induced up-regulation of protein phosphatase 2A inhibits histone modification and DNA damage repair. Hepatology 51:741–751
Durkin SS, Guo X, Fryrear KA, Mihaylova VT, Gupta SK, Belgnaoui SM, Haoudi A, Kupfer GM, Semmes OJ (2008) HTLV-1 Tax oncoprotein subverts the cellular DNA damage response via binding to DNA-dependent protein kinase. J Biol Chem 283:36311–36320
Dziurzynski K, Wei J, Qiao W, Hatiboglu MA, Kong LY, Wu A, Wang Y, Cahill D, Levine N, Prabhu S, Rao G, Sawaya R, Heimberger AB (2011) Glioma associated cytomegalovirus mediates subversion of the monocyte lineage to a tumour propagating phenotype. Clin Cancer Res 17:4642–4649
Dziurzynski K, Chang SM, Heimberger AB, Kalejta RF, McGregor Dallas SR, Smit M, Soroceanu L, Cobbs CS (2012) HCMV and gliomas symposium. Consensus on the role of human cytomegalovirus in glioblastoma. Neurol Oncol 14:246–255
Evans AR, Limp-Foster M, Kelley MR (2000) Going APE over ref-1. Mutat Res 461:83–108
Fliss PM, Brune W (2012) Prevention of cellular suicide by cytomegaloviruses. Viruses 4:1928–1949
Fortunato EA, Spector DH (2003) Viral induction of site specific chromosome damage. Rev Med Virol 13:21–37
Fortunato EA, Dell’Aquila ML, Spector DH (2000) Specific chromosome 1 breaks induced by human cytomegalovirus. Proc Natl Acad Sci U S A 97:853–858
Gamadia LE, Remmerswaal EB, Weel JF, Bemelman F, van Lier RA, Ten Berge IJ (2003) Primary immune responses to human CMV: a critical role for IFN-gamma-producing CD4+ T cells in protection against CMV disease. Blood 101:2686–2692
Gaspar M, Shenk T (2006) Human cytomegalovirus inhibits a DNA damage response by mislocalizing checkpoint proteins. Proc Natl Acad Sci U S A 103:2821–2826
Groisman IJ, Koshy R, Henkler F, Groopman JD, Alaoui-Jamali MA (1999) Downregulation of DNA excision repair by the hepatitis B virus-x protein occurs in p53-proficient and p53-deficient cells. Carcinogenesis 20:479–483
Gruhne B, Sompallae R, Masucci MG (2009) Three Epstein-Barr virus latency proteins independently promote genomic instability by inducing DNA damage, inhibiting DNA repair and inactivating cell cycle checkpoints. Oncogene 28:3997–4008
Haaf T, Golub EI, Reddy G, Radding CM, Ward DC (1995) Nuclear foci of mammalian Rad51 recombination protein in somatic cells after DNA damage and its localization in synaptonemal complexes. Proc Natl Acad Sci U S A 92:2298–2302
Hanawalt PC (2002) Subpathways of nucleotide excision repair and their regulation. Oncogene 21:8949–8956
Harkins L, Volk AL, Samanta M, Mikolaenko I, Britt WJ, Bland KI, Cobbs CS (2002) Specific localisation of human cytomegalovirus nucleic acids and proteins in human colorectal cancer. Lancet 360:1557–1563
Helleday T (2003) Pathways for mitotic homologous recombination in mammalian cells. Mutat Res 532:103–115
Hoeijmakers JH (2009) DNA damage, aging, and cancer. N Engl J Med 361:1475–1485
Hsieh P, Yamane K (2008) DNA mismatch repair: molecular mechanism, cancer, and ageing. Mech Ageing Dev 129:391–407
Jackson SP (2002) Sensing and repairing DNA double-strand breaks. Carcinogenesis 23:687–696
Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461:1071–1078
Jia L, Wang XW, Harris CC (1999) Hepatitis B virus X protein inhibits nucleotide excision repair. Int J Cancer 80:875–879
Karahalil B, Bohr VA, Wilson DM 3rd (2012) Impact of DNA polymorphisms in key DNA base excision repair proteins on cancer risk. Hum Exp Toxicol 31:981–1005
Karran P (1996) Microsatellite instability and DNA mismatch repair in human cancer. Semin. Cancer Biol 7:15–24
Krichevsky S, Pawelec G, Gural A, Effros RB, Globerson A, Yehuda DB, Yehuda AB (2004) Age related microsatellite instability in T cells from healthy individuals. Exp Gerontol 39:507–515
Kulkarni AS, Fortunato EA (2011) Stimulation of homology-directed repair at I-SceI-induced DNA breaks during the permissive life cycle of human cytomegalovirus. J Virol 85:6049–6054
Liang X, Pickering MT, Cho NH, Chang H, Volkert MR, Kowalik TF, Jung JU (2006) Deregulation of DNA damage signal transduction by herpesvirus latency-associated M2. J Virol 80:5862–5874
Lilley CE, Chaurushiya MS, Boutell C, Landry S, Suh J, Panier S, Everett RD, Stewart GS, Durocher D, Weitzman MD (2010) A viral E3 ligase targets RNF8 and RNF168 to control histone ubiquitination and DNA damage responses. EMBO J 29:943–955
Luo MH, Rosenke K, Czornak K, Fortunato EA (2007) Human cytomegalovirus disrupts both ataxia telangiectasia mutated protein (ATM)- and ATM-Rad3-related kinase-mediated DNA damage responses during lytic infection. J Virol 81:1934–1950
Mathonnet G, Lachance S, Alaoui-Jamali M, Drobetsky EA (2004) Expression of hepatitis B virus X oncoprotein inhibits transcription-coupled nucleotide excision repair in human cells. Mutat Res 554:305–318
Michaelis M, Doerr HW, Cinatl J (2009) The story of human cytomegalovirus and cancer: increasing evidence and open questions. Neoplasia 11:1–9
Michaelis M, Baumgarten P, Mittelbronn M, Driever PH, Doerr HW, Cinatl J Jr (2011) Oncomodulation by human cytomegalovirus: novel clinical findings open new roads. Med Microbiol Immunol 200:1–5
Miller-Kittler M, Sparer TE (2009) Feeling manipulated: cytomegalovirus immune manipulation. Virol J 6:4
Moorman NJ, Cristea IM, Terhune SS, Rout MP, Chait BT, Shenk T (2008) Human cytomegalovirus protein UL38 inhibits host cell stress responses by antagonizing the tuberous sclerosis protein complex. Cell Host Microbe 3:253–262
Moshous D, Callebaut I, de Chasseval R, Corneo B, Cavazzana-Calvo M, Le Deist F, Tezcan I, Sanal O, Bertrand Y, Philippe N, Fischer A, de Villartay JP (2001) Artemis, a novel DNA double-strand break repair/V(D)J recombination protein, is mutated in human severe combined immune deficiency. Cell 105:177–186
Nakai-Murakami C, Shimura M, Kinomoto M, Takizawa Y, Tokunaga K, Taguchi T, Hoshino S, Miyagawa K, Sata T, Kurumizaka H, Yuo A, Ishizaka Y (2007) HIV-1 Vpr induces ATM-dependent cellular signal with enhanced homologous recombination. Oncogene 26:477–486
Neri S, Gardini A, Facchini A, Olivieri F, Franceschi C, Ravaglia G, Mariani E (2005) Mismatch repair system and aging: microsatellite instability in peripheral blood cells from differently aged participants. J Gerontol A Biol Sci Med Sci 60:285–292
Nishiyama Y, Rapp F (1981) Enhanced capacity of DNA repair in human cytomegalovirus-infected cells. J Virol 38:164–172
Nystad M, Fagerheim T, Brox V, Fortunato EA, Nilssen O (2008) Human cytomegalovirus (HCMV) and hearing impairment: infection of fibroblast cells with HCMV induces chromosome breaks at 1q23.3, between loci DFNA7 and DFNA49—both involved in dominantly inherited, sensorineural, hearing impairment. Mutat Res 637:56–65
O’Dowd JM, Zavala AG, Brown CJ, Mori T, Fortunato EA (2012) HCMV-infected cells maintain efficient nucleotide excision repair of the viral genome while abrogating repair of the host genome. PLoS Pathog 11:e1003038
Pal S, Polyak SJ, Bano N, Qiu WC, Carithers RL, Shuhart M, Gretch DR, Das A (2010) Hepatitis C virus induces oxidative stress, DNA damage and modulates the DNA repair enzyme NEIL1. J Gastroenterol Hepatol 25:627–634
Philpott SM, Buehring GC (1999) Defective DNA repair in cells with human T-cell leukemia/bovine leukemia viruses: role of tax gene. J Natl Cancer Inst 91:933–942
Polo SE, Jackson SP (2011) Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev 25:409–433
Prost S, Ford JM, Taylor C, Doig J, Harrison DJ (1998) Hepatitis B x protein inhibits p53-dependent DNA repair in primary mouse hepatocytes. J Biol Chem 273:33327–33332
Qian Z, Xuan B, Gualberto N, Yu D (2011) The human cytomegalovirus protein pUL38 suppresses endoplasmic reticulum stress-mediated cell death independently of its ability to induce mTORC1 activation. J Virol 85:9103–9113
Raderschall EK, Stout S, Freier V, Suckow S, Schweiger HT (2002) Elevated levels of Rad51 recombination protein in tumour cells. Cancer Res 62:219–225
Ranneberg-Nilsen T, Bjørås M, Luna L, Slettebakk R, Avsnes Dale H, Seeberg E, Rollag H (2006) Human cytomegalovirus infection modulates DNA base excision repair in fibroblast cells. Virology 348:389–397
Reitsma JM, Savaryn JP, Faust K, Sato H, Halligan BD, Terhune SS (2011) Antiviral inhibition targeting the HCMV kinase pUL97 requires pUL27-dependent degradation of Tip60 acetyltransferase and cell-cycle arrest. Cell Host Microbe 9:103–114
Salsman J, Zimmerman N, Chen T, Domagala M, Frappier L (2008) Genome wide screen of three herpesviruses for protein subcellular localization and alteration of PML nuclear bodies. PLoS Pathog 4:e1000100
Salsman J, Jagannathan M, Paladino P, Chan PK, Dellaire G, Raught B, Frappier L (2012) Proteomic profiling of the human cytomegalovirus UL35 gene products reveals a role for UL35 in the DNA repair response. J Virol 86:806–820
Samanta M, Harkins L, Klemm K, Britt WJ, Cobbs CS (2003) High prevalence of human cytomegalovirus in prostatic intraepithelial neoplasia and prostatic carcinoma. J Urol 170:998–1002
Samulak D, Romanowicz-Makowska H, Smolarz B, Kulig A, Sporny S (2011) Association between Arg399Gln polymorphism of X-ray repair cross-complementing 1 (XRCC1) gene and sporadic endometrial cancer in the Polish population. Eur J Gynaecol Oncol 32:491–5
Savaryn JP, Reitsma JM, Bigley TM, Halligan BD, Qian Z, Yu D, Terhune SS (2013) Human cytomegalovirus pUL29/28 and pUL38 repression of p53-regulated p21CIP1 and caspase 1 promoters during infection. J Virol 87:2463–2474
Shen Y, Zhu H, Shenk T (1997) Human cytomegalovirus IE1 and IE2 proteins are mutagenic and mediate “hit-and-run” oncogenic transformation in cooperation with the adenovirus E1A proteins. Proc Natl Acad Sci U S A 94:3341–3345
Siew VK, Duh CY, Wang SK (2009) Human cytomegalovirus UL76 induces chromosome aberrations. J Biomed Sci 16:107
Skinner AM (2005) Turker MS (2005) Oxidative mutagenesis, mismatch repair, and aging. Sci Aging Knowl Environ 9:re3
Smolarz B, Zadrożny M, Duda-Szymańska J, Makowska M, Samulak D, Michalska MM, Mojs E, Bryś M, Forma E, Romanowicz-Makowska H (2013) RAD51 genotype and triple-negative breast cancer (TNBC) risk in Polish women. Pol J Pathol 64:39–43
Soroceanu L, Cobbs C (2011) Is HCMV a tumour promoter? Virus Res 157:193–203
Speir E, Yu ZX, Ferrans VJ, Huang ES, Epstein SE (1998) Aspirin attenuates cytomegalovirus infectivity and gene expression mediated by cyclooxygenase-2 in coronary artery smooth muscle cells. Circ Res 83:210–216
Stern-Ginossar N, Weisburd B, Michalski A, Le VT, Hein MY, Huang SX, Ma M, Shen B, Qian SB, Hengel H, Mann M, Ingolia NT, Weissman JS (2012) Decoding human cytomegalovirus. Science 338:1088–1093
Sun Y, Huang YC, Xu QZ, Wang HP, Bai B, Sui JL, Zhou PK (2006) HIV-1 Tat depresses DNA-PK(CS) expression and DNA repair, and sensitizes cells to ionizing radiation. Int J Radiat Oncol Biol Phys 65:842–850
Terhune S, Torigoi E, Moorman N, Silva M, Qian Z, Shenk T, Yu D (2007) Human cytomegalovirus UL38 protein blocks apoptosis. J Virol 81:3109–3123
Tonegawa S (1983) Somatic generation of antibody diversity. Nature 302:575–581
Trojanek J, Croul S, Ho T, Wang JY, Darbinyan A, Nowicki M, Del Valle L, Skorski T, Khalili K, Reiss K (2006) T-antigen of the human polyomavirus JC attenuates faithful DNA repair by forcing nuclear interaction between IRS-1 and Rad51. J Cell Physiol 206:35–46
Weitzman MD, Lailley CE, Chaurushiya MS (2010) Genomes in conflict: maintaining genome integrity during virus infection. Annu Rev Microbiol 61:61–81
Weller SK (2010) Herpes simplex virus reorganizes the cellular DNA repair and protein quality control machinery. PLoS Pathog 6:e1001105
Weterings E, van Gent DC (2004) The mechanism of non-homologous end-joining: a synopsis of synapsis. DNA Repair (Amst) 3:1425–1435
Wilson DM 3rd, Bohr VA (2007) The mechanics of base excision repair, and its relationship to aging and disease. DNA Repair (Amst) 6:544–559
Xiaofei E, Kowalik TF (2014) The DNA damage response induced by infection with human cytomegalovirus and other viruses. Viruses 6:2155–85
Xiaofei E, Savidis G, Chin CR, Wang S, Lu S, Brass AL, Kowalik TF (2014) A novel DDB2-ATM feedback loop regulates human cytomegalovirus replication. J Virol 88:2279–2290
Xiaofie E, Pickering MT, Debatis M, Castillo J, Lagadinos A, Wang S, Lu S, Kowalik TF (2011) An E2F1-mediated DNA damage response contributes to the replication of human cytomegalovirus. PLoS Pathog 7:e1001342
Conflict of interest
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Smolarz, B., Wilczyński, J. & Nowakowska, D. DNA repair mechanisms and human cytomegalovirus (HCMV) infection. Folia Microbiol 60, 199–209 (2015). https://doi.org/10.1007/s12223-014-0359-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-014-0359-6