Abstract
Temperately adapted perennial sorghum feedstocks have recently begun to receive increasing interest as candidate energy crops, producing significant biomass and contributing agroecological benefits including increased soil organic carbon, reduced soil erosion, reduced input requirements, and higher net energy return. Rhizomes are the primary morphological feature facilitating overwintering in Sorghum species; however, underlying physiological mechanisms governing rhizome overwintering remain poorly characterized. In this study, we investigated the composition of sorghum rhizomes from diverse germplasm before and after overwintering at two locations and three experimental environments. Significant positive correlations were found between rhizome overwintering and water-soluble carbohydrates, ethanol soluble carbohydrates, and fructan concentrations, while significant negative correlations were found between rhizome overwintering and both crude fat and starch. Near-infrared spectroscopy (NIRS) calibration equations were developed to quickly and efficiently predict the concentrations of each of these assimilates in rhizomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sorghum species are utilized worldwide for the production of grain, forage, sugar, and more recently biofuels. Sorghum bicolor (L.) Moench is the fifth most important grain crop in the world in terms of production [1]. Sorghum almum Parodi as well as Sudangrass, S. bicolor ssp. drummondii (Nees ex. Steud.) de Wet & Harlan, and its hybrids with S. bicolor have been used for decades as high biomass forage and hay crops [2, 3]. Recently, several forms of S. bicolor have been evaluated as candidates for both liquid and solid biofuel feedstocks [4–8]. This species behaves as an annual in temperate climates and as a weak perennial in tropical and subtropical climates where temperatures do not drop below freezing. Other Sorghum species, notably Sorghum halepense (L.) Pers. and Sorghum propinquum (Kunth) Hitchc., are strongly rhizomatous perennials [9].
Perennial sorghum cropping systems offer agroecological benefits not present in annual row crop production such as increased soil organic carbon, reduced soil erosion, reduced inputs requirements, and higher net energy return [10–14]. Land use change could be minimized in such systems through the utilization of marginal croplands or abandoned grasslands. In 2007, there were 614 million acres (248 million ha) of rangeland and pastureland, 36 million acres (15 million ha) of cropland used for pasture, and 37 million acres (15 million ha) of idle cropland in the USA alone [15] that could be utilized in perennial sorghum cropping systems. Improved overwintering ability would be a critical component of perennial sorghum systems and relates directly to the production of rhizomes [16, 17], modified underground stems that act as storage and propagation organs [18, 19]. Although rhizome depth of growth and derived tiller number have been implicated in overwintering ability [16, 20], factors such as rhizome assimilate composition remain poorly characterized. Furthermore, it is unknown if genetic diversity in sorghum rhizome assimilate composition exists. Carbohydrates, primarily glucose, sucrose, starch, and dextrins, vary in rhizomes throughout the growing season depending on the plant’s growth stage with levels decreasing during sprouting and seed formation [21–23]. After seed maturation, sucrose concentrations in the rhizomes increase significantly, which is believed to be beneficial for overwintering [21, 22]. Additional assimilates associated with energy storage increase in response to environmental factors. For example, varying degrees of correlation between starch content and cold tolerance have been reported [24–27], and high concentrations of sucrose have been correlated with cold tolerance in several species [24, 28]. The above findings indicate that predictive correlations between rhizome composition and overwintering ability may occur in Sorghum species.
The determination of rhizome composition requires costly wet chemistry analysis, which makes it impractical to thoroughly analyze all the genotypes necessary for genetic mapping studies and large-scale field experiments. Near-infrared spectroscopy (NIRS) provides a possible alternative to wet-chemical analysis, measuring near-infrared light absorbance and/or reflectance within sample tissues. In order to use NIRS, a calibration equation must be created for the specific tissue type and the compositional measurements desired [29]. This is accomplished by analyzing a range of samples using both wet chemistry analysis and NIRS and then correlating the results from the two analyses. Once the calibration equation is loaded into the NIRS software, compositional predictions can be made quickly at a comparatively low cost. NIRS equations have been developed for various sorghum grain, leaf, and steam components [8, 29–31], but none have been reported for sorghum rhizome composition or rhizomes in most other species. However, a calibration equation does exist for total nonstructural carbohydrates in peppermint (Mentha piperita L.) rhizomes [32]. Development of sorghum rhizome composition calibration equations would greatly facilitate high throughput phenotyping of rhizome compositional traits for both sorghum breeding and research programs.
The objectives of this study were to (1) characterize rhizome assimilate composition and its relationship to overwintering ability in a collection of S. halepense and Sorghum hybrid genotypes, (2) determine if significant genetic variation for rhizome composition exists in this germplasm, and (3) determine the usefulness of NIRS in predicting the levels of compositional assimilates in sorghum as a potential high-throughput method for determining rhizome composition.
Experimental
Plant Materials and Field Trials
Plant material used for this study consisted of nine S. halepense genotypes that were collected from several locations in Texas, USA, one genotype collected in North Carolina, USA, and one genotype from the USDA National Plant Germplasm System [33] (Table 1). These plants were propagated vegetatively during the spring of 2010 to have sufficient plant material for field planting in autumn of 2010.
Winter survival trials were conducted at Commerce (33°11′ N, 95°55′ W) and College Station (30°32′ N, 96°26′ W), Texas, USA in 2010 and 2011. At both locations, genotypes were planted in a randomized complete block design with three replications. The plots at Commerce were located on the Texas A&M University–Commerce Research Farm, and the plants were grown in columns of soil inside Polyvinyl chloride (PVC) tubes 1 m in length and 15.2 cm in diameter (see below). The tubes were placed into 92-cm deep holes that were dug with a power take-off (PTO)-driven auger attached to a tractor. The distance between soil columns at Commerce was 1.8 m to allow for the movement of the tractor between plants.
Two experimental plantings were established at the Texas A&M University Research Farm near College Station. One was similar to the plot at Commerce except that, at College Station, the tubes were submerged into the soil to a depth of only 46 cm and the distance between rows and columns within a row was 1 m. The soil level of the columns at Commerce was even with the soil level of the field, but the soil level in the columns at College Station was 46 cm above the field soil level. This placement was intended to expose the upper half of the soil columns at College Station to more extreme temperatures. For the second College Station planting, the plants were transplanted directly into the soil at a distance of 2 m between plants within a row and 2 m between rows. Because one College Station experiment was conducted in soil columns and the second was planted directly into soil in the field, they were considered as different experimental environments.
Column Construction
The soil columns were made by longitudinally cutting 3 m lengths of 15.2 cm diameter PVC pipe into two equal halves. The halves were cut into 1-m sections. At the lower end of each half-tube, three evenly spaced 0.3-cm holes were drilled and threaded with galvanized wire, which crossed at the center to create a web-like design. The two pipe halves were then placed together and secured with duct tape at locations along the length of the tube. A piece of burlap cloth was folded several times and placed on top of the wire web at the bottom of each column for the purpose of retaining the soil while allowing water percolation.
Tubes used at both locations were filled with Westwood silt loam (fine-silty, mixed, superactive, thermic Udifluventic Haplustepts) that was taken to a depth of 15.2 cm from the Texas A&M University Research Farm. The tubes were filled with the soil, tamped on the ground, and then watered overnight using a soaker hose to insure similar soil compaction across all tubes. Following watering, soil compaction caused a drop in the soil level in each tube, and soil was added to bring the soil level to ∼7.6 cm below the top of each tube resulting in a 92-cm column of soil in each tube.
Planting and Sampling
For all studies, plants were vegetatively propagated via rhizome cuttings in March of 2010. Six cuttings from each genotype (three replicates with two plants each; one for autumn sampling and one for spring sampling) were used per experiment. After a few months of growth, plants were trimmed approximately 15 cm above the soil surface and transplanted into soil columns (on 16 August) or the field environment (on 8 July). At the time of planting, tiller number varied by genotype. Plants were fertilized at 84 kg N ha−1 as ammonium sulfate and watered. Columns designated for the autumn and spring sampling at Commerce were placed in the ground on 17 September. At College Station, the tubes designated for autumn were sampled directly without placement in the ground, while the tubes designated for spring sampling were placed into the ground on 12 November.
For sampling, the tubes were removed from the ground and the tape binding the two halves of the tubes together was cut. The tubes were then opened, leaving the soil column intact. Each soil column was cut laterally into two equal halves 46 cm in length. Each half of the soil column was placed in a tub of water and the soil was washed away so rhizomes could be collected. The rhizomes were collected separately from the upper (0–46 cm) and lower (46–92 cm) halves of each column. However, only the samples taken from the upper half of each column were used for analyses because only a limited number of columns had rhizomes below 46 cm. Rhizome diameters were measured at the center of the first complete internode from the cut site. Samples were then placed into plastic bags and stored in a −80 °C freezer. Autumn sampling at Commerce took place on 22 October prior to the first freezing temperature, which occurred on 25 November. College Station columns were sampled on 10 November prior to the first freezing temperature on 27 November.
Autumn sampling of the College Station field plots (planted directly in the soil rather than in tubes) took place on 17 and 19 November. The sampling procedure for the plants in the field differed from those in tubes. Above-ground vegetation of each plant was removed before the rhizomes were collected. Using a shovel, the whole plant was excavated. After the plant crown and rhizome mass were removed, the depth of the hole was increased to 46 cm and rhizomes were collected from the material removed from the hole. Rhizomes were not found below 46 cm; therefore, samples were not collected in the 46–92 cm depth range. The lack of rhizomes below 46 cm was likely due to a hard pan that was present approximately 41 cm below the soil surface. Rhizomes were washed and placed into plastic bags and stored on ice until they were transferred into a −80 oC freezer. Since the sampling method was destructive, the remaining plants of each genotype were left in the field to be sampled the following spring. Just prior to spring rhizome sampling, several traits associated with overwintering ability and spring regrowth (the presence of new leaf material in the spring) were measured for both the field-planted and column-grown plants at both locations. For the College Station field site (not in columns) a 1-m2 quadrat centered on the primary tillers of each plant was used to count the number of green shoots and the number of leaves. These data were collected at College Station on 2 March 2011. Spring regrowth measurements were made on the plants growing in the soil column on the day of sampling, which was 1 April at College Station and 7 April at Commerce.
Spring sampling at both locations was conducted in the same manner as the previous autumn sampling, with a few exceptions. Only live rhizome tissue was collected in the spring because some rhizomes had freeze damage. Rhizomes considered “live” had a healthy/white appearance and were rigid.
Rhizome Composition Analysis
To prepare the rhizome samples for analysis, the rhizomes were placed into mesh bags and dried in a lyophilizer (Labconco, Kansas City, MO, USA) for at least 3 days at −40 °C. All fibrous root material was removed from the rhizomes. Samples were then ground to a 1-mm particle size using first a coarse grinder and then a UDY Cyclone mill (Udy Corp, Fort Collins, CO, USA). The ground tissue was placed into glass vials and stored in cold storage.
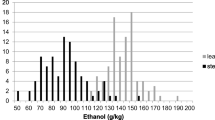
Tissue samples were analyzed by Dairy One (Utica, NY, USA) using wet chemistry analysis for crude protein (CP), crude fat (fat), starch, water soluble carbohydrates (WSCs), and ethanol-soluble carbohydrates (ESCs). WSC consists primarily of glucose, sucrose, fructose, maltose, lactose, and fructans, whereas ESC is made up of primarily glucose, sucrose, fructose, maltose, and lactose. An estimate of fructan concentration was therefore obtained by determining the concentration difference between WSC and ESC for each sample.
Assimilate Analysis via NIR
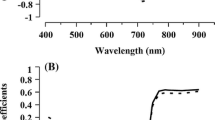
Prior to wet chemistry analysis, the samples were scanned with a Thermo Scientific Antaris II FT-NIR Analyzer (Thermo Scientific Inc., Waltham, MA, USA) with a setting of 64 coaveraged scans per sample and a resolution of 4 cm−1 between wavelengths 4,000 and 9,999 cm−1 using the OMNIC 8 software suite (Thermo Scientific Inc., Waltham, MA, USA). To insure the tissue was packed uniformly into the vials, they were placed side by side in a small box and tamped on a lab bench 15 times prior to scanning. NIRS reflectance spectral data was converted to absorbance and combined with the wet chemistry analysis data in TQ Analyst software (Thermo Scientific Inc., Waltham, MA, USA) to create prediction equations. Approximately one third of the data were used as a validation set and the other two thirds were used as a calibration set. Many different standard mathematical treatments were applied over different spectral ranges before selecting the treatment with the highest performance index. The performance index validates the NIRS curve by measuring how accurately a calibrated method can quantify independent validation samples. A value of 0 reflects no prediction while a maximal value of 100 reflects perfect prediction.
Statistical Analysis
Data were analyzed for genotype, environment, and genotype by environment (GxE) effects using a PROC MIXED statement with environment, genotype, and GxE as random variables in SAS version 9.1 [34]. The variation for each effect was then divided by the total variance to get a percentage of total variance. The two sampling dates (autumn or spring) were not comparable and analyzed separately but with otherwise identical PROC MIXED analyses including all three environments. Correlations between CP, fat, starch, WSC, ESC, and overwintering capability were determined using a PROC CORR statement. These correlation analyses were carried out on all experiments and sampling dates individually. In addition, the two soil column environments were also analyzed jointly because of their similar design, using a PROC CORR statement. Logistic regression models were created for the prediction of spring regrowth (overwintering ability) based on autumn assimilate composition data using JMP® Software, Version 9 [35].
Results and Discussion
Genotype and Genotype by Environment Effects
Both locations were exposed to subfreezing temperatures and provided suitable environments for screening. Genotypic effects on rhizome assimilate concentrations varied between and among environments and sampling dates (Table 2). Measurements of spring regrowth also show some genotype effect. No single genotype consistently produced the highest concentration of any given assimilate across all environments. Therefore, GxE effects were large for multiple assimilates in both the spring and autumn sampling dates (Table 2). The large environmental variation observed for some assimilates and sampling dates demonstrate that absolute values were not comparable across environments. From our findings, it appears that although many factors affect rhizome composition [36], screening for genetic components can identify useful diversity.
Assimilate Concentrations and Overwintering
Correlations and modeling between autumn assimilate concentrations and spring regrowth were used to estimate the effects that autumn assimilate accumulation may have on overwintering ability. Pearson correlation coefficients between regrowth and rhizome score were significant in both soil column environments, with the highest correlation coefficient being obtained when both column environments were combined (Tables 3 and 4). Plants were subjected to the most severe winter conditions at the College Station column environment because half of each column was above the soil level. This environment also had the most obvious differences between genotypes in spring regrowth. In the College Station soil column study, significant correlations were identified between spring regrowth and a number of assimilates from the autumn sampling date including starch, WSC, ESC, and fat (Table 3). Our best logistic regression model for this environment included only starch and fat, both of which were negatively correlated with spring regrowth (Table 5). WSC and ESC both had significant impacts when placed individually in a regression model meaning that their effects were unequal to zero, but their impacts were not significant when starch and fat were also included in the model. Our regression model had an R 2 of 0.49 and was significant at α = 0.001 level, indicating that all predictors in this model were correlated with post winter regrowth. Identical correlations conducted on the Commerce environment data revealed a single significant correlation between spring regrowth and autumn WSC concentration. Similar correlations and modeling approaches were used on the College Station field data; however, they did not yield significant differences. This is likely due to all field plants in that study surviving the winter resulting in lower statistical power, or it may be because above-ground biomass was removed several weeks prior to the spring sampling, and this may have caused translocation of nutrients away from rhizome tissue before harvest. Spring rhizome composition was significantly correlated with regrowth in the College Station column environment for WSC, fructans, and fat (Table 4).
One of the most notable differences between our data and that from previous studies is that we observed a negative correlation between fat content and overwintering ability. This finding is in contrast to observations on Arabidopsis above-ground growth [37] and what was hypothesized for the genus Paspalum based in part on studying rhizomes cut from plants and buried in the ground [38, 39]. Although our finding was unexpected, it is supported by significant correlations during both sampling dates for the College Station soil column studies. Most previous studies concerning the importance of higher fat content for cold tolerance have focused on actively photosynthetic tissues. These studies have used the plants' ability to continue normal metabolic functions during brief cold periods as their indicator of cold tolerance. In our study, we were concerned with nonphotosynthetic structures, rhizomes, and their ability to survive extended periods of low temperatures and produce shoots when the temperatures became warmer. It is possible that fat is beneficial for short-term cold tolerance, while being prejudicial to long term overwintering ability. In fact, if higher fat content allows a plant to continue to metabolize at lower temperatures, it may also cause the plant to use its stored energy during the winter months and deplete the plant of resources for regrowth the following year, therefore decreasing the rhizomes’ ability to successfully overwinter. Another possible explanation for our findings is that the ratio of different fatty acids may be more influential to cold tolerance than the total fat content [38, 40].
Similar to fat, starch concentrations were negatively correlated with overwintering. This is in agreement with several previous studies [24, 41, 42]. Lower starch concentrations could reflect the breakdown of starch into sugars, which may be necessary for cold acclimation [25] and would be expected to decrease the freezing potential of the cells.
Sugars, which were measured in our study using WSC (primarily glucose, sucrose, fructose, maltose, lactose, and fructans), were positively correlated with overwintering. Higher concentrations of sucrose have been found in alfalfa (Medicago sativa L.) plants that were selected for their overwintering ability [24] and similar trends were reported in centipedegrass, Eremochloa ophiruoides (Munro) Hackel [28]. Hoffman et al. [40] determined that the crowns of perennial ryegrass, Lolium perenne L., accumulated WSC when the temperature was as low as 2 °C. Others have reported that total carbohydrate accumulation in the crowns of some plant species increased freezing tolerance [43, 44]. From our findings, it appears that the presence of WSC in sorghum rhizomes may be a useful indicator in predicting the overwintering ability of a plant or genotype.
Fructans are one component of WSC, which are present in both cool and warm-season grasses [45]. ESC is a measurement of the same assimilates that are in WSC with the exception of fructans. Therefore, a determination and comparison of the carbohydrates in WSC and ESC can be used to calculate the amount of fructan present, which was significant in our study. This finding disagrees with what Pollock and Cairns [45] reported, but it suggests that genotypes that accumulate fructans may have a higher potential for over-wintering. Taken together, our data indicated that lower concentrations of fat and starch along with a higher concentration of WSC, ESC, and fructans were efficient indicators for rhizome overwintering ability within the College Station soil column environment.
Rhizome mean diameter was also significant and negatively correlated with spring regrowth when both locations were analyzed together. This was surprising, as we had expected larger rhizomes to have greater energy for regrowth. However, correlations showed that these larger rhizomes had more starch than smaller ones, which further suggests that composition plays more of a role in overwintering ability than size.
NIRS for Prediction of Overwintering-Related Rhizome Composition
NIRS analysis, if successful, could provide a less expensive and more rapid means of determining the composition of rhizomes than wet chemical analysis. Many NIRS models were fit for calibration in this study. The model that was eventually chosen incorporated all samples across all plots and locations and used Norris smoothing of five-unit segments and five-unit gaps. NIRS predictions for measured assimilates and performance index numbers (Table 6) indicate that NIRS is a robust and useful tool for predicting CP and starch as well as having potential for the prediction of WSC, ESC, and fat in sorghum rhizomes.
The NIRS model that we created will be a useful tool both for selecting winter-hardy parents in perennial sorghum breeding programs and for obtaining a better understanding of the assimilate fluxes that occur within sorghum rhizomes. In field experiments, winter freezing can often be excessively or insufficiently stringent to select a reasonable number of plants to advance, especially in the early stages of breeding programs. Selection based on composition profiles before freezing using NIRS could allow ideal selection intensity across environments. Because starch appears to be amenable to NIRS and displays the strongest correlation with spring regrowth in our most exposed environment, it would appear to be the best choice for assimilate selection. However, our data also indicate that starch concentrations are highly effected by environment and GxE (Table 2), making it less attractive than WSC and ESC. Perhaps the best, but also most expensive, approach would be to select for higher WSC and ESC while selecting for lower starch simultaneously. Another logical extension of this research would be to use a field-based NIRS system with a fiber-optic probe to take advantage of nondestructive screening of rhizomes in the field.
Conclusions
Many factors affect rhizome composition, but our findings indicate a feasibility of selecting S. halepense and Sorghum spp. genotypes with greater overwintering capacity based on lower autumn fat and starch concentrations and greater WSC, ESC, and fructan concentrations within rhizomes. We also conclude that NIRS can be used to assist in the selection of perennial parent plants with improved overwintering ability while minimizing the need for costly wet chemical analyses. Two genotypes, 09TX05 and 09TX06, preformed best in terms of overwintering and regrowth at all locations and may therefore be useful in the breeding of perennial sorghum to be used for biofuel feedstocks.
References
ICRISAT (2004) Data and information. http://www.icrisat.org/crop-sorghum.htm. 2011
Armah-Agyeman G, Loiland J, Karow R, Bean B (2002) Dryland cropping systems: Sudangrass. Oregon State Univ Ext Serv EM 8793:1–6
Pritchard AJ (1965) Inheritance of two seedling characters in Sorghum almum Parodi, and cross pollination within S. almum and between S. almum and S. halepense (L.) Pers. Aust J Agric Res 16:533–540
Rooney WL, Blumenthal J, Bean B, Mullet JE (2007) Designing sorghum as a dedicated bioenergy feedstock. Biofuels Bioprod Bioref 1:147–157
Wu X, Zhao R, Liu L, Bean S, Seib PA, Mclaren J, Madl R, Tuinstra M, Lenz M, Wang D (2008) Effects of growing location and irrigation on attributes and ethanol yields of selected grain sorghums. Cereal Chem 85:495–501
Miller AN, Ottman MJ (2010) Irrigation frequency effects on growth and ethanol yield in sweet sorghum. Agron J 102:60–70
Sattler SE, Funnell-Harris DL, Pedersen JF (2010) Efficacy of singular and stacked brown midrib 6 and 12 in the modification of lignocelluloses and grain chemistry. J Agric Food Chem 58:3611–3616
Murray SC, Sharma A, Rooney WL, Klein PE, Mullet JE, Mitchell SE, Kresovich S (2008) Genetic improvement of sorghum as a biofuel feedstock I: quantitative loci for stem sugar and grain nonstructural carbohydrates. Crop Sci 48:2165–2179
Chittenden LM, Schertz KF, Lin YR, Wing RA, Paterson AH (1994) A detailed RFLP map of Sorghum bicolor x S. propinquum, suitable for high-density mapping, suggests ancestral duplication of Sorghum chromosomes or chromosomal segments. Theor Appl Genet 87:925–933
Costanza R, d’arge R, de Groot R, Farber S, Grasso M, Hannon B, Limburg K, Naeem S, O’Neill RV, Paruelo J, Raskin RG, Sutton P, van den Belt M (1997) The value of the world’s ecosystem services and natural capital. Nature 387:253–260
Lewandowski I, Scurlock JMO, Lindvall E, Christou M (2003) The development and current status of perennial rhizomatous grasses as energy crops in the US and Europe. Biomass Bioenergy 25:335–361
Kort J, Collins M, Ditsch D (1998) A review of soil erosion potential associated with biomass crops. Biomass Bioenergy 14:351–359
McLaughlin SB, Walsh ME (1998) Evaluating environmental consequences of producing herbaceous crop for bioenergy. Biomass Bioenergy 14:317–324
Khanna M, Önal H, Dhungana B, Wander M (2010) Economics of herbaceous energy crops for electricity generation: Implications for greenhouse gas mitigation. Biomass Bioenergy 35:1474–1484
Major Uses of Land in the United States, 2007 (2007) United States Department of Agriculture/ Economic Research Service. http://www.ers.usda.gov/publications/eib-economic-information-bulletin/eib89.aspx
Paterson AH, Schertz KF, Lin YR, Liu SC, Chang YL (1995) The weediness of wild plants: molecular analysis of genes influencing dispersal and persistence of Johnsongrass, Sorghum halepense (L.) Pers. Proc Natl Acad Sci USA 92:6127–6131
Washburn JD, Murray SC, Burson BL, Klein RR, Jessup RW (2013) Targeted mapping of quantitative trait locus regions for rhizomatousness in chromosome SBI-01 and analysis of overwintering in a Sorghum bicolor × S. propinquum population. Mol Breeding 31:153–162. doi:10.1007/s11032-012-9778-8
McWhorter CG (1961) Morphology and development of Johnsongrass plants from seeds and rhizomes. Weeds 9:558–562
Monaghan N (1979) The biology of Johnson grass (Sorghum halepense). Weed Res 19:261–267. doi:10.1111/j.1365-3180.1979.tb01536.x
Warwick SI, Phillips D, Andrews C (1986) Rhizome depth: the critical factor in winter survival of Sorghum halepense (L.) Pers. (Johnson grass). Weed Res 26:381–387
Rapp KE (1948) Carbohydrate metabolism of Johnson Grass. J Am Soc Agron 40:869–873
McWhorter CG (1961) Carbohydrate metabolism of johnsongrass as influenced by seasonal growth and herbicide treatments. Weeds 9:563–568
McWhorter CG (1974) Water-soluble carbohydrates in johnsongrass. Weed Sci 22:159–163
Castonguay Y, Bertrand A, Michaud R, Laberge S (2011) Cold-induced biochemical and molecular changes in alfalfa populations selectively improved for freezing tolerance. Crop Sci 51:2132–2144. doi:10.2135/cropsci2011.02.0060
Yano R, Nakamura M, Yoneyama T, Nishida I (2005) Starch-related α-glucan/water dikinase is involved in the cold-induced development of freezing tolerance in Arabidopsis. Plant Physiol 138:837–846. doi:10.pp. 104.056374/pp.104.056374
Maier FP, Lang NS, Fry JD (1994) Freezing tolerance of three St. Augustinegrass cultivars as affected by stolon carbohydrate and water content. J Am Soc Hort Sci 119:473–476
Stoller EW, Weber EJ (1975) Differential cold tolerance, starch, sugar, protein, and lipid of yellow and purple nutsedge tubers. Plant Physiol 55:859–863. doi:10.1104/pp.55.5.859
Fry JD, Lang NS, Clifton RGP, Maier FP (1993) Freezing tolerance and carbohydrate content of low-temperature-acclimated and nonacclimated centipedegrass. Crop Sci 33:1051–1055
Murray SC (in press) Differentiation of seed, sugar, and biomass-producing genotypes in Saccharinae species. In: Paterson AH (ed) Genomics of the Saccharinae. Springer, Berlin
de Alencar Figueiredo LF, Davrieux F, Fliedel G, Rami JF, Chantereau J, Deu M, Courtois B, Mestres C (2006) Development of NIRS equations for food grain quality traits through exploitation of a core collection of cultivated sorghum. J Agric Food Chem 54:8501–8509
Murray SC, Rooney WL, Mitchell SE, Sharma A, Klein PE, Mullet JE, Kresovich S (2008) Genetic improvement of sorghum as a biofuel feedstock II: quantitative loci for stem and leaf structural carbohydrates. Crop Sci 48:2180–2193
Mitchell AR, Rechel EA, Dovel RL (1998) Three methods for determining storage carbohydrate concentration in peppermint (Mentha piperita) rhizomes. HortSci 33:754–756
Jessup RW, Whitmire DK, Farrow ZL, Burson BL (2012) Molecular characterization of non-flowering perennial Sorghum spp. hybrids. Am J Exp Ag 2:9–20
The SAS system for Windows, 9.1 edn (2003) SAS Institute, Cary
JMP®, 9 edn (2011) SAS Institute, Cary
Xiong S, Landstrom S, Olsson R (2009) Delayed harvest of reed canary grass translocates more nutrients in rhizomes. Acta Agr Scand 59:306–316
Miquel M, James D Jr, Dooner H, Browse J (1993) Arabidopsis requires polyunsaturated lipids for low-temperature survival. Proc Natl Acad Sci USA 90:6208–6212
Cyril J, Powell GL, Duncan RR, Baird WV (2002) Changes in membrane polar lipid fatty acids of seashore paspalum in response to low temperature exposure. Crop Sci 42:2031–2037
Stoller EW (1977) Differential cold tolerance of quackgrass and johnsongrass rhizomes. Weed Sci 25:348–351
Hoffman L, DaCosta M, Ebdon JS, Watkins E (2010) Physiological changes during cold acclimation of perennial ryegrass accessions differing in freeze tolerance. Crop Sci 50:1037–1047
Castonguay Y, Nadeau P, Lechasseur P, Chouinard L (1995) Differential accumulation of carbohydrates in alfalfa cultivars of contrasting winterhardiness. Crop Sci 35:509–516. doi:10.2135/cropsci1995.0011183X003500020038x
Cunningham SM, Volenec JJ (1998) Seasonal carbohydrate and nitrogen metabolism in roots of contrasting alfalfa (Medicago sativa L.) cultivars. J Plant Physiol 153:220–225. doi:10.1016/s0176-1617(98)80069-2
Livingston DP, Olien CR, Freed RD (1989) Sugar composition and freezing tolerance in barley crowns at varying carbohydrate levels. Crop Sci 29:1266–1270
Shahba MA, Qian YL, Hughes HG, Koski AJ, Christensen D (2003) Relationships of soluble carbohydrates and freeze tolerance in saltgrass. Crop Sci 43:2148–2153
Pollock CJ, Cairns AJ (1991) Fructan metabolism in grasses and cereals. Annu Rev Plant Physiol Plant Mol Biol 42:77–101
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Washburn, J.D., Whitmire, D.K., Murray, S.C. et al. Estimation of Rhizome Composition and Overwintering Ability in Perennial Sorghum spp. Using Near-Infrared Spectroscopy (NIRS). Bioenerg. Res. 6, 822–829 (2013). https://doi.org/10.1007/s12155-013-9305-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12155-013-9305-8