Abstract
Purpose
Persistent abnormal proliferation and long distant metastasis of tumors contribute to high mortality rate in non-small cell lung cancer (NSCLC) patients. Strategies that prevent NSCLC proliferation and/or metastasis have been studied but still need to be further explored. Numerous studies have proved the diversity functions of long noncoding RNAs (lncRNAs) exerted in cancer, including NSCLC. In this study, we aim to identify and investigate the role of novel lncRNAs in NSCLC progression.
Methods
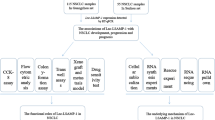
RNA sequence data were retrieved from the Cancer Genome Atlas (TCGA), differentially expressed lncRNAs (DElncRNAs) were screened out based on the R language, then real-time PCR experiment was introduced to detect the DElncRNA expression levels. A series of experiments including MTT, cell cycle, transwell, and wound healing assays were employed to explore the effect of DElncRNA MGC27382 on cell proliferation and invasion ability.
Results
We detected that DElncRNA MGC27382 is down-regulated in NSCLC tissues and cells. Overexpression of MGC27382 prevented NSCLC cell proliferation via down-regulating cyclin D1 and cyclin E. Moreover, wound healing and transwell assays indicated that the ability of cell invasion and migration could be impaired when cells were treated with MGC27382 overexpression. Further studies demonstrated that MGC27382-mediated inhibition on NSCLC progression can be impaired by LY294002, which is a frequently used inhibitor of AKT/GSK3β pathway.
Conclusion
MGC27382 is down-regulated in NSCLC. It exerts an inhibitory role in NSCLC development through suppressing the AKT/GSK3β pathway. Our results indicate that the lncRNA MGC27382 might be a tumor-suppressor gene in NSCLC. Overexpression of MGC27382 is thought to be a potential strategy for overcoming NSCLC progression.





Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Lemjabbar-Alaoui H, Hassan OU, Yang YW, et al. Lung cancer: biology and treatment options. Biochim Biophys Acta. 2015;1856:189–210.
Youlden DR, Cramb SM, Baade PD. The international epidemiology of lung cancer: geographical distribution and secular trends. J Thorac Oncol. 2008;3:819–31.
Mattick JS. The genetic signatures of noncoding RNAs. PLoS Genet. 2009;5: e1000459.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43:904–14.
Tsai MC, Manor O, Wan Y, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–93.
Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29:452–63.
Chen J, Wang R, Zhang K, et al. Long non-coding RNAs in non-small cell lung cancer as biomarkers and therapeutic targets. J Cell Mol Med. 2014;18:2425–36.
Nie W, Ge HJ, Yang XQ, et al. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016;371:99–106.
Wan L, Sun M, Liu GJ, et al. Long noncoding RNA PVT1 promotes non-small cell lung cancer cell proliferation through epigenetically regulating LATS2 expression. Mol Cancer Ther. 2016;15:1082–94.
Lu W, Zhang H, Niu Y, et al. Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial mesenchymal transition by sponging miR-150-5p. Mol Cancer. 2017;16:118.
Wang H, Niu L, Jiang S, et al. Comprehensive analysis of aberrantly expressed profiles of lncRNAs and miRNAs with associated ceRNA network in muscle-invasive bladder cancer. Oncotarget. 2016;7:86174–85.
Niu L, Deng J, Zhu F, et al. Anti-inflammatory effect of Marchantin M contributes to sensitization of prostate cancer cells to docetaxel. Cancer Lett. 2014;348:126–34.
Wang W, He X, Zheng Z, et al. Serum HOTAIR as a novel diagnostic biomarker for esophageal squamous cell carcinoma. Mol Cancer. 2017;16:75.
Wu D, Li Y, Zhang H, et al. Knockdown of Lncrna PVT1 enhances radiosensitivity in non-small cell lung cancer by sponging Mir-195. Cell Physiol Biochem. 2017;42:2453–66.
Chen W, Zhu H, Yin L, et al. lncRNA-PVT1 facilitates invasion through upregulation of MMP9 in nonsmall cell lung cancer cell. DNA Cell Biol. 2017;36:787–93.
Merchant N, Nagaraju GP, Rajitha B, et al. Matrix metalloproteinases: their functional role in lung cancer. Carcinogenesis. 2017;38:766–80.
Chen WF, Gao WD, Li QL, et al. SLIT2 inhibits cell migration in colorectal cancer through the AKT-GSK3beta signaling pathway. Int J Colorectal Dis. 2013;28:933–40.
Gao F, Huang W, Zhang Y, et al. Hes1 promotes cell proliferation and migration by activating Bmi-1 and PTEN/Akt/GSK3beta pathway in human colon cancer. Oncotarget. 2015;6:38667–80.
Lu KH, Li W, Liu XH, et al. Long non-coding RNA MEG3 inhibits NSCLC cells proliferation and induces apoptosis by affecting p53 expression. BMC Cancer. 2013;13:461.
Sun M, Liu XH, Wang KM, et al. Downregulation of BRAF activated non-coding RNA is associated with poor prognosis for non-small cell lung cancer and promotes metastasis by affecting epithelial-mesenchymal transition. Mol Cancer. 2014;13:68.
Liu Z, Sun M, Lu K, et al. The long noncoding RNA HOTAIR contributes to cisplatin resistance of human lung adenocarcinoma cells via downregualtion of p21(WAF1/CIP1) expression. PLoS ONE. 2013;8: e77293.
Weber DG, Johnen G, Casjens S, et al. Evaluation of long noncoding RNA MALAT1 as a candidate blood-based biomarker for the diagnosis of non-small cell lung cancer. BMC Res Notes. 2013;6:518.
Han L, Kong R, Yin DD, et al. Low expression of long noncoding RNA GAS6-AS1 predicts a poor prognosis in patients with NSCLC. Med Oncol. 2013;30:694.
Tseng RC, Lee SH, Hsu HS, et al. SLIT2 attenuation during lung cancer progression deregulates beta-catenin and E-cadherin and associates with poor prognosis. Cancer Res. 2010;70:543–51.
Liao K, Li J, Wang Z. Dihydroartemisinin inhibits cell proliferation via AKT/GSK3beta/cyclinD1 pathway and induces apoptosis in A549 lung cancer cells. Int J Clin Exp Pathol. 2014;7:8684–91.
Liu XH, Liu ZL, Sun M, et al. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. BMC Cancer. 2013;13:464.
Yang YR, Zang SZ, Zhong CL, et al. Increased expression of the lncRNA PVT1 promotes tumorigenesis in non-small cell lung cancer. Int J Clin Exp Pathol. 2014;7:6929–35.
Cui D, Yu CH, Liu M, et al. Long non-coding RNA PVT1 as a novel biomarker for diagnosis and prognosis of non-small cell lung cancer. Tumour Biol. 2016;37:4127–34.
Loewen G, Jayawickramarajah J, Zhuo Y, et al. Functions of lncRNA HOTAIR in lung cancer. J Hematol Oncol. 2014;7:90.
Liao M, Li B, Zhang S, et al. Relationship between LINC00341 expression and cancer prognosis. Oncotarget. 2017;8:15283–93.
Acknowledgements
This work was supported by the China Postdoctoral Science Foundation (No. 2018M630786) and the National Natural Science Foundation of China (No. 82002540).
Author information
Authors and Affiliations
Contributions
QL designed the study and drafted the manuscript. LLN, SLY, HMN, and SL were responsible for the collection and analysis of the experimental data. CCL revised the manuscript critically for important intellectual content. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study was approved by the Ethical Committee of Qilu Hospital of Shandong University and conducted in accordance with the ethical standards.
Informed consent
Yes.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Q., Li, S., Niu, L. et al. Long noncoding RNA MGC27382 inhibits proliferation and metastasis of non-small cell lung cancer cells via down-regulating AKT/GSK3β pathway. Clin Transl Oncol 23, 2548–2559 (2021). https://doi.org/10.1007/s12094-021-02658-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-021-02658-x