Abstract
Lafora disease (LD) is a fatal childhood-onset dementia characterized by the extensive accumulation of glycogen aggregates—the so-called Lafora Bodies (LBs)—in several organs. The accumulation of LBs in the brain underlies the neurological phenotype of the disease. LBs are composed of abnormal glycogen and various associated proteins, including p62, an autophagy adaptor that participates in the aggregation and clearance of misfolded proteins. To study the role of p62 in the formation of LBs and its participation in the pathology of LD, we generated a mouse model of the disease (malinKO) lacking p62. Deletion of p62 prevented LB accumulation in skeletal muscle and cardiac tissue. In the brain, the absence of p62 altered LB morphology and increased susceptibility to epilepsy. These results demonstrate that p62 participates in the formation of LBs and suggest that the sequestration of abnormal glycogen into LBs is a protective mechanism through which it reduces the deleterious consequences of its accumulation in the brain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Glycogen, a branched polymer of glucose, is found in most tissues but is particularly abundant in the liver and muscle [1]. In the brain, glycogen is present mainly in astrocytes [2,3,4], although neurons also have an active glycogen metabolism that contributes to their function [5, 6]. In mammals, glycogen is synthesized by glycogen synthase and degraded by glycogen phosphorylase. The muscular isoform of glycogen synthase (MGS) is expressed in most tissues, including the brain.
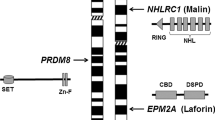
Progressive myoclonic epilepsy type 2 (EPM2, OMIM #254,780) or Lafora disease (LD) is an autosomal recessive disease characterized by severe and progressive myoclonus epilepsy, and neurodegeneration rapidly progressing to dementia and death within 5–10 years after the onset [7, 8]. LD is caused by mutations in either the EPM2A gene, which encodes laforin, a dual specificity phosphatase with a carbohydrate-binding domain, or EPM2B (also NHLRC1), which encodes malin, an E3-ubiquitin ligase. The histopathological and clinical outcomes of LD patients and mouse models of LD carrying mutations in either of these two genes are very similar, thereby indicating that malin and laforin participate in the same physiological process. The hallmark of the disease is the accumulation of cytoplasmic aggregates of poorly branched glycogen called Lafora bodies (LBs) in several tissues [9, 10]. In the brain, LBs are found in astrocytes and neurons [11,12,13]. Neuronal LBs (nLBs) typically manifest as single, large, round and juxtanuclear aggregates, while astrocytic LBs are smaller and amorphous and have a granular distribution throughout astrocytic processes [12]. We refer to these astrocytic LBs as CAL (corpora amylacea-like), since they are morphologically similar to corpora amylacea—glycogen aggregates that accumulate in normal aging [8, 14]. Blocking or reducing brain glycogen synthesis in LD mouse models prevents the progression of the disease [13, 15,16,17,18], thereby indicating that glycogen accumulation underlies the pathophysiology of LD. Furthermore, forced accumulation of glycogen in neurons leads to neuronal loss [19] while in astrocytes it induces neuroinflammation [13].
In addition to glycogen, LBs contain a number of proteins, including laforin (in malin-deficient LD), enzymes involved in glycogen metabolism such as MGS, ubiquitinated proteins, and the autophagy adaptor p62 [17, 18, 20]. The presence of ubiquitin and p62 suggests that, like other insoluble molecular aggregates characteristic of neurodegenerative diseases, LBs could be targets for autophagic clearance [17, 20,21,22,23]. In this regard, the mechanisms that drive the formation and clearance of LBs have not been identified yet. Since p62 has been shown to aggregate polyubiquitinated proteins [24], it could play a similar role in the formation of LBs. Furthermore, although glycogen accumulation underlies LD pathogenesis, it remains to be determined whether the sequestration of this polysaccharide into LBs is protective (to minimize the toxic consequences of the accumulation of abnormal glycogen) or pathogenic (LBs themselves being the toxic species). Finally, the accumulation of p62 per se is deleterious for neurons [25, 26] and other cell types [27, 28]. In this regard, p62 depletion clears nuclear inclusion bodies and increases lifespan in a model of Huntington’s disease [29].
Given all of the above, p62 may exert a neuroprotective or a neurotoxic function in the context of LD. To study the contribution of p62 to LB formation and to the pathophysiology of LD, we generated a malin knockout mouse (malinKO) [30] devoid of p62 (malinKO + p62KO). Our results demonstrate that p62 is essential for LB formation in skeletal muscle and cardiac tissue. In the brain, p62 is also involved in the formation of these aggregates. When this protein is absent in the brain, neuroinflammation is mildly enhanced and susceptibility to epilepsy is exacerbated. These observations identify p62 as a key player in the cellular protective response against glycogen aggregates.
Materials and Methods
Animal Studies
All procedures were approved by the Barcelona Science Park’s Animal Experimentation Committee and were carried out following Spanish (BOE 34/11370–421, 2013) and European Union (2010/63/EU) regulations, and The National Institutes of Health guidelines for the care and use of laboratory animals. For the generation of the malinKO + p62KO model, malinKO mice [30] were crossed with p62KO animals [23]. After weaning at 3 weeks of age, tail clippings were taken for genotyping by qPCR (performed by TransnetYX). Experiments were conducted using littermates, and males and females were included in each group. Mice were maintained on a 12/12 h light/dark cycle under specific pathogen-free conditions in the Animal Research Center (Barcelona Science Park) and allowed free access to a standard chow diet and water.
Glycogen Quantification
Mice were deeply anesthetized and decapitated. Whole brains and quadriceps were quickly removed, frozen, and pulverized in liquid nitrogen. For glycogen measurements, frozen tissue aliquots were boiled in 30% KOH for 15 min and glycogen was precipitated in 60% ethanol and then determined by an amyloglucosidase-based assay [5].
Western Blot
For western blot, lysates of frozen tissue aliquots were prepared using the following buffer: 25 mM Tris–HCl (pH 7.4), 25 mM NaCl, 1% Triton X-100, 0.1% SDS, 0.5 mM EGTA, 10 mM sodium pyrophosphate, 1 mM sodium orthovanadate, 10 mM NaF, 25 nM okadaic acid and a protease inhibitor cocktail tablet (Roche). Soluble and insoluble fractions of total homogenates were obtained as previously described [30]. Briefly, total homogenates were centrifuged at 13,000 rpm for 15 min at 4 °C. The pellet containing the insoluble fraction was resuspended in the same volume as the supernatant corresponding to the soluble fraction. Samples were loaded on 10% acrylamide gels for SDS-PAGE and transferred to Immobilon membranes (Millipore). The following primary antibodies were used: anti-MGS (3886, Cell Signaling); anti-laforin (3.5.5, kindly provided by Dr. Santiago Rodríguez de Córdoba); and anti-p62 (GP62-C, Progen). The following secondary antibodies were used: anti-rabbit and anti-mouse IgG-HRP (GE Healthcare); and anti-guinea pig HRP (Jackson Immuno Research). Proteins were detected by the ECL method (Immobilon Western Chemiluminescent HRP Substrate, Millipore), and loading control of the western blot membrane was performed using the Revert total protein stain (LI-COR Bioscience).
Histology and Immunohistochemistry
Animals were deeply anesthetized and perfused transcardially with phosphate-buffered saline (PBS) containing 4% paraformaldehyde (PBS 4% PFA). Brains, skeletal muscles and hearts were removed, post-fixed overnight with PBS 4% PFA and embedded in paraffin blocks. Periodic acid-Schiff staining (PAS) was performed using an Artisanlink Pro machine (AR16511-2 kit, Dako-Agilent).
For immunohistochemistry, 3-μm paraffin-embedded tissue sections were either dewaxed and subjected to antigen retrieval treatment with Tris–EDTA buffer pH 9 for 20 min at 97ºC using a PT Link (Dako – Agilent) or dewaxed as part of the antigen retrieval process using the Low pH EnVision™ FLEX Target Retrieval Solutions (K8005, Dako-Agilent) for 20 min at 97ºC using a PT Link (Dako – Agilent). Endogenous peroxidase was quenched with Peroxidase-Blocking Solution (S2023, Dako-Agilent). Non-specific binding was blocked using 5% of normal goat serum (16,210,064, Life technology) with 2.5% BSA (10.735.078.001, Sigma) for 60 min. Also, unspecific endogenous mouse Ig staining was blocked using the Mouse on mouse (M.O.M) Immunodetection Kit (BMK-2202, Vector Laboratories). Primary mouse IgG1 anti-GFAP (MAB360, Merck Millipore) and rabbit pAb 1 anti-IBA1 (019–19,741, WAKO) antibodies were diluted at 1:250 and 1:1000 respectively with EnVision FLEX Antibody Diluent (K800621, Dako-Agilent) and incubated overnight at 4ºC. Tissue sections were then incubated for 45 min with Polyclonal Anti-Mouse 1:100 (P0447, Dako-Agilent) or a BrightVision Poly-HRP-Anti Rabbit IgG, RTU (Immunologic, DPVR-110HRP). Antigen–antibody complexes were revealed with 3–3′-diaminobenzidine (K3468, Dako). Sections were counterstained with hematoxylin (Dako, S202084) and mounted with Toluene-Free Mounting Medium (CS705, Dako) using a Dako CoverStainer.
For immunofluorescence, endogenous peroxidase was quenched by 10 min of incubation with Peroxidase-Blocking Solution (S2023, Dako-Agilent). Non-specific binding was blocked using 5% of normal goat serum (16.210.064, Life technology) with 2.5% BSA (10.735.078.001, Sigma) for 60 min. Also, unspecific endogenous mouse Ig staining was blocked using the M.O.M Immunodetection Kit (BMK-2202, Vector Laboratories). Primary antibodies anti-MGS (15B1) (1:250, 3886 Cell Signaling), anti-GFAP (1:250, MAB360, Merck Millipore), anti-βIII-Tubulin (1:500, T86660, Sigma Aldrich), and C3d antibody (1: 100, AF2655, R&D) were incubated overnight at 4ºC.
The following secondary antibodies were used: an Alexa Fluor® 488 anti-mouse IgG (405,319, BioLegend); Alexa Fluor® 488 anti-mouse IgG1 (A21121, ThermoFisher); Alexa Fluor 568® anti-mouse IgG2b (A21144, ThermoFisher); DyLight 594 anti-rabbit (DI1094, VectorLabs); or an Alexa Fluor 647® anti-rabbit IgG (A32733, ThermoFisher), diluted at 1:500 and incubated for 60 min. Samples were stained with DAPI (D9542, Sigma) and mounted with fluorescence mounting medium (S3023 Dako). Specificity of staining was respectively confirmed by staining with rabbit IgG, polyclonal Isotype Control (ab27478, Abcam), mouse IgG1, Kappa Monoclonal (NCG01) Isotype Control (ab81032, Abcam), or a mouse IgM (PFR-03) Isotype Control (A1-10,438, ThermoFisher).
Brightfield and fluorescent images were acquired with a NanoZoomer- 2.0 HT C9600 digital scanner (Hamamatsu) equipped with a 20 × objective. For super-resolution microscopy, images were acquired in a Zeiss 880 confocal microscope equipped with Fast Airyscan and a piezo-stage. A 63 × magnification 1.40 NA oil-immersion lens with a digital zoom of 1.5 × was used. The Z-step between the stacks was set at 0.8 μm. Fast Airyscan raw data were pre-processed with the automatic setting of Zen Black.
The stainings were analyzed by the digital software analysis package QuPath [31]. For detection of the morphological features of nLBs and CAL, MGS-positive granules were identified using the Cell detection plugin (QuPath). Intensity thresholds were set for GFAP and βIII-Tubulin in the surrounding area of each MGS-positive granule. Morphology and intensity data were then exported and plotted with RStudio [32].
RT-qPCR Analyses
Total RNA of pulverized brains was prepared with the RNeasy Micro Kit (Qiagen), following the manufacturer’s instructions. Single-stranded complementary DNA was produced by reverse transcription using 1 µg of DNA-free RNA in a 20-µL reaction qScript cDNA SuperMix (Quanta bio). Quantitative polymerase chain reaction (PCR) was performed using SYBR green (Quanta bio) on the QuantStudio 6 Flex as per the manufacturer’s instructions. The ΔCt was defined as the difference between the Q-PCR cycles of the housekeeping gene and those of the target genes.
Assessment of Kainate-Induced Epilepsy
Mice were weighed and placed in individual cages to prevent contact between animals, which could startle them. They were then administered three consecutives intraperitoneal (i.p.) injections (6 mg/kg body weight) of the glutamate agonist kainic acid (KA) (Sigma) dissolved in 0.1 M PBS pH 7.4, in order to induce non‐lethal convulsive seizures. Seizure intensity after KA injections was evaluated as described previously [33,34,35] for 240 min from the first KA administration. After the first KA injections, the animals developed hypoactivity and immobility (Grade I–II). After successive injections, hyperactivity (Grade III) and scratching with mild non-convulsive seizures (Grade IV) were often observed. Some animals progressed to a whole-body convulsive seizure with loss of balance control (Grade V). Extreme behavioral manifestations such as uncontrolled hopping activity or “popcorn behavior” (blinking seizure), as well as continuous or chronic seizures (> 1’ without body movement control) were included in Grade VI. All behavioral assessments were performed blind to the experimental group (genotype) in situ and were also recorded and reanalyzed blind to the first analysis.
Statistical Analysis
Two-group hypothesis testing was evaluated using an independent sample t-test performed with the GraphPad Prism software (La Jolla, CA, USA). Two-way analysis of variation (ANOVA) was used for comparing three or more groups. Data are represented as mean ± standard error of the mean (SEM). When indicated, linear mixed-effects model was used as follows:
nLB and CAL morphology. For each morphological parameter independently, linear mixed effect models were fitted with the R package lmerTest [36] using the parameters response variable, cell type, and the interaction between cell type and genotype as covariates of interest, the experimental group and the position as adjusting factors, and both the Mouse ID and the interaction between the Mouse ID and cell type as random effect to account for non-independence among data from the same mice. Transformations: for parameter values between 0 and 1, a logit transformation was applied. Otherwise, for parameter values larger than zero, a log transformation was considered.
Q-PCR analysis at gene level. For every gene, independently, linear mixed-effects models were fitted with the R package lmerTest using the ΔCt as response variable, Genotype as covariate of interest, experimental group as adjusting factor and Mouse ID as random effect to account for the variability of technical replicates. Adjustment for multiple testing (single-step correction method) was performed using the R package multcomp [37].
Q-PCR analysis for pro- and anti-inflammatory differences. For each mouse, ΔCt levels for the measured replicates were averaged out. For each gene, the mouse group (batch) effect was also balanced out and, finally, standardization was applied to these balanced data. The average value of each genotyping condition is shown in the graphs.
To evaluate consistency between gene patterns across inflammatory groups, a linear mixed-effects model was fitted with the R package lmerTest [36] using the interaction between the genotype and the inflammatory group as fixed effects and the gene Id as random effect. Adjustment for multiple testing (single-step correction method) was performed using the R package multcomp [37].
Results
p62 Progressively Accumulates with Age in the Brain and Skeletal Muscle of MalinKO Mice
We previously described increased levels of p62 in 11-month-old malinKO brains [17, 18]. The accumulated p62 is bound to LBs, both CAL and nLBs [12]. To further characterize p62 accumulation in malinKO tissues, we performed immunofluorescence against this protein. We observed an increase in p62 in the cortex and the hippocampus of 4- and 11-month-old malinKO mice (Fig. 1A), which paralleled the accumulation of LBs. Indeed, the number of p62-positive aggregates more than doubled in the brains of 11-month-old animals compared to those aged 4 months (Fig. 1B). A massive increase in p62-positive aggregates was detected in the skeletal muscles of 11-month-old malinKO animals compared to 4-month-old counterparts (Fig. 1A, C).
Accumulation of p62 aggregates over time in malinKO mice. a Representative images of the progressive accumulation of p62 in the cortex and hippocampus of malinKO mice. DAPI: 4 ',6-diamidino-2-fenilindol; WGA: wheat germ agglutinin. Scale bar: 50 µm. b–c Quantification of the number of p62-positive LBs per area (n of particles/mm2) in the prefrontal cortex or hippocampus (B) and skeletal muscle (C). n = 7–12 mice. For comparisons between groups, one-way ANOVA was performed using Prism7 software (GraphPad). * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001, ∗ ∗ ∗ ∗ P < 0.0001. Adjusted p-values (cortex): control 4 m vs. control 11 m P = 0.9997; control 4 m vs. malinKO 4 m P = 0.7191; control 4 m vs. malinKO 11 m P = 0.0010; control 11 m vs. malinKO 4 m P = 0.7829; control 11 m vs. malinKO 11 m P = 0.0013; and malinKO 4 m vs. malinKO 11 m P = 0.0001. Adjusted p-values (hippocampus): control 4 m vs. control 11 m P > 0.9999; control 4 m vs. malinKO 4 m P = 0.9929; control 4 m vs. malinKO 11 m P = 0.0109; control 11 m vs. malinKO 4 m P = 0.9936; control 11 m vs. malinKO 11 m P = 0.0111; and malinKO 4 m vs. malinKO 11 m P = 0.0005. Adjusted p-values (skeletal muscle): control 4 m vs. control 11 m P = 0.9556; control 4 m vs. malinKO 4 m P = 0.5429; control 4 m vs. malinKO 11 m P = 0.0020; control 11 m vs. malinKO 4 m P = 0.7784; control 11 m vs. malinKO 11 m P = 0.0014; and malinKO 4 m vs. malinKO 11 m P = 0.0041
p62 Is Essential for LB Formation in Muscle and Heart Tissue But Not In the Brain
To evaluate the impact of p62 deletion on LB formation and LD progression, we generated malinKO mice devoid of p62 (malinKO + p62KO). The presence of LBs was visualized by periodic acid-Schiff staining (PAS), which specifically stains carbohydrates, and by immunofluorescence using anti-MGS antibody, since MGS is attached to LBs and can thus be used as an LB marker [13, 18].
The 11-month-old mice malinKO animals showed abundant PAS-positive LBs in all the tissues analyzed, i.e., skeletal muscle, heart, and brain, as previously described [17, 18, 30] (Fig. 2A, Supplemental Fig. 1A). Strikingly, the skeletal muscles of malinKO + p62KO mice were devoid of PAS-positive aggregates (Fig. 2A) and instead showed a diffused pattern of PAS-positive material. MGS immunostaining confirmed the absence of glycogen aggregates in the skeletal muscle of malinKO + p62KO mice (Fig. 2B). Similar results were observed in cardiac tissue, where deletion of p62 resulted in the absence of PAS-positive and MGS-positive aggregates (Supplemental Figs. 1A, B). Although devoid of LBs, malinKO + p62KO skeletal muscles showed an increase in total glycogen, as determined by biochemical quantification, similar to that seen in malinKO muscles (Fig. 2D). These results indicate that the skeletal muscles of malinKO + p62KO mice also accumulated glycogen, although not in the form of LBs.
p62 deletion prevents glycogen aggregation in skeletal muscle but not in brain. A. Histological localization of LBs. Periodic acid-Schiff staining (PAS) is shown for the indicated tissues of 11-month-old malinKO and malinKO + p62KO mice. Scale bar = 25 \(\mu\) m (skeletal muscle and heart), 200 \(\mu\) m (brain). Brain aggregates are visible in the insets. B-C. Representative immunostaining of p62 and MGS in skeletal muscle (quadriceps) and brain (cortex). Scale bar = 25 \(\mu\) m. D. Glycogen content. For comparisons between groups, one-way ANOVA followed by Holm's Multiple Comparisons Test was performed using Prism7 software (GraphPad). Results are presented as the group mean ± SEM. (n = 6 animals per group). * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001, ∗ ∗ ∗ ∗ P < 0.0001. Adjusted p-values. Brain glycogen: control vs. p62KO p = 0.4593; control vs. malinKO p < 0.0001; control vs. malinKO + p62KO p < 0.0001; and malinKO vs. malinKO + p62KO p = 0.3052. Muscle glycogen: control vs. p62KO p = 0.3238; control vs. malinKO p = 0.0358; control vs. malinKO + p62KO * p < 0.05; and malinKO vs. malinKO + p62KO p = 0.8546
In contrast, the brains of these mice showed PAS-positive aggregates (Fig. 2A). As expected, these aggregates contained MGS but not p62 (Fig. 2C). Total glycogen levels were similarly elevated in malinKO + p62KO and malinKO brains (Fig. 2D).
Accumulation of Insoluble MGS and Laforin Is Prevented in the Skeletal Muscle of MalinKO + p62KO Animals But Not in the Brain
Using western blot of total tissue homogenates, we next studied the content of proteins known to accumulate in malinKO tissues. Since LBs are insoluble aggregates that precipitate under low-speed centrifugation [30], we also examined the distribution of these proteins between the soluble and insoluble fractions, the latter corresponding to the LB-enriched fraction. Western blots confirmed the absence of p62 in tissues from p62KO and malinKO + p62KO mice. In the skeletal muscles of malinKO mice, MGS, laforin and p62 were significantly increased in the insoluble fraction (Fig. 3A, B), as previously reported [18]. However, in malinKO + p62KO skeletal muscles, MGS and laforin showed levels similar in the insoluble fraction to those of controls (Fig. 3A, B). These results were consistent with the absence of LBs in malinKO + p62KO skeletal muscles.
p62 deletion rescues accumulation of insoluble glycogen-bound proteins MGS and laforin in skeletal muscle but not in brain. A-C. Western blotting for MGS, p62 and laforin in muscle (A) and brain (C). Total protein was used as loading control. B-D. Densitometry of the western blots in skeletal muscle (B) and brain (D). For comparisons between groups, one-way ANOVA followed by Holm's Multiple Comparisons Test was performed. N = 6–9 animals/group were analyzed. Results are presented as the group mean ± SEM. * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001, ∗ ∗ ∗ ∗ P < 0.0001. Adjusted p-values for MGS (a) or laforin (b): Brain whole lysates: control vs. p62KO p = 0.057 (a), p = 0.3489 (b); control vs. malinKO p < 0.0001 (a) p < 0.0001 (b); control vs. malinKO + p62KO p = 0.0066 (a), p < 0.0001 (b); and malinKO vs. malinKO + p62KO p = 0.057 (a), p = 0.0898 (b). Brain soluble: control vs. p62KO p = 0.0143 (a), p = 0.8557 (b); control vs. malinKO p = 0.0041 (a), p = 0.7614 (b); control vs. malinKO + p62KO p = 0.0041 (a), p = 0.7614 (b); and malinKO vs. malinKO + p62KO p = 0.9128 (a), p = 0.8737 (b). Brain insoluble: control vs. p62KO p = 0.9649 (a), p = 0.9179 (b); control vs. malinKO p = 0.0013 (a), p = 0.0001 (b); control vs. malinKO + p62KO p = 0.0001 (a), p = 0.0075 (b); and malinKO vs. malinKO + p62KO p = 0.5051 (a), p = 0.1591 (b). Muscle whole lysate: control vs. p62KO p = 0.8217 (a), p = 0.7297 (b); control vs. malinKO p = 0.8217 (a), p = 0.0919 (b); control vs. malinKO + p62KO p = 0.8217 (a). p = 0.7297 (b); and malinKO vs. malinKO + p62KO p = 0.9765 (a), p = 0.1987 (b). Muscle soluble: control vs. p62KO p = 0.9597 (a), p = 0.0573 (b); control vs. malinKO p = 0.1964 (a), p = 0.0135 (b); control vs. malinKO + p62KO p = 0.1543 (a), p = 0.9727 (b); and malinKO vs. malinKO + p62KO p = 0.9597 (a), p = 0.0135 (b). Muscle insoluble: control vs. p62KO p = 0.9966 (a), p = 0.6862 (b); control vs. malinKO p = 0.0003 (a), p < 0.0001 (b); control vs. malinKO + p62KO p = 0.7838 (a), p = 0.6862 (b); and malinKO vs. malinKO + p62KO p = 0.0013 (a) p < 0.0001 (b)
In brain total homogenates, malinKO mice showed an increase in MGS and laforin, which corresponded to an increase in the insoluble fraction (Figs. 3c, d), as we previously described [18, 30]. In contrast to skeletal muscles, the brains of malinKO + p62KO mice showed similar increases in these two proteins in the insoluble fraction, consistent with the presence of LBs in this tissue.
p62 Depletion Alters the Morphology of Brain LBs
Although LBs were still present in malinKO + p62KO brains, we studied the morphology of these aggregates formed in the absence of p62. Super-resolution microscopy revealed that nLBs in malinKO + p62KO animals appeared less dense and more irregular than the typical round, compact nLBs found in malinKO brains (Fig. 4A). The morphology of astrocytic LBs (CAL), which are inherently more heterogeneous than nLBs [12] was indistinguishable between the two genotypes, both showing highly irregular shapes (Fig. 4B).
p62 deletion disrupts nLB but not CAL morphology. A-B. Representative examples of reconstructed super-resolution images of neuronal LBs (A) and astrocytic CAL (B) are shown. PFA-fixed tissue sections were incubated with anti-glycogen synthase (MGS), anti-GFAP and βIII-tubulin antibodies, nuclei were stained with Hoechst 33,342. Scale bar: 10 \(\mu\) m. C-D. Quantifications of circularity (left panels), eccentricity (middle panels) and max caliper (right panels) in neuronal LBs (C) and astrocytic CAL (D). Linear mixed model analysis was performed for statistical analysis. N = 6 animals per genotype. Adjusted p-values Circularity: astrocytic p = 0.075; neuronal p < 0.0001. Eccentricity: astrocytic p = 0.726; neuronal p < 0.0001. Max caliper: astrocytic p = 0.516; neuronal p = 0.017
We further characterized the morphology of nLBs and CAL by studying regularity parameters of MGS-stained granules in βIII-tubulin-positive/GFAP-negative areas (corresponding to nLBs) and GFAP-positive/βIII-tubulin-negative areas (corresponding to CAL). MalinKO + p62KO nLBs showed changes in circularity, eccentricity, and maximal caliper with respect malinKO nLBs (Fig. 4C, Supplementary Fig. 2). No significant differences in CAL morphology were detected between malinKO + p62KO and malinKO brains (Fig. 4D, Supplementary Fig. 2). Taken together, these results indicate that p62 is necessary for the correct packing of nLBs.
MalinKO + p62KO Mice Present Neuroinflammation
Astrogliosis, microgliosis, and increased expression of genes related to neuroinflammation are characteristic traits of malinKO brains [13, 17, 18, 38]. Given the importance of p62 in inflammatory responses [39], we studied the impact of p62 deletion on these processes. GFAP and IBA1 immunostainings showed an increase in reactive astrocytes and microglia in the hippocampi of malinKO mice (Fig. 5A, B), as previously described [13, 17, 18]. MalinKO + p62KO mice showed similar GFAP and IBA1 stainings (Fig. 5A, B), thereby indicating that the deletion of p62 alone did not significantly alter the number of reactive astrocytes or microglia. We also examined the presence of A1 astrocytes, a subset of reactive neurotoxic astrocytes whose cytoplasm accumulates the inflammatory component protein C3 [40]. MalinKO + p62KO mice showed a significant increase in reactive C3-positive astrocytes compared to control animals (Fig. 5C, D), although they did not show a significant difference with malinKO mice.
Neuroinflammation in malinKO and malinKO + p62KO mice. A. Representative immunostaining of GFAP and IBA1 in the hippocampus of the indicated experimental groups. B. Densitometry analysis of GFAP and IBA1 areas normalized to the hippocampal area. For comparisons between groups, one-way ANOVA followed by Holm's Multiple Comparisons Test was performed using Prism7 software (GraphPad). Results are presented as the group mean ± SEM. n = 5–6 animals per group. * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001. Adjusted p-values. GFAP in hippocampus: control vs. p62KO p = 0.9562; control vs. malinKO p = 0.0055; control vs. malinKO + p62KO p < 0.0028; and malinKO vs. malinKO + p62KO p = 0.7931. GFAP in cortex: control vs. p62KO p = 0.5388; control vs. malinKO p = 0.5388; control vs. malinKO + p62KO p = 0.1823; and malinKO vs. malinKO + p62KO p = 0.5388. IBA1 in hippocampus: control vs. p62KO p = 0.9719; control vs. malinKO p = 0.0175; control vs. malinKO + p62KO p = 0.0175; and malinKO vs. malinKO + p62KO p = 0.9705. IBA1 in cortex: control vs. p62KO p = 0.9620; control vs. malinKO p = 0.8390; control vs. malinKO + p62KO p = 0. 8390; and malinKO vs. malinKO + p62KO p = 0.9620 C. Representative super-resolution images of the immunofluorescence staining against C3d and GFAP in the hippocampus of a MalinKO mouse. One A1 reactive astrocyte is indicated in the arrohead. D. Quantification of the percentage of GFAP-positive cells that are C3-positive in the hippocampus. N = 7–9 mice per group. * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001. Adjusted p-values. control vs. p62KO p > 0.9999; control vs. malinKO p = 0.2539; control vs. malinKO + p62KO p = 0.0053; and malinKO vs. malinKO + p62KO p = 0.3513. E. Heat map of pro- and anti-inflammatory transcripts quantified by qPCR normalized to the housekeeper gene GAPDH. N = 6 animals per group. Linear mixed-effects models were fitted using ΔCt as response variable, the genotype as covariate of interest, the mouse experimental group replicate as adjusting factor and the mouse Id as random effect to account for variability in technical replicates. Comparisons were done independently for each gene measured. * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001. Raw and adjusted p-values are summarized in Supplementary file 3. F. dot plots of the average Z-score of the genes shown in panel A in the indicated experimental groups. Gray lines are indicative of one gene. Colored lines are the average of gray lines. The table shows the linear mixed effect model that quantified consistency in ΔCts between genes of the same inflammatory group, across the 4 genotypes. * P < 0.05, ∗ ∗ P < 0.01, ∗ ∗ ∗ P < 0.001.
We next examined the transcriptional profiles of cytokines and other mediators of the immune response associated with activated microglia. MalinKO and malinKO + p62KO mice showed a similar increase, with respect to control mice, in the expression of IL1-α, TNF-α, and C1q; cytokines; and key mediators of A1 astrocyte activation [40] (Fig. 5E). We observed a similar result in other inflammation-associated genes, including LCN2, CXCL10, CCL12, and CCL2 [38, 41]. The expression of genes involved in suppressing inflammation, like S100a10, IL10, and Arg-1A, was significantly increased in malinKO mice (Fig. 5E). Overall, malinKO mice showed the upregulation of both anti- and pro-inflammation genes, while malinKO + p62KO animals had a more marked increase of pro-inflammatory cytokines (IL12, CCL12, CD14, CCL2 and C3), and a reduced expression of the anti-inflammatory molecule Arg-1 and anti-inflammatory cytokines such as IL4 and IL13 (Fig. 5E). Next, we compared normalized ΔCt patterns across the four genotypes for gene signatures defining pro- and anti-inflammatory activity. Interestingly, malinKO + p62KO mice showed higher expression of the pro-inflammatory signature than malinKO mice, whereas this trend was reversed for the anti-inflammatory signature, with malinKO mice showing greater expression than malinKO + p62KO counterparts. This pattern was consistent for most of the genes that defined the two signatures (Fig. 5F, G). Overall, malinKO and malinKO + p62KO mice showed a similar inflammatory response, albeit modestly exacerbated in the latter.
Deletion of p62 Enhances the Epileptic Phenotype of MalinKO Mice
MalinKO mice present increased susceptibility to kainate-induced epilepsy [17, 30], a finding consistent with one of the main symptoms of LD patients. We have proposed that this pathological trait is due to the accumulation of LBs in neurons [13, 30]. Therefore, we next aimed to examine whether the change in nLB morphology in malinKO + p62KO brains was translated into a worsening of the epileptic phenotype. To that end, 5-month-old mice received three consecutive kainate injections (6 mg/kg, i.p. every 30 min) and were video-recorded for 240 min after the first injection to monitor behavior (i.e., epileptic events). MalinKO and malinKO + p62KO animals showed a significant decrease in the average onset time of the first seizure (Fig. 6A) and an increase in the number of seizures compared to control mice (Fig. 6B). Furthermore, malinKO + p62KO animals showed the highest number of all kinds of seizures per animal, and the percentage of mice that reached the most severe stage (VI) was significantly higher in this group (Fig. 6B). While mice from the four genotypes reached all severity stages, only in the malinKO and malinKO + p62KO groups all the mice reached stage IV and showed an increased proportion of mice reaching stages V and VI (Fig. 6C). Regarding the time spent in each stage, again malinKO + p62KO mice showed the highest time spent in severe stages (Fig. 6D) and the lowest in mild stages (I-III, not shown). MalinKO and malinKO + p62KO animals spent more time in stage IV, the latter group being statistically higher than the former and the only one to show significantly greater time spent in stage V when compared to control animals (Fig. 6D). In summary, the increased susceptibility to kainate-induced epilepsy of malinKO mice was further increased in malinKO + p62KO mice.
p62 deletion increases seizure susceptibility in malinKO mice. A. Onset of epileptic activity in minutes. Data are expressed as average ± SEM. N = 6 mice/group (*p < 0.05) indicate significant difference (One-way ANOVA followed by Bonferroni test; p = 0.0.0173). B. Number of seizures (mild, convulsive, or blinking) experienced per animal. Data are expressed as average ± SEM. N = 6 mice/group. Two-way ANOVA (Seizure type factor: p = 0.0003; Genotype factor: p = 0.0012; interaction: p = 0.0022). (*p < 0.05, **p < 0.01, ***p < 0.005) indicate significant difference (Kruskal–Wallis for each seizure type followed by Dunn’s test); (#p < 0.05) indicate pairwise differences between control and malinKO by Mann–Whitney U-test. C. Percentage of mice reaching seizure stages I to VI. D. Time spent in each stage (expressed in minutes) during the course of the experiment. Data are expressed as average ± SEM. N = 6 mice/group. Two-way ANOVA (Stage factor (P < 0.0001), the Genotype factor (P < 0.0001), Interaction (P < 0.0001)). (**p < 0.01, ***p < 0.005, ****p < 0.001) indicate significant pairwise differences (post-hoc Bonferroni)
Discussion
The high levels of p62 in malinKO brains [17, 18, 20] made us consider the possibility that its accumulation per se contributes to the etiopathology of LD. In this regard, it is worth noting that p62 interacts with many factors that play key roles in determining cell fate [26]. The accumulation of p62 can lead to overactivation of oxidative stress responses through the Nrf2/Keap1 pathway [42] or can co-operate with other disease-associated proteins to induce cellular toxicity [43]. Conversely, p62 depletion clears nuclear inclusion bodies and increases lifespan in a model of Huntington’s disease [29]. Strikingly, our results indicate that the accumulation of p62 per se does not underlie the etiopathology of LD, as malinKO + p62KO mice are not rescued from the characteristic pathological traits of LD but instead present exacerbated susceptibility to epilepsy.
p62 participates in the autophagic removal of protein aggregates in a process known as aggrephagy [44]. p62 contains multiple protein-interaction domains, including a ubiquitin-binding domain and an oligomerization domain, which allow it to bind to and aggregate polyubiquitinated proteins into less harmful inclusions [45]. The presence of p62 in LBs led us to hypothesize that glycogen aggregation in LD follows a similar pattern. Importantly, LBs were absent in the skeletal muscles and hearts of malinKO + p62KO mice, thereby revealing that the abnormal glycogen that is formed in LD does not aggregate by itself, as it has been generally assumed, since p62 is essential for the formation of LBs in these tissues. These results have important implications for other diseases in which glycogen aggregates accumulate in skeletal muscle and heart, including Andersen’s disease (OMIM 232,500) [46], Cori’s disease (OMIM 232,400) [47], Tarui disease (OMIM 232,800) [48], polyglucosan body myopathy-1 (OMIM 615,895) [49], and polyglucosan body myopathy-2 (OMIM 616,199) [50]. In contrast, the brains of these animals still contained LBs, indicating that the process of LB formation is tissue-specific. In this regard, it is worth noting that aggrephagy genes appear to be differentially used in a tissue-specific manner [44]. Functional redundancy of autophagy receptors [51, 52] could explain the presence of LBs in malinKO + p62KO brains, as other receptors might compensate for the absence of p62 to trigger glycogen aggregation in this tissue.
However, super-resolution analysis revealed that p62 deletion results in more irregular, less round, less compact aggregates in neurons, thereby confirming the involvement of p62 in LB formation also in this cell type. In this regard, it is interesting to note that protein aggregates rich in p62 (p62 bodies) have liquid-like properties (high sphericity) and can undergo fusion events [53]. Therefore, it is conceivable that the deletion of p62 blocks the liquid-like properties (roundness) and the fusion of insoluble glycogen aggregates into larger droplets. We have proposed that the epileptic phenotype of LD is due to the accumulation of abnormal glycogen in neurons [13, 30]. Our results show that the change in the morphology of nLBs in malinKO + p62KO mice is accompanied by an increase in susceptibility to kainate-induced epilepsy, thereby corroborating that the proper sequestration of abnormal glycogen into nLBs is essential to minimize its toxic effects in neurons.
Although p62 is present both in CAL and nLBs in the brains of malinKO mice [12], analysis of the number and morphology of astrocytic CAL did not show significant differences in malinKO + p62KO mice. However, the morphology of CAL is inherently heterogeneous [12], which could hamper the detection of changes in the parameters studied. Thus, we cannot discard that the deletion of p62 also affected CAL formation. We have recently demonstrated that the accumulation of CAL in astrocytes underlies neuroinflammation in LD [13]. Accordingly, the cytokine inflammatory program was sustained and even switched toward a potential exacerbation of the neuroinflammation in malinKO + p62KO mice. These results confirm the key role of astrocyte-driven inflammation in the pathophysiology of LD.
A longstanding question in LD is whether LBs are the toxic species themselves or whether they are formed to minimize the detrimental consequences of the accumulation of abnormal glycogen, by sequestering it into less harmful aggregates. In this regard, there is wide consensus that in proteinopathies, early stages of aggregation are responsible for cellular toxicity and neurodegeneration [54,55,56]. MalinKO + p62KO mice, in which the formation of brain LBs is altered, offer the opportunity to study intermediate states of LB formation. We observed increased susceptibility to kainate-induced epilepsy in malinKO + p62KO mice, which supports the hypothesis that immature glycogen aggregates are more harmful than mature LBs. On the basis of all these considerations and our findings, we propose a scenario in which LBs play a similar role as neurodegeneration-associated protein inclusion bodies [55, 57, 58]. Poorly branched glycogen, which cannot be degraded by glycogen phosphorylase, would be formed as a side-product of glycogen metabolism. The malin/laforin complex would serve to prevent the formation of this abnormal glycogen. Thus, in the absence of laforin or malin, it would accumulate. In this context, p62 (together with other autophagy adaptors in the case of the brain) would promote its aggregation into LBs in order to minimize the deleterious consequences of aberrant glycogen accumulation in neurons [19] and possibly in astrocytes [13] (Fig. 7).
Graphical summary of the study. Abnormal glycogen is produced as a side-product of glycogen metabolism. The malin/laforin complex prevents the formation of this abnormal polysaccharide. In the absence of laforin or malin, abnormal glycogen accumulates. In this context, p62 promotes its aggregation into LBs, to minimize the toxic consequences of its accumulation. p62 deletion correlates with an exacerbation of the inflammation and epilepsy supporting the notion that LBs are neuroprotective
Several recent articles explore possible therapies for LD based on replacing the missing genes, inhibiting glycogen synthesis or introducing enzymes that can digest LBs [18, 59,60,61,62]. Moreover, beyond LD, we have witnessed an increasing body of literature showing glycogen accumulation as a new common thread in aging, neurodegenerative diseases (including Alzheimer’s, Parkinson’s and Huntington’s disease and amyotrophic lateral sclerosis [63,64,65]) and epilepsy [18, 66,67,68,69]. Moreover, the accumulation of corpora amylacea, p62 and aggregation-prone proteins in the brain correlates with aging and neurodegeneration (8, 14, 70, 71). The accumulation of glycogen in these conditions could play an active role in disrupting cell homeostasis, causing neuroinflammation and epilepsy, as we have shown for LD. Therefore, glycogen synthesis emerges as a potential therapeutic target for aging and other neurological diseases.
Conclusions
Our study shows that p62 is necessary for LB formation in muscle and heart and that it is also important for the correct formation of LBs in the brain. The deletion of p62 worsens the susceptibility to epilepsy of LD. Our results provide an unprecedented description of a protective role of p62-directed sequestration of abnormal glycogen in LD.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- Arg-1:
-
Arginase-1
- C1q:
-
Complement component 1q
- C3:
-
Complement component 3
- CA:
-
Corpora amylacea
- CAL:
-
Corpora amylacea-like bodies
- CCL2:
-
Chemokine (C–C motif) ligand 2
- CD14:
-
Cluster of differentiation 14
- CXCL10:
-
C-X-C motif chemokine ligand 10
- CCL12:
-
Chemokine ligand 12
- EPM2a:
-
Epilepsy, Progressive Myoclonus Type 2A
- EPM2b:
-
Epilepsy, Progressive Myoclonus Type 2b
- GFAP:
-
Glial fibrillary acidic protein
- IBA1:
-
Ionized calcium-binding adapter molecule 1
- IL1- α:
-
Interleukin 1 α
- IL10:
-
Interleukin 10
- IL12:
-
Interleukin 12
- IL13:
-
Interleukin 13
- IL4:
-
Interleukin 4
- KA:
-
Kainic acid
- LBs:
-
Lafora bodies
- LCN2:
-
Lipocalin-2
- LD:
-
Lafora disease
- MGS:
-
Muscle glycogen synthase
- NHLRC1:
-
NHL Repeat Containing E3 Ubiquitin Protein Ligase 1
- nLBs:
-
Neuronal Lafora bodies
- PAS:
-
Periodic acid–Schiff
- S100A10:
-
S100 calcium-binding protein A10
- TNF-α:
-
Tumor necrosis factor α
References
Kanungo S, Wells K, Tribett T, El-Gharbawy A (2018) Glycogen metabolism and glycogen storage disorders. Annals of translational medicine 6(24):474
Cataldo AM, Broadwell RD (1986) Cytochemical identification of cerebral glycogen and glucose-6-phosphatase activity under normal and experimental conditions. II. Choroid plexus and ependymal epithelia, endothelia and pericytes. J Neurocytol 15(4):511–24
Duran J, Guinovart JJ (2015) Brain glycogen in health and disease. Mol Aspects Med 46:70–77
Duran J, Gruart A, López-Ramos JC, Delgado-García JM, Guinovart JJ (2019) Glycogen in Astrocytes and Neurons: Physiological and Pathological Aspects. Adv Neurobiol 23:311–329
Saez I, Duran J, Sinadinos C, Beltran A, Yanes O, Tevy MF et al (2014) Neurons have an active glycogen metabolism that contributes to tolerance to hypoxia. Journal of cerebral blood flow and metabolism : official journal of the International Society of Cerebral Blood Flow and Metabolism 34(6):945–955
Duran J, Gruart A, Varea O, López-Soldado I, Delgado-García JM, Guinovart JJ (2019) Lack of Neuronal Glycogen Impairs Memory Formation and Learning-Dependent Synaptic Plasticity in Mice. Front Cell Neurosci 13:374
Gentry MS, Guinovart JJ, Minassian BA, Roach PJ, Serratosa JM (2018) Lafora disease offers a unique window into neuronal glycogen metabolism. J Biol Chem 293(19):7117–7125
Cavanagh JB (1999) Corpora-amylacea and the family of polyglucosan diseases. Brain Res Brain Res Rev 29(2–3):265–295
Sakai M, Austin J, Witmer F, Trueb L (1970) Studies in myoclonus epilepsy (Lafora body form). II. Polyglucosans in the systemic deposits of myoclonus epilepsy and in corpora amylacea. Neurology. 20(2):160–76
Monaghan TS, Delanty N (2010) Lafora disease: epidemiology, pathophysiology and management. CNS Drugs 24(7):549–561
Rubio-Villena C, Viana R, Bonet J, Garcia-Gimeno MA, Casado M, Heredia M et al (2018) Astrocytes: new players in progressive myoclonus epilepsy of Lafora type. Hum Mol Genet 27(7):1290–1300
Augé E, Pelegrí C, Manich G, Cabezón I, Guinovart JJ, Duran J et al (2018) Astrocytes and neurons produce distinct types of polyglucosan bodies in Lafora disease. Glia 66(10):2094–2107
Duran J, Hervera A, Markussen KH, Varea O, López-Soldado I, Sun RC et al (2021) Astrocytic glycogen accumulation drives the pathophysiology of neurodegeneration in Lafora disease. Brain 144(8):2349–2360
Augé E, Duran J, Guinovart JJ, Pelegrí C, Vilaplana J (2018) Exploring the elusive composition of corpora amylacea of human brain. Sci Rep 8(1):13525
Turnbull J, DePaoli-Roach AA, Zhao X, Cortez MA, Pencea N, Tiberia E et al (2011) PTG depletion removes Lafora bodies and rescues the fatal epilepsy of Lafora disease. PLoS genetics. 7(4):e1002037
Pederson BA, Turnbull J, Epp JR, Weaver SA, Zhao X, Pencea N et al (2013) Inhibiting glycogen synthesis prevents Lafora disease in a mouse model. Ann Neurol 74(2):297–300
Duran J, Gruart A, Garcia-Rocha M, Delgado-Garcia JM, Guinovart JJ (2014) Glycogen accumulation underlies neurodegeneration and autophagy impairment in Lafora disease. Hum Mol Genet 23(12):3147–3156
Varea O, Duran J, Aguilera M, Prats N, Guinovart JJ (2021) Suppression of glycogen synthesis as a treatment for Lafora disease: Establishing the window of opportunity. Neurobiology of Disease 147:105173
Duran J, Tevy MF, Garcia-Rocha M, Calbó J, Milán M, Guinovart JJ (2012) Deleterious effects of neuronal accumulation of glycogen in flies and mice. EMBO Mol Med 4(8):719–729
Criado O, Aguado C, Gayarre J, Duran-Trio L, Garcia-Cabrero AM, Vernia S et al (2012) Lafora bodies and neurological defects in malin-deficient mice correlate with impaired autophagy. Hum Mol Genet 21(7):1521–1533
Deng Z, Purtell K, Lachance V, Wold MS, Chen S, Yue Z (2017) Autophagy Receptors and Neurodegenerative Diseases. Trends Cell Biol 27(7):491–504
Puri R, Suzuki T, Yamakawa K, Ganesh S (2012) Dysfunctions in endosomal-lysosomal and autophagy pathways underlie neuropathology in a mouse model for Lafora disease. Hum Mol Genet 21(1):175–184
Komatsu M, Waguri S, Koike M, Sou YS, Ueno T, Hara T et al (2007) Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131(6):1149–1163
Moscat J, Diaz-Meco MT (2009) To aggregate or not to aggregate? A new role for p62. EMBO Rep 10(8):804
Jackson KL, Lin WL, Miriyala S, Dayton RD, Panchatcharam M, McCarthy KJ et al (2017) p62 Pathology Model in the Rat Substantia Nigra with Filamentous Inclusions and Progressive Neurodegeneration. PloS one. 12(1):e0169291
Emanuele S, Lauricella M, D'Anneo A, Carlisi D, De Blasio A, Di Liberto D et al (2020) p62: Friend or Foe? Evidences for oncoJanus and neuroJanus roles. International Journal of Molecular Sciences 21(14):5029
Rusten TE, Stenmark H (2010) p62, an autophagy hero or culprit? Nat Cell Biol 12(3):207–209
Kenific CM, Debnath J (2015) Cellular and metabolic functions for autophagy in cancer cells. Trends Cell Biol 25(1):37–45
Kurosawa M, Matsumoto G, Kino Y, Okuno M, Kurosawa-Yamada M, Washizu C et al (2015) Depletion of p62 reduces nuclear inclusions and paradoxically ameliorates disease phenotypes in Huntington’s model mice. Hum Mol Genet 24(4):1092–1105
Valles-Ortega J, Duran J, Garcia-Rocha M, Bosch C, Saez I, Pujadas L et al (2011) Neurodegeneration and functional impairments associated with glycogen synthase accumulation in a mouse model of Lafora disease. EMBO Mol Med 3(11):667–681
Bankhead P, Loughrey MB, Fernández JA, Dombrowski Y, McArt DG, Dunne PD et al (2017) QuPath: Open source software for digital pathology image analysis. Sci Rep 7(1):16878
Team R (2020) RStudio: Integrated Development for R. RStudio, PBC, Boston, MA [Available from: http://www.rstudio.com/
Rangel A, Madroñal N, Gruart A, Gavín R, Llorens F, Sumoy L et al (2009) Regulation of GABA(A) and glutamate receptor expression, synaptic facilitation and long-term potentiation in the hippocampus of prion mutant mice. PloS one. 4(10):e7592
Rangel A, Burgaya F, Gavín R, Soriano E, Aguzzi A, Del Río JA (2007) Enhanced susceptibility of Prnp-deficient mice to kainate-induced seizures, neuronal apoptosis, and death: Role of AMPA/kainate receptors. J Neurosci Res 85(12):2741–2755
Carulla P, Bribián A, Rangel A, Gavín R, Ferrer I, Caelles C et al (2011) Neuroprotective role of PrPC against kainate-induced epileptic seizures and cell death depends on the modulation of JNK3 activation by GluR6/7-PSD-95 binding. Mol Biol Cell 22(17):3041–3054
Kuznetsova A, Brockhoff PB, Christensen RHB (2017) lmerTest Package: Tests in Linear Mixed Effects Models 2017 82(13):26
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom J Biometrische Zeitschrift 50(3):346–363
Duran J, Brewer MK, Hervera A, Gruart A, Del Rio JA, Delgado-García JM et al (2020) Lack of Astrocytic Glycogen Alters Synaptic Plasticity but Not Seizure Susceptibility. Mol Neurobiol 57(11):4657–4666
Schimmack G, Schorpp K, Kutzner K, Gehring T, Brenke JK, Hadian K et al (2017) YOD1/TRAF6 association balances p62-dependent IL-1 signaling to NF-κB. eLife 6:e22416
Liddelow SA, Guttenplan KA, Clarke LE, Bennett FC, Bohlen CJ, Schirmer L et al (2017) Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541(7638):481–487
Lahuerta M, Gonzalez D, Aguado C, Fathinajafabadi A, García-Giménez JL, Moreno-Estellés M et al (2020) Reactive Glia-Derived Neuroinflammation: a Novel Hallmark in Lafora Progressive Myoclonus Epilepsy That Progresses with Age. Mol Neurobiol 57(3):1607–1621
Komatsu M, Kurokawa H, Waguri S, Taguchi K, Kobayashi A, Ichimura Y et al (2010) The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nat Cell Biol 12(3):213–223
Kalogeropulou AF, Zhao J, Bolliger MF, Memou A, Narasimha S, Molitor TP et al (2018) P62/SQSTM1 is a novel leucine-rich repeat kinase 2 (LRRK2) substrate that enhances neuronal toxicity. Biochem J 475(7):1271–1293
Zheng H, Yuan C, Zhang H, Chen Y (2020) The tissue- and developmental stage-specific involvement of autophagy genes in aggrephagy. Autophagy 16(4):589–599
Yao TP (2010) The role of ubiquitin in autophagy-dependent protein aggregate processing. Genes Cancer 1(7):779–786
Bruno C, Cassandrini D, Assereto S, Akman HO, Minetti C, Di Mauro S (2007) Neuromuscular forms of glycogen branching enzyme deficiency. Acta Myol 26(1):75–78
Lucchiari S, Santoro D, Pagliarani S, Comi GP (2007) Clinical, biochemical and genetic features of glycogen debranching enzyme deficiency. Acta Myol 26(1):72–74
Raben N, Sherman JB (1995) Mutations in muscle phosphofructokinase gene. Hum Mutat 6(1):1–6
Nilsson J, Schoser B, Laforet P, Kalev O, Lindberg C, Romero NB et al (2013) Polyglucosan body myopathy caused by defective ubiquitin ligase RBCK1. Ann Neurol 74(6):914–919
Malfatti E, Nilsson J, Hedberg-Oldfors C, Hernandez-Lain A, Michel F, Dominguez-Gonzalez C et al (2014) A new muscle glycogen storage disease associated with glycogenin-1 deficiency. Ann Neurol 76(6):891–898
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL et al (2015) The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524(7565):309–314
Sánchez-Martín P, Saito T, Komatsu M (2019) p62/SQSTM1: “Jack of all trades” in health and cancer. FEBS J 286(1):8–23
Sun D, Wu R, Zheng J, Li P, Yu L (2018) Polyubiquitin chain-induced p62 phase separation drives autophagic cargo segregation. Cell Res 28(4):405–415
Bucciantini M, Giannoni E, Chiti F, Baroni F, Formigli L, Zurdo J et al (2002) Inherent toxicity of aggregates implies a common mechanism for protein misfolding diseases. Nature 416(6880):507–511
Arrasate M, Mitra S, Schweitzer ES, Segal MR, Finkbeiner S (2004) Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431(7010):805–810
Haass C, Selkoe DJ (2007) Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer’s amyloid beta-peptide. Nat Rev Mol Cell Biol 8(2):101–112
Mahul-Mellier AL, Burtscher J, Maharjan N, Weerens L, Croisier M, Kuttler F et al (2020) The process of Lewy body formation, rather than simply α-synuclein fibrillization, is one of the major drivers of neurodegeneration. Proc Natl Acad Sci USA 117(9):4971–4982
Ramdzan YM, Trubetskov MM, Ormsby AR, Newcombe EA, Sui X, Tobin MJ et al (2017) Huntingtin Inclusions Trigger Cellular Quiescence, Deactivate Apoptosis, and Lead to Delayed Necrosis. Cell Rep 19(5):919–927
Ahonen S, Nitschke S, Grossman TR, Kordasiewicz H, Wang P, Zhao X et al (2021) Gys1 antisense therapy rescues neuropathological bases of murine Lafora disease. Brain 144(10):2985–2993
Gumusgoz E, Guisso DR, Kasiri S, Wu J, Dear M, Verhalen B et al (2021) Targeting Gys1 with AAV-SaCas9 Decreases Pathogenic Polyglucosan Bodies and Neuroinflammation in Adult Polyglucosan Body and Lafora Disease Mouse Models. Neurotherapeutics 18(2):1414–1425
Nitschke S, Chown EE, Zhao X, Gabrielian S, Petković S, Guisso DR et al (2021) An inducible glycogen synthase-1 knockout halts but does not reverse Lafora disease progression in mice. The Journal of biological chemistry. 296:100150
Brewer MK, Uittenbogaard A, Austin GL, Segvich DM, DePaoli-Roach A, Roach PJ et al (2019) Targeting Pathogenic Lafora Bodies in Lafora Disease Using an Antibody-Enzyme Fusion. Cell Metab 30(4):689-705.e6
Huang L, Hollingsworth RI, Castellani R, Zipser B (2004) Accumulation of high-molecular-weight amylose in Alzheimer’s disease brains. Glycobiology 14(5):409–416
Dodge JC, Treleaven CM, Fidler JA, Tamsett TJ, Bao C, Searles M et al (2013) Metabolic signatures of amyotrophic lateral sclerosis reveal insights into disease pathogenesis. Proc Natl Acad Sci USA 110(26):10812–10817
Rai A, Singh PK, Singh V, Kumar V, Mishra R, Thakur AK et al (2018) Glycogen synthase protects neurons from cytotoxicity of mutant huntingtin by enhancing the autophagy flux. Cell Death Dis 9(2):201
Dalsgaard MK, Madsen FF, Secher NH, Laursen H, Quistorff B (2007) High glycogen levels in the hippocampus of patients with epilepsy. Journal of cerebral blood flow and metabolism : official journal of the International Society of Cerebral Blood Flow and Metabolism 27(6):1137–1141
Streichenberger N, Ryvlin P, Guénot M, Sindou M, Kopp N, Mauguière F (2001) Polyglucosan bodies and temporal lobe epilepsy: an incidental finding or more? Clin Neuropathol 20(4):172–175
Abubakr A, Wambacq I, Donahue JE, Zappulla R (2005) The presence of polyglucosan bodies in temporal lobe epilepsy: its role and significance. J Clin Neurosci 12(8):911–914
Cloix JF, Hévor T (2009) Epilepsy, regulation of brain energy metabolism and neurotransmission. Curr Med Chem 16(7):841–853
Rohn TT (2015) Corpora amylacea in neurodegenerative diseases: cause or effect? International Journal of Neurology and Neurotherapy 2(3):031
Sinadinos C, Valles-Ortega J, Boulan L, Solsona E, Tevy MF, Marquez M et al (2014) Neuronal glycogen synthesis contributes to physiological aging. Aging Cell 13(5):935–945
Acknowledgements
We thank Anna Adrover and Vanessa Fernandez for technical assistance and IRB Barcelona’s Advanced Digital Microscopy Facility, Histopathology Facility and Bioinformatics and Biostatistics Facility for technical support. Thanks also go to Nikos Giakoumakis for his kind support with super-resolution microscopy and Adrià Caballé for support with statistical analysis. Tanya Yates for correcting the English manuscript. p62KO mice were kindly provided by Dr. Maria Sonegas (Spanish National Cancer Research Centre-CNIO, Madrid, Spain) with permission from Prof. Tetsuro Ishii (University of Tsukuba, Japan).
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This study was supported by grants from the Spanish Ministry of Science, Innovation, and Universities (MCIU/FEDER/AEI) (BFU2017-84345-P to JJG and JD and PID2020-118699 GB-I00 to JD), the CIBER de Diabetes y Enfermedades Metabólicas Asociadas (ISCIII, Ministerio de Ciencia e Innovación), and a grant from the National Institutes of Health (NIH NINDS P01NS097197) to JJG and JD. This research was supported by PRPSEM Project with ref. RTI2018-099773-B-I00 from (MCIU/FEDER/AEI), the CERCA Program, and the Commission for Universities and Research of the Department of Innovation, Universities, and Enterprise of the Generalitat de Catalunya (SGR2017-648) to JADR. IRB Barcelona and IBEC are the recipients of a Severo Ochoa Award of Excellence from MINECO (Government of Spain). AH was supported by the Severo Ochoa Program at IBEC. MKB is a recipient of a fellowship of the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement No. 75451.
Author information
Authors and Affiliations
Contributions
PP and JD designed and performed the main experiments and analyzed the data. AH and JADR conducted the in vivo experiments. OV, ILS, MKB, AG, MA, and NP conducted additional experiments. PP wrote the manuscript. JD and JJG designed the project and edited the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All procedures were approved by the Barcelona Science Park’s Animal Experimentation Committee and were carried out in accordance with the European Community Council Directive and National Institutes of Health guidelines for the care and use of laboratory animals.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (PDF 3492 KB)
Supplementary Figure 1. p62 deletion prevents glycogen aggregation in heart. A. Histological localization of LBs in heart. Periodic acid-Schiff staining (PAS) is shown for the indicated tissues of 11-month-old malinKO and 11-month-old malinKO+p62KO littermates. Scale bar=25 µm. B. Representative immunostaining of p62 and MGS in heart. Scale bar=25 µm Supplementary Figure 2. Additional morphology parameters of neuronal LBs and astrocytic CAL. Quantifications of the indicated morphological parameters in neuronal LBs and astrocytic CAL. A linear mixed model analysis was performed for statistical analysis. N=6 animals per genotype. Adjusted p-values. Area: astrocytic p=0.657; neuronal p=0.269. Perimeter: astrocytic p=0.952; neuronal p=0.592. MGS mean intensity: astrocytic p=0.771; neuronal p=0.388. Centroid Xm: astrocytic p=0.392; neuronal p=0.959. Centroid Ym: astrocytic p=0.646; neuronal p=0.809. Min caliper: astrocytic p=1; neuronal p=0.162. Supplementary File 3. Raw and adjusted p-values of the pairwise comparisons in Figure 7A. Supplementary File 4. SYBR green primers used in Figure 7.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pellegrini, P., Hervera, A., Varea, O. et al. Lack of p62 Impairs Glycogen Aggregation and Exacerbates Pathology in a Mouse Model of Myoclonic Epilepsy of Lafora. Mol Neurobiol 59, 1214–1229 (2022). https://doi.org/10.1007/s12035-021-02682-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02682-6