Abstract
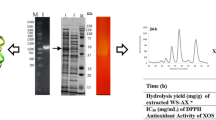
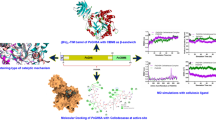
The filamentous fungus Sclerotinia sclerotiorum produces a complete set of cellulolytic enzymes. We report here the purification and the biochemical characterization of a new β-glucosidase from S. sclerotiorum which belongs to the family 3 of glycoside hydrolases and that was named as SsBgl3. After two size-exclusion chromatography steps, purified protein bands of 80 and 90 kDa from SDS-PAGE were subjected to a mass spectrometry analysis. The results displayed four peptides from the upper band belonging to a polypeptide of 777 amino acids having a calculated molecular weight of 83.7 kDa. Biochemical analysis has been carried out to determine some properties. We showed that this SsBgl3 protein displayed both β-glucosidase and exoglucanase activities with optimal activity at 55 °C and at pH 5. The transglycosylation activity was investigated using gluco-oligosaccharides TLC analysis. The molecular modeling and comparison with different crystal structures of β-glucosidases showed that SsBgl3 putative protein present three domains. They correspond to an (α/β)8 domain TIM barrel, a five-stranded α/β sandwich domain (both of which are important for active-site organization), and a C-terminal fibronectin type III domain. Enzyme engineering will be soon investigated to identify the key residues for the catalytic reactions.





Similar content being viewed by others
References
Gruno, M., Valjamae, P., Pettersson, G., & Johansson, G. (2004). Inhibition of the Trichoderma reesei cellulases by cellobiose is strongly dependent on the nature of the substrate. Biotechnology and Bioengineering, 86, 503–511.
Kaur, J., Chadha, B. S., Kumar, B. A., Kaur, G., & Saini, H. S. (2007). Purification and characterization of ß-glucosidase from Melanocarpus sp. MTCC 3922. Electronic Journal of Biotechnology, 10(2), 260–270.
Wilson, D. B. (2009). Cellulases and biofuels. Current Opinion in Biotechnology, 20, 295–299.
Sukumaran, R. K., Surender, V. J., Sindhu, R., Binod, P., et al. (2010). Lignocellulosic ethanol in India: Prospects, challenges and feedstock availability. Bioresource Technology, 101, 4826–4833.
Vlasenko, E., Schulein, M., Cherry, J., & Xu, F. (2010). Substrate specificity of family 5, 6, 7, 9, 12, and 45 endoglucanases. Bioresource Technology, 101, 2405–2411.
Bhatia, Y., Mishra, S., & Bisaria, V. S. (2002). Microbial beta-glucosidases: Cloning, properties, and applications. Critical Reviews in Biotechnology, 22, 375–407.
Shaikh, F. A., & Withers, S. G. (2008). Teaching old enzymes new tricks: Engineering and evolution of glycosidases and glycosyl transferases for improved glycoside synthesis. Biochemistry and Cell Biology, 86, 169–177.
Saibi, W., Amouri, B., & Gargouri, A. (2007). Purification and biochemical characterization of a transglucosilating β-glucosidase of Stachybotrys strain. Applied Microbiology and Biotechnology, 77(2), 293–300.
Smaali, I., Maugard, T., Limam, F., Legoy, M. D., & Marzouki, N. (2007). Efficient synthesis of gluco-oligosaccharides and alkyl-glucosides by transglycosylation activity of β-glucosidase from Sclerotinia sclerotiorum. World Journal of Microbiology & Biotechnology, 23(1), 145–149.
Han, Y., & Chen, H. (2008). Characterization of beta-glucosidase from corn stover and its application in simultaneous saccharification and fermentation. Bioresource Technology, 99, 6081–6087.
Jeya, M., Joo, A. R., Lee, K. M., Tiwari, M. K., et al. (2010). Characterization of beta-glucosidase from a strain of Penicillium purpurogenum KJS506. Applied Microbiology and Biotechnology, 86, 1473–1484.
Pontoh, J., & Low, N. H. (2002). Purification and characterization of beta-glucosidase from honey bees (Apis mellifera). Insect Biochemistry and Molecular Biology, 32, 679–690.
Saloheimo, M., Nakari-Setala, T., Tenkanen, M., & Penttila, M. (1997). cDNA cloning of a Trichoderma reesei cellulase and demonstration of endoglucanase activity by expression in yeast. European Journal of Biochemistry/FEBS, 249, 584–591.
Sorensen, A., Lubeck, M., Lubeck, P. S., & Ahring, B. K. (2013). Fungal beta-glucosidases: A bottleneck in industrial use of lignocellulosic materials. Biomolecules, 3, 612–631.
Varghese, J. N., Hrmova, M., & Fincher, G. B. (1999). Three-dimensional structure of a barley beta-d-glucan exohydrolase, a family 3 glycosyl hydrolase. Structure, 7, 179–190.
Pozzo, T., Pasten, J. L., Karlsson, E. N., & Logan, D. T. (2010). Structural and functional analyses of beta-glucosidase 3B from Thermotoga neapolitana: A thermostable three-domain representative of glycoside hydrolase 3. Journal of Molecular Biology, 397, 724–739.
Kentaro, S., Jun-Ichi, S., Young-Woo, N., Toru, N., Shuji, T., Takayoshi, W., et al. (2013). Crystal structures of glycoside hydrolase family 3 beta-glucosidase 1 from Aspergillus aculeatus. Biochemical Journal, 452(2), 211–221.
Mandels, M., & Weber, J. (1969). The production of cellulases. Advances in Chemistry Series, 95, 391–414.
Deshpande, M. V., Eriksson, K. E., & Pettersson, L. G. (1984). An assay for selective determination of exo-1,4,-beta-glucanases in a mixture of cellulolytic enzymes. Analytical Biochemistry, 138, 481–487.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Laemmli, U. K., Molbert, E., Showe, M., & Kellenberger, E. (1970). Form-determining function of the genes required for the assembly of the head of bacteriophage T4. Journal of Molecular Biology, 49, 99–113.
Blum, H., Beier, H., & Gross, B. (1987). Improved silver staining of plant proteins RNA and DNA in polyacrylamide gels. Electrophoresis, 8, 93–99.
Bordoli, L., & Schwede, T. (2012). Automated protein structure modeling with SWISS-MODEL Workspace and the Protein Model Portal. Methods in Molecular Biology, 857, 107–136.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 426–428.
Smaali, M. I., Michaud, N., Marzouki, N., Legoy, M. D., & Maugard, T. (2004). Comparison of two beta-glucosidases for the enzymatic synthesis of beta-(1-6)-beta-(1-3)-gluco-oligosaccharides. Biotechnology Letters, 26, 675–679.
Mouelhi, R., Abidi, F., Galai, S., & Marzouki, M. N. (2014). Immobilized Sclerotinia sclerotiorum invertase to produce invert sugar syrup from industrial beet molasses by-product. World Journal of Microbiology & Biotechnology, 30, 1063–1073.
Chahed, H., Ezzine, A., Mlouka, A. B., Hardouin, J., et al. (2014). Biochemical characterization, molecular cloning, and structural modeling of an interesting beta-1,4-glucanase from Sclerotinia sclerotiorum. Molecular Biotechnology, 56, 340–350.
Ellouze, O., Mejri, M., Smaali, I., Limam, F., & Marzouki, M. N. (2007). Induction, properties and application of xylanase activity from Sclerotinia sclerotiorum S2 fungus. Journal of Food and Biochemistry, 31, 96–107.
Park, T. H., Choi, K. W., Park, C. S., Lee, S. B., et al. (2005). Substrate specificity and transglycosylation catalyzed by a thermostable beta-glucosidase from marine hyperthermophile Thermotoga neapolitana. Applied Microbiology and Biotechnology, 69, 411–422.
Parry, N. J., Beever, D. E., Owen, E., Vandenberghe, I., et al. (2001). Biochemical characterization and mechanism of action of a thermostable beta-glucosidase purified from Thermoascus aurantiacus. The Biochemical Journal, 353, 117–127.
Nakkharat, P., & Haltrich, D. (2006). Purification and characterisation of an intracellular enzyme with beta-glucosidase and beta-galactosidase activity from the thermophilic fungus Talaromyces thermophilus CBS 236.58. Journal of Biotechnology, 123, 304–313.
Pitson, S. M., Seviour, R. J., & McDougall, B. M. (1997). Purification and characterization of an extracellular beta-glucosidase from the filamentous fungus Acremonium persicinum and its probable role in beta-glucan degradation. Enzyme and Microbial Technology, 21, 182–190.
Seidle, H. F., Marten, I., Shoseyov, O., & Huber, R. E. (2004). Physical and kinetic properties of the family 3 beta-glucosidase from Aspergillus niger which is important for cellulose breakdown. The Protein Journal, 23, 11–23.
Harhangi, H. R., Steenbakkers, P. J., Akhmanova, A., Jetten, M. S., et al. (2002). A highly expressed family 1 beta-glucosidase with transglycosylation capacity from the anaerobic fungus Piromyces sp. E2. Biochimica et Biophysica Acta, 1574, 293–303.
Christakopoulos, P., Goodenough, P. W., Kekos, D., Macris, B. J., et al. (1994). Purification and characterisation of an extracellular beta-glucosidase with transglycosylation and exo-glucosidase activities from Fusarium oxysporum. European Journal of Biochemistry/FEBS, 224, 379–385.
Yang, S., Jiang, Z., Yan, Q., & Zhu, H. (2008). Characterization of a thermostable extracellular beta-glucosidase with activities of exoglucanase and transglycosylation from Paecilomyces thermophila. Journal of Agricultural and Food Chemistry, 56, 602–608.
Fujita, Y., Ito, J., Ueda, M., Fukuda, H., & Kondo, A. (2004). Synergistic saccharification, and direct fermentation to ethanol, of amorphous cellulose by use of an engineered yeast strain codisplaying three types of cellulolytic enzyme. Applied and Environmental Microbiology, 70(2), 1207–1212.
Yamada, R., Nakatani, Y., Ogino, C., & Kondo, A. (2013). Efficient direct ethanol production from cellulose by cellulase-and cellodextrin transporter-co-expressing Saccharomyces cerevisiae. AMB Express, 3(1), 1–7.
Lima, M. A., Oliveira-Neto, M., Kadowaki, M. A., Rosseto, F. R., et al. (2013). Aspergillus niger beta-glucosidase has a cellulase-like tadpole molecular shape: Insights into glycoside hydrolase family 3 (GH3) beta-glucosidase structure and function. The Journal of Biological Chemistry, 288, 32991–33005.
Kalyani, D., Lee, K. M., Tiwari, M. K., Ramachandran, P., et al. (2012). Characterization of a recombinant aryl beta-glucosidase from Neosartorya fischeri NRRL181. Applied Microbiology and Biotechnology, 94, 413–423.
Chen, P., Fu, X., Ng, T. B., & Ye, X. Y. (2011). Expression of a secretory beta-glucosidase from Trichoderma reesei in Pichia pastoris and its characterization. Biotechnology Letters, 33, 2475–2479.
Smaali, M. I., Gargouri, M., Limmam, F., Maugard, T., Legoy, M. D., & Marzouki, N. (2004). A β-glucosidase from Sclerotinia sclerotiorum: Biochemical characterization and use in oligosaccharides synthesis. Applied Biochemistry and Biotechnology, 112, 63–78.
Decker, C. H., Visser, J., & Schreier, P. (2000). Beta-glucosidases from five black Aspergillus species: Study of their physico-chemical and biocatalytic properties. Journal of Agricultural and Food Chemistry, 48, 4929–4936.
Jager, S., Brumbauer, A., Feher, E., Reczey, K., & Kiss, L. (2001). Production and characterization of beta-glucosidases from different Aspergillus strains. World Journal of Microbiology & Biotechnology, 17, 455–461.
Korotkova, O. G., Semenova, M. V., Morozova, V. V., Zorov, I. N., et al. (2009). Isolation and properties of fungal beta-glucosidases. Biochemistry. Biokhimiia, 74, 569–577.
Riou, C., Salmon, J. M., Vallier, M. J., Gunata, Z., & Barre, P. (1998). Purification, characterization, and substrate specificity of a novel highly glucose-tolerant beta-glucosidase from Aspergillus oryzae. Applied and Environmental Microbiology, 64, 3607–3614.
Wiseman, A. (1982). Fungal and other beta-d-glucosidases—Their properties and application. Enzyme Microbiology and Biotechnology, 4, 73–78.
Smaali, M. I., Gourgouri, M., Limam, F., Fattouch, S., Maugard, T., Legoy, M. D., & Marzouki, N. (2003). Production, purification, and biochemical characterization of two β-glucosidases from Sclerotinia sclerotiorum. Applied Biochemistry and Biotechnology, 111, 29–39.
Zhou, W., Irwin, D. C., Escovar-Kousen, J., & Wilson, D. B. (2004). Kinetic studies of Thermobifida fusca Cel9A active site mutant enzymes. Biochemistry, 43, 9655–9663.
Karkehabadi, S., Helmich, K. E., Kaper, T., Hansson, H., Mikkelsen, N. E., Gudmundsson, M., et al. (2014). Biochemical characterization and crystal structures of a fungal family 3 β-Glucosidase, Cel3A from Hypocrea jecorina. Journal of Biological Chemistry, 289(45), 31624–31637.
Brzobohaty, B., Moore, I., Kristoffersen, P., Bako, L., et al. (1993). Release of active cytokinin by a beta-glucosidase localized to the maize root meristem. Science, 262, 1051–1054.
Easen, A. (1993). Beta-glucosidases. Biochemistry and molecular biology. Washington, DC: American Chemical Society.
de la Mata, I., Castillon, M. P., Dominguez, J. M., Macarron, R., & Acebal, C. (1993). Chemical modification of beta-glucosidase from Trichoderma reesei QM 9414. Journal of Biochemistry, 114, 754–759.
Bisaria, V. S., & Mishra, S. (1989). Regulatory aspects of cellulase biosynthesis and secretion. Critical Reviews in Biotechnology, 9, 61–103.
Tomme, P., Warren, R. A., & Gilkes, N. R. (1995). Cellulose hydrolysis by bacteria and fungi. Advances in Microbial Physiology, 37, 1–81.
Singhania, R. R., Patel, A. K., Sukumaran, R. K., Larroche, C., & Pandey, A. (2013). Role and significance of beta-glucosidases in the hydrolysis of cellulose for bioethanol production. Bioresource Technology, 127, 500–507.
Acknowledgments
The Laboratory of Protein Engineering and Bioactive Molecules (LIP-MB) in the National Institute of Applied Sciences and Technology of Tunis, University of Carthage financed this work. The Tunisian Ministry of High Education, Scientific Research and Technology is gratefully acknowledged for the financial support of the training program. The laboratory of polymers, biopolymers and surfaces (PBS-UMR 6270) of Rouen is also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no commercial or financial conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chahed, H., Ezzine, A., Mlouka, M.A.B. et al. A Novel Three Domains Glycoside Hydrolase Family 3 from Sclerotinia sclerotiorum Exhibits β-Glucosidase and Exoglucanase Activities: Molecular, Biochemical, and Transglycosylation Potential Analysis. Mol Biotechnol 57, 993–1002 (2015). https://doi.org/10.1007/s12033-015-9892-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-015-9892-z