Abstract
Leaf senescence is associated with changes in proteomics. In this study, to quantitatively analyze changes in protein abundance related to leaf senescence in cotton (Gossypium hirsutum L.), we employed a proteomic approach utilizing iTRAQ and physiological assays throughout the senescence of leaf tissue in cotton grown under typical field conditions. Physiological tests showed leaf chlorophyll content and photosynthetic rates decreased significantly throughout the aging process. A total of 195 differentially abundant proteins (DAPs) throughout leaf senescence were identified by mass spectrometry. Of these, 91 (47%) proteins were upregulated, while the remaining 104 (53%) were downregulated. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses indicated that the expression of genes in several pathways potentially associated with aging changed throughout senescence, including metabolic pathways, photosynthesis, pyruvate metabolism, nitrogen metabolism, and diterpenoid biosynthesis. Our findings provide a deeper understanding of aging in plants as well as fundamental data describing leaf senescence in cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton (Gossypium hirsutum L.) is the world’s remarkable oilseed source and is also a primary natural fiber source (Guo et al. 2017). Leaves are the major photosynthetic organ of cotton plants, and they are essential to plant growth and biomass accumulation, fundamental covariates of agricultural yields. Leaf senescence is a process that increases plant productivity and is caused by a wide range of signal crosstalk that harmonizes environmental factors with the age-related internal mechanisms of plants (Allu et al. 2014; Quirino et al. 2000). However, premature senescence of leaves is a serious cause of crop yield reduction (Balazadeh et al. 2010). This phenomenon shortens the period of photosynthesis of plants and reduces the accumulation of photosynthetic products, resulting in reduced yield and quality (Liu et al. 2016a). Premature senescence can decrease fiber quality and fiber yields, with substantial negative impacts (Dong et al. 2006).
Some senescent plants grown under optimum conditions have higher chlorophyll content prior to their senescence and thus have prolonged lifespans. Chlorophyll degrading enzyme activity increases during leaf senescence, and photosynthetic pigments (i.e., carotenoids, lutein, and chlorophyll) fall faster; at the same time, some genes with integral roles in photosynthesis are downregulated, leading to a decrease in leaf photosynthetic capacity (Wu et al. 2012). These changes cause the leaves to turn yellow, which is the morphological sign of aging (Lim et al. 2007).
The initiation and development of leaf senescence are accompanied by a large number of changes in gene expression, and the activation of a series of new genes is necessary for the onset of aging (Gepstein 2004; Zhang et al. 2010). In plants, hundreds of aging-related genes have been identified (Bhalerao et al. 2003; Andersson et al. 2004; Gepstein 2004; van der Graaff et al. 2006; Zhang et al. 2010). In addition, transcriptase analyses have revealed the complexity of regulation of leaf senescence (He et al. 2001). Multiple studies have also identified senescence-related genes by assaying transcriptomes, transcription factors and plant hormones in senescing leaves (Kong et al. 2013; He et al. 2001). However, few studies have focused on aging leaf proteomics.
Proteomes differ distinctly among tissues and cells, and the expression and modification of proteins are also affected by environmental factors and developmental stages. In recent years, with the development of ultra-high-performance liquid chromatography (UPLC) and mass spectrometry, the advantages of the isobaric tags for relative and absolute quantitation (iTRAQ) approach have become quite obvious. This approach has become a vital research technology for studying the molecular mechanisms of plant leaf protein responses, and iTRAQ can be used in high-throughput comparative proteomics analyses conducted across different development stages (Qin et al. 2016).
In this study, we investigated proteomic changes throughout the senescence of photosynthetically active leaves on the main stems of cotton plants grown under typical field conditions. The iTRAQ method was used to assess proteomic changes in leaves and to identify differentially expressed proteins, throughout the aging process. The bioinformatics method is used to analyze the differentially expressed proteins (DAPs) and reveal their biological functions. Further characterization of these DAPs will clarify the signaling pathways activated by cotton in response to and as a part of senescence.
Materials and methods
Plant materials
The field experiments were conducted in Baoding (38°49′N, 115°44′E), Hebei, China, during the 2016 growing season. The experimental area is located within the warm-temperate arid zone with a continental climate. ‘Lumianyan 28’ (Gossypium hirsutum L.), a commercial cotton cultivar that is widely cultivated in the Yellow River Basin of China, was used exclusively in this study.
Seeds were sown on April 25, 2016, and plants were harvested in late September 2016. Samples for tests were given priority to assay leaf stem function (using samples from the third main-stem leaf from the terminal bud). Samples were collected on August 6, August 14, and September 1, at leaf ages of 18 days, 26 days, and 44 days, which are hereafter referred to as M1, M2, and M3, respectively. Immediately after collection, all tissue samples were frozen in liquid nitrogen and stored at − 80 °C.
Pigment determination
The third main-stem leaves from the terminal bud in field-grown plants were assayed for pigment concentration. Chlorophyll was extracted from 0.1 g fresh leaf samples using 95% acetone and measured at 645 and 663 nm with a UV spectrophotometer (Shimadzu UV-2450 Kyoto, Japan) (Lichtenthaler 1987).
Leaf gas-exchange measurement
The third main-stem leaves were subjected to gas-exchange analysis with the LI-6400XT Portable Photosynthesis System (Li-Cor Inc. Lincoln, NE, USA) to determine net photosynthesis. During the measurements, the determination conditions were as previously described by Pu et al. (2015).
Protein extraction
To extract proteins, the prepared leaf samples were first ground into dry powder in a mortar with liquid nitrogen. Then, the dry powder was dissolved in 200 μL of triethylammonium bicarbonate (TEAB) lysis buffer (pH 8.0, 8 M urea/100 mM TEAB) and ultrasonicated for 15 min. Then, centrifuge at 10,000 × g for 20 min, submerged and incubated the supernatant for 2 h by the addition of four volumes of cold acetone containing 10 mM dithiothreitol (DTT). The mixture was then centrifuged at 12,000 rpm and 4 °C for 20 min, the supernatant was discarded, and it was mixed with 800 μL of acetone at 56 °C to degrade the disulfide bond in the extracted protein. Next, the mixture was centrifuged again by the above centrifugation method and dried under vacuum. The collected dry precipitate was dissolved in 100 μL of TEAB lysis buffer (Liao and Huang 2011), and then, the total protein concentration was measured using the classical Bradford method (Bradford 1976).
The stored protein sample was reduced with 10 mM DTT for 30 min at 56 °C, then alkylated with 50 mM iodoacetamide in the dark for 30 min at 50 °C, diluted fivefold, and finally, trypsin digested with 100 mM TEAB (Zhang et al. 2018).
Digestion and desalting
One hundred micrograms of each protein sample were dissolved in lysis buffer and diluted with 50 mM NH4HCO3 (500 μL) to reduce and alkylate the sample, followed by the addition of 2 μg trypsin, and incubated at 37 °C overnight to complete protein digestion. Next, an equal volume of 0.1% formic acid (FA) was added to the above solution to complete the acidification. Peptide samples were purified using a Strata-X C18 column (Phenomenex Inc., Torrance, CA, USA), first activated with methanol, equilibrated three times with 0.1 mL FA, then twice with 0.1% FA + 5% acetonitrile (ACN), and then eluted with 1 mL of 0.1% FA + 80% CAN. The eluted peptide was dried with a vacuum concentrator and then redissolved into 20 μL of 0.5 M TEAB buffer solution for peptide labeling.
Labeling and fractionation
The iTRAQ Reagent-8 plex Multiplex kit (AB SCIEX UK Limited, Warrington, UK) was employed to label the prepared samples. Reagents 113, 114, and 115 were used to label the M1 samples, and 118, 119, and 121 were used for the M3 samples. Next, the labeled samples were fractionated by HPLC (Thermo DIONEX Ultimate 3000 BioRS, Thermo Fisher Scientific, Waltham, MA, USA) using a Durashell C18 column (5 μm, 100 Å, 4.6 × 250 mm) to obtain 12 fractions.
LC–MS/MS analysis
We performed LC–ESI–MS/MS data analysis with the AB SCIEX nano-LC–MS/MS system (Triple TOF 5600 plus, AB, Foster City, CA, USA). In this study, the method described by Yang et al. (2018) was applied for sample operation and condition settings. The quadrupole isolation window for the TripleTOF 5600 + mass spectrometer was 0.6–0.8 Da.
iTRAQ protein identification, quantification, and bioinformatic analysis
ProteinPilot software v4.5 (SCIEX, Framingham, MA, USA) was used to analyze raw MS/MS file data. In protein identification, we integrated Paragon algorithm 2 into the ProteinPilot, was employed in queries against the NCBI Gossypium database (consisting of 182,674 items, as of January 2016). The parameters used for analyses were those that were previously described by Pu et al. (2015). For functional annotation, all DAPs were searched against the Gene Ontology (GO) (Conesa et al. 2005), as well as the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases (Kanehisa and Goto 2000).
qPCR analysis
The third main-stem leaf samples subjected to iTRAQ analysis were also used for RNA preparation. Total RNA quantitative RT-PCR (qPCR) was performed as previously described (Chu et al. 2012). The qPCR was performed on an ABI7500 real-time system. All PCR primers were designed using Primer Express 3.0 and are shown in Table S1. Relative expression-level evaluation was conducted using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Results
Pigment content and photosynthesis assays
Chlorophyll content is an important index of leaf senescence (Zhang et al. 2010). We investigated the chlorophyll content of the third main-stem leaves from the terminal apex at leaves aged 18 days, 26 days, and 44 days (Fig. 1) to characterize progress in leaf senescence. Leaf chlorophyll content considerably decreased with increasing leaf age. The total Chl contents at 26 days and 44 days were reduced by approximately 10.27% and 37.52%, respectively (p <0.05).
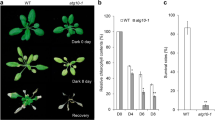
Photosynthetic performance of third main-stem leaves collected from cotton plants grown in field plots at three different developmental stages. a Total chlorophyll content. Data are means of nine biological replicates ± SE. b Photosynthetic rate. Data are means of six biological replicates ± SE. Different letters at the top of the bars in each plot indicate significant differences (p < 0.05) between treatments
The trend in photosynthetic rate was consistent with that of chlorophyll content, decreasing with leaf age. The photosynthetic rates at 26 days and 44 days were reduced by approximately 20.84% and 35.47%, respectively (p <0.05).
The senescence of the leaves collected in 18 days, 26 days, and 44 days, corresponding to full maturity, medium-term senescence, and late senescence was determined according to changes in photosynthetic rate and chlorophyll content.
Quantitative identification of cotton leaf proteins using iTRAQ
To further understand the molecular mechanism of cotton leaf senescence in field plots, the proteomic changes in third main-stem leaves were separately assessed for leaves aged at 18 days, 26 days, and 44 days. In addition, the number of proteins as well as their functional properties and associated pathways were analyzed.
The total spectra were comprised of 323,662 elements, based on three replicates. From this, 16,463 of the identified spectra matched known peptides. Ultimately, 16,463 peptides and 3882 proteins were identified, with 2705 proteins (69.68%) containing at least two unique peptides (Fig. 2a). A total of 1177 proteins matched one identified peptide, 918 matched 2, 148 matched more than 10, and the remainder matched 3–9 (Fig. 2a). Using the method of percentage of matched amino acids from the identified peptides, the protein sequence coverage of the particular protein was obtained (Fig. 2b), yielding an average coverage of 15.63%.
Protein differences throughout leaf development
In the present research, any proteins with more than a 1.2-fold difference that was statistically significant were designated as significant DAPs. Accordingly, the number of DAPs was 195 (Table S2); among these, 91 (47%) were upregulated, while 104 (53%) were downregulated.
To understand the functional categories of DAPs, we performed a GO enrichment analysis on them. Classify DAPs based on biological processes, molecular functions, and cellular components (Fig. 3, Table S3), which included 23 biological process, 8 cellular component, and 10 molecular function categories. The main biological functional categories represented were generation of precursor metabolites and energy, ribosome biogenesis, carbohydrate catabolic process, photosynthesis, and the photosynthetic electron transport chain. The cellular component categories represented were plastid, chloroplast, organelle, thylakoid, cytoplasm, plastid stroma, and photosynthetic membrane. DAPs were classified into binding, phosphoenolpyruvate carboxykinase activity, and lytic enzyme activity, depending on their molecular functional properties.
Signal pathway analyses can help to clarify the biological functions of DAPs to determine the main biochemical pathways and signal transduction pathways through which proteins respond to age. Accordingly, these DAPs were further analyzed by the KEGG database and found to be mainly enriched in metabolic pathways (53.4%), photosynthesis (8.5%), pyruvate metabolism (6.8%), nitrogen metabolism (4.2%), diterpenoid biosynthesis (3.4%), carotenoid biosynthesis (3.4%), and galactose metabolism (3.4%; Table 1).
Complementation of iTRAQ data with qPCR of selected candidates
From among all 195 DAPs, 10 were selected for transcriptional analysis (Fig. 4). Among these proteins, a majority showed consistent mRNA and protein expression patterns. However, two proteins exhibited contrasting expression patterns. This result may be due to differences in post-transcriptional, translational, or post-translational regulation. While most of the changes agreed with respect to sign, our results confirmed that mRNA transcript abundance does not correlate perfectly with protein abundance (Liu et al. 2016b). The results indicate generally parallel but flexible associations between mRNA and protein expression profiles in aging-associated proteins.
Comparative analysis of RNA and protein levels of selected differentially accumulated proteins as revealed by qPCR and iTRAQ. The wild-type control data are represented by a relative expression value of 1.0. Values are mean ± SD (n = 3) in cde1 leaf data. A1E Aldose 1-epimerase, Htr4, Histone H3.3, GS1 glutamine synthetase nodule isozyme-like, OEE3 Oxygen-evolving enhancer 3, chloroplastic, KynB Kynurenine formamidase, Rrl21A 60S ribosomal protein L21-1, Csp41B PREDICTED: chloroplast stem-loop binding protein of 41 kDa b, chloroplastic, PetC cytochrome b6/f complex subunit IV (chloroplast), PsaB photosystem I P700 apoprotein A2 (chloroplast), Pe pectinesterase
Discussion
Proteomic methods for global expression analyses and protein identification in plants have become highly efficient in recent years (Qin et al. 2016). Thus, iTRAQ technology and LC–MS/MS have been employed together widely, demonstrating their proven value in discovery-based proteomics. Accordingly, iTRAQ has become a major tool for both qualitative and quantitative protein researches. In the present study, by comparing the relative expression patterns (in M3 versus M1 leaf samples) of the proteins in the third main-stem leaves, 195 DAPs were identified. Of these 195 DAPs, 91 were up-regulated and 104 were downregulated. Through the KEGG analysis of DAPs, the following key signaling pathways were identified as significantly enriched for involvement in leaf senescence: metabolic pathways, photosynthesis, pyruvate metabolism, and nitrogen metabolism.
Reduced Chl biosynthetic protein levels underlie Chl deficiency in leaves
Leaves are the primary photosynthetic organ of plants (Evans and Von 2011; Makino 2011). Although a critical and likely adaptive phenomenon, leaf senescence may limit yields and biomass accumulations owing to its associated reduction in photosynthesis. The pigment chlorophyll plays an important role in photosynthesis due to its effects on light capture and energy transfer (Pu et al. 2015). As such, it is vital to the growth of plants and thus crop yield. The present study showed that the Chl a + b content of M3 leaves was significantly reduced relative to M1 leaves.
Magnesium chelatase (or Mg-protoporphyrin IX chelatase) is an enzyme that catalyzes the magnesium insertion step of Chl synthesis and consists of three subunits (ChlH, ChlD, and ChlI). We observed consistent reductions of some important Chl biosynthesis-related proteins in the M3-stage leaves, and the magnesium-chelatase subunit chlI (ChlI) levels were also significantly reduced; the fold change of ChlI between the M3 and M1 stages was 0.574.
This protein is part of the metabolism of porphyrin-containing compounds and is involved in the chlorophyll biosynthesis pathway. ChlI is also a subunit of Mg chelatase, mainly responsible for redox regulation, and its decrease leads to a decrease in subsequent Chl biosynthesis. This protein is involved in the chlorophyll biosynthesis pathway, which is part of porphyrin-containing compound metabolism. ChlI is also the primary redox-regulated subunit of Mg chelatase, and its decreases cause an eventual subsequent decrease in Chl biosynthesis. Similar results were obtained in a study of chloroplast deficiencies in rice (Zhang et al. 2006). Overall, leaf senescence is associated with the decay of chlorophyll synthesis components, which thus result in the failure to effectively synthesize chlorophyll and thereby reduces chlorophyll content over time.
Downregulation of the photosynthesis pathway
Senescence is an extremely complex biochemical process involving the expression of a number of aging-related genes and signaling pathways (Breeze et al. 2011; van der Graaff et al. 2006; Shah et al. 2013). We observed that photosynthetic capacity was diminished over the course of senescence. Our study identified ten proteins in the photosynthesis pathway, one of the most important pathways associated with senescence, and most photosystem I (PSI) and photosystem II (PSII) proteins were downregulated by 0.5–1.6-fold.
In addition, five of these proteins were in the PSII complex: PsbB, PsbD, PsbO, PsbP, and PsbQ. PsbB encodes the D2 protein in the PSII reaction center. D2 and D1 constitute two heterologous dimers involved in the electron transfer of most cofactors and pigment molecules. The two proteins are localized within the thylakoid membrane, and their expression was downregulated over the course of leaf senescence. Inhibition of light capture by leaves in photosynthesis resulted in the inhibition of the expression of these proteins. PsbO, PsbP, and PsbQ constitute the oxygen-releasing complexes in higher plants, which collectively play an important role as stabilizing proteins in the oxygen-releasing activity of PSII. In this study, PsbO, PsbP, and PsbQ were upregulated during the aging process, which may indicate that these three proteins protect against the photosynthetic oxygen release system during aging, thus ensuring the absorption of light energy. PsaA (photosystem IP700 apoprotein A1) and PsaB (photosystem IP700 apoprotein A2) are concentrated in the PSI complex, and as leaves aged, their expression decreased as well. This phenomenon indicates that photosynthetic electron transport in senescent leaves is gradually inhibited, thus decreasing photosynthesis. Cytb6 f performs electron transfer between PSII and PSI light responsive complexes, which together modulate the state transition of the photosystem by activating a protein kinase (Hu et al. 2011). Pet D (cytochrome, b6/f, complex subunit IV) and PetA (cytochrome, f) within Cytb6 f are both downregulated, thus restricting electron transport. Overall, senescence impacts the capture of light energy and decreases electron transport function, thus decreasing the photosynthetic rate.
Nitrogen metabolism pathway
Nitrogen cycling in leaves can cause chlorophyll degradation, thus decreasing photophosphorylation and photosynthesis while inducing leaf senescence (Zhou et al. 2010). In addition, glutamine synthetase (GS) increases in association with leaf senescence (Yin and Chai 1997). In this study, the expression of GS increased as the leaves aged, suggesting that the high expression of GS may increase the nitrogen use of crops while reducing nitrogen loss (Yu et al. 2014). Glutamate synthase (GOGAT) is the key enzyme involved in the assimilation of NO3−, and its expression decreased in this study. Increases in GS expression and decreases in GOGAT expression both induce excessive glutamine production in plant. Accordingly, as glutamine is involved in nitrogen transport, the free movement of nitrogen increases throughout leaf senescence.
Pyruvate metabolism pathway
Pyruvate, a metabolite produced by the glycolysis of cytoplasmic glucose, plays an important role in cell metabolism. Examination of the pyruvate metabolism pathway revealed that six proteins related to energy metabolism were downregulated, while two proteins related to reproductive process and cell apoptosis were upregulated.
Sphingolipid metabolism pathway
Sphingolipid metabolism is associated with apoptosis (Chipuk et al. 2012). In this study, the expression of two proteins, β-galactosidase (β-Gal, EC3.2.1.23) and α-galactosidase (α-Gal, EC3.2.1.22), changed throughout leaf senescence. β-Gal is an enzyme related to cell-wall degradation. The removal of β-galactosidase residues from the non-reductive end of its branched chain can degrade pectin and hemicelluloses as well as glycoproteins and glycolipids. This destabilizes some cell-wall components, causing the cell wall to swell and soften. These changes both release stored energy for the rapid growth and development of plants as well as promote cell-wall degradation during senescence, causing the hydrolysis of polysaccharides, glycoproteins, and galactose residues at the ends of galactose lipids during polysaccharide and cell-wall component metabolism, thus releasing free galactose. The downregulation of β-Gal expression may reduce energy storage during plant growth and development, while α-Gal is essential for promoting seed germination and maturation, leaf development and senescence, and fruit development and maturation (Sun et al. 2015). In this study, over the course of senescence, the expression of α-Gal was upregulated, suggesting that α-Gal promotes leaf senescence.
Conclusions
In summary, the iTRAQ comparative proteomics approach was used to identify 195 DAPs between senesced leaves (44 day) relative to mature leaves (18 day). Of these, 91 proteins (47%) were upregulated, while the remaining 104 (53%) were downregulated. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses indicated that the enrichment of several pathways potentially associated with aging changed throughout senescence, including metabolic pathways, photosynthesis, pyruvate metabolism, nitrogen metabolism, and diterpenoid biosynthesis. These results may provide a deeper understanding of aging in plants as well as fundamental data describing leaf senescence in cotton. Further genetic functional analyses are needed to elucidate the molecular mechanisms that regulate chlorophyll metabolism and photosynthesis throughout leaf senescence.
Author contribution statement
Conceived and designed the experiments: LLT, LCD, and BZY. Performed the experiments: LAC, CHJ, and WM. Analyzed the data: ZYJ, SHCH, WXD, and LYCH. Wrote the paper: LLT.
References
Allu AD, Soja AM, Wu A, Szymanski J, Balazadeh S (2014) Salt stress and senescence: identification of cross-talk regulatory components. J Exp Bot 65(14):3993. https://doi.org/10.1093/jxb/eru173
Andersson A, Keskitalo J, Sjodin A, Bhalerao R, Sterky F, Wissel K et al (2004) A transcriptional timetable of autumn senescene. Genome Biol 5(4):1–13. https://doi.org/10.1186/gb-2004-5-4-r24
Balazadeh S, Siddiqui H, Allu AD, Matallanaramirez LP, Caldana C, Mehrnia M et al (2010) A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J 62(2):250–264. https://doi.org/10.1111/j.1365-313X.2010.04151.x
Bhalerao R, Keskitalo J, Sterky F, Erlandsson R, Björkbacka H, Birve SJ et al (2003) Gene expression in autumn leaves. Plant Physiol 131(2):430–442. https://doi.org/10.1104/pp.012732
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Breeze E, Harrison E, Mchattie S, Hughes L, Hickman R, Hill C et al (2011) High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 23(3):873–894. https://doi.org/10.1105/tpc.111.083345
Chipuk JE, Mcstay GP, Bharti A, Kuwana T, Clarke CJ, Siskind LJ et al (2012) Sphingolipid metabolism cooperates with BAK and BAX to promote the mitochondrial pathway of apoptosis. Cell 148(5):988–1000. https://doi.org/10.1016/j.cell.2012.01.038
Chu P, Chen H, Zhou Y, Li Y, Ding Y, Jiang L et al (2012) Proteomic and functional analyses of Nelumbo nucifera annexins involved in seed thermotolerance and germination vigor. Planta 235(6):1271–1288. https://doi.org/10.1007/s00425-011-1573-y
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Dong H, Li W, Tang W, Li Z, Zhang D, Niu Y (2006) Yield, quality and leaf senescence of cotton grown at varying planting dates and plant densities in the Yellow River Valley of China. Field Crops Res 98(2–3):106–115. https://doi.org/10.1016/j.fcr.2005.12.008
Evans JR, Von CS (2011) Enhancing photosynthesis. Plant Physiol 155(1):19. https://doi.org/10.1104/pp.110.900402
Gepstein S (2004) Leaf senescence-not just a ‘wear and tear’ phenomenon. Genome Biol 5(3):212. https://doi.org/10.1186/gb-2004-5-3-212
Guo Y, Dou L, Evans O, Pang C, Wei H, Song M et al (2017) Identification of GT factors in response to stresses and leaf senescence in Gossypium hirsutum L. J Plant Growth Regul 36(1):22–42. https://doi.org/10.1007/s00344-016-9619-9
He Y, Tang W, Swain JD, Green AL, Jack TP, Gan S (2001) Networking senescence-regulating pathways by using Arabidopsis enhancer trap lines. Plant Physiol 126(2):707–716. https://doi.org/10.1104/pp.126.2.707
Hu F, Huang JL, Qin F, Yue CL, Wang GX (2011) Progress in chloroplast thylakoid membrane and membrane proteins. Chinese bulletin of life sciences 23(3):291–298
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Kong X, Luo Z, Dong H, Eneji AE, Li W, Lu H (2013) Gene expression profiles deciphering leaf senescence variation between early- and late-senescence cotton lines. PLoS One 8(7):e69847. https://doi.org/10.1371/journal.pone.0069847
Liao JL, Huang YJ (2011) Evaluation of protocols used in 2-D electrophoresis for proteome analysis of young rice caryopsis. Genomics Proteomics Bioinform 09(6):229–237. https://doi.org/10.1016/S1672-0229(11)60026-0
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148(1):350–382. https://doi.org/10.1016/0076-6879(87)48036-1
Lim PO, Kim HJ, Gil Nam H (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136. https://doi.org/10.1146/annurev.arplant.57.032905.105316
Liu ZX, Yu C, Wang ZW, Xie YH, Sang XC, Yang ZL et al (2016a) Phenotypic characterization and fine mapping of mps1, a premature leaf senescence mutant in rice (Oryza sativa L.). J Integr Agric 15(9):1944–1954. https://doi.org/10.1016/s2095-3119(15)61279-5
Liu R, Wang Y, Qin G, Tian S (2016b) iTRAQ-based quantitative proteomic analysis reveals the role of the tonoplast in fruit senescence. J Proteomics 146:80–89. https://doi.org/10.1016/j.jprot.2016.06.031
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Makino A (2011) Photosynthesis, grain yield, and nitrogen utilization in rice and wheat. Plant Physiol 155(1):125. https://doi.org/10.1104/pp.110.165076
Pu C, Gui XY, Yang Q, Li NZ, Cheng Z, Feng QZ et al (2015) iTRAQ-based quantitative proteomics analysis of Brassica napus leaves reveals pathways associated with chlorophyll deficiency. J Proteomics 113:244–259. https://doi.org/10.1016/j.jprot.2014.10.005
Qin J, Zhang J, Liu D, Yin C, Wang F, Chen P et al (2016) iTRAQ-based analysis of developmental dynamics in the soybean leaf proteome reveals pathways associated with leaf photosynthetic rate. Mol Genet Genomics Mgg 291(4):1595. https://doi.org/10.1007/s00438-016-1202-3
Quirino Betania F, Noh YooSun, Himelblau Edward et al (2000) Molecular aspects of leaf senescence. Trends Plant Sci 5(7):278–282. https://doi.org/10.1016/S1360-1385(00)01655-1
Shah ST, Pang C, Fan S, Song M, Arain S, Yu S (2013) Isolation and expression profiling of GhNAC transcription factor genes in cotton (Gossypium hirsutum L.) during leaf senescence and in response to stresses. Gene 531(2):220–234. https://doi.org/10.1016/j.gene.2013.09.007
Sun JJ, Liu YJ, Qin Q, Sui SZ, Li MY (2015) Cloning and transcriptional expression analysis of α-Galactosidase gen from Chimonanthus praecox. J Southwest Univ Nat Sci Ed 37(1):25–32. https://doi.org/10.13718/j.cnki.xdzk.2015.01.004
Van der Graaff E, Schwacke R, Schneider A, Desimone M, Flügge U-I, Kunze R (2006) Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol 141(2):776–792. https://doi.org/10.1104/pp.106.079293
Wu A, Allu AD, Garapati P, Siddiqui H, Dortay H, Zanor M-I et al (2012) JUNGBRUNNEN1, a reactive oxygen species—responsive NAC transcription factor, regulates longevity in Arabidopsis. Plant Cell 24(2):482–506. https://doi.org/10.1105/tpc.111.090894
Yang Y, Ma L, Zeng H, Chen LY, Zheng Y, Li CX et al (2018) iTRAQ-based proteomics screen for potential regulators of wheat (Triticum aestivum L.) root cell wall component response to Al stress. Gene 675(30):301–311. https://doi.org/10.1016/j.gene.2018.07.008
Yin LP, Chai XQ (1997) Changes of gIutamine synthesis and proteolyase in different wheat speicies during the leaves nature senescence (in Chinese with english abstract). J Shanxi Teach Univ Nat Sci Ed 1:46–49
Yu JL, Zhu ZK, Zhang ZH, Shu JB, Yang C, Song HX et al (2014) Influence of glutamine synthase and glutamate synthase on N reutilization in Brassica Napus under different nitrogen condition (in Chinese with english abstract). Crops 6:81–85
Zhang H, Li J, Yoo JH, Yoo SC, Cho SH, Koh HJ et al (2006) Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol 62(3):325. https://doi.org/10.1007/s11103-006-9024-z
Zhang A, Lu Q, Yin Y, Ding S, Wen X, Lu C (2010) Comparative proteomic analysis provides new insights into the regulation of carbon metabolism during leaf senescence of rice grown under field conditions. J Plant Physiol 167(16):1380–1389. https://doi.org/10.1016/j.jplph.2010.05.011
Zhang X, Liu Q, Zhou W, Li P, Alolga RN, Qian Z et al (2018) A comparative proteomic characterization and nutritional assessment of naturally- and artificially-cultivated Cordyceps sinensis. J Proteomics 181(2018):24–35. https://doi.org/10.1016/j.jprot.2018.03.029
Zhou Q, Zhao CP, Cao CX, Jiang QJ, Jiang HD (2010) Effects of N dressing ratio on carbon and nitrogen transport and on grain yield of Lolium multiflorum. Acta Prataculturae Sinica 19(4):47–53
Acknowledgements
The authors wish to acknowledge Prof. Kai Xiao in the Key Laboratory of Crop Growth Regulation of Hebei Province, Baoding 071001 Hebei Province help with support and discussion. The authors are grateful to the anonymous reviewers for their valuable comments and suggestions. This work was supported by the National Key R and D Program of China (2018YFD0100306) and National Natural Science Foundation of China (Nos. 31171495, 31301270 and 31571610).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Wojtaszek.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2019_2921_MOESM2_ESM.xlsx
Supplementary material 2 S2 Table. Functional description and protein expression patterns in M3 and M1 leaf samples. (XLSX 41 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liu, L., Li, A., Chen, J. et al. iTRAQ-based quantitative proteomic analysis of cotton (Gossypium hirsutum L.) leaves reveals pathways associated throughout the aging process. Acta Physiol Plant 41, 144 (2019). https://doi.org/10.1007/s11738-019-2921-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-019-2921-y