Abstract
Phalaenopsis, an epiphytic crassulacean acid metabolism (CAM) plant, requires moderate variations of day/night temperatures for flowering. In this study, changes in chlorophyll content, chlorophyll fluorescence, sugar components, titratable acidity and soluble protein content in Phalaenopsis leaves during flowering were observed. Comparative proteomic analysis of Phalaenopsis leaves in the vegetative and flowering phase was performed for the first time using iTRAQ (isobaric tags for relative and absolute quantification). A total of 126 proteins were differentially expressed in Phalaenopsis leaves. Analysis of potential functions revealed that the major categories of predicted function of the up-regulated proteins were protein destination (27 %), photosynthesis (15.9 %), primary metabolism (14.3 %) and defense (12.7 %) in the flowering phase, while the major categories of predicted function of the down-regulated proteins were protein destination (33.3 %), primary metabolism (20.6 %), transportation (14.3 %) and signal transduction (11.1 %). Proteome profile analysis indicated that the proteome changes were consistent with changes in sugar and protein metabolites. Some novel proteins were differentially expressed, most of which were identified as signaling proteins, including 14-3-3 proteins, fibrillin, rapid alkalinization factors (RALF), the Ras-related protein RABB1c, calreticulin and calmodulin. Histone, importin alpha, multidrug resistance proteins and the ABC transporters were also differentially expressed. These results provide insights into the mechanisms that regulate flowering in complex flowering plants.


Similar content being viewed by others
References
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309(5737):1052–1056. doi:10.1126/science.1115983
Ahn CS, Lee JH, Reum Hwang A, Kim WT, Pai HS (2006) Prohibitin is involved in mitochondrial biogenesis in plants. Plant J 46(4):658–667. doi:10.1111/j.1365-313X.2006.02726.x
Baker NR (2008) Chlorophyll fluorescence: a probe of photosynthesis in vivo. Annu Rev Plant Biol 59:89–113. doi:10.1146/annurev.arplant.59.032607.0927
Bernier G, Périlleux C (2005) A physiological overview of the genetics of flowering time control. Plant Biotechnol J 3(1):3–16. doi:10.1111/j.1467-7652.2004.00114.x
Bhasin M, Reinherz EL, Reche PA (2006) Recognition and classification of histones using support vector machine. J Comput Biol 13(1):102–112. doi:10.1089/cmb.2006.13.102
Blanco-Rivero A, Shutova T, Román MJ, Villarejo A, Martinez F (2012) Phosphorylation controls the localization and activation of the lumenal carbonic anhydrase in Chlamydomonas reinhardtii. PLoS One 7(11):e49063. doi:10.1371/journal.pone.0049063
Bönisch C, Hake SB (2012) Histone H2A variants in nucleosomes and chromatin: more or less stable? Nucleic Acids Res 40(21):10719–10741. doi:10.1093/nar/gks865
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Cao Y, Dai Y, Cui S, Ma L (2008) Histone H2B monoubiquitination in the chromatin of FLOWERING LOCUS C regulates flowering time in Arabidopsis. Plant Cell 20(10):2586–2602. doi:10.1105/tpc.108.062760
Ceusters J, Borland AM, Ceusters N, Verdoodt V, Godts C, De Proft MP (2010) Seasonal influences on carbohydrate metabolism in the CAM bromeliad Aechmea ‘Maya’: consequences for carbohydrate partitioning and growth. Ann Bot 105(2):301–309. doi:10.1093/aob/mcp275
Chen D, Guo B, Hexige S, Zhang T, Shen D, Ming F (2007) SQUA-like genes in the orchid Phalaenopsis are expressed in both vegetative and reproductive tissues. Planta 226(2):369–380. doi:10.1007/s00425-007-0488-0
Chen WH, Tseng YC, Liu YC, Chuo CM, Chen PT, Tseng KM, Yeh YC, Ger MJ, Wang HL (2008) Cool-night temperature induces spike emergence and affects photosynthetic efficiency and metabolizable carbohydrate and organic acid pools in Phalaenopsis aphrodite. Plant Cell Rep 27(10):1667–1675. doi:10.1007/s00299-008-0591-0
Chevalier D, Morris ER, Walker JC (2009) 14-3-3 and FHA domains mediate phosphoprotein interactions. Annu Rev Plant Biol 60:67–91. doi:10.1146/annurev.arplant.59.032607.092844
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. doi:10.1093/bioinformatics/bti610
Corbesier L, Coupland G (2006) The quest for florigen: a review of recent progress. J Exp Bot 57(13):3395–3403. doi:10.1093/jxb/erl095
Demchenko KN, Voitsekhovskaja OV, Pawlowski K (2014) Plasmodesmata without callose and calreticulin in higher plants-open channels for fast symplastic transport? Front. Plant Sci 5:74. doi:10.3389/fpls.2014.00074
Derks A, Schaven K, Bruce D (2015) Diverse mechanisms for photoprotection in photosynthesis. Dynamic regulation of photosystem II excitation in response to rapid environmental change. Biochim Biophys Acta 1847(4–5):468–485. doi:10.1016/j.bbabio.2015.02.008
Dietz KJ, Jacob S, Oelze ML, Laxa M, Tognetti V, de Miranda SM, Baier M, Finkemeier I (2006) The function of peroxiredoxins in plant organelle redox metabolism. J Exp Bot 57(8):1697–1709. doi:10.1093/jxb/erj160
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38(Web Server issue):W64–W70. doi:10.1093/nar/gkq310
Ghosh A, Komar AA (2015) Eukaryote-specific extensions in ribosomal proteins of the small subunit: structure and function. Translation (Austin) 3(1):e999576. doi:10.1080/21690731.2014.999576
Gitelson AA, Gritz Y, Merzlyak MN (2003) Relationships between leaf chlorophyll content and spectral reflectance and algorithms for non-destructive chlorophyll assessment in higher plant leaves. J Plant Physiol 160(3):271–282. doi:10.1078/0176-1617-00887
Guo WJ, Lee N (2006) Effect of leaf and plant age and day/night temperature on net CO2 uptake in Phalaenopsis amabilis var. formosa. J Am Soc Hort Sci 131(3):320–326
Harmut A, Lichtenthaler K (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Method Enzymol 148:350–383. doi:10.1016/0076-6879(87)48036-1
He Y (2012) Chromatin regulation of flowering. Trends Plant Sci 17(9):556–562. doi:10.1016/j.tplants.2012.05.001
Hsiao YY, Pan ZJ, Hsu CC, Yang YP, Hsu YC, Chuang YC, Shih HH, Chen WH, Tsai WC, Chen HH (2011) Research on orchid biology and biotechnology. Plant Cell Physiol 52(9):1467–1486. doi:10.1093/pcp/pcr100
Jack T (2004) Molecular and genetic mechanisms of floral control. Plant Cell. 16(Suppl):S1–S17. doi:10.1105/tpc.017038
Jing X, Hou P, Lu Y, Deng S, Li N, Zhao R, Sun J, Wang Y, Han Y, Lang T, Ding M, Shen X, Chen S (2015) Overexpression of copper/zinc superoxide dismutase from mangrove Kandelia candel in tobacco enhances salinity tolerance by the reduction of reactive oxygen species in chloroplast. Front Plant Sci. 6:23. doi:10.3389/fpls.2015.00023
Kataoka K, Sumitomo K, Fudano T, Kawase K (2003) Changes in sugar content of Phalaenopsis leaves before floral transition. Sci Hortic 102:121–132. doi:10.1016/j.scienta.2003.12.006
Khan MR, Ai XY, Zhang JZ (2014) Genetic regulation of flowering time in annual and perennial plants. Wiley Interdiscip Rev RNA 5(3):347–359. doi:10.1002/wrna.1215
Kim KY, Park SW, Chung YS, Chung CH, Kim JI, Lee JH (2004) Molecular cloning of low-temperature-inducible ribosomal proteins from soybean. J Exp Bot 55(399):1153–1155. doi:10.1093/jxb/erh125
Komatsu S, Han C, Nanjo Y, Altaf-Un-Nahar M, Wang K, He D, Yang P (2013) Label-free quantitative proteomic analysis of abscisic acid effect in early-stage soybean under flooding. J Proteome Res 12(11):4769–4784. doi:10.1021/pr4001898
Kong DL, Yan N, Hu H (2006) Effects of flowering on photosynthesis and allocation of assimilation product in two Cypripedium (Orchidaceae) species. Acta Botanica Yunnanica. 28(6):639–644 (in Chinese)
Krause GH, Weiss E (1991) Chlorophyll fluorescence and photosynthesis: the basics. Ann Rev Plant Physiol 42:313–349. doi:10.1007/978-1-61779-237-3_16
Li DM, Lü FB, Zhu GF, Sun YB, Liu HL, Liu JW, Wang Z (2014) Molecular characterization and functional analysis of a Flowering locus T homolog gene from a Phalaenopsis orchid. Genet Mol Res 13(3):5982–5994. doi:10.4238/2014.August.7.14
Lin MJ, Hsu BD (2004) Photosynthetic plasticity of Phalaenopsis in response to different light environments. J Plant Physiol 161(11):1259–1268. doi:10.1016/j.jplph.2004.05.009
Mattoo RU, Goloubinoff P (2014) Molecular chaperones are nanomachines that catalytically unfold misfolded and alternatively folded proteins. Cell Mol Life Sci 71(17):3311–3325. doi:10.1007/s00018-014-1627-y
Murphy E, De Smet I (2014) Understanding the RALF family: a tale of many species. Trends Plant Sci 19(10):664–671. doi:10.1016/j.tplants.2014.06.005
Newton LA, Runkle ES (2009) High-temperature inhibition of flowering of Phalaenopsis and Doritaenopsis orchids. HortScience 44(5):1271–1276
Omerovic J, Laude AJ, Prior IA (2007) Ras proteins: paradigms for compartmentalised and isoform-specific signaling. Cell Mol Life Sci 64(19–20):2575–2589. doi:10.1007/s00018-007-7133-8
Orelle C, Carlson ED, Szal T, Florin T, Jewett MC, Mankin AS (2015) Protein synthesis by ribosomes with tethered subunits. Nature 524(7563):119–124. doi:10.1038/nature14862
Paradiso R, De Pascale S (2014) Effects of plant size, temperature, and light intensity on flowering of Phalaenopsis hybrids in mediterranean greenhouses. Sci World J 2014:420807. doi:10.1155/2014/420807
Paradiso R, Maggio A, De Pascale S (2012) Moderate variations of day/night temperatures affect flower induction and inflorescence development in Phalaenopsis. Sci Hortic 139:102–107. doi:10.1016/j.scienta.2012.03.007
Perochon A, Aldon D, Galaud JP, Ranty B (2011) Calmodulin and calmodulin-like proteins in plant calcium signaling. Biochimie 93(12):2048–2053. doi:10.1016/j.biochi.2011.07.012
Pnueli L, Gutfinger T, Hareven D, Ben-Naim O, Ron N, Adir N, Lifschitz E (2001) Tomato SP-interacting proteins define a conserved signaling system that regulates shoot architecture and flowering. Plant Cell 13(12):2687–2702. doi:10.1105/tpc.010293
Pollet B, Vanhaecke L, Dambre P, Lootens P, Steppe K (2011) Low night temperature acclimation of Phalaenopsis. Plant Cell Rep 30(6):1125–1134. doi:10.1007/s00299-011-1021-2
Purwestri YA, Ogaki Y, Tamaki S, Tsuji H, Shimamoto K (2009) The 14-3-3 protein GF14c acts as a negative regulator of flowering in rice by interacting with the florigen Hd3a. Plant Cell Physiol 50(3):429–438. doi:10.1093/pcp/pcp012
Ramesh K, Chandrasekaran B (2002) Chlorophyll dynamics in rice (Oryza sativa) before and after flowering based on SPAD (chlorophyll) meter monitoring and its relation with grain yield. J Agron Crop Sci 188:102–105. doi:10.1046/j.1439-037X.2002.00532.x
Shilov IV, Seymour SL, Patel AA, Loboda A, Tang WH, Keating SP, Hunter CL, Nuwaysir LM, Schaeffer DA (2007) The Paragon Algorithm, a next generation search engine that uses sequence temperature values and feature probabilities to identify peptides from tandem mass spectra. Mol Cell Proteom 6(9):1638–1655. doi:10.1074/mcp.T600050-MCP200
Shrestha R, Gómez-Ariza J, Brambilla V, Fornara F (2014) Molecular control of seasonal flowering in rice, Arabidopsis and temperate cereals. Ann Bot. 114(7):1445–1458. doi:10.1093/aob/mcu032
Singh DK, McNellis TW (2011) Fibrillin protein function: the tip of the iceberg? Trends Plant Sci 16(8):432–441. doi:10.1016/j.tplants.2011.03.014
Singh AK, Kumar R, Pareek A, Sopory SK, Singla-Pareek SL (2012) Overexpression of rice CBS domain containing protein improves salinity, oxidative, and heavy metal tolerance in transgenic tobacco. Mol Biotechnol 52(3):205–216. doi:10.1007/s12033-011-9487-2
Srikanth A, Schmid M (2011) Regulation of flowering time: all roads lead to Rome. Cell Mol Life Sci 68(12):2013–2037. doi:10.1007/s00018-011-0673-y
Stone SL (2014) The role of ubiquitin and the 26S proteasome in plant abiotic stress signaling. Front Plant Sci 5:135. doi:10.3389/fpls.2014.00135
Vizcaíno JA, Csordas A, del-Toro N, Dianes JA, Griss J, Lavidas I, Mayer G, Perez-Riverol Y, Reisinger F, Ternent T, Xu QW, Wang R, Hermjakob H (2016) 2016 update of the PRIDE database and related tools. Nucleic Acids Res 44(D1):D447–D456
Wang Y, Lin S, Song Q, Li K, Tao H, Huang J, Chen X, Que S, He H (2014) Genome-wide identification of heat shock proteins (Hsps) and Hsp interactors in rice: Hsp70s as a case study. BMC Genom 15(1):344. doi:10.1186/1471-2164-15-344
Wiśniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6(5):359–362. doi:10.1038/nmeth.1322
Xu SP, Zhu XS, Li C, Ye QS (2014) Effects of CO2 enrichment on photosynthesis and growth in Gerbera jamesonii. Sci Hortic 177:77–84. doi:10.1016/j.scienta.2014.07.022
Yuan XY, Jiang SH, Wang MF, Ma J, Zhang XY, Cui B (2014) Evaluation of internal control for gene expression in Phalaenopsis by quantitative real-time PCR. Appl Biochem Biotechnol 173(6):1431–1445. doi:10.1007/s12010-014-0951-x
Acknowledgments
This work was supported by the Key Technology Project of Henan Province (14B180036) and the Zhengzhou Natural Science Project (141PPTGG420). We thank Dr. Liebo Shu for assistance in proteomic analysis (Shanghai Bo Yuan Biotechnology Co., LTD, China).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Hajduch.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2016_2196_MOESM1_ESM.doc
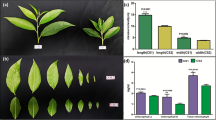
Supplementary material 1. Five phases for sampling. A: Vegetative growth (mature, no spiking plants, VG); B: Spike emergence (spiking with four nodes and on flower bud, SE); C: Floral bud differentiation (spiking with seven nodes and 2 mm floral bud at the sixth node, FBD); D: Mature flower bud (the first flower bud with the diameter of 7 mm, MFB); E: Flowering phase (FP). The youngest fully expanded leaves were collected in the middle region of leaves excepting the midribs for metabolites analysis. The leaves from the vegetative phase (A) and the flowering phase (E) were used for differential protein analysis (DOC 12124 kb)
Rights and permissions
About this article
Cite this article
Yuan, XY., Xu, SP., Liang, F. et al. Comparative proteomic analysis of Phalaenopsis leaves in the vegetative and flowering phase. Acta Physiol Plant 38, 175 (2016). https://doi.org/10.1007/s11738-016-2196-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-016-2196-5