Abstract
The main objective of this study was to analyze the differences in profiles of RNase activities from triticale embryos (Triticosecale, cv. Ugo) between dormant and non-dormant caryopses and to determine the influence of exogenous abscisic acid (ABA) on the activities of these enzymes. The major RNase from the examined tissue was detected following SDS-PAGE, with substrate-based gel assay, described by Yen and Green (Plant Physiol 97:1487–1493, 1991). The activities of enzymes were characterized according to their pH optima, ion dependence, EDTA sensitivity and DNase activity. In embryos with arrested growth (in a natural way by dormancy or artificially by ABA treatment), the activity of two enzymes—24 and 27 kDa—belonging to class I RNases was completely inhibited, whereas that of two other RNases of this family—23 and 25 kDa—was detectable. However, the activity of the class I ribonucleases (enzymes responsible for cellular Pi release) was very low. Moreover, in contrast with non-dormant caryopses, imbibing embryos of dormant or ABA-treated seeds contained 13- and 14-kDa enzymes. These enzymes have not been classified so far, and their specific properties are different from the generally accepted properties of ribonucleolytic enzymes. In addition to the above results, the Pi content in the analyzed samples was determined by the Ames (Methods Enzymol 8:115–118, 1966) method. The results suggest a very low and constant level of inorganic phosphate in dormant samples as well as an evidently decreasing Pi content in embryos under the influence of ABA treatment. The inhibition of the class I RNases activity induced by abscisic acid implies that one of the roles of ABA in seed dormancy may consist in arresting the catabolic release of Pi, which results in retarding the embryo’s growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Seed dormancy is a state in which viable seeds are unable to germinate even under optimal conditions (water, light, temperature, oxygen). Dormancy of cereal crops at harvest is usually a desirable trait because it prevents preharvest sprouting, i.e. precocious germination of grains in the head prior to harvest (Gubler et al. 2005; Fang and Chu 2008). The ability to maintain a dormant state protects seeds against considerable damage to grain quality. Seed dormancy is known to be regulated by a wide range of hormones, among which the most important is abscisic acid (Kermode 2005). Many genetic and physiological studies have shown that abscisic acid (ABA) is responsible for the induction and maintenance of dormancy and the suppression of precocious germination of seeds. Moreover, seed dormancy depends on both ABA content (equally, ABA:GA balance is very important) and ABA sensitivity (Benech-Arnold et al. 1999; Romagosa et al. 2001; Finkelstein et al. 2008; Schramm et al. 2010). It is well known that one of the major actions of ABA in preventing precocious germination of seeds is the inhibition of certain reserve mobilizing enzymes (Robertson et al. 1989; Gómez-Cadenas et al. 1999, 2001; Bethke et al. 2006).
Cereal grain germination involves coordinated action of endosperm and embryonic tissues to mobilize storage reserves of the starchy endosperm (carbohydrates, proteins, lipids, phosphate store: phytin, RNA and DNA). This mobilization is induced by hydrolases secreted from the aleurone layer and embryo. Alongside other main hydrolases (especially amylases, but also proteases, lipases and phytases), ribonucleases are essential enzymes for seed germination. Secretion of RNases and nucleases from the embryo and aleurone into the endosperm during germination is thought to cause the mobilization of RNA and DNA stored in cereal seeds. The decay of the starchy endosperm nucleic acid in turn provides a fraction of the phosphate needed by the growing embryo (Ritchie et al. 2000). An increase in the activity of RNases and other nucleases has also been observed in plants as a response to a variety of stresses, including sucrose or phosphate starvation (Bariola et al. 1994; Gallie et al. 2002). On the contrary, the activity of ribonucleases is inhibited under ABA treatment (Spanò et al. 2008).
The ribonucleases of higher plant cells have been grouped into four major classes (Yen and Green 1991; Green 1994). Class I RNases are highly RNA specific, have molecular masses of 20–25 kDa, a pH optimum of 5–5.5 and low sensitivity to EDTA. Similarly, class II RNases are strictly RNA specific and show low EDTA sensitivity, they have molecular masses of 17–21 kDa and a pH optimum of 6–7. The members of class III RNases are endonucleases (31–35 kDa) capable of hydrolyzing phosphodiester bonds in RNA or DNA (called also group I nucleases, Dohnalek et al. 2011), exhibiting high sensitivity to EDTA and optimum activity at pH of 5–6.5. Finally, class IV RNases are exonucleases (100 kDa) digesting both RNA and DNA, showing optimum activity at pH of 7–9 and high EDTA sensitivity.
The aim of this work was to find differences in the activity of RNases from triticale embryos of dormant and non-dormant caryopses and to evaluate the influence of exogenously applied ABA on these hydrolases. The enzyme activities were characterized according to the specific properties described above. In addition, we estimated the orthophosphate content in embryos from dormant, non-dormant and ABA-treated caryopses.
Materials and methods
Plant material and germination conditions
The experiments were conducted on cv. Ugo triticale (Triticosecale) caryopses, supplied by the Plant Cultivation Station in Strzelce. One batch of grains was stored at −20°C directly after harvesting (dormant caryopses). Another one was kept at room temperature (21°C) for 2 months to break dormancy (after-ripened, non-dormant caryopses) and then stored at −20°C.
Ribonucleases were isolated from the embryonic tissue of dormant and non-dormant triticale caryopses after their imbibition or germination, respectively. Non-dormant caryopses were also treated with ABA to determine the influence of the hormone on changes in the RNase profile. In the first step of the experiment, caryopses were surface disinfected in 0.5% sodium hypochlorite for 20 min and washed with sterilized water. Twenty intact grains were placed on two layers of Whatman paper no 1 (Whatman, Maidstone, Kent, UK) in a Petri dish (∅ 9 cm) containing 15 ml distilled water or 400 μM ABA and incubated for 72 h in the darkness at 20°C.
The content of phosphorus in triticale embryos or germs from dormant, non-dormant and ABA-treated (10, 50, 100 and 400 μM ABA) caryopses was detected every 24 h of imbibition/germination, which was conducted for 7 days.
Extraction of RNases
Ribonucleases were isolated from plant tissue by homogenization of separated embryos/germs in extraction buffer (10 μl/100 mg tissue) at room temperature. Extraction buffer contained 150 mM citric acid–Na2HPO4 (pH 3) and 0.1 mM PMSF. Insoluble material was removed five times by centrifugation at 27,000g for 10 min at 4°C, and soluble extracts were stored at −70°C.
Detection of RNases according to Yen and Green (1991)
The main ribonucleases in triticale embryos/germs were identified by SDS-PAGE procedure, consisting of four main steps (Fig. 1). Firstly, a gel was filled with high-molecular mass RNA so that it would not flow out during electrophoresis. Secondly, RNases were renatured by removal of SDS from the gel while rinsing it with isopropanol (isopropanol in turn was rinsed from the gel with preincubation buffer). Thirdly, the gel was incubated at the pH and temperature appropriate for the RNases. Finally, the gel was negative stained with toluidine blue, which dyed the RNA contained in it while the presence and activity of ribonucleases were evidenced by sites (bands) in the gel which remained colourless.
SDS-PAGE
Before electrophoresis, extracts were directly thawed and clarified by centrifugation at 27,000g for 10 min at 4°C. The protein content in each supernatant was measured according to the method of Bradford (1976). Subsequently, proteins were denatured at room temperature by mixing 1 part of each protein extract with 1 part of 2× sample-loading buffer (2% [w/v] SDS, 10% [v/v] glycerol and 0.025% [w/v] bromophenol blue in 50 mM Tris–HCl buffer, pH 6.8). About 25 μg of the tested proteins and 5 μl standard proteins (prestained SDS-PAGE standards, low range, Bio-Rad) were loaded on the stacking gel consisting of 4.5% [w/v] acrylamide, 0.12% [w/v] bisacrylamide, 0.063 M Tris (pH 6.8), 0.08% [w/v] TEMED and 0.08% [w/v] APS. Separation of enzymes was conducted in the separating gel composed of 11.3% [w/v] acrylamide, 0.3% [w/v] bisacrylamide, 0.46 M Tris–HCl (pH 9), 2.4 mg/ml torula yeast RNA (Sigma), 0.08% [w/v] TEMED and 0.08% [w/v] APS. Electrophoresis was run in a mini-apparatus for electrophoresis (Mini PROTEAN II, Bio-Rad) at constant voltage 200 V for 45 min in running buffer composed of 27.5 mM Tris–HCl, 1.4% [w/v] glycine and 0.1% [w/v] SDS.
Rinsing, incubating, staining and drying of gels
Having completed the electrophoresis, the gels were rinsed twice in 25% [v/v] isopropanol in 0.001 M Tris HCl for 10 min so as to remove SDS. In turn, isopropanol was removed by rinsing the gels twice in preincubation buffer (2 μM ZnCl2 in 0.01 M Tris–HCl) for 10 min. Once SDS and isopropanol have been removed, the gels were incubated in 0.1 M Tris–HCl at 51°C for 50 min to create optimum conditions for the activity of ribonucleases (higher or lower temperatures depress the activity of most RNases). When the incubation was completed, the gels were rinsed in 0.01 M Tris–HCl for 10 min to remove low-molecular products of RNA digestion, and then stained with 0.2% [w/v] toluidine blue solution (Sigma) in 0.01 M Tris–HCl for 10 min. The gels were destained for 10 min and twice for 20 min by rinsing in 0.01 M Tris–HCl. Finally, the polyacrylamide gels were fixed in 10% [v/v] glycerol in 0.01 M Tris–HCl and dried between sheets of cellophane. The Tris–HCl buffers used for rinsing, incubating, staining and fixing of the gels had pH equal to 7, which is optimum for many RNases. All the steps were carried out at room temperature except the incubation.
Modifications
To determine properties of ribonucleases, in some experiments the conditions set for electrophoresis were modified by:
-
removal of zinc ions from the preincubation buffer or increasing their concentration up to 200 μM ZnCl2 in 0.01 M Tris–HCl buffer,
-
addition of disodium versene to the incubation buffer (1 and 10 mM EDTA-Na in 0.01 M Tris–HCl),
-
change in the reaction of all applied Tris–HCl buffers from pH 7 to the range of 3 to 9,
-
addition of ssDNA from herring sperm (Sigma) to the gel matrix (0.6 mg/ml) instead of RNA while simultaneously changing the pH of the Tris–HCl buffer used to prepare the ssDNA gel from 9 to 8.7.
Estimation of inorganic phosphate by the Ames (1966) method
Inorganic phosphorus was determined in embryos or germs isolated from dormant, non-dormant and ABA-treated caryopses (10, 50, 100 and 400 μM ABA). For this aim, 400 mg of plant tissue was homogenized in a cold mortar in 6 ml 10% of ice-cold TCA. The homogenate was centrifuged at 6,000g for 10 min. The supernatant was rinsed twice with 6 ml 5% TCA. Next, combined supernatants were centrifuged again and incubated for 15 min on ice. After filtering, the solutions were used for determination of inorganic phosphorus. Each extraction step was carried out at 4°C.
1.4 ml Ames reagent was added to 0.6 ml of the extract prepared as above and incubated in a water bath at 45°C for 20 min. Ames reagent was produced immediately before use by mixing 1 part of 10% ascorbic acid with 6 parts of 0.42% ammonium molybdate in 1 N H2SO4. The determination of Pi was based on two reactions: (1) formation of ammonium phosphomolybdate from orthophosphate and ammonium molybdate in acid environment, and (2) reduction of phosphomolybdic acid by the reducing agents contained in Ames reagent to blue molybdenum oxides. The presence of molybdenum oxides was detected at 820 nm against a control sample (containing Ames reagent and distilled water instead of the plant extract). The concentration of phosphorus in the analyzed samples was read from the standard curve made with a solution of KH2PO4 containing Pi from 0.1 to 2 μg. The concentration of phosphorus was expressed in mg/g dry matter of the analyzed tissue.
Results
Differences in the RNase activities
Using the methods described above, we identified five ribonucleases in germs of non-dormant caryopses (control sample): three class I RNases (20, 24, 27 kDa) and two class III RNases (33, 35 kDa). In addition, we found that during the first step of germination of non-dormant caryopses, two class III RNases (33 and 35 kDa) and one class I RNase (20 kDa) were synthesized in embryos or germs. The activity of the other class I RNases (with masses of 24 and 27 kDa) started to appear with an increasing level of tissue hydration beginning 24 h after caryopses germination (Fig. 2).
The results show that in embryos with arrested growth (in a natural way by dormancy or artificially by ABA treatment), the activity of class I RNases (24 and 27 kDa) was completely inhibited, whereas the three other RNases of this family (20, 23 and 25 kDa) could be detected. However, the activity of class I ribonucleases, enzymes responsible for cellular Pi release was very low. Moreover, in contrast with non-dormant caryopses, imbibing embryos of dormant or ABA-treated caryopses contained 13 and 14 kDa enzymes. These enzymes have not been classified so far. Besides, their specific properties were different from the generally accepted properties of nucleolytic enzymes. Similar to the control, in the dormant and ABA-treated samples we found the same two enzymes of class III RNases—33 and 35 kDa proteins (Figs. 3, 4, 5).
Properties of RNases
pH optima
Changes in the activity of ribonucleases were observable by applying buffers of different pH (3–9) while rinsing, incubating and staining gels (Fig. 3).
Class III RNases (33 and 35 kDa) extracted from embryos of dormant and ABA-treated caryopses showed optimum activity at pH 6, whereas the activity of the control enzymes (with the same masses) was the highest at pH 7.
The main RNases of class I (involved in Pi release) from dormant and ABA-treated samples (23 and 25 kDa) had the same optimum activity (pH 5) as RNases from the control (20, 24, 27 kDa) with the exception of the 20-kDa enzyme, whose optimum activity was at pH 4. In contrast to the control, activities of all class I RNases from dormant or ABA-treated tissue were lower at pH 6, and completely inhibited at pH 7.
Ribonucleases with masses of 13 and 14 kDa, characteristic for dormant and ABA-treated samples, showed the highest activity at pH 8.
Ion dependence
The addition of low concentrations of ZnCl2 (2 μM) in the preincubation buffer enhanced the activity of all RNases, whereas the presence of higher concentrations of ZnCl2 (20 and 200 μM) in the preincubation buffer inhibited the activity of enzymes except for two ribonucleases with masses of 13 and 14 kDa, whose activity was increased (Fig. 4).
EDTA sensitivity
To find out whether the activity of RNases raised by ZnCl2 is sensitive to chelation of bivalent ions, gels preincubated in 2 μM ZnCl2 were subsequently incubated in 1 or 10 mM solution of EDTA-Na. Only two ribonucleases with masses of 33 and 35 kDa, were sensitive to EDTA-Na. The activity of the other enzymes was greatly enhanced by both 1 and 10 mM EDTA-Na (Fig. 5).
In the control sample, disodium versene most strongly increased the activity of RNases with masses of 20 and 27 kDa and induced the 24-kDa enzyme (RNases with optimum activity at pH 5).
In dormant and ABA-treated samples, 1 mM EDTA-Na induced ribonucleases with masses of 23 and 25 kDa (whose activity optimum was at pH 5), whereas 10 mM EDTA-Na induced the 20-kDa ribonuclease (with optimum activity at pH 4). Furthermore, disodium versene enhanced the activity of RNases with masses of 13 and 14 kDa.
DNase activity
The 33 and 35 kDa RNases also showed DNase activity (Fig. 6) and are therefore nucleases hydrolyzing phosphodiester bonds in both RNA and DNA.
Changes in the content of inorganic phosphorus
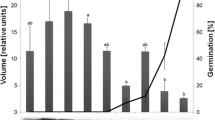
Differences in the content of Pi in embryo/germ between dormant, non-dormant and ABA-treated caryopses are shown in Fig. 7. In the dormant sample, a constant, low level of inorganic phosphorus was observed throughout the whole period of imbibition of caryopses. On the other hand, the content of Pi was distinctly decreasing in non-dormant caryopses under the influence of exogenously applied ABA (the observed changes were ABA dose dependent).
Discussion
The experiments presented in this paper have enabled us to identify the major RNases from triticale embryos and to show differences in the enzyme profile under natural seed dormancy and changes induced by exogenous ABA. RNases were identified according to the distinguishing properties of these enzymes.
It is assumed that the activity of plant RNases is adversely affected by zinc ions (Nguyen et al. 1988). Yen and Green (1991) proved that it was true only under the effect of high Zn2+ concentrations, whereas low concentrations of zinc ions (2 μM ZnCl2) stimulate the activity of some ribonucleases. In this research, the activity of all identified enzymes increased under the influence of 2 μM ZnCl2. A positive effect of zinc ions on the activity of the 13 and 14 kDa RNases (from embryos of dormant and ABA-treated caryopses) was also observed at high concentrations of ZnCl2 (20 and 200 μM).
It is commonly assumed that many nucleases responsible for hydrolytic degradation of ssDNA are zinc metalloproteins (Desai and Shankar 2003). Here we show that the 33 and 35-kDa enzymes were the only ones which showed both RNase and DNase activity, which was enhanced in the presence of zinc ions and completely inhibited by chelation of these ions with EDTA. Yen and Green (1991) suggest that Zn2+ ions can cause both an increase in the activity of ribonucleases and their renaturation. Therefore, in this paper, in accordance with the method suggested by Yen and Green (1991), ions of this metal were added to preincubation buffer during the ongoing renaturation of enzymes. This phenomenon explains why enzymes insensitive to chelation are activated by Zn2+ ions. Based on the above assumption, we concluded that the 33 and 35-kDa (class III) enzymes identified in this study are probably metalloproteins, which share both attributes: Zn2+ dependence and EDTA sensitivity, whereas the activity of the other enzymes, insensitive to EDTA, increased only under the positive effect of Zn2+ on their renaturation.
During their experiments on the effect of monovalent cations on the activity of ribonucleases, Yen and Green (1991) concluded that the presence of NaCl and KCl in buffers used for preparation, rinsing, preincubation and incubation of gels caused a small albeit positive effect on the isolated enzymes. Interesting results have been obtained in our study resulting from the application of EDTA-Na. EDTA, the chelator of divalent ions, formed complexes with zinc cations, leaving Na+ and Cl− ions in the solution. Thus, the application of EDTA-Na during the incubation of gels enabled us to determine the effect of Na+ on the RNase activity. We therefore concluded that all the enzymes, except class III riboucleases (sensitive to EDTA), were activated by Na+ cations. The effect was the most distinct in the case of class I RNases.
Exogenous ABA strongly inhibited the activity of class I ribonucleases and, in addition, caused complete inhibition of the activity of two enzymes (24 and 27 kDa) in embryos, in which two other RNases with masses of 23 and 25 kDa appeared instead. However, the activity of the two latter hydrolases (similarly to that of another enzyme with mass of 20 kDa belonging to the same class) was low. Exactly identical relationships were found in embryos of dormant caryopses. Because of the similar properties, acid RNases with masses of 23, 24, 25 and 27 kDa probably make up a group of isoenzymes. Moreover, we consider the possibility of the conversion of the 24 and 27 kDa RNases to the 23 and 25 kDa enzymes.
Among ribonucleases, a non-uniform group of enzymes is distinguished, known as S-like RNases. The polymorphism of genes which code them (differences in the DNA sequence and position of loci in chromosomes) shapes not only the species- or cultivar-specific variability but also the tissue specificity (Ma and Oliveira 2000; Kock et al. 2006; MacIntosh et al. 2010). Acid RNases PD2 (from almond tree) and RSH1 (from barley) of the mass 25 kDa have been classified as belonging to this group of enzymes (Ma and Oliveira 2000; Gausing 2000). Results of numerous studies confirm that S-like RNases are synthesized in response to environmental stress, including phosphate starvation (Bariola et al. 1999; Ma and Oliveira 2000; Kock et al. 2006). In response to a decreased concentration of Pi in a cell, class I RNases, which are for instance responsible for the release of Pi during RNA degradation, are accumulated in the vacuole (Chen et al. 2000). Bariola et al. (1999) concluded that Arabidopsis in response to phosphorus deficit produces RNS1 (S-like acid ribonuclease) present in the vacuole. The same authors report that a low content of RNS1 is strongly correlated with a high level of anthocyanins.
It is well known that caryopses of red wheat are more resistant to sprouting than those of white wheat and that an increase in the synthesis of anthocyanins is associated, inter alia with phosphorus deficiency. Quick et al. (1997) proved that embryos of dormant caryopses of the common wild oat (Avena fatua L.) contained less Pi, whose level started to increase during post-harvest maturation. In dormant seeds, positive correlation has been shown between the content of anthocyanins and their sensitivity to ABA (Himi et al. 2002). Bewley (1997) reports that in mutants of Arabidopsis, which lack the genes ABI3 (responsible for seeds’ sensitivity to ABA), the synthesis of anthocyanin biosynthesis pathway enzymes is likewise halted.
In contrast Bethke et al. (1996) found positive correlation between the content of GA3 (the phytohormone responsible for germination and acting as an antagonist to ABA) and the level of Pi. These authors suggest that in response to an increased content of GA3 in non-dormant caryopses, starch becomes degraded, the pH of vacuolar sap becomes more acidic and, in consequence, phytase—an enzyme that can break down phytin (a main storage form of phosphorus)—is activated. Moreover, a change in pH in vacuoles from neutral (6.6–7.0) to acid (5.8 and less) favours activation of class I ribonucleases, causing further release of Pi. An increase in the content of phosphorus in non-dormant caryopses is the final event which contributes to increased metabolism of cells and consequently to the growth and development of an embryo. Furthermore, Quick et al. (1997) report that the effect of exogenous Pi on metabolism of carbohydrates resembles that of GA3.
Based on this work and that of others (Chalker-Scott 1999; Rubio et al. 2001; Akihiro et al. 2005; McKibbin et al. 2006; Badone et al. 2010) we present a hypothesis on the role of ABA in regulating the dormancy mechanism of caryopses. This hypothesis states that by inhibiting the activity of class I ribonucleases, ABA helps to inhibit the release of inorganic phosphorus, which may favour both retardation of the embryo’s growth and accumulation of starch granules and anthocyanins. This hypothesis is in accord with the assumptions made by other researchers, who suggest that ABA stimulates synthesis of the α-amylase inhibitor and enzymes of the metabolic pathway of anthocyanins (Robertson et al. 1989; Hattori et al. 1992; White and Rivin 2000; Furtado et al. 2003).
Abbreviations
- ABA:
-
Abscisic acid
- APS:
-
Ammonium persulfate
- DNase:
-
Deoxyribonuclease
- EDTA-Na:
-
Disodium ethylenediaminetetraacetate (disodium versene)
- GA:
-
Gibberellic acid
- Pi :
-
Orthophosphate
- PMSF:
-
Phenylmethylsulfonyl fluoride
- RNase:
-
Ribonuclease
- SDS-PAGE:
-
Sodium dodecyl sulphate polyacrylamide gel electrophoresis
- ssDNA:
-
Single-stranded DNA
- TCA:
-
Trichloroacetic acid
- TEMED:
-
N,N,N′,N′-tetramethyl-ethylenediamine
References
Akihiro T, Mizuno K, Fujimura T (2005) Gene expression of ADP-glucose pyrophosphorylase and starch contents in rice cultured cells are cooperatively regulated by sucrose and ABA. Plant Cell Physiol 46(6):937–946
Ames BN (1966) Assay of inorganic phosphate, total phosphate and phosphatase. Methods Enzymol 8:115–118
Badone FC, Cassani E, Landoni M, Doria E, Panzeri D, Lago C, Francesca M, Nielsen E, Pilu R (2010) The low phytic acid1-241 (lpa1-241) maize mutation alters the accumulation of anthocyanin pigment in the kernel. Planta 231:1189–1199
Bariola PA, Howard CJ, Taylor CB, Verburg MT, Jaglan VD, Green PJ (1994) The Arabidopsis ribonuclease gene RNS1 is tightly controlled in response to phosphate limitation. Plant J 6(5):673–685
Bariola PA, MacIntosh GC, Green PJ (1999) Regulation of S-like ribonuclease levels in Arabidopsis. Antisense inhibition of RNS1 or RNS2 elevates anthocyanin accumulation. Plant Physiol 119:331–342
Benech-Arnold RL, Giallorenzi MC, Frank J, Rodriguez V (1999) Termination of hull-imposed dormancy in developing barley grains is correlated with changes in embryonic ABA levels and sensitivity. Seed Sci Res 9:39–47
Bethke PC, Hillmer S, Jones RL (1996) Izolation of intact protein vacuoles from barley aleurone—identification of aspartic and cysteine proteases. Plant Physiol 110:521–529
Bethke PC, Hwang Y, Zhu T, Jones RL (2006) Global patterns of gene expression in the aleurone of wild-type and dwarf1 mutant rice. Plant Physiol 140:484–498
Bewley JD (1997) Seed germination and dormancy. Plant Cell 9:1055–1066
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Chalker-Scott L (1999) Environmental significance of anthocyanins in plant. Stress Responses Photochem Photobiol 70(1):1–9
Chen DL, Delatorre CA, Bakker A, Abel S (2000) Conditional identification of phosphate-starvation-response mutants in Arabidopsis thaliana. Planta 211:13–22
Desai NA, Shankar V (2003) Single-strand-specific nucleases. FEMS Microbiol Rev 26:457–491
Dohnalek J, Koval T, Lipovova P, Podzimek T, Matousek J (2011) Structure analysis of group I plant nucleases. J Synchrotron Rad 18(1):29–30
Fang J, Chu C (2008) Abscisic acid and the pre-harvest sprouting in cereals. Plant Signal Behav 3(12):1046–1048
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
Furtado A, Henry R, Scott K, Meech S (2003) The promoter of the asi gene directs expression in the maternal tissues of the seed in transgenic barley. Plant Mol Biol 52:787–799
Gallie DR, Chang S-C, Young TE (2002) Induction of RNase and nuclease activity in cultured maize endosperm cells following sucrose starvation. Plant Cell Tissue Organ Cult 68:163–170
Gausing K (2000) A barley gene (rsh1) encoding a ribonuclease S-like homologue specifically expressed in young light-grown leaves. Planta 210:574–579
Gómez-Cadenas A, Verhey SD, Holappa LD, Shen Q, Ho THD, Walker-Simmons MK (1999) An abscisic acid-induced protein kinase, PKABA1, mediates abscisic acid-suppressed gene expression in barley aleurone layers. Proc Natl Acad Sci USA 96:1767–1772
Gómez-Cadenas A, Zentella R, Walker-Simmons MK, Ho THD (2001) Gibberellin/abscisic acid antagonism in barley aleurone cells: site of action of the protein kinase PKABA1 in relation to gibberellin signaling molecules. Plant Cell 13:667–679
Green PJ (1994) The ribonucleases of higher plants. Annu Rev Plant Physiol Plant Mol Biol 45:415–421
Gubler F, Millar AA, Jacobsen JV (2005) Dormancy release, ABA and pre-harvest sprouting. Curr Opin Plant Biol 8:183–187
Hattori T, Vasil V, Rosenkrans L, Hannah LC, McCarty DR, Vasil IK (1992) The Viviparous-1 gene and abscisic acid activate the CI regulatory gene for anthocyanin biosynthesis during seed maturation in maize. Genes Dev 6:609–618
Himi E, Mares DJ, Yanagisawa A, Noda K (2002) Effect of grain colour gene (R) on grain dormancy and sensitivity of the embryo to abscisic acid (ABA) in wheat. J Exp Bot 53(374):1569–1574
Kermode AR (2005) Role of abscisic acid in seed dormancy. J Plant Growth Regul 24:319–344
Kock M, Stenzel I, Zimmer A (2006) Tissue-specific expression of tomato ribonuclease LX during phosphate starvation-induced root growth. J Exp Bot 57(14):3717–3726
Ma R-C, Oliveira MM (2000) The RNase PD2 gene of almond (Prunus dulcis) represents an evolutionarily distinct class of S-like RNase genes. Mol Gen Genet 263:925–933
MacIntosh GC, Hillwig MS, Meyer A, Flagel L (2010) RNase T2 genes from rice and the evolution of secretory ribonucleases in plants. Mol Genet Genomics 283:381–396
McKibbin RS, Muttucumaru N, Paul MJ, Powers SJ, Burrell MM, Coates S, Purcell PC, Tiessen A, Geigenberger P, Halford NG (2006) Production of high-starch, low-glucose potatoes through over-expression of the metabolic regulator SnRK1. Plant Biotechnol J 4:409–418
Nguyen IT, Palcic MM, Hadziyev D (1988) Characterization of cell-wall-bound nuclease and ribonuclease from potato tuber. Agric Biol Chem 52:957–965
Quick WA, Hsiao AI, Hanes JA (1997) Dormancy implications of phosphorus levels in developing caryopses of wild oats (Avena fatua L.). J Plant Growth Regul 16:27–34
Ritchie S, Swanson AJ, Gilroy S (2000) Physiology of the aleurone layer and starchy endosperm during grain development and early seedling growth: new insights from cell and molecular biology. Seed Sci Res 10:193–212
Robertson M, Walker-Simmons M, Munro D, Hill RD (1989) Induction of α-amylase inhibitor synthesis in barley embryos and young seedlings by abscisic acid and dehydration stress. Plant Physiol 91:415–420
Romagosa I, Prada D, Moralejo MA, Sopena A, Munoz P, Casas AM, Swanston JS, Molina-Cano JL (2001) Dormancy, ABA content and sensitivity of a barley mutant to ABA application during seed development and after ripening. J Exp Bot 52:1499–1506
Rubio V, Linhares F, Solano R, Martín AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Schramm EC, Abellera JC, Strader LC, Campbell KG, Steber CM (2010) Isolation of ABA-responsive mutants in allohexaploid bread wheat (Triticum aestivum L.): drawing connections to grain dormancy, preharvest sprouting, and drought tolerance. Plant Sci 179:620–629
Spanò C, Buselli R, Grilli I (2008) Dormancy and germination in wheat embryos: ribonucleases and hormonal control. Biol Plant 52(4):660–667
White CN, Rivin CJ (2000) Gibberellins and seed development in maize. II. Gibberellin synthesis inhibition enhances abscisic acid signaling in cultured embryos. Plant Physiol 122:1089–1097
Yen Y, Green PJ (1991) Identification and properties of the major ribonucleases of Arabidopsis thaliana. Plant Physiol 97:1487–1493
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Abe.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Szypulska, E., Weidner, S. ABA may affect the dormancy in triticale caryopses by inhibition of embryo class I RNases, enzymes involved in Pi release. Acta Physiol Plant 34, 877–884 (2012). https://doi.org/10.1007/s11738-011-0885-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11738-011-0885-7