Abstract
We introduce the Robust Logistic Zero-Sum Regression (RobLZS) estimator, which can be used for a two-class problem with high-dimensional compositional covariates. Since the log-contrast model is employed, the estimator is able to do feature selection among the compositional parts. The proposed method attains robustness by minimizing a trimmed sum of deviances. A comparison of the performance of the RobLZS estimator with a non-robust counterpart and with other sparse logistic regression estimators is conducted via Monte Carlo simulation studies. Two microbiome data applications are considered to investigate the stability of the estimators to the presence of outliers. Robust Logistic Zero-Sum Regression is available as an R package that can be downloaded at https://github.com/giannamonti/RobZS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Over the past decade, the interest in understanding the importance of the role of the microbiome in human health has increased, especially in studies concerning the association of a medical status with the microbial communities, providing new ways to classify individuals, and to predict their disease risks (Qin et al. 2010). This growing interest is motivated by the diffuse use of high-throughput sequencing technologies, such as the approach based on sequencing of 16S ribosomal RNA gene, which is ever-present in all bacterial genomes, or the approach based on shotgun metagenomic sequencing. The resulting sequencing reads are vectors of bacterial taxa abundances, that generally are clustered into operational taxonomic units (OTUs) at different taxonomic levels. The analysis of these data is a statistical and computational challenge as they are typically high-dimensional, sparse, zero inflated due to the presence of many rare taxa, and compositional (Gloor et al. 2017). In fact, the total sequence read counts of the subjects can vary significantly from sample to sample, so that the data should be normalized before the analysis. For a given sample, the resulting microbiome dataset is essentially a compositional matrix, in which each row contains information on relative OTUs. A common normalization is to standardize each row to sum up to one.
This paper considers logistic regression analysis of microbiome compositional data, with the aim to identify the bacterial taxa that are associated with a dichotomous response, such as a medical status of interest. The goal is twofold: to classify the subjects on the basis of the estimated model, and to perform variable selection, namely to select the most relevant taxa associated to the response of interest. Standard logistic regression should not be implemented due to the unit sum normalization of the covariates; they are in fact totally collinear.
Several methods to perform regression with compositional explanatory variables are available in the literature: Aitchison and Bacon-Shone (1984) proposed the linear log-contrast model for continuous response applying the log-ratio transformation Aitchison (1982) to compositional covariates. The critical point of this proposal is the arbitrariness in the choice of a reference taxon, but also the estimation results become unstable when the number of predictors by far exceeds the number of observations.
In the high-dimensional setting, Lin et al. (2014) considered variable selection in the context of regression with compositional covariates for continuous response by imposing a zero-sum constraint on the regression coefficients and an \(\ell _1\) penalty to the likelihood function. Lu et al. (2019) extended the zero-sum model to the generalized linear regression framework, while Zacharias et al. (2017) applied an elastic-net regularization to the logistic zero-sum model.
The penalized logistic regression performs stable estimation and avoids overfitting, but, since it is based on the maximum likelihood method, it suffers from outliers, producing unreliable classification results. A robust approach could overcome this disadvantage. However, in the high dimensional setting it is arduous or even impossible for the practitioner to identify outliers or observations that deviate somehow from an underlying model. Therefore, outliers need to be automatically identified and downweighted in the estimation procedure of a robust estimator. Some robust procedures are already available in the literature. Among others, Avella-Medina and Ronchetti (2017) proposed a robust penalized quasi-likelihood estimator for generalized linear models, Park and Konishi (2016) suggested a robust penalized logistic regression based on a weighted likelihood methodology, and Kurnaz et al. (2018) adopted a trimmed elastic-net estimator for linear and logistic regression. However, none of these options satisfy the zero-sum constraint.
This paper presents a Robust Logistic Zero-Sum Regression (RobLZS) model with compositional explanatory variables. The RobLZS method attains robustness by minimizing a trimmed sum of deviances. The suggested method can be applied in various fields of research, such as in biostatistics, but also in medicine, economics, ecology, demography, psychology and many more.
The rest of this paper is organized as follows. Section 2 presents the regression methods for compositional covariates and fleshes out our proposed robust estimator. Section 3 shows a Monte Carlo simulation to investigate the performance of RobLZS with respect to other competing estimators, Sect. 4 presents results from an analysis of two real microbiome studies, and Sect. 5 concludes.
2 Sparse logistic regression models with compositional covariates
In the usual linear regression setup, a response variable \(Y_i \in \mathbb {R}\) is connected to a vector of covariates \( \mathbf {X} _i\in \mathbb {R}^p\) by a linear model \(\mathbb {E}(Y_i| \mathbf {X}_i= \mathbf {x}_i)=\beta _0+ \mathbf {x}_i^{\mathrm {T}} \varvec{\beta }\), \((i=1, \ldots ,n)\), with the regression coefficients \( \varvec{\beta } \in \mathbb {R}^p\).
To take into account the compositional nature of the covariates vector, we can assume that each vector \( \mathbf {x}_i\) lies in the unit simplex \( {\mathscr {S}} ^p=\{ \mathbf {x}_i=(x_{i1}, \ldots x_{ip})^{\mathrm {T}}: x_{ij}>0, \text { for } j=1, \ldots , p,\,\) \(\text { and } \sum _{j=1}^px_{ij}=1\}\). The standard log-contrast model by Aitchison and Bacon-Shone (1984) is defined as \(\mathbb {E}(Y_i| \mathbf {Z}^p_i= \mathbf {z}_i^p)=\beta _0+ \mathbf {z}_i^{p\mathrm {T}} \varvec{\beta }_{\setminus p}\), where \( \mathbf {Z}_i^p \in \mathbb {R}^{n \times (p-1)}\) is the log-ratio design matrix, with \(z_{ij}^p=\log \big (\frac{x_{ij}}{x_{ip}}\big )\), p denotes the reference component, and \( \varvec{\beta }_{\setminus p}=(\beta _1, \ldots , \beta _{p-1})\) is the vector of \((p-1)\) coefficients. Lin et al. (2014) reformulated the linear log-contrast model in a symmetric form introducing linear constraints on the coefficients,
where \( \mathbf {z}_i=\log ( \mathbf {x}_i)\) are log-transformed covariates. For the sake of simplicity, and without loss of generality, we assume that the intercept \(\beta _0\) is zero, although our formal justification will allow for an intercept.
Model (1) exempts us from choosing the reference component, as it was necessary in the aforementioned standard log-contrast model by Aitchison and Bacon-Shone (1984) , while gaining interpretability.
Note that the zero-sum constraint in (4) is crucial for an estimator of regression coefficients to fulfill the desirable properties of compositional data analysis, namely the scale invariance, the permutation invariance, and the subcompositional coherence properties (Aitchison 1986). The scale invariance property guarantees that the regression coefficient \( \varvec{\beta }\) is independent from an arbitrary scaling of the basis count from which the composition is obtained, i.e. \(\log (\delta \mathbf {x}_i)^{\mathrm {T}} \varvec{\beta }=\log ( \mathbf {x}_i)^{\mathrm {T}} \varvec{\beta }\), for any constant \(\delta \). The permutation invariance property, i.e. the estimator is unchanged if we permute the columns of \( \mathbf {Z}\) and the elements of \( \varvec{\beta }\) in the same way, derives directly from the symmetric form of (1). The subcompositional coherence states that the regression coefficients \( \varvec{\beta }\) remain unaffected by correctly excluding some or all of the zero components (Lin et al. 2014). It is important to remember that each coefficient \(\beta _j\) should be interpreted in the context of the other non-zero coefficients. Because of the zero-sum constraint, the regression coefficients split up the full composition of regressors into two subsets of variables: taxa with a positive regression coefficient and those with a negative coefficient. Therefore, the fitted regression model depicts the relationship, or balance, between these groups of parts.
Note that applying the standard tool kit of linear regression analysis to the standard log-contrast model does not guarantee solutions that are permutation invariant, due to its asymmetric form.
Model (4) could be extended to the generalized linear model (GLM) framework, in which the density function of the outcome is a member of the exponential family
where \(\nabla \) denotes the gradient. In case of binary outcome, a two-class logistic regression model is often used, and thus we have
with the corresponding log-likelihood
where \(d( \mathbf {z}_i^{\mathrm {T}} \varvec{\beta },y_i)=-y_i\mathbf {z}_i^{\mathrm {T} } \varvec{\beta }+ A( \mathbf {z}_i^{\mathrm {T}} \varvec{\beta })\) is the deviance for the ith component. In the high-dimensional setting, when \(n \ll p\), a sparse solution for the estimation of the parameter \( \varvec{\beta }\) can be obtained by using a penalized negative log-likelihood. Thus, the penalized estimate of \( \varvec{\beta }\) is the solution of the optimization problem
and it is called the Logistic Zero-Sum (LZS) estimator. \(P_{\alpha }( \varvec{\beta })\) is the elastic-net regularization penalty (Zou and Hastie 2005), defined as
where \(\alpha \in [0,1]\) and \( \lambda \in [0, \infty )\) are the tuning parameters: \(\alpha \) balances the \(\ell _2\) and \(\ell _1\) penalizations, while \(\lambda \) controls the sparsity of the solution.The zero-sum constraint carries interpretation benefit in penalized regression, where each regression coefficient represents the effect of a variable on the outcome, adjusting for all other selected variables.
Lu et al. (2019) imposed a lasso penalty (setting \(\alpha =1\) in \(P_{\alpha }( \varvec{\beta })\)) to the estimator (4), while Zacharias et al. (2017) considered an elastic-net regularization, and they adopted a coordinate descent algorithm to fit logistic elastic-nets with zero-sum constraints. Bates and Tibshirani (2019) showed a link between the model (1) and the model that includes as covariates the log of all pairwise ratios, suggesting a different interpretation of the linear log-contrast model.
2.1 The RobLZS estimator
The estimator for \( \varvec{\beta }\) in (4) is based on the maximum log-likelihood method, where every observation enters the log-likelihood function with the same weight. Thus, the estimator is not robust against the presence of outliers, which can lead to unreliable classification results. Commonly, outliers in logistic regression can be classified into leverage points, which are deviating points in the space of the covariates, vertical outliers, which are mislabeled observations in the response, or outliers in both spaces (Nurunnabi and West 2012).
We consider here a penalized maximum trimmed likelihood estimator, an analog for the generalized linear model of the sparse least trimmed squares (LTS) estimator for robust high-dimensional linear models (Alfons et al. 2013; Neykov et al. 2014; Kurnaz et al. 2018). We call our proposal the Robust Logistic Zero-Sum estimator (hereafter indicated by the acronym RobLZS).
The RobLZS estimator is a penalized minimum divergence estimator, as it uses a trimmed sum of deviances. The elastic-net penalty is considered in the penalization, which enables variable selection and estimation at the same time, and effectively deals with the existence issue of the estimator in case of non-overlapping groups (Albert and Anderson 1984; Friedman et al. 2010). In the estimation process, only the best subset of h observations with the smallest deviances are considered. Then a system of robustness weights is computed within the algorithm, in a similar way as for the robust weighted Bianco-Yohai (BY) estimator for logistic regression (Bianco and Yohai 1996). The final estimator is computed by considering all the observations in the sample, but with weights assigned according to their outlyingness.
The algorithm to obtain \( \hat{ \varvec{\beta }}_{\text {RobLZS}}\) is detailed in Sect. 2.2. The selection of the tuning parameters \(\alpha \) and \(\lambda \) will be discussed in Sect. 2.3, and an extensive Monte Carlo simulation study, reported in Sect. 3, demonstrates the robustness of the estimator in presence of data outliers, suggesting that the RobLZS estimator is an effective tool for the classification task as well as for variable selection.
2.2 Algorithm
The proposed algorithm is conform to the fast-LTS algorithm (Rousseeuw and Van Driessen 2006), which has been extended to the high-dimensional setting (Alfons et al. 2013).
For a fixed combination of the tuning parameters \(\alpha \) and \(\lambda \), the objective function of the RobLZS estimator has the form
based on a subsample of observations, where H is an outlier-free subset of the set of all indexes \(\{1, 2,\ldots , n\}\), and |H| denotes the cardinality of set H, with \(|H|=h \le n\), and \(P_{\alpha }( \varvec{\beta })\) is the elastic-net regularization penalty as in (4). For each subsample given by the set H we can obtain \(\hat{ \varvec{\beta }}_H\) as
The optimal solution \(\hat{ \varvec{\beta }}_{opt}\) is given by,
where
hence, \(\hat{ \varvec{\beta }}_{opt}\) is obtained as the LZS estimator applied to the optimal subset of \(h \le n\) observations which lead to the smallest penalized sum of deviances, where the zero-sum constraint needs to be preserved.
The optimal subset \(H_{opt}\) is obtained by using a modification of the fast-LTS algorithm, based on iterated concentration steps (C-steps) (Rousseeuw and Van Driessen 2006) on diverse initial subsets, which we describe in the following.
At iteration \(\kappa \), let \(H_{\kappa }\) denote a certain subsample with \(|H_{\kappa }|= h ={\lfloor }\xi (n+1){\rfloor }\), \(\xi \in [0.5, 1]\) with \(1-\xi \) the trimmed portion, and \({\lfloor }.{\rfloor }\) means rounding down to the nearest integer. In this article we choose \(\xi =0.75\), thus \((1-\xi )\%=\) 25% is an initial guess of the maximum outlier proportion in the sample.
Let \(\hat{ \varvec{\beta }}_{H_{\kappa }}\) be the coefficients of the corresponding zero-sum fit, see Model (4). After computing the deviances \(d( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_{\kappa }},y_{i})\), for \(i=1, \ldots , n\), the subsample \(H_{\kappa +1}\) for iteration \(\kappa +1\) is defined as the set of indices corresponding to the h smallest deviances. These indexes are subsequently intended to point at outlier-free observations, and their group composition should be in the same proportion as for the whole (training) data set. Thus, let \(n_0\) and \(n_1\) be the numbers of observations in the two groups, with \(n=n_0+n_1\). Then \(h_0= {\lfloor }\xi (n_0+1){\rfloor }\) and \(h_1=h-h_0\) define the group sizes in each h-subset. A new h-subset is created with the \(h_0\) indexes with the smallest deviances \(d( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_{\kappa }},y_{i}=0)\) and with the \(h_1\) indexes with the smallest deviances \(d( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_{\kappa }},y_{i}=1)\).
Let \(\hat{ \varvec{\beta }}_{H_{\kappa +1}}\) denote the coefficients of the LZS fit based on the subset \(H_{\kappa +1}\). It is straightforward to derive that
We can see that a C-step results in a decrease of the objective function, and that the algorithm iteratively converges to a local optimum in a finite number of steps. In order to increase the chance to approximate the global optimum, a large number of random initial subsets \(H_0\) of size h for any sequence of C-steps should be used. Each initial subset \(H_0\) is obtained through a search with elemental subsets of size 4, two from each group, as suggested by Kurnaz et al. (2018). This elemental subset is used to grow the likelihood, and such a small subset of observations has a higher chance to be outlier-free.
For a fixed combination of the tuning parameters \(\lambda \ge 0\) and \(\alpha \in [0,1]\), the implemented algorithm is as follows:
-
1.
Draw s (we choose \(s=500\) to increase the chance to get the global minimum) random initial elemental subsamples \(H_s^{el}\) of size 4, and let \(\hat{ \varvec{\beta }}_{H_s^{el}}\) be the corresponding estimated coefficients.
-
2.
For all s subsets and estimated coefficients \(\hat{ \varvec{\beta }}_{H_s^{el}}\), the deviances \(d( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_s^{el}},y_{i})\) are computed for all observations \(i=1, \ldots , n\). Then two C-steps are carried out, starting with the h-subset defined by the indexes of smallest values of the deviances.
-
3.
Retain only the best \(s_1=10\) subsets of size h, and for each subsample perform C-steps until convergence. To identify the best h-subsets we compute robust deviances for all n observations, using the weighted Bianco-Yohai robust logistic regression approach (Bianco and Yohai 1996) as implemented by Croux and Haesbroeck (2003). In this approach, the deviance function has been replaced by a function \(\varphi _{BY}\) to downweight outliers, which significantly improved the classification and prediction (Croux and Haesbroeck 2003). Also here, the deviances \(d( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_s^{el}},y_{i})\), for \(i=1, \ldots , n\), are substituted in the objective function (5) with the functions \(\varphi _{BY}( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_s^{el}},y_{i})\): the smallest values of \(\varphi _{BY}\) are assigned to correct classified observations, which are positive predicted scores \(\eta _i\) corresponding to an observation with \(y_i=1\), and negative predicted scores \(\eta _i\) related to an observation with \(y_i=0\). A desirable subset is the one with the smallest sum of \(\varphi _{BY}( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_{H_s^{el}},y_{i})\); in other words, a subset in which the two groups are highly separated. Finally, the subset with the smallest sum \(\varphi _{BY}( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}_H,y_{i})\) for all \(i \in H\) forms the best index set. Note that this robust criterion is more tolerant to single observations with a score with wrong sign compared to the non-robust deviances, and thus there is a stronger focus on obtaining an h-subset where most of the points are clearly separated.
We consider a warm start strategy (Friedman et al. 2010) to reduce the computational cost of the algorithm, which, in principle, should be computed for each possible combination of the tuning parameters. The warm start is based on the intuition that, for a particular combination of \(\alpha \) and \(\lambda \), the best h-subset from step 3 may also be advisable for another couple of tuning parameters which is adjacent of this \(\alpha \) and/or \(\lambda \), thus the step 1 should be performed only once. A further reweighting step, that downweights outliers detected by \(\hat{ \varvec{\beta }}_{opt}\) given in (6), is considered to increase the efficiency of the proposed estimator. We consider outliers as observations with Pearson residuals larger than a certain quantile of the standard normal distribution. Since the RobLZS estimator is biased due to regularization, it is necessary to center the residuals. Denote \(r_i^s\) as the Pearson residuals,
where \(\mu (\hat{ \varvec{\beta }}_{opt},\mathbf {z}_i)\) and \(\nu (\hat{ \varvec{\beta }}_{opt},\mathbf {z}_i)\) are respectively the fitted mean and fitted variance function of the response variable. The Pearson residuals, which are commonly used in practice in the context of generalized linear models, are normally distributed under small dispersion asymptotic conditions (Dunn and Gordon 2018). Then the binary weights are defined by
where \(\Phi \) is the cumulative distribution function of the standard normal distribution. A typical choice for \(\delta \) is 0.0125, so that 2.5% of the observations are expected to be flagged as outliers in the normal model.
The RobLZS estimator is defined as
where \(n_w=\sum _{i=1}^nw_i\) is the sum of weights, \(\alpha _{opt}\) is the optimal parameter obtained considering the optimal subset \(H_{opt}\), whereas the tuning parameter \(\lambda _{upd}\) is obtained by a 5-fold cross-validation procedure. This update of the tuning parameter \(\lambda \) is necessary, because with a bigger number of observations also the sum of deviances changes compared to (6), and thus the weight for the penalty needs to be adapted.
Robust Logistic Zero-Sum Regression has been available as an R package that can be downloaded at https://github.com/giannamonti/RobZS.
2.3 Parameter selection
To select the optimal combination \((\alpha _{opt},\lambda _{opt})\) of the tuning parameters \(\alpha \in [0,1]\) and \(\lambda \in [\varepsilon \cdot \lambda _{Max},\,\lambda _{Max}]\), with \(\varepsilon >0\), leading to the optimal subset \(H_{opt}\), a repeated K-fold cross-validation (CV) procedure (Hastie et al. 2001), on each best h-subset, on a two-dimensional surface is adopted, with \(K=5\).
In K-fold cross-validation the data are split into folds \(V_1,\ldots ,V_K\) of approximately equal size in which the two classes are represented in about the same proportions as in the complete dataset. We leave out the part \(V_k\), where k is the fold index, \(k\in \{1,\ldots ,K\}\), train the model on the observations with index \(i\notin V_k\) of the other \(K-1\) parts (combined), and then obtain predictions for the left-out kth part. Note that we only consider samples of size h at this stage which are supposed to be outlier-free, and thus the derived prediction error criterion is robust.
As criterion we use the mean of the deviances (MD),
The chosen couple \((\alpha _{opt},\lambda _{opt})\), over a grid of values \(\alpha \in [0,1]\) and \(\lambda \in [\varepsilon \cdot \lambda _{Max},\,\lambda _{Max}]\), is the one giving the smallest CV error in (9). Here, \(\lambda _{Max}\) is an estimate of the parameter \(\lambda \) that leads to a model with full sparsity, see Kurnaz et al. (2018) for details. In the simulations we considered 41 equally spaced values for \(\alpha \), and a grid of 40 values for \(\lambda \).
3 Simulations
In the following simulation studies we are comparing the performance of the RobLZS estimator to other competing sparse estimators. In particular, we considered the Lasso (the regular least absolute shrinkage and selection operator) (Tibshirani 1994), the logistic Zero-Sum (LZS) estimator (Altenbuchinger et al. 2017; Zacharias et al. 2017), and the robust EN(LTS) estimator for logistic regressions (Kurnaz et al. 2018), denoted by RobLL in the following. In order to compare with the Lasso solution, we have set the parameter \(\alpha \) equal to 1 for the methods involving elastic-net penalties. The LZS estimator preserves the zero-sum constraint, but is not robust to the presence of outliers. RobLL is robust, but does not preserve the zero-sum constraint. The Lasso is neither robust, nor does it preserve the zero-sum constraint, while the RobLZS has both properties.
3.1 Sampling schemes
We generate the covariate data,
inspired by the true bacterial abundances in a microbiome analysis (Lin et al. 2014; Shi et al. 2016), as follows.
First an \(n/2\times p\) data matrix \( \mathbf {W}_1=[w_{ij}]_{1\le i \le n/2;\, 1\le j \le p }\) is generated by sampling from a multivariate log-normal distribution \(\ln N_p( \varvec{\theta }_1,\varvec{\Sigma })\), with \( \varvec{\theta }_1=(\theta _{11},\ldots , \theta _{1p})^T=(1,1, \ldots , 1)^T\). Then, independently, another \(n/2\times p\) data matrix \( \mathbf {W}_2=[w_{ij}]_{1\le i \le n/2;\, 1\le j \le p }\) is generated by sampling from a multivariate log-normal distribution \(\ln N_p( \varvec{\theta }_2,\varvec{\Sigma })\), with mean parameter \( \varvec{\theta }_2=(\theta _{21},\ldots , \theta _{2p})^T\) set as \(\theta _{2j}=3\), for \(j=1,\ldots ,5\), and \(\theta _{2j}=1\) otherwise to allow some OTUs to be more abundant than others. The correlation structure of the predictors is defined by \(\varvec{\Sigma }=[\Sigma _{ij}]_{1\le i,j \le p}=\rho ^{|i-j|}\), with \(\rho \)=0.2 to mimic the correlation between different taxa. We get the \(n\times p\) data matrix \( \mathbf {W}=[w_{ij}]_{1\le i \le n;\, 1\le j \le p }=\bigg [\begin{array}{c} \mathbf {W}_1\\ \mathbf {W}_2\end{array}\bigg ]\), and finally the log-compositional design matrix \( \mathbf {Z}=[z_{ij}]_{1\le i \le n;\, 1\le j \le p }\) is obtained by the transformation
The first n/2 values of the binary response were set to 0, and the last n/2 entries were set to 1. Thus, the response values \(y_i\), for \(i=1,\ldots ,n\), directly reflect the grouping structure entailed by the different centers of the matrices \( \mathbf {W}_1\) and \( \mathbf {W}_2\).
The true parameter \( \varvec{\beta }=(\beta _j)_{1\le j \le p}\) is set to \(\beta _1=\beta _3=\beta _5=\beta _{11}=\beta _{13}=-0.5\), \(\beta _2=1\), \(\beta _{16}=1.5\), and \(\beta _j=0\) for \(j \in \{1, \ldots , p\} \setminus \{1,2,3,5,11,13,16\}\), the intercept is set to \(\beta _0=-1\).
The observations \(\mathbf {z}_i\), \(i=1,\ldots ,n/2\), of the covariates for \(y_i=0\), are arranged according to increasing values of \(\mu ( \varvec{\beta },\mathbf {z}_i)\) in the design matrix \( \mathbf {Z}\). This is because in the various contamination schemes we will modify a proportion of the first entries of this group, and thus these are observations with the poorest fit to that group.
The two robust estimators are calculated taking \(\xi =3/4\) for an easy comparison. This means that n/4 is an initial guess of the maximal proportion of outliers in the data. For each replication, we choose the optimal tuning parameter \(\lambda _{opt}\) as described in paragraph 2.3, with a repeated 5-fold CV procedure and a suitable sequence of 40 values between \(\varepsilon \cdot \lambda _{Max}\) and \(\lambda _{Max}\), with \(\varepsilon >0\), used to adjust this range.
Different sample size/dimension combinations \((n, p)=(50,\, 30),\, (100,\, 200)\) and \((100,\, 1000)\) are considered, thus a low-high dimensional setting (\(n>p\)), a moderate-high dimensional setting (\(n<p\)), and a high-dimensional setting (\(n\ll p\)). The simulations are repeated 100 times for each setting to keep computation costs reasonably low.
For each of the three simulation settings we applied the following contamination schemes:
-
Scenario A. (Clean) No contamination.
-
Scenario B. (Lev) Leverage points: we replace the first \(\gamma \)% (with \(\gamma =10\) or 20) of the observations by values coming from a p-dimension log-normal distribution with mean vector \(\theta _j=3\), for \(j=1,\ldots ,5\), and \(\theta _j=0.5\) otherwise, and a correlation equal to 0.9 for each pair of variable components, then the resulting log-compositional design matrix \( \mathbf {Z}\) is obtained by normalizing the true abundances.
-
Scenario C. (Vert) Vertical outliers: we assign to the first \(\gamma \)% (with \(\gamma =10\) or 20) of the observations the wrong class membership.
-
Scenario D. (Both) Horizontal and Vertical outliers: this is a more extreme situation in which each outlier has both types of contaminations, combining scenarios B and C.
Below we present the simulation results for \(\gamma =10\)%; similar results have been obtained for \(\gamma =20\)%, and they are reported in Sect. 1 of the Supporting Information (SI) for the sake of completeness.
3.2 Performance measures
To evaluate the prediction performance of the proposed sparse method, in comparison to the other models, we consider several measures. For this purpose, an independent test sample of size n without outliers was generated in each simulation run.
To quantify the prediction error in the whole range of the predictive probabilities we used three different measures (Cessie and Houwelingen 1992):
-
Mean prediction error, defined as
$$\begin{aligned} \text {MPE}=\frac{1}{n} \sum _{i=1}^n\bigg (y_i^{\star }-\mu (\hat{ \varvec{\beta }}, \mathbf {z}_i^{\star })\bigg )^2 , \end{aligned}$$(10)where \(y_i^{\star }\) and \( \mathbf {z}_i^{\star }\) denote the response and the covariate vector of the test set data, respectively, n is the total number of data points in the test set, \(\hat{ \varvec{\beta }}\) is the parameter estimate derived from the training data, and the prediction \(\mu (\hat{ \varvec{\beta }}, \mathbf {z}_i^{\star })\) is a probability that the response is equal to 1 based on the parameter estimates.
-
Mean absolute error, defined as
$$\begin{aligned} \text {MAE}=\frac{1}{n} \sum _{i=1}^n\bigg |y_i^{\star }-\mu (\hat{ \varvec{\beta }}, \mathbf {z}_i^{\star })\bigg | . \end{aligned}$$(11) -
Logarithmic loss (or minus log-likelihood error), defined as
$$\begin{aligned} \text {ML}=-\frac{1}{n} \sum _{i=1}^n\bigg \{y_i^{\star }\log \big (\mu (\hat{ \varvec{\beta }}, \mathbf {z}_i^{\star })\big )+(1-y_i^{\star })\log \big (1-\mu (\hat{ \varvec{\beta }}, \mathbf {z}_i^{\star })\big )\bigg \} . \end{aligned}$$(12)
Different statistics based on the accuracy matrix are used to evaluate the ability of the estimators in discriminating the true binary outcome:
-
Sensitivity (Se): the true positive rate, in other words the proportion of actual positives that are correctly identified.
-
Specificity (Sp): true negative rate, or \(1 - \)false positive rate, thus the proportion of actual negatives that are correctly identified.
-
AUC: the proportion of area below the receiver operating characteristics (ROC) curve.
Concerning sparsity, the estimated models are evaluated by the number of false positives (FP) and the number of false negatives (FN), defined as
where here positives and negatives refer to nonzero and zero coefficients, respectively.
3.3 Simulation results
We report averages (mean) and standard deviations (sd) of the performance measures defined in the previous section over all 100 simulation runs, for each method and for the different contamination schemes. In the following tables, the best values (of “mean”) among the different methods are presented in bold. Tables 1, 2, and 3 show the predictive performance of the different methods in the different scenarios and sample size/dimension combinations. Table 4 shows the corresponding selection performances.
The results for Scenario A (no contamination) show that all methods have comparable performance in terms of Sensitivity, Specificity and AUC. This is different in the contaminated scenarios, and the difference gets more pronounced with growing dimension. For instance, in Scenario B the AUC is quite comparable for the different methods in the lower-dimensional case, but there is a big difference between the non-robust and the robust methods in the high-dimensional case; the latter methods attain about the same AUC as in the uncontaminated case. Scenario C shows an advantage of the non-robust methods for the Sensitivity, but a drawback for Specificity, such that the AUC for the robust methods gets higher values (even more in higher dimension). Similar conclusions can be drawn from Scenario D.
For the prediction measures MPE, MAE and ML, the results in the uncontaminated case are again quite comparable, with only a slight performance loss of the robust methods. This is also based on the application of a reweighting step at the end, which gains efficiency for the estimator. For the contamination scenarios one can see a similar trend towards better results for the robust methods with increasing dimension. Generally, the RobLZS attains usually the best results for the MAE, for Scenario B even by far the best results. It is interesting to see that the LZS estimator achieves quite poor results in Scenario D. However, it can also be seen that the Lasso estimator surprisingly delivers relatively good results in this setting. One should be aware that leverage points might not have such strong effects here because of the normalization of the observations to sum 1. Moreover, it is worth mentioning that Lasso and its robust counterpart (RobLL) do not preserve the zero-sum constraint of the coefficients, thus they lead in any case not to an appropriate solution for compositional data, and are reported here only for benchmarking purposes.
In terms of the selection properties presented in Table 4, one can see similar performance of all methods in all settings for the false negative rate (FN). RobLZS shows slightly better results in Scenarios B and D. For the false positive rate (FP) one can see clearer differences between the methods, again more pronounced when the dimensionality of the covariates increases: Scenario A leads to clearly higher values for LZS and RobLZS, similar in Scenario C; the methods RobLL and LZS are preferable in Scenario B, while RobLL is the clear winner in Scenario D (at least in higher dimension). As mentioned above, only the methods LZS and RobLZS fulfill the sum-zero constraint of the regression coefficients, and thus the other methods do not result in log-contrasts. Moreover in this contest, the omission of important variables is usually more problematic than the inclusion of unimportant variables with shrinkage coefficients.
Overall, the proposed RobLZS estimator performs remarkably well in all simulation settings, in the uncontaminated case as well as in presence of outliers. It tends to slightly less sparsity, thus including more of the non-relevant variables, but shows excellent performance with identifying the truly relevant ones. The classification performance is excellent, and the precision measures reveal clear advantages over the non-robust methods in case of contamination. Moreover, the standard deviations of RobLZS for the AUC are almost always smaller than for the non-robust methods in the contaminated scenarios and for all considered sample sizes, showing a high stability of the estimations over 100 simulations.
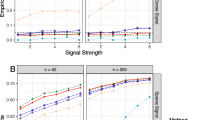
More simulation results are presented in the Supporting Information (SI): Sect. 1 presents results for 20% contamination, Sect. refsec:method shows results for gradually increasing contamination from zero to 30%, and Sect. 3 compares LZS and RobLZS by making use of the elastic-net penalty.
4 Applications to microbiome data
We illustrate the performance of our proposed estimator by applying it to two datasets related to human microbiome data: the first one is related to inflammatory bowel diseases (IBD) (Morgan et al. 2012), and the second one is concerned with an application to Parkinson’s disease (PD) (Dong et al. 2020). The two microbiome datasets were preprocessed by filtering out OTUs which had more than 90% zeros. The remaining zero counts were then replaced by a pseudo-count value 0.5 to allow for the logarithmic transformation.
The original IBD dataset consists of microbiome data with 81 samples for investigating the association between the gut microbiota and a chronic and relapsing inflammatory condition known as Crohn’s disease, with 19 healthy and 62 IBD affected individuals . The dimension of the microbiome data set originally was \(n \times p = 81 \times 367\), and after preprocessing the final number of OTUs is \(p=95\).
For the original PD dataset we have dimension \(n \times p = 327 \times 4707\), and after preprocessing the resulting microbiome data consists of \(p=1016\) final OTUs.
For a fair investigation of the prediction performance of the four sparse estimators, a 5-fold cross-validation procedure was repeated 20 times, resulting in 100 fitted models for each sparse regression method. In the training set, the parameter selection follows the one described in the simulation section.
4.1 Results for the IBD data
Accuracy measures such as Sensitivity (true positive rate), Specificity (false positive rate) and AUC were used to assess the classification performance of the different methods. The AUC represents a trade-off between Sensitivity and Specificity. The results are presented as boxplots in Fig. 1. The RobLZS estimator shows a lower Sensitivity but a higher Specificity, resulting in an AUC that is higher on average than for the other estimators.
Since it is not known which variables should be selected by the models, we can only compare the regression coefficients and the resulting model sparsity for the four different methods. Figure 2 presents the regression coefficients as average over all models derived from the repeated CV. The horizontal axis (Index) corresponds to the variable number. The general picture is that all methods more or less are conform with the zero and non-zero coefficients. For RobLL we observe for some variables much higher coefficients.
The sparsity of the repeated CV models is compared in Fig. 3, by showing the proportion of models (out of all 100) which have resulted in at least the number of zero coefficients indicated by the horizontal axis. One can see that the classical methods Lasso and LZS lead to a comparable sparsity; RobLZS results in less sparsity, and RobLL is much less sparsity. From the simulations we know that RobLZS has slightly better performance to identify the correct variables, but (depending on the outlier configuration) it tends to include also non-relevant variables in the model.
An important issue is to investigate if there are outliers in the data set. Outliers can only be reliably identified with the robust procedure. We thus apply RobLZS to the complete data set and show in Fig. 4 (left) a plot of the scores \( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}\) versus the deviances. Red color indicates the identified outliers with large deviances, blue color is for regular observations. As a comparison we also show the corresponding scores and deviances from the non-robust LZS estimator (pink crosses), which leads to much smaller deviances in general. A further comparison of the scores from the RobLZS and the LZS estimator is shown in the right plot, with plot symbol according to the class variable (healthy/disease), and color according to the outlyingness information from RobLZS. The outliers are exclusively originating from IBD affected individuals, and their scores are very different (for the robust method) from the scores of the other individuals in this group. One can assume that these persons have some common feature, being different from the remaining IBD affected people.
IBD data: the left plot shows the deviances against the scores \( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}\); the blue/red squares refer to the RobLZS method, outliers are in red, and the pink crosses refer to the non-robust LZS method. The right plot shows again the scores from both estimators, with symbol color according to the outlyingness information from RobLZS, and symbol according to the class (color figure online)
4.2 Results for the PD data
Figure 5 shows boxplots of the values for Sensitivity, Specificity and AUC from all \(20\times 5\) models from repeated CV. We can see a similar picture as for the IBD data, with lower sensitivity for RobZS compared to the other estimators, but higher Specificity and overall a slightly higher (average) AUC.
Figure 6 (left) compares the resulting average regression coefficients, for better readability now only for the estimators LZS and RobLZS. One can see that both estimators are in agreement for bigger values of the coefficients (and for the sign). Some differences are for smaller values, but again the sign is mostly in agreement. The right plot shows the obtained sparsity for all estimators, and we can draw the same conclusions as for the IBD data: Lasso and LZS are very similar, RobLL leads to less sparsity, and RobLZS is in between.
Similar to the results shown in Fig. 4 for the IBD data, Fig. 7 shows the scores against the deviances when LZS and RobLZS are applied to the complete PD data set. RobLZS leads to much higher (absolute) values of the scores, but also to clearly higher deviances, with several outliers indicated in red. For these data, the outliers are not separated from the remaining data, which is also shown in the right plot with a direct comparison of the scores for the classical and the robust procedure. The indicated outliers are observations for which the sign of the RobLZS scores corresponds to the wrong group label. Thus, these observations have a data structure which differs from that of the majority in the group, and this is the reason why they are downweighted by the robust method.
PD data: The left plot shows the deviances against the scores \( \mathbf {z}_{i}^{\mathrm {T}}\hat{ \varvec{\beta }}\); the blue/red squares refer to the RobLZS method, outliers are in red, and the pink crosses refer to the non-robust LZS method. The right plot shows again the scores from both estimators, with symbol color according to the outlyingness information from RobLZS, and symbol according to the class (color figure online)
Since the RobLZS estimator is able to identify outliers, one can also compute a robustified version of the accuracy measures, where the identified outliers are excluded. Thus, Sensitivity, Specificity and AUC are only computed based on the regular observations which are not indicated to be outliers. This is done in Table 5 for both example data sets. Here, for simplicity, the estimators LZS and RobLZS are only applied once to the complete data set, and from this fit the measures are computed. It then can be seen that the non-outlier version (column “without out.”) of the accuracy measures for RobLZS leads to excellent in-sample fit.
One could also compare the accuracy measures with LZS when the outliers identified by RobLZS have been removed. This comparison, however, is not really appropriate, because when only applying the method LZS, one would not get any (reliable) outlier information. Nevertheless, we obtain the following results for LZS without outliers for the IBD data: Se\(=1.000 \), Sp\(=0.421 \), and AUC\(=0.711\). For the PD data we obtain: Se=0.957, Sp=0.772, AUC=0.865.
5 Conclusions
A new robust estimator called RobLZS for sparse logistic regression with compositional covariates has been introduced. Due to an elastic-net penalty with an intrinsic variable selection property it can deal with high-dimensional covariates. The compositional aspect is considered with a log-contrast model, which leads to a zero-sum constraint on the regression coefficients. Robustness of the estimator is achieved by trimming, where the trimming proportion has to be selected according to an initial guess of the maximum fraction of outliers in the data. We recommend a trimming proportion of about 25%, thus using about 3/4 of the observations, which should be reasonable in practice to protect against outliers, and also leads to higher efficiency of the initial estimator (Sun et al. 2020). The efficiency of the estimator is further increased by a reweighting step for the computation of the final estimator, where the information from all observations that correspond to the model is considered. This reweighting builds on the approximate normal distribution of the Pearson residuals, see (7), which might be problematic in a high-dimensional sparse data setting with a low number of observations. Indeed, our simulations for the uncontaminated case revealed that the proportion of identified outliers is somewhat higher (around 4-5% instead of the intended 2.5%). However, the reweighted estimator still improved the estimator without reweighting, and thus this option seems reasonable.
We have proposed an algorithm to compute the estimator, and R code for its computation has been made publicly available at https://github.com/giannamonti/RobZS. The iterative algorithm successively minimizes the objective function by carrying out so-called C-steps, which have been used also in the context of other robust estimators (Rousseeuw and Van Driessen 2006). In simulation studies we have compared the estimator with its non-robust counterpart, as well as with Lasso regression and a robustified Lasso estimator, which cannot appropriately handle compositional covariates. The RobLZS estimator works reasonably well under uncontaminated data, delivering results which are similar for the non-robust counterpart. Under contamination one obtains a classifier that is usually better or much better than the non-robust version, but it tends to produce less sparsity by adding more of the non-relevant variables.
The applications to real compositional microbiome data sets also revealed the advantages of the RobLZS estimator, whose classification accuracy is remarkably excellent. For practitioners, the most important advantage might be the ability of the procedure to identify outliers, thus observations that strongly deviate from the model, being aware of the unreliable results obtained from non-robust procedures in presence of outliers. The reasons for outlyingness can be manyfold, it could be mislabeled observations, but also individuals with a different multivariate data structure. In the context of the data sets used here, investigating those outliers in more detail may lead to relevant conclusions about the health status of the persons.
References
Aitchison J (1982) The statistical analysis of compositional data. J R Stat Soc Series B Stat Methodol 44(2):139–177
Aitchison J (1986) The statistical analysis of compositional data. Chapman and Hall, London
Aitchison J, Bacon-Shone J (1984) Log contrast models for experiments with mixtures. Biometrika 71(2):323–330
Albert A, Anderson JA (1984) On the existence of maximum likelihood estimates in logistic regression models. Biometrika 71(1):1–10
Alfons A, Croux C, Gelper S (2013) Sparse least trimmed squares regression for analyzing high-dimensional large data sets. Ann Appl Stat 7(1):226–248
Altenbuchinger M, Rehberg T, Zacharias HU, Stämmler F, Dettmer K, Weber D, Hiergeist A, Gessner A, Holler E, Oefner PJ, Spang R (2017) Reference point insensitive molecular data analysis. Bioinformatics 33(2):219–226
Avella-Medina M, Ronchetti E (2017) Robust and consistent variable selection in high-dimensional generalized linear models. Biometrika 105(1):31–44
Bates S, Tibshirani R (2019) Log-ratio lasso: scalable, sparse estimation for log-ratio models. Biometrics 75(2):613–624
Bianco AM, Yohai VJ (1996) Robust statistics, data analysis, and computer intensive methods. In: Rieder H (ed) Honor of Peter Hubers 60th Birthday, chap Robust Estimation in the Logistic Regression Model. Springer, New York, pp 17–34
Cessie SL, Houwelingen JCV (1992) Ridge estimators in logistic regression. J R Stat Soc C-Appl 41(1):191–201
Croux C, Haesbroeck G (2003) Implementing the Bianco and Yohai estimator for logistic regression. Comput Stat Data Anal 44(1):273–295
Dong M, Li L, Chen M, Kusalik A, Xu W (2020) Predictive analysis methods for human microbiome data with application to Parkinsons disease. PloS One 15(8):e0237779
Dunn PK, Gordon KS (2018) Generalized linear models with examples in R. Springer, New York
Friedman J, Trevor H, Tibshirani R (2010) Regularization paths for generalized linear models via coordinate descent. J Stat Softw 33(1):1–22
Gloor GB, Macklaim JM, Pawlowsky-Glahn V, Egozcue JJ (2017) Microbiome datasets are compositional: and this is not optional. Front Microbiol 8:2224
Hastie T, Tibshirani R, Friedman J (2001) The elements of statistical learning. Springer Series in Statistics, Springer, New York Inc
Kurnaz FS, Hoffmann I, Filzmoser P (2018) Robust and sparse estimation methods for high-dimensional linear and logistic regression. Chemom Intell Lab Syst 172:211–222
Lin W, Shi P, Feng R, Li H (2014) Variable selection in regression with compositional covariates. Biometrika 101(4):785–797
Lu J, Shi P, Li H (2019) Generalized linear models with linear constraints for microbiome compositional data. Biometrics 75(1):235–244
Morgan XC, Tickle TL, Sokol H, Gevers D, Devaney KL, Ward DV, Reyes JA, Shah SA, LeLeiko N, Snapper SB, Bousvaros A, Korzenik J, Sands BE, Xavier RJ, Huttenhower C (2012) Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol 13(9)
Neykov NM, Filzmoser P, Neytchev PN (2014) Ultrahigh dimensional variable selection through the penalized maximum trimmed likelihood estimator. Stat Pap 55(1):187–207
Nurunnabi A, West G (2012) Outlier detection in logistic regression: a quest for reliable knowledge from predictive modeling and classification. In: 2012 IEEE 12th international conference on data mining workshops, pp 643–652
Park H, Konishi S (2016) Robust logistic regression modelling via the elastic net-type regularization and tuning parameter selection. J Stat Comput Simul 86(7):1450–1461
Qin J, Li R, Raes J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65
Rousseeuw PJ, Van Driessen K (2006) Computing LTS regression for large data sets. Data Min Knowl Discov 12(1):29–45
Shi P, Zhang A, Li H (2016) Regression analysis for microbiome compositional data. Ann Appl Stat 10(2):1019–1040
Sun H, Cui Y, Gao Q, Wang T (2020) Trimmed lasso regression estimator for binary response data. Stat Probab Lett 159:108679
Tibshirani R (1994) Regression shrinkage and selection via the lasso. J R Stat Soc Series B Stat Methodol 58:267–288
Zacharias HU, Rehberg T, Mehrl S, Richtmann D, Wettig T, Oefner PJ, Spang R, Gronwald W, Altenbuchinger M (2017) Scale-invariant biomarker discovery in urine and plasma metabolite fingerprints. J Proteome Res 16(10):3596–3605
Zou H, Hastie T (2005) Regularization and variable selection via the elastic net. J R Stat Soc Series B Stat Methodol 67(2):301–320
Acknowledgements
Research financially supported by the Italian Ministry of University and Research, FAR (Fondi di Ateneo per la Ricerca) 2019. We greatly acknowledge the DEMS Data Science Lab for supporting this work by providing computational resources.
Funding
Open access funding provided by Universitá degli Studi di Milano - Bicocca within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Declaration
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Monti, G.S., Filzmoser, P. Robust logistic zero-sum regression for microbiome compositional data. Adv Data Anal Classif 16, 301–324 (2022). https://doi.org/10.1007/s11634-021-00465-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11634-021-00465-4