Abstract
A reliable and simple Agrobacterium-mediated transformation system for the unicellular green algae model organism Chlamydomonas reinhardtii has been developed. The protocol has been successfully employed with both neomycin phosphotransferase II (nptII) and the phleomycin resistance (bleI) genes coupled with the selective agents paromomycin and zeocin, respectively. A set of binary vectors were assembled that carry the selectable marker cassettes under control either of the Rbcs2 alone or fused to the HSP270A leader sequence, PsaD, or ß-tubulin2 promoters. The corresponding T-DNA elements also harbored a cassette with a codon-optimized version of yellow fluorescence protein (YFP) under control of the Rbcs2 promoter in which the YFP open reading frame was interrupted with the first intron of Rbcs2 to prevent expression in Agrobacterium tumefaciens. The resultant binary vectors were introduced into A. tumefaciens strain C58C1/pMP90, and the derived transconjugants were used for transformation studies with the walled C. reinhardtii strain CC124. Estimated transformation frequencies ranged from 0.09 to 2.86 colonies per 106 cells inoculated. Molecular characterizations on a subset of the transgenic lineages revealed that most of the transgenic events harbored single locus insertions. Moreover, sequencing of captured junction fragments about the T-DNA insertion site showed that minimal disruption of the C. reinhardtii genome occurred. However, the transgenic lineages often harbored truncated T-DNA regions within the non-selectable marker gene cassettes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The unicellular alga Chlamydomonas reinhardtii has a long history as a model system for study of the fundamental underpinnings of core biological processes. These include photosynthesis, flagellar assembly and mechanics (Harris 2001), heavy metal transport and compartmentalization (Hanikenne 2003), epigenetic regulation of gene expression (Jeong et al. 2002; van Dijk et al. 2005; Casas-Mollano et al. 2008), and a platform to understand intracellular carbon flux changes under nutrient sufficient and depleted environments (Miller et al. 2010; Msanne et al. 2012). Genetic engineering of cells is a powerful tool for both functional genomics programs and translation of foundational discoveries into application. In this regard, C. reinhardtii is rather unique given that all three of its genetic compartments are competent for genetic manipulation (Kindle 1990; Kindle et al. 1991; Sizova et al. 1996; Bateman and Purton 2000; Remacle et al. 2006). These genetic engineering tools, coupled with the wealth of other genomic resources and databases (Vallon and Dutcher 2008; Dal’Molin et al. 2011), including a draft of the C. reinhardtii genome (Merchant et al. 2007), have greatly enhanced the attractiveness of this model algal system.
The introduction of transgenes into the nuclear compartment of C. reinhardtii, while rather robust, has been historically hampered by poor expression of transgene(s) (Fischer and Rochaix 2001; Shao and Bock 2008) and by a tendency of the resultant transformants to carry large genetic deletions about the junction of the inserted foreign allele (Dent et al. 2005). The former limits the potential of C. reinhardtii as a feedstock for biological products (Franklin and Mayfield 2004), and the latter can complicate hypothesis testing in functional genomics programs when phenotyping insertional mutant populations. Most protocols communicated for nuclear transformation of C. reinhardtii implement a direct DNA delivery system, employing electroporation, glass beads, silicon carbide whiskers, or microprojectile bombardment (Dunahay 1993; Neupert et al. 2012). In higher plants’ nuclear transformation, implementing direct DNA delivery techniques often leads to complex, multi-copy transgenic alleles, which can be mitigated by utilizing Agrobacterium-based protocols (Cheng et al. 2001; Dai et al. 2001; Shou et al. 2004). To this end, the objectives of the studies communicated herein were to develop a dependable Agrobacterium-mediated transformation protocol for C. reinhardtii and genotype a subset of the derived transgenic lineages to gauge the utility for both biotechnology applications and use in functional genomics efforts with respect to direct DNA methods.
Materials and methods
Vector Constructions
A C. reinhardtii codon-optimized version of the selectable marker gene neomycin phosphotransferase II (nptII) coupled with the 5ʹ leader sequence derived from ribosomal protein L13 (rpl13) along with a codon-optimized version of the visual marker yellow fluorescence protein (YFP) fused to the 5ʹ untranslated region of RbcS2 was used in the transformation studies. The YFP open reading frame (ORF) was interrupted by the first intron of the C. reinhardtii RbcS2 gene to block translation within A. tumefaciens cells. Both nptII and YFP genes were commercially synthesized (GenScript Co. Piscataway, NJ). In some experiments, a second selectable marker gene, ble (Stevens et al. 1996), was tested. The synthetic nptII gene was assembled into three expression cassettes regulated by either the C. reinhardtii RbcS2 promoter alone or fused with the HSP70A element (Sizova et al. 2001), the PsaD promoter (Fischer and Rochaix 2001), or the ß2-tubulin promoter (Davies and Grossman 1994) while the single ble selectable marker was placed under control of the RbcS2 promoter.
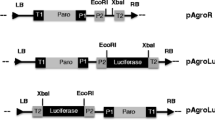
A total of seven binary vectors were assembled (Fig. 1). The respective nptII, ble, and YFP cassettes were subcloned into the base binary plasmid pPZP202 (Hajdukiewicz et al. 1994), and the resultant vectors were designated as pPTN1127, pPTN1129, pPTN1130, pPTN1133, pPTN1170, and pPTN1176 (Fig. 1). The first three harbored just the nptII cassette under control of one of the three regulatory elements RbcS2 (pPTN1127), PsaD (pPTN1129), or ß2-tubulin (pPTN1130). The plasmid pPTN1133 carried an nptII cassette under control of the RbcS2-HSP70A regulatory element and a YFP cassette regulated by the ß2-tubulin promoter. The plasmid pPTN1170 carried the same YFP and nptII cassettes as pPTN1133 plus the ble selectable marker while vector pPTN1176 harbored the two selectable marker cassettes, nptII and ble (Fig. 1). The binary plasmids were mobilized into A. tumefaciens nopaline strain C58C1/pMP90 (Koncz and Schell 1986) via tri-parental mating, and the derived transconjugants were used in the transformation studies for C. reinhardtii (Fig. 2).
Diagrammatic representation of T-DNA elements used in the study. Binary vector names along with approximate size, in parentheses, of the corresponding T-DNA are listed underneath the bidirectional arrow. RB and LB indicate right border and left border elements, respectively. Rbcs2 Pro and Rbcs2 terms indicate Rbcs2 promoter and terminator, respectively. PsaD and ß2-Tub Pro refer to PsaD and ß2-tubulin promoters. HSP70A indicates 5ʹUTR of HSP70A gene. Note: Sizes of the respective elements are not drawn to scale.
PCR analyses for integration of plasmid DNA into Chlamydomonas genome in the microprojectile-derived and Agrobacterium-mediated transgenic events derived from binary vector pPTN1133. (a) Diagram of PCR amplification along the T-DNA and a segment of pPTN1133 plasmid backbone. (b) PCR results of using corresponding primer sets.
Agrobacterium-Mediated Transformation
The A. tumefaciens transconjugants were grown in YEP medium with the appropriate antibiotics to an OD650 of 0.8 to 1.0. The cultures were centrifuged in 50-mL conical tubes, and the bacterial pellet was suspended to the same OD650 in co-cultivation medium composed of low salt, supplemented with 1% glucose, buffered with 20 mM MES (pH 5.4), plus 200 µM acetosyringone. The bacterial cultures were placed on a shaker for 1 h and subsequently placed on ice in the dark until use.
C. reinhardtii cells, strain CC124, were cultured in TAP medium under photosynthetic active radiance (PAR) of 40 to 50 µmol m−2 s−1 at room temperature until cell densities reached OD750 of approximately 0.5 at which time the cells were harvested by centrifugation and re-suspended to a concentration of 2 × 108 cells mL−l. An aliquot of 200 µL of the algae culture was spread on co-culture medium solidified with 0.8% agar (Sigma Cas No. 9002–18-0), and the plates were subsequently air-dried in a laminar flow hood for 30 min. Following the dry-down step, the algal lawn was overlaid with 200 µL of the A. tumefaciens inoculum, and plates were co-cultured in the dark at 24 °C for 48 h. At the completion of the co-cultivation step, the cells were scraped off the plate and washed 2 times in TAP medium supplemented with 100 mg L−1 each of cefotaxime and carbenicillin. The cells were then plated onto delay medium (TAP supplemented with the cefotaxime and carbenicillin) to counter-select against A. tumefaciens for 15 h under PAR of 40 to 50 µmol m−2 s−1, after which the cells were collected and re-plated onto TAP selection plates supplemented with cefotaxime and carbenicillin and with paromomycin at 15 mg L−1 or zeocin at 7.5 mg L−1 for nptII and ble selectable marker cassettes, respectively. The selection plates were maintained at room temperature under light (40 to 50 µmol m−2 s−1 PAR) for 7 d prior to counting resistant colonies.
Microprojectile-Mediated Transformation
Microprojectile bombardments were conducted using the PDS-1000He Particle Delivery System (Bi-Rad Lab., Hercules, CA). Plasmid DNA (1.0 mg mL−1) was precipitated onto 25.0 µL of 0.6 µm gold particles suspended in 50% glycerol (60.0 mg mL−1) by the sequential addition of 2.5 µL plasmid prep, 25 µL of 2.5 M CaCl2, and 10 µL 0.1 M spermidine with continuous vortexing for 3 min. The coated particles were subsequently washed with 140 µL of 70% ethanol followed by a second wash with 100% ethanol and finally suspended in 24.0 µL of 100% ethanol. Each particle preparation was partitioned onto two macrocarriers. The algal plates were prepared as described above with the exception that the medium used was full-strength TAP supplemented with the antibiotic regime of carbenicillin and cefotaxime to limit contamination potential. The plates were positioned 9 cm from the launch assembly. Following bombardment, cells were collected in 5 mL TAP plus carbenicillin and cefotaxime, centrifuged and re-suspended in 15 mL of TAP plus antibiotics, and placed in a 50-mL Erlenmeyer flask under dim light (40 to 50 µmol m−2 s−1 PAR) with no shaking for 15 h. The cultures were subsequently collected and plated onto selection plates as described above.
Characterization of Transgenic C. reinhardtii Events
Putative resistant colonies were re-cultured onto fresh selection plates for an additional 10 d and subsequently bulked-up in 250 mL cultures for down-stream characterization. Monitoring for expression of the selectable marker gene nptII was carried out using a commercially available ELISA kit (Agdia® Corp. Elkhart, IN, Cat# 73000/0480). Imaging for expression of the YFP visual marker was conducted using a Nikon 1A confocal laser microscope system (Nikon Instruments, Melville, NY) with dual scanning at 488 nm laser excitation and 500 to 550 nm emission. Fluorescence from chloroplast was monitored with 641 nm laser excitation and 662 to 737 nm emission. Images were captured at 100 × magnification.
Total genomic DNA was isolated from C. reinhardtii cells using either a modified CTAB (Murray and Thompson 1980) protocol or a method adapted from Dellaporta et al. (1983). Southern blot analyses were carried out on approximately 10 µg of genomic DNA digested with Bam HI (Fig. 3) or Not I (Fig. 4), and the resultant fragments were separated on a 0.8% agarose gel. The resultant gel was blotted onto a nylon membrane (Bio-Rad, Hercules, CA, Cat#162–0196). Probes used in the hybridization step were prepared by random prime labeling incorporating 32P-radiolabelled dCTP using Prime-It II kit (Stratagene Corp., La Jolla, CA, Cat# 300385). Hybridizations and washing steps were carried out as previously described (Eckert et al. 2006). For analyses of DNA integration in the transgenic events of both Agrobacterium and microprojectile derivatives, a series of primer sets (Supplementary Table S1) were used to amplify various segments of the T-DNA and the binary plasmid backbone sequence (Fig. 2).
Southern blot analyses on transgenic Chlamydomonas reinhardtii events. (a) Microprojectile-derived transgenic events obtained from delivery of binary vector pPTN1133 hybridized with nptII ORF. (b) Blot in a stripped and re-hybridized with YFP ORF. (c) Blot in b stripped and re-hybridized with aadA ORF. (d) Agrobacterium-mediated transgenic events derived from binary vector pPTN1133 probed with nptII ORF. (e) Blot in (d) stripped and re-hybridized with YFP ORF. (f) Blot in e stripped and re-hybridized with aadA ORF. Lane + indicates 50 pg of pPTN1133. Molecular marker is in kb. Genomic DNA was digested with BamHI. (g) Diagram of pPTN1133 showing BamHI sites and relative probe binding locations (red bars corresponding to nptII, YFP, and aadA).
Southern blot analysis using junction fragment probes. Total genomic DNA isolated from five selected Agrobacterium-mediated transgenic Chlamydomonas reinhardtii events (pPTN1133) designated A-01, A-05, A-07, A-49, and A-50. WT lanes refer to wild-type CC124 total genomic DNA. Lanes designated with + refer to 50 pg of plasmid DNA harboring the corresponding cloned junction fragment. Genomic DNA was digested with NotI. Probes used in the respective hybridization of blots correspond to junction fragments obtained from DNA proximal to the LB region of the selected transgenic events.
DNA junction fragments about the insertion sites of the introduced transgene(s) were captured using either inverse PCR (Ochman et al. 1988) or thermal asymmetric interlaced (TAIL) PCR (Liu et al. 1995) methods. Amplified DNA fragments were cloned into pCR2.1-TOPO (Invitrogen Cat# 45–0641) and sequenced (Eurofins MWG Operon). The sequence junction fragments were blast searched (www.phytozome.net). As a confirmatory step and to analyze the integrity of the transgenic allele insertion, the junction fragments were subsequently re-amplified directly from the transgenic events with primer set designs based on the DNA sequence captured from the initial TAIL or inverse PCR reactions.
Results and Discussion
Evaluation of Synthetic nptII and YFP Transgenes in C. reinhardtii
To confirm functionality of the synthetic codon-optimized versions of the selectable and visual marker genes, microprojectile bombardment of intermediate pUC-based plasmids, designated pPTN1111 and pPTN1106, which harbor nptII and YFP cassettes, respectively, and are present in the binary vector pPTN1133 (Fig. 1) as a transgene stack, was introduced into C. reinhardtii. Based on recovery of paromomycin-resistant colonies under a selection pressure of 15 mg L−l that were also positive by ELISA assay, the estimated transformation frequency was 3.4 CFU per million cells. Co-bombardment transformations using pPTN1111 and pPTN1106 resulted in an estimated transformation frequency of 2.5 CFU per million cells, translating to a co-expression rate of 3.5% with expression of YFP observed in 0.09 CFU per million cells. The co-expression frequency was significantly lower than what was observed in higher plants when delivering multiple transgenic cassettes into cells via biolistics (Gordon-Kamm et al. 1990; Chen et al. 1998, 2006; Tang et al. 1999).
Several possibilities, including genetic and epigenetic, can account for the poor co-expression frequencies often encountered with Chlamydomonas transformation (Cerutti et al. 1997; Fischer and Rochaix 2001). Molecular analyses employing 12 PCR primer sets that spanned the T-DNA element of pPTN1133, including regions just outside the respective border elements, were conducted on 17 of the transgenic lines not co-expressing YFP derived from these microprojectile-mediated transformations. The results revealed that within these selected lines eight of the 17 appeared to have an intact YFP cassette with 13 missing some portion of the binary vector backbone (Fig. 2); the latter rarely occurred with direct DNA delivery into the nuclear compartment of higher plants. These data suggested that approximately 50% of the transgenic lineages not co-expressing YFP were due to genetic truncations in the transgenic allele.
Agrobacterium-Mediated Transformation of C. reinhardtii
To assess the ability of A. tumefaciens to transform and subsequently characterize the complexity of the integration sites of transgenic alleles introduced into the C. reinhardtii genome via Agrobacterium-mediated transformation, a facile protocol was designed based on known parameters critical for the bacterium to transfer T-DNA elements through the type IV secretion system of the organism into a host genome. These parameters include low pH and reduced phosphorus coupled with the presence of a phenolic inducer of the two-component regulatory system of the secretion apparatus, acetosyringone, during the co-cultivation of the A. tumefaciens with the algae cells (Stachel et al. 1985; Fullner and Nester 1996; McCullen and Binns 2006). Secondly, the protocol implements an antibiotic counter selection scheme to impede A. tumefaciens multiplication while maintaining growth rates of the C. reinhardtii. Hence, the protocol provided a microenvironment conducive for A. tumefaciens during the co-cultivation step without compromising growth of C. reinhardtii during the selection phase of the transformation protocol.
A set of binary vectors were assembled that carry a synthetic nptII cassette regulated by either the Rbcs2, PsaD, or ß2-tubulin promoter alone or stacked with a YFP and/or ble cassettes (Fig. 1). The base binary vector backbone, pPZP202 (Hajdukiewicz et al. 1994), that was used, along with the disarmed nopaline strain of A. tumefaciens, C58C1/pMP90 (Koncz and Schell 1986), is a standard component in transformation systems often used for higher plants. The transformation protocol was evaluated on the walled strain, C. reinhardtii CC-124. To mitigate confounding results due to bacterial expression of YFP, the ORF was synthesized with the Rbcs intron from C. reinhardtii.
A summary of the Agrobacterium-mediated transformations carried out is listed in Table 1. The results indicated comparable transformation frequencies between the Agrobacterium-mediated delivery and microprojectile bombardment introduction of the respective plasmids, pPTN1133 and pPTN1131, with the latter vector identical to the former except that the YFP cassette was under control of the PsaD rather than the ß2-tubulin promoter (Fig. 1). Observed mean transformation frequencies, calculated based on antibiotic resistant colonies per million cells, ranged from 0.09 to 2.86 (Table 1). Importantly, the respective transformations tabulated in Table 1 were conducted at different times by individuals from different laboratories thereby reflecting the reliability, repeatability, and critically the translatability of this Agrobacterium-mediated transformation protocol for C. reinhardtii.
Molecular Characterization of Transgenic Cell Lineages of C. reinhardtii
Southern blot analyses on a subset of the transgenic cell lineages derived from both microprojectile bombardment and Agrobacterium-mediated transformation methods with the binary vector pPTN1133 (Fig. 1) are shown in Fig. 3. With respect to the former transformation method, the whole circular plasmid was delivered into the cells. The data collected from the hybridization analyses revealed a higher number of multi-locus insertions with the direct DNA approach as compared to Agrobacterium-mediated events (Fig. 3). Surprisingly, though, a relatively high percentage of single locus events (13/17 = 76%) was also observed with microprojectile bombardment, which was atypical in nuclear transformation in higher plants with direct DNA methods.
Co-transformation-based genotyping via PCR of the non-selectable YFP cassette was detected in approximately 59% (10/17) of the microprojectile bombardment events and 42% (5/12) of the Agrobacterium-mediated events (Fig. 2b). Presence of the plasmid backbone outside of the T-DNA was monitored by re-probing the blots with the bacterial selectable marker gene, aadA (Fig. 2c). The data revealed that the aadA marker was integrated in seven of the 17 microprojectile events and absent in all the Agrobacterium-mediated events (Fig. 2c). Hence, unlike what is typically observed in nuclear higher plant transgenic events derived from direct DNA methods (Cheng et al. 2001; Svitashev et al. 2002; Makarevitch et al. 2003), a significant number of transgenic events lack integration of the backbone regions of delivered plasmids. Genotyping by PCR (Fig. 2) of 12 selected Agrobacterium-mediated transgenic lineages, not co-expressing YFP, suggested that a lack of co-expression in eight of the lines was due to truncations in the YFP cassette (Fig. 2).
Junction fragments about the foreign alleles introduced through Agrobacterium-mediated transformation were captured using either TAIL-PCR (Liu et al. 1995) or inverse PCR (Ochman et al. 1988) methodology. The integrity of the junctions, map positions, and gene models in which the T-DNA elements reside from the genotyped events are summarized in Table 2. The junction fragment sequences blast searched against the C. reinhardtii genome (www. Phytozome.net) suggested that, analogous to T-DNA integration in higher plant nuclear compartments, integration was essentially a random event. Among the junction fragments analyzed, five were predicted to interrupt genes (Table 2). Importantly, the analyses indicated that the junction fragments about the site of T-DNA integration are essentially intact (Table 2). This was unlike other reports studying direct DNA integration in the C. reinhardtii nuclear genome where large deletions near the site of integration are often observed (Dent et al. 2005). This attribute was important to consider when implementing transformation approaches for insertional- and/or activation-tagging for functional genomics studies with C. reinhardtii.
The junction fragments isolated from five events were used as probes in Southern blot analysis to visualize molecular weight shifts in the hybridization signals between wild-type CC124 and transgenic lineages as an additional confirmation that the junctions captured were linked to the T-DNA. As can be seen in Fig. 4, in each case the expected upshift in molecular weight of the hybridization signal is observed, supporting the linkage of the captured DNA junction element with the T-DNA.
The junction fragment sequence information about the T-DNA alleles allowed the present study to gather deeper genotypic data on the integrity of the T-DNA element integrated (Fig. 5) in the genome of a subset of the transgenic events produced via Agrobacterium-mediated transformation. A visual representation of the T-DNA elements integrated in nine of the transgenic events derived from Agrobacterium-mediated transformation (pPTN1133; Fig. 1) is shown in Fig. 5. Each of the integrated T-DNAs has significant truncations proximal to the right border element where the visual marker YFP cassette resides, which supports the PCR genotyping results (Fig. 2). This lack of foreign locus integrity in turn can negatively impact co-expression of transgenes in C. reinhardtii. For example, Agrobacterium-mediated transformations conducted with A. tumefaciens transconjugants carrying pPTN1133 and pPTN1170 were monitored for co-expression of YFP. The results revealed that among 54 transgenic pPTN1113 lines selected on paromomycin and monitored for co-expression of YFP, only one was expressing YFP (Fig. 6), translating to a co-expression frequency of 1.9%. Among 51 transgenic pPTN1170 lines selected on paromomycin, one was identified as co-expression YFP (2.0% co-expression); and across 54 transgenic pPTN1170 lines selected on zeocin, none was positive for YFP expression. Employing a dual selection approach, supplementing selection plates with both paromomycin and zeocin, transformation frequencies were drastically reduced; but among three transgenic pPTN1170 dual selected transgenic events, one was positive for YFP expression (Fig. 6).
Integrity of T-DNA elements present in selected Agrobacterium-mediated transgenic events. A diagrammatic view of the T-DNA element of the binary vector pPTN1133 is shown at the bottom. Transgenic event designations are listed along the left side of the figure. The number in parenthesis next to the event designation refers to the base pair deletion within the T-DNA proximal to the RB element. The number in parenthesis along the right side corresponds to the base pair deletion proximal to the LB element.
Confocal images of Agrobacterium-mediated transgenic Chlamydomonas reinhardtii events expressing YFP. Panels labeled WT, pPTN1133, and pPTN1170 refer to wild-type CC124 cells, an Agrobacterium-mediated transgenic event carrying T-DNA element of binary vector pPTN1133 selected on paromomycin, and a dual selected, paromomycin and zeocin, event carrying the T-DNA of binary vector pPTN1170, respectively. YFP* column of images is false colored green. Red panel images display autofluorescence emanating from chlorophyll, while merged panel images are a stacked image of corresponding YFP* and red images.
Using a co-bombardment strategy to simultaneously introduce YFP and nptII cassettes into C. reinhardtii, co-expression of YFP in nptII positive cell lineages ranged from 0.7 up to 3.9% across nine independent co-transformation set-ups. Therefore, unlike in higher plants, co-expression of non-selected transgene(s) was relatively low. Hence, while co-transformation occurred at a rate of over 40% using either microprojectile bombardment or Agrobacterium-mediated gene transfer, integrity of the transgenic allele appeared to be a significant factor impacting co-expression of the non-selectable transgenic cassette (Figs. 2 and 3).
A previous report communicating an Agrobacterium-mediated transformation system for C. reinhardtii using the octopine strain of A. tumefaciens LBA4404 (Hoekema et al. 1983) revealed a 50-fold increase in transformation frequency as compared to the direct DNA delivery using glass beads (Kumar et al. 2004). This protocol utilized the binary vector pCAMBIA1304 which harbors a hygromycin selectable marker along with a fusion visual marker cassette coupling ß-glucuronidase (GUS) and green florescent protein (GFP). Both cassettes were under control of the viral 35S CaMV promoter element (Benfey and Chua 1990), and the GC content of the transgenes was not optimized for C. reinhardtii. These two features of pCAMBIA1304 were not conducive for a reliable transformation system for C. reinhardtii. The 35S CaMV promoter, although reportedly functional in C. reinhardtii (Tang et al. 1995), has relatively poor activity in this alga (Day et al. 1990) and other algae cells (Walker et al. 2005). Codon optimization of transgenes has been shown to be a very important parameter for translational efficiency in C. reinhardtii (Fuhrmann et al. 1999; Shao and Bock 2008). Hence, this may account for the apparent non-repeatability of this earlier Agrobacterium-mediated transformation protocol in the C. reinhardtii scientific community, which is reflected by the minimal number of subsequent communications further exploiting an Agrobacterium-mediated transformation system to manipulate the C. reinhardtii nuclear genome (Pratheesh et al. 2012, 2014).
To indirectly evaluate the effectiveness of the 35S CaMV promoter in C. reinhardtii, a comparative transformation study was conducted using a binary vector harboring standard higher plant nptII cassette under control of the 35S CaMV promoter and pPTN1133 in which nptII is under control of the Rbcs2-HSP70 promoter. Across three independent microprojectile bombardment experiments, no paromomycin-resistant colonies were recovered with the 35S-CaMV::nptII cassette while a mean paromomycin-resistant CFU per million cells of 0.13 ± 0.3 was obtained with the Rbcs2-HSP70::nptII cassette (pPTN1133). This outcome supported the importance of proper designs of transgenic alleles for successful recovery of C. reinhardtii transformants.
More recently, Mini et al. (2018) did a comparative study between a direct DNA-based and Agrobacterium-mediated transformation of C. reinhardtii. In this report, the researchers utilized an nptII allele from Streptomyces rimosus (Sizova et al. 1996, 2001) as the selectable marker rather than the E. coli version typically used in transformation systems of higher plants (Fraley et al. 1983). They also evaluated both walled and wall-less strains of C. reinhardtii, and two A. tumefaciens strains, the octopine strain LBA4404 (Hoekema et al. 1983) and nopaline strain C58C1 (Koncz and Schell 1986). The outcomes reported showed that both Agrobacterium strains could transform the wall-less strain (cw15) with equal efficiency; however, only the nopaline strain was able to transform the walled strain (CW15). These results also confirmed the importance of proper designs of the genetic elements for successful and reliable Agrobacterium-mediated transformation of this model alga, including proper selection of promoter elements, codon optimization of the open reading frames, and a proper co-culturing microenvironment (Mini et al. 2018).
Key attributes of the Agrobacterium-mediated transformation protocol described herein include (1) comparable transformation efficiencies with direct DNA methods when using a walled cell strain; (2) no autolysin pretreatment required; (3) robust application, as it has been successfully conducted in independent laboratories; and (4) junction fragments about the transgenic allele are essentially intact. This latter attribute is of paramount importance for insertional- and/or activation-tagging approaches to gain insight on the underlying biology governing this model alga and for the delivery and expression of transgenic alleles. The presence of large deletions around the foreign allele, which is common when using direct DNA methods for nuclear transformation of C. reinhardtii (Dent et al. 2005), makes it difficult to link genotype to phenotype in a mutant collection and to create cell lines expressing a transgenic allele without collateral damage about the locus in which the transgenic allele resides. The only direct DNA protocol communicated for C. reinhardtii that possesses the attribute of intact junction fragments required the combination of electroporation with linearized DNA containing no vector backbone (Gonzalez-Ballester et al. 2011). This method, while reliable, adds labor and cost in the development of the mutagenized population of C. reinhardtii.
The drawback of the Agrobacterium-mediated transformation protocol outlined herein and by others (Mini et al. 2018) is the observation that the non-selectable transgenic allele is often truncated (Fig. 5). Historically, lack of co-expression of non-selected transgenic cassettes in the C. reinhardtii nuclear compartment has been attributed to epigenetic phenomena (Cerutti et al. 1997), improper regulatory elements controlling gene(s) of interest (Fischer and Rochaix 2001), or GC content issues requiring codon optimization for efficient translation (Shao and Bock 2008). While each of these parameters clearly can have an impact on co-expression frequencies and levels, genotyping of the transgenic C. reinhardtii events generated via microprojectile bombardment and Agrobacterium-mediated transformation methods suggests deletions of the non-selected transgenic cassette are the predominate underlying cause for lack of co-expression. Indeed, taking a mutant screening approach, C. reinhardtii isolates were identified in which the presence of intact non-selected transgenic cassettes was improved, which translated to higher co-expression frequencies (Neupert et al. 2009). Hence, research investments in DNA repair mechanisms of C. reinhardtii (Vlček et al. 2008) may provide insights into steps to improve upon this parameter along with informing future genetic designs that permit for more precision in genetic variation at a locus introduced by genome editing reagents (Sizova et al. 2021).
Successful genome editing in C. reinhardtii has been achieved through deployment of CRISPR-Cas-mediated approaches (Ghribi et al. 2020) with reports communicating CRISPR reagents that incorporate endonucleases from organisms Lachnospiraceae bacterium ND2006 (Ferenczi et al. 2017) and Streptococcus pyogenes (Akella et al. 2021), CRISPR/Cas12a and CRISPR/Cas9 systems, respectively, being effective in creating targeted INDELs in the genome (Ghribi et al. 2020). The editing reagents have been introduced as DNA elements, both transiently (Guzmán-Zapata et al. 2019) and as a stable transgenic allele (Park et al. 2020), or as in vitro assembled ribonucleoproteins (Ferenczi et al. 2017).
Interestingly, frequencies of precise knock-ins were rather high (> 10%) when the assembled ribonucleoprotein complex was co-delivered with either small single-stranded DNA (Ferenczi et al. 2017; Akella et al. 2021) or doubled-stranded expression cassette templates (Kim et al. 2020). These reports opened the possibility that an improved facile knock-in method for C. reinhardtii can be developed by coupling an Agrobacterium-mediated reagent delivery approach with a retron-based in vivo synthesis of single-stranded multi-copy DNA templates by employing Cas9 Retron precise Parallel Editing via homologY (CRISPEY) editing reagents (Sharon et al. 2018; Zhao et al. 2022). The CRISPEY editing reagents can be assembled into a binary vector and the T-DNA cargo introduced either as a stable expressing transgenic allele, using a common A. tumefaciens strain like C58C1/pMP90 (Koncz and Schell 1986), or transiently expressed using a novel mutant strain (Gelvin and Lee 2021) that makes an integrative deficient T-strand complex that is translocated to the algal cell, but fails to form a stable transgenic allele.
Conclusion
A reliable and simple Agrobacterium-mediated delivery was described that provides workable transformation frequencies for the algae model organism C. reinhardtii. The major attribute of the Agrobacterium-mediated method for C. reinhardtii was the integrity of the genomic junction fragments. The drawback of the approach for transformation of this algae, however, was the low co-expression frequency of the non-selectable transgenic allele, which is primarily due to truncations in the T-DNA.
References
Akella S, Ma X, Bacova R, Harmer ZP, Kolackova M, Wen X, Wright DA, Spalding MH, Weeks DP, Cerutti H (2021) Co-targeting strategy for precise, scarless gene editing with CRISPR/Cas9 and donor ssODNs in Chlamydomonas. Plant Physiol 187:2637–2655
Bateman JM, Purton S (2000) Tools for chloroplast transformation in Chlamydomonas: expression vectors and a new dominant selectable marker. Mol Gen Genet 263:404–410
Benfey PN, Chua NH (1990) The cauliflower mosaic virus 35s promoter: combinatorial regulation of transcription in plants. Science 250:959–966
Casas-Mollano JA, Jeong BJ, Xu J, Moriyama H, Cerutti H (2008) The MUT9p kinase phosphorylates histone H3 threonine 3 and is necessary for heritable epigenetic silencing in Chlamydomonas. Proc Natl Acad Sci USA 105:6486–6491
Cerutti H, Johnson AM, Gillham NW, Boynton JE (1997) Epigenetic silencing of a foreign gene in nuclear transformants of Chlamydomonas. Plant Cell 9:925–945
Chen L, Marmey P, Taylor NJ, Brizard JP, Espinoza C, D’Cruz P, Huet H, Zhang S, de Kochko A, Beachy RN, Fauquet C (1998) Expression and inheritance of multiple transgenes in rice plants. Nat Biotechnol 10:1060–1064
Chen R, Matsui K, Ogawa M, Oe M, Ochiai M, Kawashima H, Sakuradani E, Shimizu S, Ishimoto M, Hayashi M, Murooka Y, Tanaka Y (2006) Expression of ∆6, ∆5 desaturase and GLELO elongase genes from Mortierella alpina for the production of arachidonic acid in soybean [Glycine max (L.) Merrill] seeds. Plant Sci 170:399–406
Cheng ZQ, Huang XQ, Wu R (2001) Comparison of biolistic and Agrobacterium-mediated transformation methods on transgene copy number and rearrangement frequency in rice. Acta Bot Sinica 43:826–833
Dai S, Zheng P, Marmey P, Zhang S, Tian W, Chen S, Beachy RN, Fauquet C (2001) Comparative analysis of transgenic rice plants obtained by Agrobacterium-mediated transformation and particle bombardment. Mol Breeding 7:25–33
Dal’Molin CG, Quek L, Palfreyman RW, Nielsen LK (2011) AlgaGEM- a genome-scale metabolic reconstruction of algae based on the Chlamydomonas reinhardtii genome. BMC Genomics 12(Suppl 14):S15
Davies JP, Grossman AR (1994) Sequences controlling transcription of the Chlamydomonas reinhardtii ß2-tubulin gene after deflagellation and during the cell cycle. Mol Cell Biol 14:5165–5174
Day A, Debuchy R, van Dillewijn J, Purton S, Rochaix JD (1990) Studies on the maintenance and expression of cloned DNA fragments in the nuclear genome of the green alga Chlamydomonas reinhardtii. Physiol Plant 78:254–260
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: Version II. Plant Mol Biol Reptr 1:19–21
Dent RM, Haglund CM, Chin BL, Kobayashi MC, Niyogi KK (2005) Functional genomics of eukaryotic photosynthesis using insertional mutagenesis of Chlamydomonas reinhardtii. Plant Physiol 137:545–556
Dunahay G (1993) Transformation of Chlamydomonas reinhardtii with silicon carbide whiskers. Biotechniques 15:457–460
Eckert H, LaVallee B, Schweiger BJ, Kinney AJ, Cahoon EB, Clemente T (2006) Co-expression of the borage ∆6 desaturase and the Arabidopsis ∆15 desaturase results in high accumulation of stearidonic acid in the seeds of transgenic soybean. Planta 224:1050–1057
Ferenczi A, Pyott DE, Xipnitou A, Molnar A (2017) Efficient targeted DNA editing and replacement in Chlamydomonas reinhardtii using Cpf1 ribonucleoproteins and single-stranded DNA. Proc Natl Acad Sci USA 114:13567–13572
Fischer N, Rochaix JD (2001) The flanking regions of PsaD drive efficient gene expression in the nucleus of the green alga Chlamydomonas reinhardtii. Mol Genet Genomics 265:888–894
Fraley RT, Rogers SG, Horsch RB, Sanders PR, Flick JS, Adams SP, Bittner ML, Brand LA, Fink CL, Fry JS, Galluppi GR, Goldberg SB, Hoffmann NL, Woo SC (1983) Expression of bacterial genes in plant cells. Proc Natl Acad Sci USA 80:4803–4807
Franklin SE, Mayfield SP (2004) Prospects for molecular farming in the green alga Chlamydomonas reinhardtii. Curr Opin Plant Biol 7:159–165
Fuhrmann M, Oertel W, Hegemann P (1999) A synthetic gene coding for the green fluorescent protein (GFP) is a versatile reporter in Chlamydomonas reinhardtii. Plant J 19:353–361
Fullner KJ, Nester EW (1996) Temperature affects the T-DNA transfer machinery of Agrobacterium tumefaciens. J Bacteriol 178:1498–1504
Gelvin SB, Lee LY (2021) Methods and composition for transferring T-DNA into plant. USA Patent Application Number 17(494):977
Ghribi M, Nouemssi SB, Meddeb-Mouelhi F, Desgagné-Penix I (2020) Genome editing by CRISPR-Cas: a game change in the genetic manipulation of Chlamydomonas. Life 10:295
Gonzalez-Ballester D, Pootakham W, Mus F, Yang W, Catalanotti C, Magneschi L, de Montaigu A, Higuera JJ, Prior M, Galván A, Fernandez E, Grossman AR (2011) Reverse genetics in Chlamydomonas: a platform for isolating insertional mutants. Plant Methods 7:1–13
Gordon-Kamm WJ, Spencer TM, Mangano ML, Adams TR, Daines RJ, Start WG, O’Brien JV, Chambers SA, Adams WR, Willets NG, Rice TB, Mackey CJ, Krueger RW, Kausch AP, Lemaux PG (1990) Transformation of maize cells and regeneration of fertile transgenic plants. Plant Cell 2:603–618
Guzmán-Zapata D, Sandoval-Vargas JM, Macedo-Osorio KS, Salgado-Manjarrez E, Castrejón-Flores JL, Oliver-Salvador MDC, Durán-Figueroa NV, Nogué F, Badillo-Corona JA (2019) Efficient editing of the nuclear APT reporter gene in Chlamydomonas reinhardtii via expression of a CRISPR-Cas9 module. Int J Mol Sci 20:1247
Hajdukiewicz P, Svab Z, Maliga P (1994) The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol Biol 25:989–994
Hanikenne M (2003) Chlamydomonas reinhardtii as a eukaryotic photosynthetic model for studies of heavy metal homeostasis. New Phytol 159:331–340
Harris EH (2001) Chlamydomonas as a model organism. Annu Rev Plant Physiol Plant Mol Biol 52:363–406
Hoekema A, Hirsch PR, Hooykaas PJJ, Schilperoort R (1983) A binary plant vector strategy based on separation of vir- and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303:179–180
Jeong BJ, Wu-Scharf D, Zhang C, Cerutti H (2002) Suppressors of transcriptional transgenic silencing in Chlamydomonas are sensitive to DNA-damaging agents and reactive transposable elements. Proc Natl Acad Sci USA 99:1076–1081
Kim J, Lee S, Baek K, Jin ES (2020) Site-specific knock-out and on-site heterologous gene overexpression in Chlamydomonas reinhardtii via a CRISPR-Cas9-mediated knock-in method. Front Plant Sci 11:306. https://doi.org/10.3389/fpls.2020.00306
Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 87:1228–1232
Kindle KL, Richards KL, Stern DB (1991) Engineering the chloroplast genome: techniques and capabilities for chloroplast transformation in Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 88:1721–1725
Koncz C, Schell J (1986) The promoter of TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gen Genet 204:383–396
Kumar SV, Misquitta RW, Reddy VS, Rao BJ, Rajam MV (2004) Genetic transformation of the green alga-Chlamydomonas reinhardtii by Agrobacterium tumefaciens. Plant Sci 166:731–738
Liu YG, Mitsukawa N, Oosumi T, Whittier RF (1995) Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J 8:457–463
Makarevitch I, Svitashev SK, Somers DA (2003) Complete sequence analysis of transgene loci from plants transformed via microprojectile bombardment. Plant Mol Biol 52:421–432
McCullen CA, Binns AN (2006) Agrobacterium tumefaciens plant cell interactions and activities required for interkingdom macromolecule transfer. Ann Rev Cell Dev Biol 22:101–127
Merchant SS, Prochnik SE, Vallon O, Harris EH, Karpowicz SJ, Witman GB, Terry A, Salamov A, Fritz-Laylin LK, Marechal-Drouard L, Marshall WF, Qu LH, Nelson DR, Sanderfoot AA, Spalding MH, Kapitonov VV, Ren Q, Ferris P, Lindquist E, Shapiro H, Lucas SM, Grimwood J, Schmutz J, Cardol P, Cerutti H, Chanfreau G, Chen CL, Cognat V, Croft MT, Dent R, Dutcher S, Fernandez E, Fukuzawa H, Gonzalez-Ballester D, Gonzalez-Halphen D, Hallmann A, Hanikenne M, Hippler M, Inwood W, Jabbari K, Kalanon M, Kuras R, Lefebvre PA, Lemaire SD, Lobanov AV, Lohr M, Manuell A, Meier I, Mets L, Mittag M, Mittelmeier T, Moroney JV, Moseley J, Napoli C, Nedelcu AM, Niyogi K, Novoselov SV, Paulsen IT, Pazour G, Purton S, Ral JP, Riano-Pachon DM, Riekhof W, Rymarquis L, Schroda M, Stern D, Umen J, Willows R, Wilson N, Zimmer SL, Allmer J, Balk J, Bisova K, Chen CJ, Elias M, Gendler K, Hauser C, Lamb MR, Ledford H, Long JC, Minagawa J, Page MD, Pan J, Pootakham W, Roje S, Rose A, Stahlberg E, Terauchi AM, Yang P, Ball S, Bowler C, Dieckmann CL, Gladyshev VN, Green P, Jorgensen R, Mayfield S, Mueller-Roeber B, Rajamani S, Sayre RT, Brokstein P, Dubchak I, Goodstein D, Hornick L, Huang YW, Jhaveri J, Luo Y, Martinez D, Ngau WC, Otillar B, Poliakov A, Porter A, Szajkowski L, Werner G, Zhou K, Grigoriev IV, Rokhsar DS, Grossman AR (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318:245–250
Miller R, Wu G, Deshpande RR, Vieler A, Gärtner K, Li X, Moellering ER, Zäuner S, Cornish AJ, Liu B, Bullard B, Sears BB, Kuo MH, Hegg EL, Shachar-Hill Y, Shiu SH, Benning C (2010) Changes in transcript abundance in Chlamydomonas reinhardtii following nitrogen deprivation predict diversion of metabolism. Plant Physiol 154:1737–1752
Mini P, Demurtas OC, Valentini S, Pallara P, Aprea G, Ferrante P, Giuliano G (2018) Agrobacterium-mediated and electroporation-mediated transformation of Chlamydomonas reinhardtii: a comparative study. BMC Biotechnol 18:11. https://doi.org/10.1186/s12896-018-0416-3
Msanne J, Xu D, Konda AR, Casas-Mollano JA, Awada T, Cahoon EB, Cerutti H (2012) Metabolic and gene expression changes triggered by nitrogen deprivation in the photoautotrophically grown microalgae Chlamydomonas reinhardtii and Coccomyxa sp. C-169. Phyochemistry 75:50–59
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight DNA. Nucleic Acid Res 8:4321–4326
Neupert J, Karcher D, Bock R (2009) Generation of Chlamydomonas strains that efficiently express nuclear transgenes. Plant J 57:1140–1150
Neupert J, Shao N, Lu Y, Bock R (2012) Genetic transformation of the model green alga Chlamydomonas reinhardtii. In Transgenic plants: methods and protocols (Dunwell, J.M. and Wetten, A.C. eds). New York, NY: Springer Science + Business, LLC, 35–47.
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120:621–623
Park RV, Asbury H, Miller SM (2020) Modification of a Chlamydomonas reinhardtii CRISPR/Cas9 transformation protocol for use with widely available electroporation equipment. MethodsX 7:100855
Pratheesh PT, Shonima GM, Jiji T, Abraham CI, Muraleedhara KG (2012) Study on efficacy of different Agrobacterium tumefaciens strains in genetic transformation of microalga Chlamydomonas reinhardtii. Adv Appl Sci Res 3:2679–2686
Pratheesh PT, Vineetha M, Kurup GM (2014) An efficient protocol for the Agrobacterium-mediated genetic transformation of microalga Chlamydomonas reinhardtii. Mol Biotechnol 56:507–515
Remacle C, Cardol P, Coosemans N, Gaisne M, Boonefoy N (2006) High-efficiency biolistic transformation of Chlamydomonas mitochondria can be used to insert mutations in complex I genes. Proc Natl Acad Sci USA 103:4771–4776
Shao N, Bock R (2008) A codon-optimized luciferase from Gaussia princeps facilitates the in vivo monitoring of gene expression in the model alga Chlamydomonas reinhardtii. Curr Genet 53:381–388
Sharon E, Chen SAA, Khosla NM, Smith JD, Pritchard JK, Fraser HB (2018) Functional genetic variants revealed by massively parallel precise genome editing. Cell 175:544–557
Shou H, Frame BR, Whitham SA, Wang K (2004) Assessment of transgenic maize events produced by particle bombardment or Agrobacterium-mediated transformation. Mol Breed 13:201–208
Sizova IA, Fuhrmann M, Hegemann P (2001) A Streptomyces rimosus aphVIII gene coding for a new type of phosphotransferase provides stable antibiotic resistance to Chlamydomonas reinhardtii. Gene 17:221–229
Sizova I, Kelterborn S, Verbenko V, Kateriya S, Hegemann P (2021) Chlamydomonas POLQ is necessary for CRISPR/Cas9-mediated gene targeting. G3 11(7), https://doi.org/10.1093/g3journal/jkab114.
Sizova IA, Lapina TV, Frolova ON, Alexandrova NN, Akopiants KE, Danilenko VN (1996) Stable nuclear transformation of Chlamydomonas reinhardtii with a Streptomyces rimosus gene as the selectable marker. Gene 181:13–18
Stachel SE, Messens E, Van Montagu M, Zambryski P (1985) Identification of the signal molecules produced by wounded plant cells that activate T-DNA transfer in Agrobacterium tumefaciens. Nature 318:624–629
Stevens DR, Rochaix JD, Purton S (1996) The bacterial phleomycin resistance gene ble as a dominant selectable marker in Chlamydomonas. Mol Gen Genet 251:23–30
Svitashev SK, Pawlowski WP, Makarevitch I, Plank DW, Somers DA (2002) Complex transgene locus structures implicate multiple mechanisms for plant transgene rearrangement. Plant J 32:433–445
Tang DK, Qiao SY, Wu M (1995) Insertion mutagenesis of Chlamydomonas reinhardtii by electroporation and heterologous DNA. Biochem Mol Biol Int 36:1025–1035
Tang K, Tinjuangjun P, Xu Y, Sun X, Gatehouse JA, Ronald PC, Qi H, Lu X, Christou P, Kohli A (1999) Particle-bombardment-mediated co-transformation of elite Chinese rice cultivars with genes conferring resistance to bacterial blight and sap-sucking insect pests. Planta 208:552–563
Vallon O, Dutcher S (2008) Treasure hunting in the Chlamydomonas genome. Genetics 179:3–6
van Dijk K, Xu H, Cerutti H (2005) Epigenetic silencing of transposons in the green alga Chlamydomonas reinhardtii. In: Hammann C (ed) Nucleic acids and molecular biology (Nellen, W. Springer-Verlag, Berlin, pp 159–178
Vlček D, Ševčovičová A, Sviežená B, Gálová E, Miadoková E (2008) Chlamydomonas reinhardtii: a convenient model system for the study of DNA repair in photoautotrophic eukaryotes. Curr Genet 53:1–22
Walker TL, Purton S, Becker DK, Collet C (2005) Microalgae as bioreactors. Plant Cell Rep 24:629–641
Zhao B, Chen SAA, Lee J, Fraser HB (2022) Bacterial retrons enable precise gene editing in human cells. The CRISPR J 5:31–39
Acknowledgements
Special thanks go out to Christian Elowsky for assistance in confocal imaging and Ed Cahoon for making growth facility space available for cultures. Please note: All plasmids communicated in this manuscript are freely available upon request.
Funding
This work was supported with funds provided by National Science Foundation EPSCoR (Nebraska Center for Algal Biology and Biotechnology) and the Nebraska Research Initiative.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Quach, T.N., Sato, S.J., Behrens, M.R. et al. A facile Agrobacterium-mediated transformation method for the model unicellular green algae Chlamydomonas reinhardtii. In Vitro Cell.Dev.Biol.-Plant 59, 671–683 (2023). https://doi.org/10.1007/s11627-023-10389-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11627-023-10389-7