Abstract
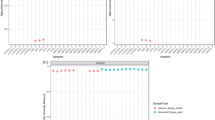
The disposal of hazardous crude oil tank bottom sludge (COTBS) represents a significant waste management burden for South Mediterranean countries. Currently, the application of biological systems (bioremediation) for the treatment of COTBS is not widely practiced in these countries. Therefore, this study aims to develop the potential for bioremediation in this region through assessment of the abilities of indigenous hydrocarbonoclastic microorganisms from Libyan Hamada COTBS for the biotreatment of Libyan COTBS-contaminated environments. Bacteria were isolated from COTBS, COTBS-contaminated soil, treated COTBS-contaminated soil, and uncontaminated soil using Bushnell Hass medium amended with Hamada crude oil (1 %) as the main carbon source. Overall, 49 bacterial phenotypes were detected, and their individual abilities to degrade Hamada crude and selected COBTS fractions (naphthalene, phenanthrene, eicosane, octadecane and hexane) were evaluated using MT2 Biolog plates. Analyses using average well colour development showed that ~90 % of bacterial isolates were capable of utilizing representative aromatic fractions compared to 51 % utilization of representative aliphatics. Interestingly, more hydrocarbonoclastic isolates were obtained from treated contaminated soils (42.9 %) than from COTBS (26.5 %) or COTBS-contaminated (30.6 %) and control (0 %) soils. Hierarchical cluster analysis (HCA) separated the isolates into two clusters with microorganisms in cluster 2 being 1.7- to 5-fold better at hydrocarbon degradation than those in cluster 1. Cluster 2 isolates belonged to the putative hydrocarbon-degrading genera; Pseudomonas, Bacillus, Arthrobacter and Brevundimonas with 57 % of these isolates being obtained from treated COTBS-contaminated soil. Overall, this study demonstrates that the potential for PAH degradation exists for the bioremediation of Hamada COTBS-contaminated environments in Libya. This represents the first report on the isolation of hydrocarbonoclastic bacteria from Libyan COTBS and COTBS-contaminated soil.





Similar content being viewed by others
References
Adetutu E, Bird C, Kadali K, Bueti A, Shahsavari E, Taha M, Patil S, Sheppard P, Makadia T, Simons K (2014) Exploiting the intrinsic hydrocarbon-degrading microbial capacities in oil tank bottom sludge and waste soil for sludge bioremediation. Int J Environ Sci Technol. doi:10.1007/s13762-014-0534-y
Aleer S, Adetutu EM, Makadia TH, Patil S, Ball AS (2011) Harnessing the hydrocarbon-degrading potential of contaminated soils for the bioremediation of waste engine oil. Water Air Soil Pollut 218:121–130
Arun A, Raja PP, Arthi R, Ananthi M, Kumar KS, Eyini M (2008) Polycyclic aromatic hydrocarbons (PAHs) biodegradation by basidiomycetes fungi, Pseudomonas isolate, and their cocultures: comparative in vivo and in silico approach. Appl Biochem Biotechnol 151:132–142
Benedek T, Vajna B, Táncsics A, Márialigeti K, Lányi S, Máthé I (2013) Remarkable impact of PAHs and TPHs on the richness and diversity of bacterial species in surface soils exposed to long-term hydrocarbon pollution. World J Microbiol Biotechnol 29:1989–2002
Bento FM, Camargo FA, Okeke BC, Frankenberger WT (2005) Comparative bioremediation of soils contaminated with diesel oil by natural attenuation, biostimulation and bioaugmentation. Bioresour Technol 96:1049–1055
Bhattacharyya JK, Shekdar AV (2003) Treatment and disposal of refinery sludges: Indian scenario. Waste Manag Res 21:249–261
Bochner B (1989) Sleuthing out bacterial identities. Nature 339:157–158
Brito E, Duran R, Guyoneaud R, Goñi-Urriza M, Garcia De Oteyza T, Crapez MA, Aleluia I, Wasserman JC (2009) A case study of in situ oil contamination in a mangrove swamp (Rio De Janeiro, Brazil). Mar Pollut Bull 58:418–423
Bushnell L, Haas H (1941) The utilization of certain hydrocarbons by microorganisms. J Bacteriol 41:653–673
Cerqueira VS, Hollenbach EB, Maboni F, Vainstein MH, Camargo FAO, Peralba MCR, Bento FM (2011) Biodegradation potential of oily sludge by pure and mixed bacterial cultures. Bioresour Technol 102:11003–11010
Chaerun SK, Tazaki K, Asada R, Kogure K (2004) Bioremediation of coastal areas 5 years after the Nakhodka oil spill in the Sea of Japan: isolation and characterization of hydrocarbon-degrading bacteria. Environ Int 30:911–922
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet 12:499–510
Ferrari M, Neirotti E, Albornoz C, Mostazo M, Cozzo M (1996) Biotreatment of hydrocarbons from petroleum tank bottom sludges in soil slurries. Biotechnol Lett 18:1241–1246
Gallego JLR, Garcia-Martinez MJ, Llamas JF, Belloch C, Peláez AI, Sanchez J (2007) Biodegradation of oil tank bottom sludge using microbial consortia. Biodegradation 18:269–281
Haack SK, Garchow H, Klug MJ, Forney LJ (1995) Analysis of factors affecting the accuracy, reproducibility, and interpretation of microbial community carbon source utilization patterns. Appl Environ Microbiol 61:1458–1468
Hamamura N, Ward DM, Inskeep WP (2013) Effects of petroleum mixture types on soil bacterial population dynamics associated with the biodegradation of hydrocarbons in soil environments. FEMS Microbiol Ecol 85:168–178
Jacques RJ, Okeke BC, Bento FM, Teixeira AS, Peralba MC, Camargo FA (2008) Microbial consortium bioaugmentation of a polycyclic aromatic hydrocarbons contaminated soil. Bioresour Technol 99:2637–2643
Jain PK, Gupta VK, Pathak H, Lowry M, Jaroli D (2010) Characterization of 2 T engine oil degrading indigenous bacteria, isolated from high altitude (Mussoorie), India. World J Microbiol Biotechnol 26:1419–1426
Jones J (1970) Studies on freshwater bacteria: effect of medium composition and method on estimates of bacterial population. J Appl Microbiol 33:679–686
Kadali KK, Simons KL, Skuza PP, Moore RB, Ball AS (2012) A complementary approach to identifying and assessing the remediation potential of hydrocarbonoclastic bacteria. J Microbiol Methods 88:348–355
Kaufman L, Rousseeuw PJ (2009) Finding groups in data: an introduction to cluster analysis. Wiley, New Jersey
Kebria DY, Khodadadi A, Ganjidoust H, Badkoubi A, Amoozegar M (2009) Isolation and characterization of a novel native Bacillus strain capable of degrading diesel fuel. Int J Environ Sci Tech 6:435–442
Kniggendorf A-K, Gaul TW, Meinhardt-Wollweber M (2011) Hierarchical cluster analysis (HCA) of microorganisms: an assessment of algorithms for resonance Raman spectra. Appl Spectrosc 65:165–173
Kriipsalu M, Marques M, Maastik A (2008) Characterization of oily sludge from a wastewater treatment plant flocculation-flotation unit in a petroleum refinery and its treatment implications. J Mater Cycles Waste Manag 10:79–86
Kubota K, Koma D, Matsumiya Y, Chung S-Y, Kubo M (2008) Phylogenetic analysis of long-chain hydrocarbon-degrading bacteria and evaluation of their hydrocarbon-degradation by the 2, 6-DCPIP assay. Biodegradation 19:749–757
Kumar M, León V, Materano ADS, Ilzins OA, Luis L (2008) Biosurfactant production and hydrocarbon-degradation by halotolerant and thermotolerant Pseudomonas sp. World J Microbiol Biotechnol 24:1047–1057
Li P, Sun T, Stagnitti F, Zhang C, Zhang H, Xiong X, Allinson G, Ma X, Allinson M (2002) Field-scale bioremediation of soil contaminated with crude oil. Environ Eng Sci 19:277–289
Machín-Ramírez C, Okoh A, Morales D, Mayolo-Deloisa K, Quintero R, Trejo-Hernández M (2008) Slurry-phase biodegradation of weathered oily sludge waste. Chemosphere 70:737–744
Maioli OL, Rodrigues KC, Knoppers BA, Azevedo DA (2010) Polycyclic aromatic and aliphatic hydrocarbons in Mytella charruana, a bivalve mollusk from Mundaú Lagoon, Brazil. Microchem J 96:172–179
Makadia TH, Adetutu EM, Simons KL, Jardine D, Sheppard PJ, Ball AS (2011) Re-use of remediated soils for the bioremediation of waste oil sludge. J Environ Manag 92:866–871
Mardis ER (2008) The impact of next-generation sequencing technology on genetics. Trends Genet 24:133–141
Margesin R, Labbé D, Schinner F, Greer CW, Whyte LG (2003) Characterization of Hydrocarbon-degrading microbial populations in contaminated and pristine alpine soils. Appl Environ Microbiol 69:3085–3092
Martínková L, Uhnáková B, Pátek M, Nešvera J, Křen V (2009) Biodegradation potential of the genus Rhodococcus. Environ Int 35:162–177
Morozova O, Marra MA (2008) Applications of next-generation sequencing technologies in functional genomics. Genomics 92:255–264
Nwuche C, Ugoji E (2008) Effects of heavy metal pollution on the soil microbial activity. Int J Environ Sci Tech 5:409–414
Osborn AM, Moore ER, Timmis KN (2000) An evaluation of terminal‐restriction fragment length polymorphism (T‐RFLP) analysis for the study of microbial community structure and dynamics. Environ Microbiol 2:39–50
Palmroth MR, Langwaldt JH, Aunola TA, Goi A, Puhakka JA, Tuhkanen TA (2006) Treatment of PAH‐contaminated soil by combination of Fenton’s reaction and biodegradation. J Chem Technol Biotechnol 81:598–607
Peng RH, Xiong AS, Xue Y, Fu XY, Gao F, Zhao W, Tian YS, Yao QH (2008) Microbial biodegradation of polyaromatic hydrocarbons. FEMS Microbiol Rev 32:927–955
Perfumo A, Banat I, Canganella F, Marchant R (2006) Rhamnolipid production by a novel thermophilic hydrocarbon-degrading Pseudomonas aeruginosa AP02-1. Appl Microbiol Biotechnol 72:132–138
Roy S, Hens D, Biswas D, Biswas D, Kumar R (2002) Survey of petroleum-degrading bacteria in coastal waters of Sunderban Biosphere Reserve. World J Microbiol Biotechnol 18:575–581
Sheppard PJ, Adetutu EM, Makadia TH, Ball AS (2011) Microbial community and ecotoxicity analysis of bioremediated, weathered hydrocarbon-contaminated soil. Soil Res 49:261–269
Sopeña F, Laiz L, Morillo E, Sanchez-Trujillo MA, Villaverde J, Jurado V, Saiz-Jimenez C (2013) Phenanthrene biodegradation by Pseudomonas xanthomarina isolated from an aged contaminated soil. CLEAN–Soil Air Water. doi:10.1002/clen.201300247
Tang J, Wang R, Niu X, Wang M, Chu H, Zhou Q (2010) Characterisation of the rhizoremediation of petroleum-contaminated soil: effect of different influencing factors. Biogeosciences 7:3961–3969
Varanasi P, Fullana A, Sidhu S (2007) Remediation of PCB contaminated soils using iron nano-particles. Chemosphere 66:1031–1038
Verma S, Bhargava R, Pruthi V (2006) Oily sludge degradation by bacteria from Ankleshwar, India. Int Biodeterior Biodegrad 57:207–213
Vézie C, Rapala J, Vaitomaa J, Seitsonen J, Sivonen K (2002) Effect of nitrogen and phosphorus on growth of toxic and nontoxic Microcystis strains and on intracellular microcystin concentrations. Microb Ecol 43:443–454
Vitte I, Duran R, Jézéquel R, Caumette P, Cravo-Laureau C (2011) Effect of oxic/anoxic switches on bacterial communities and PAH biodegradation in an oil-contaminated sludge. Environ Sci Pollut Res 18:1022–1032
Wang Q, Zhang S, Li Y, Klassen W (2011) Potential approaches to improving biodegradation of hydrocarbons for bioremediation of crude oil pollution. J Environ Prot 2:47–55
Whang L-M, Liu P-WG, Ma C-C, Cheng S-S (2008) Application of biosurfactants, rhamnolipid, and surfactin, for enhanced biodegradation of diesel-contaminated water and soil. J Hazard Mater 151:155–163
Xu JL, Zhang J, Huang TL, Xi AL (2013) Comparative bioremediation of oil contaminated soil by natural attenuation, biostimulation and bioaugmentation. Adv Mater Res 777:258–262
Yuste L, Corbella ME, Turiégano MJ, Karlson U, Puyet A, Rojo F (2000) Characterization of bacterial strains able to grow on high molecular mass residues from crude oil processing. FEMS Microbiol Ecol 32:69–75
Zhang X, Li J, Huang Y, Thring R (2010) Surfactant enhanced biodegradation of petroleum hydrocarbons in oil refinery tank bottom sludge. J Can Pet Technol 49:34–39
Zhao T, Tang X, Wang D, Gan X, Hua R (2011) Isolation and identification of malathion-degrading strain of Brevundimonas diminuta. Electrical and Control Engineering (ICECE), 2011 International Conference on, 2011. IEEE, 1600-1602
Acknowledgments
This work was supported by the Libyan Ministry of Higher Education and Science Research. The authors thank the Environmental and Natural Resources Engineering, Azzawiya University, Libya, the management of Azzawiya oil refinery, Libya, and brothers Ali Mansour, Mohamad Ahfeda and Nassradden Souf for supplying soil and COTBS samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Robert Duran
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Figure. 1a Chromatogram of soil extract obtained from treated COTBS contaminated soil. Figure. 1 b: Chromatogram of COTBS. (PPTX 363 kb)
Rights and permissions
About this article
Cite this article
Mansur, A.A., Adetutu, E.M., Kadali, K.K. et al. Assessing the hydrocarbon degrading potential of indigenous bacteria isolated from crude oil tank bottom sludge and hydrocarbon-contaminated soil of Azzawiya oil refinery, Libya. Environ Sci Pollut Res 21, 10725–10735 (2014). https://doi.org/10.1007/s11356-014-3018-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-014-3018-1