Abstract
Liriodendron tulipifera L., a member of Magnoliaceae in the order Magnoliales, has been used extensively as a reference species in studies on plant evolution. However, genomic resources for this tree species are limited. We constructed cDNA libraries from ten different types of tissues: premeiotic flower buds, postmeiotic flower buds, open flowers, developing fruit, terminal buds, leaves, cambium, xylem, roots, and seedlings. EST sequences were generated either by 454 GS FLX or Sanger methods. Assembly of almost 2.4 million sequencing reads from all libraries resulted in 137,923 unigenes (132,905 contigs and 4,599 singletons). About 50% of the unigenes had significant matches to publically available plant protein sequences, representing a wide variety of putative functions. Approximately 30,000 simple sequence repeats were identified. More than 97% of the cell wall formation genes in the Cell Wall Navigator and the MAIZEWALL databases are represented. The cinnamyl alcohol dehydrogenase (CAD) homologs identified in the L. tulipifera EST dataset showed different expression levels in the ten tissue types included in this study. In particular, the LtuCAD1 was found to partially recover the stiffness of the floral stems in the Arabidopsis thaliana CAD4 and CAD5 double mutant plants, of the LtuCAD1 in lignin biosynthesis. L. tulipifera genes have greater sequence similarity to homologs from other woody angiosperm species than to non-woody model plants. This large-scale genomic resour"HistryDatesce will be instrumental for gene discovery, cDNA microarray production, and marker-assisted breeding in L. tulipifera, and strengthen this species' role in comparative studies.






Similar content being viewed by others
References
Albert VA, Soltis DE, Carlson JE, Farmerie WG, Wall PK, Ilut DC et al (2005) Floral gene resources from basal angiosperms for comparative genomics research. BMC Plant Biol 5:5
Allen E, Xie Z, Gustafson AM, Carrington JC (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121:207–221
Altschul SF, Gish W, Miller EW, Myiers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Axtella MJ, Bartel DP (2005) Antiquity of microRNAs and their targets in land plants. Plant Cell 17:1658–1673
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Bae K, Byun J (1987) Screening of leaves of higher plants for antibacterial action. Kor J Pharmacog 8:1
Barakat A, Bagniewska-Zadworna A, Choi A, Plakkat U, DiLoreto D, Yellanki P, Carlson J (2009) The cinnamyl alcohol dehydrogenase gene family in Populus: phylogeny, organization, and expression. BMC Plant Biol 9:26
Berlin A, Maximenko V, Bura R, Kang KY, Gilkes N, Saddler J (2005) A rapid microassay to evaluate enzymatic hydrolysis of lignocellulosic substrates. Biotechnol Bioeng 93:880–886
Beck DE (1990) Liriodendron tulipifera L. yellow-poplar. In: Burns RM, Honkala BH (tech. coords.) Silvics of North America: 1. Conifers; 2. Hardwoods. Agriculture Handbook 654, USDA, Forest Service, Washington, DC, 2:877
Carlson JE, Leebens-Mack JH, Wall PK, Zahn LM, Mueller LA, Landherr LL, Hu Y, Ilut DC, Arrington JM, Choirean S, Becker A, Field D, Tanksley SD, Ma H, dePamphilis CW (2006) EST database for early flower development in California poppy (Eschscholzia californica Cham., Papaveraceae) tags over 6,000 genes from a basal eudicot. Plant Mol Biol 62:351–369
Çelen I, Harper D, Labbé N (2008) A multivariate approach to the acetylated poplar wood samples by near infrared spectroscopy. Holzforschung 62:189–196
Chanderbali AS, Yoo M-J, Zahn LM, Brockington SF, Wall PK, Gitzendanner MA, Albert VA, Leebens-Mack J, Altman NS, Ma H, dePamphilis DW, Soltis DE, Soltis PS (2010) Conservation and canalization of gene expression during angiosperm diversification accompany the origin and evolution of the flower. PNAS 107:22570–22575
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Chevreux B, Pfisterer T, Drescher B, Driesel AJ, Müller WEG, Wetter T, Suhai S (2004) Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res 14:1147–1159
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 24:P3
Desfeux C, Clough SJ, Bent AF (2000) Female reproductive tissues are the primary target of Agrobacterium-mediated transformation by the Arabidopsis floral-dip method. Plant Physiol 123:895–904
Dsouza M, Larsen N, Overbeek R (1997) Searching for patterns in genomic data. Trends Genet 13:497–498
Girke T, Lauricha J, Tran H, Keegstra K, Raikhel NV (2004) The cell wall navigator database: a systems-based approach to organism-unrestricted mining of protein families involved in cell wall metabolism. Plant Physiol 136:3003–3008
Guillaumie S, San Clemente H, Deswarte C, Martinez Y, Lapierre C, Murigneux A, Barrière Y, Pichon M, Goffner D (2007) MAIZEWALL: database and developmental gene expression profiling of cell wall biosynthesis and assembly in maize. Plant Physiol 143:339–363
Hernandez R, Davalos JF, Sonti SS, Kim Y, Moody RC (1997) Strength and stiffness of reinforced yellow-poplar glued laminated beams. Res Pap FPL-RP-554. US Department of Agriculture, Forest Service, Forest Products Laboratory, Madison
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protoc 4:44–57
Hunt D (1998) Magnolias and their allies. International Dendrology Society & Magnolia Society, pp 304
Hwang SS, Lee SJ, Kim HK, Ka JO, Kim KJ, Song HG (2008) Biodegradation and saccharification of wood chips of Pinus strobus and Liriodendron tulipifera by white rot fungi. J Microbiol Biotechnol 18:1819–1826
Jansen RK, Cai Z, Raubeson LA, Daniell H, dePamphilis CW, Leebens-Mack J, Müller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee S-B, Peery R, McNeal JR, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. PNAS 104:19369–19374
Jackson S, Rounsley S, Purugganan M (2006) Comparative sequencing of plant genomes: choices to make. Plant Cell 18:1100–1104
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Karlin S, Campbell AM, Mrázek J (1998) Comparative DNA analysis across diverse genomes. Ann Rev Gen 32:185–225
Kim KD, Lee EJ (2005) Potential tree species for use in the restoration of unsanitary landfills. Environ Manage 36:1–14
Klugh KR, Cumming JC (2003) Variation in organic acid exudates among mycorrhizal species colonizing Liriodendron tulipifera L. (yellow poplar) in the presence of aluminum. Ecological Society of America Annual Meeting Abstracts 88:186
Koo B-W, Min B-C, Park N, Eom C, Yeo H, Ryu K-O, Choi I-G (2008) Chemical and physical characterizations of Liriodendron tulipifera on growth periods. Proceedings of the 30th Symposium on Biotechnology for Fuels and Chemicals, 4–7 May, New Orleans, LA, USA
Koo B-W, Park N, Yeo H, Lee S-Y, Kim H-Y, Kim H, Choi I-G (2009) Organosolv pretreatment of Liriodendron tulipifera with acid and alkali catalysts. Proceedings of the 31st Symposium on Biotechnology for Fuels and Chemicals, 3–6 May, San Francisco, CA, USA
Kuhl JC, Cheung F, Yuan Q, Martin W, Zewdie Y, McCallum J, Catanach A, Rutherford P, Sink KC, Jenderek M, Prince JP, Town CD, Havey MJ (2004) A unique set of 11,008 onion expressed sequence tags reveals expressed sequence and genomic differences between the monocot orders Asparagales and Poales. Plant Cell 16:114–125
Kumpatla S, Mukhopadhyay S (2005) Mining and survey of simple sequence repeats in expressed sequence tags of dicotyledonous species. Genome 48:985
Lafayette PR, Eriksson KE, Dean JF (1999) Characterization and heterologous expression of laccase cDNAs from the lignifying xylem of yellow-poplar (Liriodendron tulipifera). Plant Mol Biol 40:23–35
Li X, Wu HX, Dillon SK, Southerton SG (2009) Generation and analysis of expressed sequence tags from six developing xylem libraries in Pinus radiata D. Don. BMC Genomics 10:41
Li X, Wu HX, Southerton SG (2010) Comparative genomics reveals conservative evolution of the xylem transcriptome in vascular plants. BMC Evol Biol 10:190
Liang H, Barakat A, Schlarbaum SE, Carlson JE (2011) Organization of the chromosome region harboring a FLORICAULA/LEAFY gene in Liriodendron. Tree Genet Genomes 7:373–384
Liang H, Barakat A, Schlarbaum SE, Mandoli DS, Carlson JE (2010) Comparison of gene order of GIGANTEA loci in yellow-poplar, monocots and eudicots. Genome 53:533–544
Liang H, Carlson JE, Leebens-Mack JH, Wall PK, Mueller LA, Buzgo M et al (2008) An EST database for Liriodendron tulipifera L. floral buds: the first EST resource for functional and comparative genomics in Liriodendron. Tree Genet Genomes 4:419–433
Liang H, Fang EG, Tomkins JP, Luo M, Kudrna D, Kim HR et al (2007) Development of a BAC library for yellow-poplar (Liriodendron tulipifera) and the identification of genes associated with flower development and lignin biosynthesis. Tree Genet Genomes 3:215–225
Marchler-Bauer A, Panchenko AR, Shoemaker BA, Thiessen PA, Geer LY, Bryant SH (2002) CDD: a database of conserved domain alignments with links to domain three-dimensional structure. Nucl Acids Res 30:281–283
McInerney J (1998) GCUA (General Codon Usage Analysis). Bioinformatics 14:372–373
Merkle SA, Hoey MT, Watson-Pauley BA, Schlarbaum SE (1993) Propagation of Liriodendron hybrids via somatic embryogenesis. Plant Cell Tissue Organ Cult 34:191–198
Moody RC, Hernandez R, Davalos JF, Sonti SS (1993) Yellow poplar timber beam performance. Res. Pap. FPL-RP-520. Department of Agriculture, Forest Service, Forest Products Laboratory, Madison
Moon MK, Oh HM, Kwon BM, Baek NI, KimSH KJS, Kim DK (2007) Farnesyl protein transferase and tumor cell growth inhibitory activities of lipiferolide isolated from Liriodendron tulipifera. Arch Pharm Res 30:299–302
Moore MJ, Bell CD, Soltis PS, Soltis DE (2007) Using plastid genome-scale data to resolve enigmatic. PNAS 104:19363–19368
Murray EE, Lotzer J, Eberle M (1989) Codon usage in plant genes. Nucl Acid Res 17:477–498
Parks CR, Wendel JF (1990) Molecular divergence between Asian and North American species of Liriodendron (Magnoliaceae) with implications for interpretation of fossil floras. Am J Bot 77:1243–1256
Poinar HN, Schwarz C, Qi J, Shapiro B, Macphee RD, Buigues B, Tikhonov A, Huson DH, Tomsho LP, Auch A et al (2006) Metagenomics to paleogenomics: large-scale sequencing of mammoth DNA. Science 311:392–394
Qiu YL, Dombrovska O, Lee J, Li L, Whitlock BA, Bernasconi-Quadroni F, Rest JS, Davis CC, Borsch T, Hilu KW, Renner SS, Soltis DE, Soltis PS, Zanis MJ, Cannone JJ, Gutell RR, Powell M, Savolainen V, Chatrou LW, Chase MW (2005) Phylogenetic analyses of basal angiosperms based on nine plastid, mitochondrial, and nuclear genes. Intl J Plant Sci 166:815–842
Ranocha P, Chabannes M, Chamayou S, Danoun S, Jauneau A, Boudet AM, Goffner D (2002) Laccase down-regulation causes alterations in phenolic metabolism and cell wall structure in poplar. Plant Physiol 129:145–155
Rengel D, Clemente HS, Servant F, Ladouce N, Paux E, Wincker P, Couloux A, Sivadon P, Grima-Pettenati J (2009) A new Eucalyptus genomic resource dedicated to wood formation: a comprehensive survey. BMC Plant Biol 9:36
Ronse de Craene L, Soltis DE, Soltis PS (2003) Evolution of floral structures in basal angiosperms. Intl J Plant Sci 164:S329–S363
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols in the series Methods in Molecular Biology. Humana, Totowa, pp 365–386
Sibout R, Eudesb A, Mouilleb G, Polletc B, Lapierrec C, Jouaninb L, Séguin A (2005) CINNAMYL ALCOHOL DEHYDROGENASE-C and -D are the primary genes involved in lignin biosynthesis in the floral stem of Arabidopsis. Plant Cell 17:2059–2076
Sibout R, Eudes A, Pollet B, Goujon T, Mila I, Granier F, Séguin A, Lapierre C, Jouanin L (2003) Expression pattern of two paralogs encoding cinnamyl alcohol dehydrogenases in Arabidopsis: isolation and characterization of the corresponding mutants. Plant Physiol 132:848–860
Soltis PS, Soltis DE, Chase MW, Endress PK, Crane PR (2004) The diversification of flowering plants. In: Cracraft J, Donoghue MJ (eds) Assembling the tree of life. Oxford University Press, Oxford, pp 154–167
Soltis DE, Soltis PS, Chase MW, Endress P (2005) Phylogeny, evolution, and classification of flowering plants. Sinauer Associates, Sunderland
Soltis DE, Ma H, Frohlich MW, Soltis PS, Albert VA, Oppenheimer DG, Altman NS, dePamphilis CD, Leebens-Mack J (2007) The floral genome: an evolutionary history of gene duplication and shifting patterns of gene expression. Trends Plant Sci 12:358–367
Wall PK, Leebens-Mack J, Müller KF, Field D, Altman NS, de Pamphilis CW (2008) PlantTribes: a gene and gene family resource for comparative genomics in plants. Nucleic Acids Res 36:D970–D976
Wei ZX, Wu ZY (1993) Pollen ultrastructure of Liriodendron and its systematic significance. Acta Bot Yunnanica 15:163–166
Williams RS, Feist WC (2004) Durability of yellow-poplar and sweetgum and service life of finishes after long-term exposure. For Prod J 54:96–101
Xiang Q, Lee LYY, Torget RW (2004) Kinetics of glucose decomposition during dilute-acid hydrolysis of lignocellulosic biomass. Appl Biochem Biotechnol 114:1127–1138
Xu M, Li H, Zhang B (2006) Fifteen polymorphic simple sequence repeat markers from expressed sequence tags of Liriodendron tulipifera. Mol Ecol Notes 6:728–730
Xu M, Sun Y, Li H (2010) EST-SSRs development and paternity analysis for Liriodendron spp. New For 40:361–382
Zahn LM, Kong H, Leebens-Mack JH, Kim S, Soltis PS, Landherr LL, Soltis D, dePamphilis CW, Ma H (2005) The evolution of the SEPALLATA subfamily of MADS-box genes: a pre-angiosperm origin with multiple duplications throughout angiosperm history. Genetics 169:2209–2223
Zahn LM, Leebens-Mack J, de Pamphilis CW, Ma H, Theissen G (2006) To B or not to B a flower: the role of DEFICIENS and GLOBOSA orthologs in the evolution of the angiosperms. J Hered 96:225–240
Zhang K, Qian Q, Huang Z, Wang Y, Li M, Hong L, Zeng D, Gu M, Chu C, Cheng C (2006) GOLD HULL and INTERNODE2 encodes a primarily multifunctional cinnamyl-alcohol dehydrogenase in rice. Plant Physiol 140:972–983
Acknowledgments
We thank Stephan Schuster and Lynn Tomsho for their assistance in 454 sequencing, Yi Hu for RNA isolations, Denis S. Diloreto for seedlings, Stephen Ficklin for the mining of SSRs, and Xinguo Li for providing the pure xylem unigenes for Populus, loblolly pine, and white spruce. This study was mainly supported by the National Science Foundation grant, Ancestral Angiosperm Genome project (Award # DBI-0638595, PI: dePamphilis). A National Institute of Food and Agriculture, USDA grant to HL (project number SC-1700324, technical contribution No. 5832 of the Clemson University Experiment Station) contributed the sequencing of a one half 454 plate.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by R. Sederoff
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
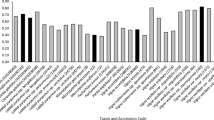
Cumulative codon usage in Liriodendron tulipifera transcriptome (DOC 57 kb)
Online Resource 2
Dinucleotide frequencies (DOC 31 kb)
Online Resource 3
(TXT 0 kb)
Online Resource 4
(PDF 25,536 kb)
Online Resource 5
(TXT 5,654 kb)
Online Resource 6
(TXT 15 kb)
Online Resource 7
Conserved miRNAs identified in Liriodendron tulipifera cDNA unigenes (TXT 4 kb)
Online Resource 8
Liriodendron genes potentially targeted by the identified miRNAs. The targets having hits in the Cell Wall Navigator and/or Maize Wall databases are highlighted (TXT 57 kb)
Online Resource 9
Liriodendron tulipifera unigenes with significant hits in the Cell Wall Navigator database (TXT 389 kb)
Online Resource 10
Liriodendron tulipifera unigenes with significant hits in the Maize Wall database (TXT 619 kb)
Rights and permissions
About this article
Cite this article
Liang, H., Ayyampalayam, S., Wickett, N. et al. Generation of a large-scale genomic resource for functional and comparative genomics in Liriodendron tulipifera L.. Tree Genetics & Genomes 7, 941–954 (2011). https://doi.org/10.1007/s11295-011-0386-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-011-0386-2