Abstract
Underutilised crops suffer from under-investigation relative to more mainstream crops, but often possess improved stress tolerance and/or nutrition, making them potentially important for breeding programmes in the context of climate change and an expanding human population. Developing basic genome resources for underutilised crops may therefore catalyse analyses to facilitate their use, through improved understanding of population structure, phylogeny, candidate genes, and linkage mapping. We carried out nuclear and plastid genome sequencing and assembly for five underutilised legumes: jack bean, sword bean, Kersting’s groundnut, moth bean, and zombi pea. Using only ‘off-the-shelf’, free-to-use bioinformatic tools, we also developed a simple but effective pipeline to identify thousands of markers, which could be applied in other species. We assembled 53–68% of the genome and 73–95% of the gene space in the five legumes. The assemblies were fragmented but nevertheless useful for identifying between 34,000—60,000 microsatellites. Examination of 32 markers in zombi pea revealed 16 primer pairs which amplified in at least half of the eight accessions tested and were polymorphic. We also present nuclear genome size estimates for 17 legume taxa (12 for the first time), comprising the above five species as well as other domesticated legume species and crop wild relatives. We aim for the newly developed markers and genome size estimates presented here to be useful for the research community by aiding genomic and population genetic studies for these taxa, and to provide information on approaches that can be applied for investigating other important yet underutilised crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Climate change and a rapidly increasing population underlie major concerns over food security (Schmidhuber and Tubiello 2007; FAO 2016). Currently, there is an overreliance on a limited number of staple crops, with 90% of the global basic nutritional intake being derived from only fifteen crops (FAO 2020). This is leading to an increasingly homogenous food supply dominated by globally important cereal and oil crops, along with a decline of other cereal and tuber crops (Khoury et al. 2014). Though substantial efforts have been made to create climate- and disease-resistant varieties of staple crops that are also high yielding, these have a limited capacity to adapt to different growing conditions, and productivity gains from crop improvement are likely reaching a plateau (Grassini et al. 2013). Current agricultural practices, such as excessive fertiliser use, are unsustainable and major changes in food production will be needed to eradicate hunger and malnutrition whilst coping with a rising population and the changing climate conditions. Greater crop biodiversity is crucial for the development of stable crop production systems that harvest sufficient food (Campbell et al. 2016; Massawe et al. 2016; Renard and Tilman 2019).
Underutilised crops are domesticated, semi-domesticated, or wild species or varieties of plants that may be significant in local production systems, but are currently overlooked in research, breeding, and policy making (Chivenge et al. 2015; Massawe et al. 2015; Hunter et al. 2019; Li et al. 2020). Many of these species were previously widely grown but have fallen into disuse for a range of economic, agronomic, and cultural reasons. Globally, there are hundreds of underutilised plants that could offer alternative food sources; it is therefore important to investigate these species and identify solutions for negative attributes that may reduce their popularity, such as growing and storage difficulties and the presence of antinutrients (Li and Siddique 2020).
Introducing a new crop into standard farming practices is challenging; it requires dedicated breeding programmes to establish these cultivars and make them economically viable. In developed countries, where crop breeding is mostly conducted by private companies rather than public communities, underutilised crops are rarely considered, and breeding programmes carried out by universities tend to have low impact on farming practices. This lack of investment slows the development of true-breeding, high-yielding cultivars of potential value for global food systems, while the continued high research interest and dissemination of major crops results in minor crop varieties and the associated indigenous knowledge being lost (Chivenge et al. 2015; Khoury et al. 2022). Improving an underutilised crop through breeding not only rescues the threatened crop varieties, but may also further increase interest and encourage investment by private and public sectors (Li et al. 2020), thus efforts to make these crops more marketable would be beneficial (Stamp et al. 2012).
Generating molecular markers for use in determining relationships between different populations and varieties, generating genetic maps, and identifying genomic regions controlling adaptive (or maladaptive) traits will help us begin to understand the genetics and attributes of underutilised crops and provide valuable resources to inform molecular breeding techniques (Hodel et al. 2016a; Chapman 2019). The advent of high-throughput sequencing (HTS) and accompanying bioinformatic tools have reduced the cost and time required for assembling genome-scale sequences (e.g., whole genome assemblies and transcriptomes) and developing markers such as SSRs (i.e., simple sequence repeats or microsatellites) (Hodel et al. 2016b). These approaches have thus been adopted for hundreds of plant species in the last decade, including several underutilised crops (e.g., Chapman 2015; Vatanparast et al. 2016; Sathyanarayana et al. 2017; Singh et al. 2020). Furthermore, even draft quality genome assemblies can serve additional purposes, such as identifying genes of interest (e.g., loci associated with drought tolerance) and examining sequence evolution across taxa (Fisher et al. 2022).
Legumes (Fabaceae) comprise the second most important group of crops after cereals (Poaceae) and have a large impact on nutritional security due to their high protein content. Their ability to fix atmospheric nitrogen also means that they can often grow on poor soils without additional fertiliser input (Kebede 2021). Given the environmental and economic cost of synthetic fertilisers, the expansion of legume production could have wide benefits for farmers and the environment. In addition, several underutilised legumes are perennial, leading to an improved soil stability.
Here we present draft nuclear and plastid genomes and genome-derived SSR markers for five species of underutilised legumes with no available genomic data in public repositories as of time of publishing: jack bean (Canavalia ensiformis (L.) DC.), sword bean (Canavalia gladiata (Jacq.) DC.), Kersting’s groundnut (Macrotyloma geocarpum (Harms) Maréchal & Baudet), moth bean (Vigna aconitifolia (Jacq.) Marechal), and zombi pea (Vigna vexillata (L.) A. Rich). These are all nutrient rich and possess characteristics that give resistance to various environmental conditions under which staple and well-known legumes may struggle, as explained below.
Jack bean and sword bean are closely related climbing legumes in the genus Canavalia. The two species are similar in growth habit, but distinguished by seed colour and hilum length (Moteetee 2016). Jack bean is typically an annual plant native to South and Central America, with uses in both human diets and animal fodder. Young green pods are consumed as a vegetable in various parts of Asia; it is also farmed on a modest scale in non-Asian nations and can be grown in marginal soils with low water availability due to its deep roots. Sword bean is a perennial species that originated in East Asia and is now grown throughout the tropics (Haq 2011). It is generally considered a hardy species with potentially useful agricultural traits (Ekanayake et al. 2000; Haq 2011), including a deep root system that confers resilience to drought and waterlogging. Both species contain various antinutrients, including lectins, tannins, and protease inhibitors (Vadivel et al. 2010; Doss et al. 2011) that must be removed using specific preparation methods before safe consumption in high quantities. Nevertheless, jack bean and sword bean have potential to be used more widely in human nutrition as their protein quality is comparable to staple legume crops and they are rich in macro- and micronutrients. For example, jack bean contains a high quantity of protein (23–34%) derived from several (but not all) essential amino acids, and is additionally a good source of carbohydrates (ca. 55%) and micronutrients (Akpapunam and Sefa-Dedeh 1997). Due to its low-fat content (0.2% in fresh seeds), jack bean may represent a viable source of nutrition for those seeking a low-fat, high-protein fortified diet. Trials have demonstrated that jack bean can be incorporated in popular dishes typically made from other legumes (Karoli et al. 2017). Sword bean is used in various parts of Asia as a substitute for mashed potatoes or broad beans, the leaves have the potential to cure ailments such as skin rashes and constipation. The two species are phylogenetically distinct from many staple legumes, being found in the subtribe Diocleinae and therefore divergent from chickpea (Cajaninae), soybean (Glycininae) and the dozens of Vigna and Phaseolus crop species (Phaseolinae) (Kajita et al. 2001). This placement makes their investigation important from the standpoint of understanding the phylogenetics of the legumes more broadly.
Kersting’s groundnut, widely grown in West Africa, is a geocarpic legume, i.e. pods develop underground, the same growth habit observed in peanuts and Bambara groundnut. It is drought resistant and higher in essential minerals and the amino acids lysine, arginine and methionine than other legumes (Assogba et al. 2016; Ayenan and Ezin 2016). Kersting’s groundnut has a palatable taste and has potential for use in infant food formulations, as a source of iron for anaemic patients, as well as helping prevent malnutrition (Garcia-Casal et al. 2018). Despite these benefits, Kersting’s groundnut, especially as a food, is becoming increasingly neglected (Chivenge et al. 2015)—yield is low relative to other similar crops, harvesting the underground pods is physically demanding, and the seeds are highly susceptible to Bruchid infestations post-harvest, leading to large losses. Therefore, the beans must be sold quickly and cannot be relied upon during food shortages (Chivenge et al. 2015). Nevertheless, it is used in rituals by some groups, which is helping to conserve the species. Although relatively understudied, recent genetic analyses have (1) revealed the relationships between Kersting’s groundnut and two other underutilised crops in the genus (i.e. Macrotyloma axillare and M. uniflorum (Fisher et al. 2022)), and (2) demonstrated significant population structure but low genetic diversity overall in Kersting’s groundnut (Kafoutchoni et al. 2021).
Moth bean and zombi pea are species in the genus Vigna, which encompasses multiple well-known food crops such as cowpea, adzuki bean, mung bean and dozens of minor legume crops (Delgado-Salinas et al. 2011). Moth bean has been identified as a potentially significant crop for the future due to its high content of digestible protein and micronutrients (especially iron and zinc), nutraceutical properties (as an anti-diabetic, antioxidant and anti-hypertensive), and medicinal properties (as a pain reliever) (Adsule 1996; Bhadkaria et al. 2022). Moth bean is grown primarily in the arid regions of India and Pakistan, and its high drought tolerance highlights its potential for cultivation in parts of Africa and other water-limited areas, especially under the predicted future climate (Baath et al. 2018). Nevertheless, it remains as an underutilised crop due to the low yield of existing cultivars and difficulty of harvesting (mowers cannot be used due to the shape of the plants therefore sickles are typically used). Moth bean also contains antinutrients, but these can be reduced via traditional processing methods.
Zombi pea is a perennial climbing legume of African origin that is mainly used for food, forage, and erosion control (Karuniawan et al. 2006). There are two domesticated forms: a seed form that is grown in Africa, and a tuberous root form found in a small region around Indonesia (Dachapak et al. 2018). Wild populations of the species are found throughout the tropics and subtropics, where they may be harvested for local use (Amkul et al. 2020). Zombi pea is used primarily as a backup crop due to its reliability during periods of both high and low rainfall, when more popular crops such as sweet potato and cassava cannot be grown (Karuniawan et al. 2006). Moreover, the tubers tend to be high in protein compared to traditional root crops (Chandel 1972, cited in (Dachapak et al. 2017). The wide distribution of zombi pea and its resilience to varying water availability could make it an excellent candidate crop for future improvement (Karuniawan et al. 2006).
Despite their current status as underutilised, the five focus crops of this study represent agronomic and nutritional traits of interest for future-proofing the global food supply. To aid in future genomics-based investigations, we generated short-read HTS data for one accession of each of the five crops, which we used to identify nuclear genome SSR markers and to produce draft plastid genomes. In addition, we estimated the genome size of these and 12 other underutilised legume crops and wild relatives. This information is critical for undertaking successful whole-genome assemblies by enabling the amount of sequencing required to capture the entire genome at a given coverage depth to be estimated, and later serving as a benchmark to evaluate assembly completeness. Additionally, genome size may provide insights into crop resilience, as this trait has long-recognized impacts at the nuclear, cellular, and whole-plant level, which ultimately can play a role in influencing where and how plants grow (reviewed in Pellicer et al. 2018). We anticipate that the genomic resources presented here will be used to address diverse research questions. In addition, by using off-the-shelf, free-to-use bioinformatic tools, our pipeline should encourage other researchers to adopt this strategy for other underutilised species.
Materials and methods
Genome size measurements
Genome size (expressed as 1C-values, the DNA content in an un-replicated gametic nucleus) was measured for 31 individuals representing 17 legume taxa, using a one-step flow cytometry procedure. Approximately 1 cm2 fresh, mature leaf tissue was co-chopped in a petri dish with the internal standard Solanum lycopersicum L. “Stupiké polní rané”, 958.44 Mb/1C (Doležel et al. 1992) using a new razor blade in 1 ml of buffer. A further 1 ml of buffer was added to the sample and the contents gently mixed. Samples were prepared using General Purpose Buffer (GPB) (Loureiro et al. 2007), Galbraith’s buffer (Galbraith et al. 1983) or Cystain PI OxProtect (Sysmex UK Ltd), depending on the individual species to obtain the lowest coefficient of variation (CV) (Pellicer et al. 2021a) (see Supplementary Table 1). Polyvinyl pyrrolidone (PVP) and β-mercaptoethanol were also added to the GPB and Galbraith buffer as described (Pellicer et al. 2021b) to improve the quality of the flow histograms.
The sample was then passed through a 30 μm nylon filter, stained with 100 μl propidium iodide (1 mg/ml) and incubated on ice for 15 min. One sample was prepared from each individual and three replicates were run, recording up to 1000 nuclei per fluorescence peak using a Sysmex CyFlow Space (Sysmex Europe GmbH, Norderstedt, Germany) flow cytometer fitted with a 100 mW green solid state laser. The resulting histograms were analysed with the Windows™-based FlowMax software (v. 2.9 2014, Sysmex GmbH) and the average of each sample was used to estimate the genome size.
Nuclear genome assembly
Two accessions of the five species selected for genome sequencing were grown in the greenhouse at the University of Southampton, and one was selected for DNA extraction and sequencing (Table 1). DNA extraction was carried out using a modified CTAB method (Doyle and Doyle 1990); DNA quality and quantity were estimated using agarose gel electrophoresis and NanoDrop 2000 (Thermo Scientific, UK). Samples were treated with RNase and sent to Novogene (UK) for sequencing on a HiSeq X10. Using as references the genome sizes available in the Kew Plant DNA C-values database (Pellicer and Leitch 2020) and other literature, we aimed to achieve ≥ 10X coverage and therefore generated different amounts of data for each species depending on their genome size. The amounts varied from 6 Gb for jack bean (based on a genome size of another Canavalia species, C. rosea, estimated at 267 Mb/1C (Lin et al. 2021) to 11 Gb for the zombi pea (based on a previously recorded genome size of 587 Mb/1C for the species (Pellicer and Leitch 2020).
Raw sequencing data in native format (fastq) was quality checked using Trimmomatic (Bolger et al. 2014) and cleaned using the settings ILLUMINACLIP 2:30:10, LEADING:5, TRAILING:5, SLIDINGWINDOW:4:15. In Trimmomatic, we also excluded from consideration both unpaired reads and those shorter than 72 bp.
We performed nuclear genome assembly using ABySS ver. 2.0.2 (Jackman et al. 2017) and for all species a range of kmer options from 64 to 120 in steps of 8 were examined. For each species, we used the largest N50 value in combination with total assembly length to select a single assembly for SSR marker development, although we acknowledge that other metrics can be used to identify the ‘best’ assembly. However, given that we are aiming to generate a draft assembly for marker development, we feel this is appropriate and intuitive to implement across the five species. We assessed gene space coverage using BUSCO ver5.3.2 (Benchmarking Universal Single-Copy Orthologs; (Simao et al. 2015) to determine the proportion of eudicot conserved genes (eudicots_odb10 gene set) present in each of the five assemblies and whether these recovered loci were complete (in single or multiple copy), fragmented or missing.
Plastid genome assembly
Plastid genomes (plastomes) were assembled using NOVOplasty ver. 4.1 (Dierckxsens et al. 2016) based on 15 M paired-end reads. The plastid gene rbcL for each of the five species was downloaded from GenBank (Accession numbers U74238, MW960581, LC375226, MH391992 and KX087537) and used as the ‘seed’ sequence. Plastomes were annotated using GeSeq (Tillich et al. 2017) and visualised using OGDRAW (Greiner et al. 2019).
Marker identification
Simple sequence repeat (SSR) markers were exclusively identified from contigs > 500 bp in length, a cut-off used to avoid potential problems associated with primer design for SSRs in short contigs. Markers were identified using MISA (Thiel et al. 2003; http://pgrc.ipk-gatersleben.de/misa/) with a minimum of eight repeat units required to identify dinucleotide repeats, six for trinucleotide repeats, and four for tetra-, penta- and hexanucleotide repeats.
The quality of the identified markers was estimated by synthesising and testing 32 primer pairs on eight different accessions of the zombi pea, representing two individuals from four varietal groups (V. vexillata var. vexillata, var. angustifolia (Schumach. & Thonn.) Baker, var. ovata (E.Mey.) B.J.Pienaar and var. macrosperma Maréchal, Mascherpa & Stainier; Table 2). There were 12, 12 and 8 primers spanning dinucleotide, trinucleotide and tetranucleotide repeats, respectively. Loci for primer design were selected randomly from those with a moderate length repeat unit (20mer for di- and trinucleotide repeats and 8-10mer for tetranucleotide repeats). We followed previously published methods (Yang et al. 2018; Chapman 2019) for DNA extraction, PCR and genotyping.
Results
Genome size variation
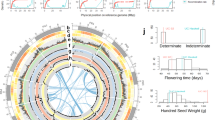
The genome size values for the 17 legume taxa ranged between 372 Mb/1C in Macrotyloma uniflorum and 802 Mb/1C in Vigna vexillata var. macrosperma, representing a ~ twofold variation (Fig. 1; Online Resource 1). These genomes were relatively small and narrowly distributed compared to 920 estimates for Fabaceae (mean = 2034 Mb/1C, ~ 98-fold variation) and 10,770 angiosperms (mean = 5020 Mb/1C, ~ 2440-fold variation). Apparent intraspecific variation was relatively low in the nine species for which we measured more than one individual, ranging from 0 (in Pachyrhizus erosus) to 151 Mb (in Vigna subterranea), although the number of replicates was also low (i.e., 2–8 individuals). The range in CV values across all estimates was 3.03–5.25%.
Genome size estimates of 31 legume accessions comprising 17 taxa, obtained using flow cytometry. For exact values see Supplementary Table 1. The five individuals selected for genome sequencing are shown in black. * This estimate should be treated as preliminary as it was impossible to obtain a clean extraction using GPB, Galbraith, or Cystain PI OxProtect
Nuclear and plastid genome assembly
We obtained between 26.2 and 46.5 M raw reads for each of the five individuals sequenced (Table 1). Raw reads have been deposited in the NCBI SRA under bioproject PRJNA882244. After quality control and trimming, ca. 97% of the reads were retained. We achieved the desired ≥ 10X coverage for Kersting’s groundnut (11.5X) and moth bean (14.9X), but lower coverage for jack bean and sword bean (ca. 6X), and zombi pea (8.4X).
After assembling the data using multiple kmer settings we aimed to select the assembly with the highest contig N50 and total assembly length. The assembly with the highest N50 was not necessarily the longest length assembly (Online Resource 2; Online Resource 3), hence we made a compromise. For jack bean and sword bean we used kmer = 64 and for the other species we used kmer = 80. For the five selected assemblies, the lowest N50 (less than 5 kb) was determined for the three species with the lowest coverage (jack bean, sword bean and zombi pea). For Kersting’s groundnut and moth bean the N50 was 15.5 and 11.5 kb respectively (Online Resource 2). Across the five species, there was a negative correlation between genome size and N50 (Pearson’s correlation ρ = − 0.890, P = 0.043). Based on the assembly length from ABySS and the genome size measurement from flow cytometry, we estimate that we assembled 53.0% to 68.4% of the (haploid) genome for the five species. Nevertheless, the BUSCO analysis based on 2326 conserved eudicot genes suggests that we sequenced 73–95% of the gene space in these species (Fig. 2). Therefore, while the assemblies are incomplete and fragmented, a large portion of the gene space is covered. Nearly half of the BUSCOs were complete and single copy in the two Canavalia species, and this rose to 70% in the zombi pea and > 85% in both Kersting’s groundnut and moth bean. Assemblies with the larger N50 had a greater proportion of complete BUSCO loci (Pearson’s correlation ρ = 0.888, P = 0.044).
Results of BUSCO (Benchmarking Universal Single-Copy Orthologs) analysis to assess gene space coverage. Genes identified in the five genomes were compared to 2326 conserved genes from the eudicots and are reported as complete—single copy, C(S); complete—duplicated, C(D); fragmented (F); or missing (M)
The plastid genomes assembled from these five species ranged in size from 149,321 bp (zombi pea) to 158,144 bp (jack bean). All exhibited the expected quadripartite structure (Online Resource 4).
Marker identification
We identified SSRs in each of the genome assemblies (after removing short contigs < 500 bp). The number of SSRs varied from 34,092 (moth bean) to 59,756 (sword bean) (Fig. 3). This equated to one SSR per 6.6, 6.8, 7.5, 9.1, 11.7 kb in Kersting’s groundnut, sword bean, jack bean, moth bean and zombi pea, respectively. Dinucleotide repeat SSRs predominated in four species (46–60% of SSRs) with the exception being zombi pea where trinucleotide repeat SSRs were most common (39% of SSRs). Penta- and hexanucleotide repeat SSRs were the least common summing to less than 10% of all SSRs across all five species (Fig. 3). To estimate the level of success in our primer design, we tested 32 markers in a panel of eight zombi pea accessions (Table 2). Of these, 17 amplified in more than half the individuals and all but one of these (i.e., 16) were polymorphic. The number of alleles at these 16 loci ranged from 2–6 (average 4.1).
Discussion
An important first step in understanding underutilised crops, which are often stress tolerant and/or nutritionally beneficial, is to quantify their genetic variation (Somta et al. 2011; Robotham and Chapman 2015; Dachapak et al. 2017; Minnaar-Ontong et al. 2021; Sserumaga et al. 2021), identify close relatives (Yang et al. 2018; Fisher et al. 2022), and ascertain variation within ‘inbred’ lines or varieties (Ho et al. 2016). Microsatellites (or SSRs) are molecular markers commonly used for this purpose, and their development is now relatively routine once genetic, genomic and/or transcriptomic resources are available (Hodel et al. 2016a; Chapman 2019). In this study we selected five underutilised legume crops with scant genomic resources and used low level genome sequencing (6–14X) to identify SSR markers that will be useful for both investigating the population genetic variation within these crops and for genetic mapping of quantitative traits.
We identified between ca. 34,000 and 60,000 SSRs per species. Our approach to assembling the genome was not designed to optimise the assembly (except that a range of kmers were used) and the number of reads used was relatively small. We therefore assume that some potential SSR markers could be from misassembled genome contigs and hence will not translate into usable markers. However, we do not believe this to be a significant problem; when we tested 32 SSR markers in eight zombi pea accessions, 16 amplified across at least half of the samples and were polymorphic. We also note that we did not optimise the PCR conditions, so further markers may amplify if we did this. Our results are similar to a previous study examining SSR design from de novo assembled genomes using small numbers of reads (similar to the approach employed here) which demonstrated that 14 of 18 SSR markers amplified in at least 50% (and on average 95%) of 12 individuals in a panel representing cultivated tomato and its wild relative, and all were polymorphic (Chapman 2019). Nevertheless, it should also be noted that even markers designed from whole genome assemblies and enriched for markers that should amplify across divergent accessions do not always amplify (e.g., Bhattarai et al. 2021).
The newly designed markers here should be of immediate use in their respective crops. Previous genetic mapping analyses of zombi pea and moth bean have used SSR markers from related legumes (Dachapak et al. 2018; Yundaeng et al. 2019). In both, hundreds of markers were screened because markers from the target species were not available. Of these, between 13.6 to 79.2% of markers amplified in moth bean (Yundaeng et al. 2019) and 19.2 to 51.4% in zombi pea (Dachapak et al. 2018). Whilst SSR markers from related species are clearly valuable, amplification success decreases as phylogenetic distance increases, and so having SSR markers developed from these focal species will hopefully be of immediate use. Population genetic variation in Kersting’s groundnut has recently been investigated using HTS-derived markers (DArT-Seq™; Kafoutchoni et al. 2021), but as far as we are aware, SSR markers have not been developed, and these are expected to be more cost-effective where a smaller number of markers would suffice. For jack bean and sword bean, population genetic variation has not been examined as far as we are aware; hence our work will potentially stimulate the investigation of, and ultimately investment in, these crops.
Whole genome sequence data, even when only assembled into partial drafts, can also provide valuable genetic information of use to researchers in addition to molecular markers such as SSRs. For example, we recently used a draft assembly of the perennial horse gram genome to identify gene orthologues across the legume family and test for positive selection (Fisher et al. 2022). This study revealed a number of gene ontology categories that appeared to be under positive selection in the family, as well as contributing towards identifying lineage-specific genes (Fisher et al. 2022).
Finally, the estimated genome sizes of several underutilised legume crops and related wild relatives provide useful information for researchers planning full genome sequencing of these taxa as the genome size is useful for estimating the amount of sequencing data needed to achieve the required coverage and hence the budget and time required for sequencing and assembling a genome (e.g., Li and Harkess 2018). Moreover, genome size has long-recognised impacts at the nuclear-, cellular- and organism-level that in turn plays a role in setting the thresholds on the plasticity of an organism’s functional traits independently of the information encoded in the DNA (Suda et al. 2015). These so-called “nucleotypic” effects are of considerable ecological significance as they influence where, when and how plants grow (reviewed in Greilhuber and Leitch 2013; Faizullah et al. 2021), and may therefore provide useful insights for crop plants concerning resilience and adaptive potential.
Going forward, we hope that by demonstrating the ease at which a partial and draft genome can be sequenced and assembled using publicly available scripts and limited bioinformatic experience, we encourage others to adopt similar practices for other species. Application of these methods to the species studied here, as well as other underutilised crops, could be an important step in rescuing and improving threated crop varieties, and to increasing the resilience of our food production systems In the face of climate change.
Data Availability
Raw reads have been deposited in the NCBI SRA under bioproject PRJNA882244. Assemblies and SSR lists are available from figshare (10.6084/m9.figshare.22006886) or from the corresponding author upon request. Genome sizes will be added to the next release of the Kew Plant DNA C-values database (https://cvalues.science.kew.org/).
References
Adsule RN (1996) Moth bean (Vigna aconitifolia (Jacq.) Maréchal). In: Nwokolo E, Smartt J (eds) Food and feed from legumes and oilseeds. Springer US, Boston, pp 203–205
Akpapunam MA, Sefa-Dedeh S (1997) Jack bean (Canavalia ensiformis): nutrition related aspects and needed nutrition research. Plant Foods Hum Nutr 50:93–99
Amkul K, Somta P, Laosatit K, Wang L (2020) Identification of QTLs for Domestication-Related Traits in Zombi Pea [Vigna vexillata (L.) A. Rich], a Lost Crop of Africa. Front Genet 11:803
Assogba P, Ewedje EEBK, Dansi A, Loko YL, Adjatin A, Dansi M, Sanni A (2016) Indigenous knowledge and agro-morphological evaluation of the minor crop Kersting’s groundnut (Macrotyloma geocarpum (Harms) Maréchal et Baudet) cultivars of Benin. Genet Resour Crop Evol 63:513–529
Ayenan MAT, Ezin VA (2016) Potential of Kersting’s groundnut [Macrotyloma geocarpum (Harms) Maréchal & Baudet] and prospects for its promotion. Agric Food Secur 5:10
Baath G, Northup B, Gowda P, Turner K, Rocateli A (2018) Mothbean: a potential summer crop for the southern great plains. Am J Plant Sci 9:1391–1402
Bhadkaria A, Narvekar DT, Gupta N, Khare A, Bhagyawant SS (2022) Moth bean (Vigna aconitifolia (Jacq.) Marechal) seeds: a review on nutritional properties and health benefits. Discov Food 2:18
Bhattarai G, Shi A, Kandel DR, Solís-Gracia N, da Silva JA, Avila CA (2021) Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions. Sci Rep 11:9999
Bolger A, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Campbell BM, Vermeulen SJ, Aggarwal PK, Corner-Dolloff C, Girvetz E, Loboguerrero AM, Ramirez-Villegas J, Rosenstock T, Sebastian L, Thornton PK et al (2016) Reducing risks to food security from climate change. Glob Food Sec 11:34–43
Chapman MA (2015) Transcriptome sequencing and marker development for four underutilized legumes. Appl Plant Sci 3:1400111
Chapman MA (2019) Optimizing depth and type of high-throughput sequencing data for microsatellite discovery. Appl Plant Sci 7:e11298
Chivenge P, Mabhaudhi T, Modi AT, Mafongoya P (2015) The potential role of neglected and underutilised crop species as future crops under water scarce conditions in Sub-Saharan Africa. Int J Environ Res Public Health 12:5685–5711
Dachapak S, Somta P, Poonchaivilaisak S, Yimram T, Srinives P (2017) Genetic diversity and structure of the zombi pea (Vigna vexillata (L.) A. Rich) gene pool based on SSR marker analysis. Genetica 145:189–200
Dachapak S, Tomooka N, Somta P, Naito K, Kaga A, Srinives P (2018) QTL analysis of domestication syndrome in zombi pea (Vigna vexillata), an underutilized legume crop. PLoS ONE 13:e0200116
Delgado-Salinas A, Thulin M, Pasquet R, Weeden N, Lavin M (2011) Vigna (Leguminosae) sensu lato: the names and identities of the American segregate genera. Am J Bot 98:1694–1715
Dierckxsens N, Mardulyn P, Smits G (2016) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45:e18–e18
Doležel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol Plant 85:625–631
Doss A, Pugalenthi M, Vadivel VG, Subhashini G, Anitha Subash R (2011) Effects of processing technique on the nutritional composition and antinutrients content of underutilized food legume Canavalia ensiformis L. DC. Int Food Res J 18:965–970
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ekanayake S, Jansz ER, Nair BM (2000) Literature review of an underutilized legume: Canavalia gladiata L. Plant Foods Hum Nutr 55:305–321
Faizullah L, Morton JA, Hersch-Green EI, Walczyk AM, Leitch AR, Leitch IJ (2021) Exploring environmental selection on genome size in angiosperms. Trends Plant Sci 26:1039–1049
FAO (2016) The state of food and agriculture: Climate change, agriculture and food security. Food and Agriculture Organization of the United Nations, Rome
FAO (2020) FAOSTAT database collections Food and Agriculture Organization of the United Nations. Italy, Rome
Fisher D, Reynolds I, Chapman MA (2022) The perennial horse gram (Macrotyloma axillare) genome, phylogeny, and selection across the Fabaceae. In: Chapman MA (ed) Underutilised crop genomes. Springer
Galbraith DW, Harkins KR, Maddox JM, Ayres NM, Sharma DP, Firoozabady E (1983) Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220:1049–1051
Garcia-Casal MN, Peña-Rosas JP, De-Regil LM, Gwirtz JA, Pasricha SR (2018) Fortification of maize flour with iron for controlling anaemia and iron deficiency in populations. Cochrane Database Syst Rev 12:Cd010187
Grassini P, Eskridge KM, Cassman KG (2013) Distinguishing between yield advances and yield plateaus in historical crop production trends. Nat Commun 4:2918
Greilhuber J, Leitch IJ (2013) Genome Size and the Phenotype. In: Greilhuber J, Dolezel J, Wendel JF (eds) Plant genome diversity volume 2: physical structure, behaviour and evolution of plant genomes. Springer, Vienna, pp 323–344
Greiner S, Lehwark P, Bock R (2019) OrganellarGenomeDRAW (OGDRAW) version 1.3.1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res 47:W59-w64
Haq N (2011) Underutilized food legumes: potential for multipurpose uses. CABI International, Cambridge
Ho WK, Muchugi A, Muthemba S, Kariba R, Mavenkeni BO, Hendre P, Song B, Deynze AV, Massawe F, Mayes S (2016) Use of microsatellite markers for the assessment of bambara groundnut breeding system and varietal purity before genome sequencing. Genome 59:427–431
Hodel RGJ, Gitzendanner MA, Germain-Aubrey CC, Liu XX, Crowl AA, Sun M, Landis JB, Segovia-Salcedo MC, Douglas NA, Chen SC et al (2016a) A new resource for the development of SSR markers: millions of loci from a thousand plant transcriptomes. Appl Plant Sci 4:1600024
Hodel RGJ, Segovia-Salcedo MC, Landis JB, Crowl AA, Sun M, Liu XX, Gitzendanner MA, Douglas NNA, Germain-Aubrey CC, Chen SC et al (2016b) The report of my death was an exaggeration: a review for researchers using microsatellites in the 21st century. Appl Plant Sci 4:1600025
Hunter D, Borelli T, Beltrame DMO, Oliveira CNS, Coradin L, Wasike VW, Wasilwa L, Mwai J, Manjella A, Samarasinghe GWL et al (2019) The potential of neglected and underutilized species for improving diets and nutrition. Planta 250:709–729
Jackman SD, Vandervalk BP, Mohamadi H, Chu J, Yeo S, Hammond SA, Jahesh G, Khan H, Coombe L, Warren RL et al (2017) ABySS 2.0: resource-efficient assembly of large genomes using a Bloom filter. Genome Res 27:768–777
Kafoutchoni KM, Agoyi EE, Agbahoungba S, Assogbadjo AE, Agbangla C (2021) Genetic diversity and population structure in a regional collection of Kersting’s groundnut (Macrotyloma geocarpum (Harms) Maréchal & Baudet). Genet Resour Crop Evol 68:3285–3300
Kajita T, Ohashi H, Tateishi Y, Bailey C, Doyle JJ (2001) rbcL and Legume Phylogeny, with particular reference to Phaseoleae, Millettieae, and Allies. Syst Bot 26:515–536
Karoli N, Sumari JO, Marealle H (2017) Utilization of jack beans (Canavalia ensiformis) for human consumption in Tanzania. Int J Agric Food Secur 3:39–49
Karuniawan A, Iswandi A, Kale PR, Heinzemann J, Grüneberg WJ (2006) Vigna vexillata (L.) A. Rich. Cultivated as a root crop in bali and timor. Genet Resour Crop Evol 53:213–217
Kebede E (2021) Contribution, utilization, and improvement of legumes-driven biological nitrogen fixation in agricultural systems. Front Sustain Food Syst 5:767998
Khoury CK, Bjorkman AD, Dempewolf H, Ramirez-Villegas J, Guarino L, Jarvis A, Rieseberg LH, Struik PC (2014) Increasing homogeneity in global food supplies and the implications for food security. Proc Natl Acad Sci USA 111:4001–4006
Khoury CK, Brush S, Costich DE, Curry HA, de Haan S, Engels JMM, Guarino L, Hoban S, Mercer KL, Miller AJ et al (2022) Crop genetic erosion: understanding and responding to loss of crop diversity. New Phytol 233:84–118
Li FW, Harkess A (2018) A guide to sequence your favorite plant genomes. Appl Plant Sci 6:e1030
Li X, Siddique KHM (2020) Future smart food: harnessing the potential of neglected and underutilized species for Zero Hunger. Matern Child Nutr 16(Suppl 3):e13008
Li X, Yadav R, Siddique KHM (2020) Neglected and underutilized crop species: the key to improving dietary diversity and fighting hunger and malnutrition in Asia and the Pacific. Front Nutr 7:593711–593711
Lin R, Zheng J, Pu L, Wang Z, Mei Q, Zhang M, Jian S (2021) Genome-wide identification and expression analysis of aquaporin family in Canavalia rosea and their roles in the adaptation to saline-alkaline soils and drought stress. BMC Plant Biol 21:333
Loureiro J, Rodriguez E, Doležel J, Santos C (2007) Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann Bot 100:875–888
Massawe F, Mayes S, Cheng A (2016) Crop diversity: an unexploited treasure trove for food security. Trends Plant Sci 21:365–368
Massawe FJ, Mayes S, Cheng A, Chai HH, Cleasby P, Symonds R, Ho WK, Siise A, Wong QN, Kendabie P et al (2015) The potential for underutilised crops to improve food security in the face of climate change. Procedia Environ Sci 29:140–141
Minnaar-Ontong A, Gerrano AS, Labuschagne MT (2021) Assessment of genetic diversity and structure of Bambara groundnut [Vigna subterranea (L.) verdc.] landraces in South Africa. Sci Rep 11:7408
Moteetee AN (2016) Canavalia (Phaseoleae, Fabaceae) species in South Africa: naturalised and indigenous. S Afr J Bot 103:6–16
Pellicer J, Leitch IJ (2020) The Plant DNA C-values database (release 7.1): an updated online repository of plant genome size data for comparative studies. New Phytol 226:301–305
Pellicer J, Hidalgo O, Dodsworth S, Leitch IJ (2018) Genome size diversity and its impact on the evolution of land plants. Genes (basel) 9:88
Pellicer J, Powell RF, Leitch IJ (2021a) The application of flow cytometry for estimating genome size, ploidy level endopolyploidy, and reproductive modes in plants. In: Besse P (ed) Molecular plant taxonomy methods in molecular biology. Humana, New York, pp 325–362
Pellicer J, Powell RF, Leitch IJ (2021b) The application of flow cytometry for estimating genome size, ploidy level endopolyploidy, and reproductive modes in plants. Methods Mol Biol 2222:325–361
Renard D, Tilman D (2019) National food production stabilized by crop diversity. Nature 571:257–260
Robotham O, Chapman MA (2015) Population genetic analysis of hyacinth bean (Lablab purpureus (L.) Sweet, Leguminosae) indicates an East African origin and variation in drought tolerance. Genet Resour Crop Evol 64:139–148
Sathyanarayana N, Pittala RK, Tripathi PK, Chopra R, Singh HR, Belamkar V, Bhardwaj PK, Doyle JJ, Egan AN (2017) Transcriptomic resources for the medicinal legume Mucuna pruriens: de novo transcriptome assembly, annotation, identification and validation of EST-SSR markers. BMC Genomics 18:409
Schmidhuber J, Tubiello FN (2007) Global food security under climate change. Proc Natl Acad Sci USA 104:19703–19708
Simao FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212
Singh D, Singh CK, Tribuvan KU, Tyagi P, Taunk J, Tomar RSS, Kumari S, Tripathi K, Kumar A, Gaikwad K et al (2020) Development, characterization, and cross species/genera transferability of novel EST-SSR Markers in Lentil, with their molecular applications. Plant Mol Biol Rep 38:114–129
Somta P, Chankaew S, Rungnoi O, Srinives P (2011) Genetic diversity of the Bambara groundnut (Vigna subterranea (L.) Verdc.) as assessed by SSR markers. Genome 54:898–910
Sserumaga JP, Kayondo SI, Kigozi A, Kiggundu M, Namazzi C, Walusimbi K, Bugeza J, Molly A, Mugerwa S (2021) Genome-wide diversity and structure variation among lablab Lablab purpureus (L.) sweet accessions and their implication in a Forage breeding program. Genet Resour Crop Evol 2997–3010
Stamp P, Messmer R, Walter A (2012) Competitive underutilized crops will depend on the state funding of breeding programmes: an opinion on the example of Europe. Plant Breed 131:461–464
Suda J, Meyerson LA, Leitch IJ, Pyšek P (2015) The hidden side of plant invasions: the role of genome size. New Phytol 205:994–1007
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422
Tillich M, Lehwark P, Pellizzer T, Ulbricht-Jones ES, Fischer A, Bock R, Greiner S (2017) GeSeq—versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45:W6–W11
Vadivel V, Doss A, Pugalenthi M (2010) Evaluation of nutritional value and protein quality of raw and differentially processed sword bean [Canavalia gladiata (Jacq.) DC.] seeds. Afr J Food Agric Nutr Dev 10:2850–2865
Vatanparast M, Shetty P, Chopra R, Doyle JJ, Sathyanarayana N, Egan AN (2016) Transcriptome sequencing and marker development in winged bean (Psophocarpus tetragonolobus; Leguminosae). Sci Rep 6:29070
Yang S, Grall A, Chapman MA (2018) Origin and diversification of winged bean (Psophocarpus tetragonolobus (L.) DC.; Fabaceae) a multi-purpose underutilised legume. Am J Bot 105:888–897
Yundaeng C, Somta P, Amkul K, Kongjaimun A, Kaga A, Tomooka N (2019) Construction of genetic linkage map and genome dissection of domestication-related traits of moth bean (Vigna aconitifolia), a legume crop of arid areas. Mol Genet Genomics 294:621–635
Acknowledgements
This work was supported by the use of the IRIDIS High Performance Computing Facility at the University of Southampton.
Funding
We are grateful to the University of Southampton for funding the sequencing which represents undergraduate projects for the first five authors. MAC was funded by the Natural Environment Research Council (NE/S002022/1) and MSG by a Future Leader Fellowship from Royal Botanic Gardens, Kew.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Online Resource 1 – Accessions and genome size estimates.
Online Resource 2 – Assembly metrics. N:500, number of contigs greater than 500 bp; N50, length (bp) of the shortest contig for which longer and equal length contigs cover at least 50% of the assembly; sum, total assembly length (Mb). Highlighted rows indicate the assemblies used for SSR identification.
Online Resource 3 – Assembly metrics for the five species across multiple kmers. The x-axis labels give the three-letter code for the species followed by the kmer length (Cen = Canavalia ensiformis; Cgl = C. gladiata; Mge = Macrotyloma geocarpum; Vac = Vigna aconitifolia; Vve = V. vexillata).
Online Resource 4 – Plastid genome assemblies of five underutilised legume species. Cen, Canavalia ensiformis (jack bean); Cgl, Canavalia gladiata (sword bean); Mge, Macrotyloma geocarpum (Kersting’s groundnut); Vac, Vigna aconitifolia (moth bean); and Vve, Vigna vexillata (zombi pea).
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diakostefani, A., Velissaris, R., Cvijanovic, E. et al. Genome resources for underutilised legume crops: genome sizes, genome skimming and marker development. Genet Resour Crop Evol 71, 427–438 (2024). https://doi.org/10.1007/s10722-023-01636-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01636-2