Abstract
RNA interference (RNAi) plays an important role in regulating gene expression in eukaryotes. Previously, we generated Arabidopsis and tobacco plants expressing double-stranded RNA (dsRNA) targeting a cotton bollworm (Helicoverpa armigera) P450 gene, CYP6AE14. Bollworms fed on transgenic dsCYP6AE14 plants showed suppressed CYP6AE14 expression and reduced growth on gossypol-containing diet (Mao et al., in Nat Biotechnol 25: 1307–1313, 2007). Here we report generation and analysis of dsRNA-expressing cotton (Gossypium hirsutum) plants. Bollworm larvae reared on T2 plants of the ds6-3 line exhibited drastically retarded growth, and the transgenic plants were less damaged by bollworms than the control. Quantitative reverse-transcription polymerase chain reaction (RT-PCR) showed that the CYP6AE14 expression level was reduced in the larvae as early as 4 h after feeding on the transgenic plants; accordingly, the CYP6AE14 protein level dropped. These results demonstrated that transgenic cotton plants expressing dsCYP6AE14 acquired enhanced resistance to cotton bollworms, and that RNAi technology can be used for engineering insect-proof cotton cultivar.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crop plants are widely cultivated and suffer from attacks by insect herbivores (Gassmann et al. 2009; Wittstock et al. 2004; Christou et al. 2006; Ferry et al. 2006). Cotton bollworm (Helicoverpa armigera) is one of the most destructive agricultural pests, causing severe yield loss in crop production. Host plants of this generalist lepidopteran include not only cotton but also other crop species. In the past three decades, transgenic technology has been developed to generate insect-proof plants to reduce yield loss and pesticide utilization (Bale et al. 2008; Kos et al. 2009). Engineering crop plants to express genes coding for insecticidal crystalline proteins from Bacillus thuringiensis (Bt), so-called Cry toxins, has achieved great success both economically and ecologically (Qaim and Zilberman 2003; Wu et al. 2008). However, with lasting cultivation of Bt crops, increasing insect resistance to transgenic crops and outbreaks of nontarget pests were reported (Bravo and Soberon 2008; Gahan et al. 2001; Tabashnik et al. 2008; Lu et al. 2010), which calls for new approaches.
Since the discovery that ingested dsRNA could trigger RNAi in the nematode (Caenorhabditis elegans) (Fire et al. 1998), this gene-silencing approach has become a valuable tool in functional genomics (Cronin et al. 2009; Wesley et al. 2001). Injection of siCoo2-RNA into pea aphids led to Coo2 suppression and caused significant mortality of insects fed on host plant (Mutti et al. 2008). When the diamondback moth, Plutella xylostella, was fed synthetic dsRNA against CYP6BG1, expression of the P450 gene was suppressed, resulting in reduced larval resistance to the insecticide permethrin (Bautista et al. 2009). It has been reported that plants could be armed with dsRNA to fend off insect pests, as transgenic plants producing dsRNAs against selected insect genes showed suppressive effects on cotton bollworm (Mao et al. 2007) and western corn rootworm (Baum et al. 2007) gene expression. Because the introduced dsRNA can be highly specific to target insects, this approach limits the adverse effects on nontarget organisms. RNAi-based gene regulation has been reported in different insect orders, including Lepidoptera, Hemiptera, Coleoptera, Diptera, and Hymenoptera (Huvenne and Smagghe 2010), which makes it possible to develop RNAi technology for control of various insect pests (Gordon and Waterhouse 2007).
Many plant secondary metabolites are toxic to or repel insects, enabling host plants to escape from insect herbivores (Gatehouse 2002). To counteract plant defenses, insects have developed adaptive mechanisms, which often involve a set of genes whose products metabolize the chemicals from plants (Wittstock et al. 2004). Most cotton cultivars accumulate gossypol and related sesquiterpene aldehydes in both aerial tissues and roots, and these phytoalexins form a chemical arsenal against herbivorous (Meng et al. 1999; Tan et al. 2000; Du et al. 2004; Stipanovic et al. 2006). Previously, we isolated a P450 monooxygenase gene, CYP6AE14, from Helicoverpa armigera; expression of CYP6AE14 was induced by gossypol, and its expression level was correlated with larval growth when gossypol was present in the diet. When bollworms were fed on transgenic Arabidopsis plants producing dsRNA against CYP6AE14 (dsCYP6AE14), expression of CYP6AE14 was suppressed; after transferring to a gossypol-containing diet, the larvae showed decreased tolerance to gossypol (Mao et al. 2007). Therefore, if cotton plants are engineered to express dsCYP6AE14, they may be better protected from bollworms due to the presence of gossypol in cotton plants. Here we report the generation of transgenic dsCYP6AE14 cotton plants, which indeed acquired enhanced resistance to cotton bollworms.
Materials and methods
Plant and insect culture
Cotton plants (Gossypium hirsutum cv. R15) were grown in greenhouse under 28–30°C, 60–80% relative humidity. Young leaves with the same condition and developing bolls at about 9 days post anthesis (9 DPA) were used for insect feeding tests.
Cotton bollworm (Helicoverpa armigera) eggs were obtained from Nanjing Agricultural University and reared as previously described (Peng et al. 2005). For each feeding experiment, synchronous larvae were selected, weighed individually, and divided into groups; each group contained 20–30 individuals. After feeding on different diets for indicated days, larvae were weighed, and midguts were taken for further analysis. Statistical data analysis was performed using Student′s t-test in Excel software.
Construction of vectors and cotton plant transformation
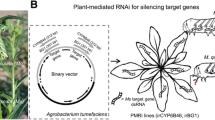
The dsRNA construct pBI121-dsCYP6AE14 as described by Mao et al. (2007) contained a 35S promoter, a sense fragment of CYP6AE14 complementary DNA (cDNA) from +472 to +940, a 120-nucleotide intron of Arabidopsis RTM1 gene (Johansen and Carrington 2001), the CYP6AE14 fragment in antisense orientation, and a NOS terminator (Fig. 1a). The CYP6AE14 fragment (469 bp) was obtained by PCR amplification of H. armigera cDNA clones with primers GIPF (5′-GAAGATTTTCTCGATAAGGAAG-3′) and GIPR (5′-ATATAAAGCACTGTGCCACTAAG-3′). Binary vectors harboring the desired construct were transferred into Agrobacterium tumefaciens strain LBA4404 by electroporation, followed by transformation of cotyledon and hypocotyl explants from Gossypium hirsutum cv. R15 (Li et al. 2002). After the stages of callus induction, proliferation, embryogenic callus induction, embryo differentiation, and finally plantlet regeneration, the plantlets were transferred to pots in greenhouse for further growth. Transgenic plants were screened by kanamycin selection and further confirmed by PCR for presence of the neomycin phosphotransferase II (NptII) gene and RNA gel blot for presence of the dsCYP6AE14 transcripts.
Effect of T1 transgenic cotton on larvae growth. a The dsRNA construct pBI121-dsCYP6AE14 contained a 35S promoter, a sense fragment of CYP6AE14 cDNA from +472 to +940, a 120-nucleotide intron of Arabidopsis RTM1 gene (Johansen and Carrington 2001), the CYP6AE14 fragment in antisense orientation, and a NOS terminator. b northern blot detection of dsRNA homologues to CYP6AE14 in the leaves of transgenic (ds1-ds6) and nontransgenic control (R15) plants. c, d Net weight increase of larvae reared on leaves of T1 transgenic cotton plants. Third-instar larvae previously grown on artificial diet were transferred to nontransgenic (R15) or T1 transgenic cotton plant leaves for 4 days, respectively. Values are means ± standard deviation (SD). *P < 0.05; **P < 0.01
T1 seeds were germinated and rescreened to determine transgenic lines using PCR assay (NptII gene detection), and RNA and DNA gel blot assays. Transgenic plants were denoted using the format ds6-3, where “ds” represents a positive transgenic line, “6” is an independent primary transformant, and “3” is the number of the T1 progeny of that plant.
RNA analysis
Total RNAs were isolated from H. armigera by Trizol reagent (Invitrogen, Carlsbad, CA) or from cotton as described (Wu et al. 2002). The RNAs were separated on 1.0% denaturing agarose gel and transferred to Hybond-N+ filter membrane (Amersham Pharmacia Biotech, Uppsala). For small RNAs, the RNA samples, 40 μg per lane, were loaded on a TBE-urea gel (15%); after electrophoresis, they were electroblotted onto the Hybond-N+ membrane. The membranes were ultraviolet (UV) cross-linked and hybridized with ExpressHyb solutions (Clontech, Palo Alto, CA). Probes were obtained by PCR using primers as described for vector construction. The probes were randomly labeled with 32P-dCTP using the Prime-a-Gene labeling system (Promega, Madison, WI). For RT-PCR, first-strand cDNA was prepared using the ReverTra Ace kit (TOYOBO, Osaka). Real-time RT-PCR (qRT-PCR) was performed on a Bio-Rad iCycler with iQ SYBR Green Supermix (Bio-Rad), following a two-step protocol: 95°C for 3 min, 40 cycles of denaturation at 95°C for 20 s and annealing/extension at 60°C for 20 s.
DNA analysis
Genomic DNA was isolated as described (Benbouza et al. 2006). Transgenic lines were determined by PCR detection of presence of NptII. The NptII gene was detected using primers kanF: 5′-GGCGATACCGTAAAGCACGAGGAA-3′ and kanR: 5′-GCTATGACTGGGCACAACAGACAAT-3′, respectively, generating a 680-bp fragment.
Genomic DNA (20 µg) was digested with indicated enzyme for 16 h, separated on 0.8% agarose gel, and transferred onto Hybond N1 membrane (Amersham Pharmacia Biotech, Uppsala). DNA gel blot analysis of G. hirsutum cv. R15 and transgenic cottons was carried out using probes that were obtained by PCR using primers as described for vector construction.
Protein analysis
The rabbit antiserum against a CYP6AE14 fragment (150–311 amino acid residues) was raised, and the antibody was purified by binding with Protein A-Sepharose CL6B (Sigma, St. Louis, MO), followed by selective elution of IgG with 50 mM glycine, pH 3.0, 0.5 mM NaCl, neutralized with 1 M Tris/HCl to pH 7.0, and used at 1:500 dilution.
Total proteins of the midgut of H. armigera were extracted and loaded onto a 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) gel (20 µg proteins per lane). After electrophoresis, the proteins were electrotransferred to a Hybond-C membrane (Amersham, Buckinghamshire, UK). Blots were incubated with the primary antibody for 1 h, than incubated with horseradish peroxidase (HRP)-conjugated anti-rabbit antiserum as the secondary antibody for 45 min. The blot was developed using enhanced chemiluminescence (ECL) detection solution (Tiangen, China) and exposed to X-ray films (Kodak, Japan).
Analysis of sesquiterpene aldehydes
Total sesquiterpene aldehydes were quantitated with a phloroglucinol/HCl assay (Liu et al. 1999). A total of 500 mg sample was immersed in liquid nitrogen, ground to a fine powder, and extracted with 70% acetone for 30 min. After centrifugation, an equal volume of reagent (1% phloroglucinol, 2 N HCl in 95% ethanol) was added to the acetone extract, followed by incubation at 55°C for 5 min. Absorbance at 555 nm was measured immediately. Standard curve was prepared with gossypol (Sigma).
Results
Generation of transgenic cotton plants producing dsCYP6AE14
To examine if RNAi technology could be applied directly to cotton plants for enhanced insect resistance, we transferred the dsCYP6AE14 construct into cotton plants (Gossypium hirsutum cv. R15). Twenty-eight primary (T0) transgenic cotton lines were obtained. Transcripts of dsCYP6AE14 in T1 generation plants were examined by northern blotting. Of the 20 individual plants from five T1 lines, at least 6 (ds1-2, ds2-3, ds3-3, ds4-1, ds4-3, and ds6-3) showed relatively high expression level of the dsRNA (Fig. 1a). When placed on the control or the transgenic dsCYP6AE14 cotton leaves for 4 days, second-instar larvae exhibited retarded growth on dsCYP6AE14 leaves. Among these lines, ds6-3 exhibited the most obvious adverse effect on larval growth (Fig. 1b, c). T2 generation plants of the ds6-3 line were then generated. In a PCR assay, all of the T2 plants showed positive signals of NptII, a marker gene used for screening transgenic plants (Supplementary Fig. 1). RNA blot showed that the dsCYP6AE14 transcripts were present in all of the T2 plants examined (Fig. 2a). Southern blot indicated that there were two copies of dsCYP6AE14 in ds6-3 and the T2 plants (Fig. 2b). The ds6-3 line was then used for further investigation.
northern and Southern blots of dsCYP6AE14 in T1 and T2 lines of ds6-3. a northern blot to detect dsRNA of CYP6AE14 in leaves of T2 plants of ds6-3. b Southern blot assay to determine copies of CYP6AE14 in T1 and T2 plants of ds6-3. Genomic DNA of ds6-3 and three individuals of T2 generation (ds6-3-2, ds6-3-3, and ds6-3-4) were digested by enzymes as indicated
CYP6AE14 expression in bollworms was suppressed by dsCYP6AE14 cotton
To see if the RNAi signal could be transmitted into bollworm midgut cells during ingestion of the transgenic cotton tissues, second-instar larvae, previously reared on artificial diets, were transferred to leaves of the T2 ds6-3 plants, and RNA gel blotting was performed to detect the small RNA (sRNA) fragments of CYP6AE14. The sRNAs of CYP6AE14 were detected in midgut of the bollworms at day 3 post transferring to the transgenic leaf (Fig. 3a). CYP6AE14 gene expression was then examined by both qRT-PCR and RNA blot at different intervals. The data show that CYP6AE14 expression level was decreased as early as 4 h post transferring, and the suppression was evident at 8 and 16 h (Fig. 3b, c). Consistent with the gene expression level decrease, the CYP6AE14 protein level drastically dropped in larvae reared on T2 ds6-3 leaves for 3 days (Fig. 3d). These data demonstrate that CYP6AE14 expression in the cotton bollworm midgut could be suppressed by transgenic cotton leaves expressing dsCYP6AE14.
Suppression of CYP6AE14 expression in larvae fed on ds6-3 T2 plants. a northern blot of the small RNAs of CYP6AE14 in midgut of third-instar larvae fed on nontransgenic control R15 (lane 2) and ds6-3 T2 plants (lane 3) for 3 days. Lane 1: total RNAs from ds6-3 T2 plants as positive control. b and c northern blot (b) and qRT-PCR (c) analysis of CYP6AE14 transcripts in midgut of second-instar larvae fed on control (R15) or ds6-3 T2 (ds) plants for indicated time. d western blot detection of CYP6AE14 proteins in midgut of second-instar larvae fed on nontransgenic control R15 or ds6-3 T2 plants for 3 days. Values are means ± SD
Transgenic dsCYP6AE14 cotton plants showed enhanced protection from bollworms
To detect the effects of T2 plants on bollworm growth, second-instar larvae reared on leaves of the wild-type and T2 ds6-3 plants were weighed, respectively. Larvae on the ds6-3 plant leaves exhibited obvious growth retardation, with only 60% weight increase in comparison with those on the wild-type leaves, after assaying for 4 days; growth retardation became more evident when the assay was extended to 6 and 10 days (Fig. 4a, b). It was reported that high concentrations of gossypol in diets would inhibit bollworm growth (Mao et al. 2007). We then measured gossypol equivalents in the wild-type and the transgenic cotton leaves. Quantitative analysis of gossypol with a phloroglucinol/HCl assay (Liu et al. 1999) showed that gossypol equivalents was slightly lower in the ds6-3 than in the wild-type leaves, but the difference was not significant (Fig. 4c). Therefore, different growth rates of the larvae on the control and transgenic ds6-3 leaves were not due to variations of gossypol content. Because the P450 enzyme of CYP6AE14 plays a key role in the bollworm response to gossypol-containing diet (Mao et al. 2007), we assumed that downregulation of CYP6AE14 would impair gossypol detoxification in bollworms, thus resulting in accumulation of higher concentrations of gossypol in the larvae midgut. We found that gossypol equivalent was indeed higher (about 1.4-fold) in larvae fed on the ds6-3 leaves compared with those fed on the wild-type leaves (Fig. 4d).
Effect of ds6-3 T2 plants on larvae growth. a Net weight increase of larvae fed on leaves of nontransgenic control R15 (blue) or ds6-3 T2 (ds) plants (red) for indicated days. b Images of larvae that were fed on leaves of nontransgenic control R15 or ds6-3 T2 plants for 4 and 6 days, respectively. c Gossypol equivalents in leaves of nontransgenic control R15 or ds6-3 T2 plants. d Gossypol equivalents in midgut of the larvae fed on leaves of nontransgenic control R15 and ds6-3 T2 plants, respectively, for 6 days. Values are means ± SD. *P < 0.05; **P < 0.01
We have demonstrated that bollworms from the dsCYP6AE14 cotton had lower growth rates and higher gossypol equivalents in midgut. Retarded growth could be caused by antifeedant effect of gossypol, resulting in less consumption of cotton tissues by the larvae. Alternatively, the consumption was not changed but the higher gossypol absorption inhibited bollworm growth. Leaf consumption assay was thus performed. Second-instar larvae were transferred to ds6-3 leaves; during the first 3 days of the assay, ingestion of the ds6-3 leaves was reduced by 21% (Fig. 5a). When measurement of larval leaf intake was extended to another 3-day period (days 4–6), a greater difference (44%) was observed (Fig. 5b).
Transgenic cotton plants were less damaged by bollworms than the control. Second-instar larvae were divided into 2 groups. Each group contained 40 individual larvae and were fed on control (R15) and ds6-3 T2 (ds) leaves with similar conditions. a Consumption of leaves of control and ds6-3 T2 plants by second-instar larvae for the first 3 days. b Leaf consumption from the 4th to 6th day was recorded. Values are means ± SD. *P < 0.05. c Image of larvae on cotton boll. Larvae previously reared on leaves of nontransgenic control R15 or ds6-3 T2 plants for 10 days were transferred to cotton boll for another day
Damage of bolls by bollworms causes severe quality and production losses of cotton. To test the protection of bolls by the RNAi transgenic cotton, larvae that previously lived on leaves from the wild-type and ds6-3 leaves for 10 days were then transferred to bolls (about 9 days post anthesis, DPA). We found that, after 1 day, the larvae from the wild-type leaves burrowed into bolls and consumed the contents, whereas the larvae from the ds6-3 leaves chewed only shallow gouges in the boll surface, whereas the boll contents were almost intact (Fig. 5c). Together, these data indicate that the transgenic dsCYP6AE14 cotton plants were less damaged by cotton bollworms in comparison with the untransformed control.
Discussion
RNAi-triggered gene suppression through uptake of dsRNAs has been reported for many insect species, including those of Coleoptera, Diptera, Hemiptera, Hymenoptera, Isoptera, Lepidoptera, and Orthoptera (Huvenne and Smagghe 2010). It was shown that bacteria could be used to express dsRNA; after feeding a lepidopteran pest (Spodoptera exigua) with a bacteria-mixed diet, the target gene was silenced (Tian et al. 2009). To counteract plant nematodes, viruses were modified for dsRNA production in plant. RNAi was then observed in nematodes reared on the virus-infected plants (Valentine et al. 2007; Dubreuil et al. 2009). For sucking insects, RNA interference was successfully achieved by feeding insects with sugar solutions containing dsRNA synthesized in vitro (Bautista et al. 2009; Walshe et al. 2009; Zhou et al. 2008). Each of these approaches, however, suffers from certain limitations, such as low uptake efficiency, high cost, and inapplicability for field control of pests. Bacteria-based RNAi can be a convenient platform for functional genomics of some insect species, but this technology cannot produce plant with improved insect resistance. Viruses are highly efficient in triggering dsRNA production in plant, but the dsRNA production is not stable in offspring, and the heterogeneity in RNAi efficiency between virus-infected plants may limit its utilization. In this investigation, the introduced dsRNA of CYP6AE14 was stably expressed not only in T1 but also in T2 generation, and enhanced resistance to bollworms was observed in both generations, suggesting that transgenic plant-based RNAi may be a feasible strategy for generating insect-proof plants.
Bt crops have been grown worldwide and offer a high degree of protection (Qaim and Zilberman 2003; Wu et al. 2008). At present, Bt-based strategies seem more effective than RNAi-based technology for agricultural pest control; however, recent reports of resistance to Bt toxins existing in field populations of insects (Bravo and Soberon 2008; Gahan et al. 2001; Tabashnik et al. 2008) call for new approaches. Utilization of RNAi in pest control offers an alternative. On the other hand, Bt insecticide proteins have no or little effects on sap-sucking homopteran pests such as aphids, leafhoppers, and whitefly (Price and Gatehouse 2008). Since gene suppression by ingested dsRNA has been reported in aphids (Whyard et al. 2009), and RNAi knockdown of a salivary transcript led to lethality in the pea aphid (Mutti et al. 2006, 2008), RNAi-based technology also may be a promising approach for protection of plants from sucking pests.
Growth inhibition of cotton bollworms due to ingestion of dsCYP6AE14-expressing plants occurred only when gossypol was present in diet, demonstrating that genes involved in detoxification or defense reactions against plant secondary metabolites can serve as targets of pest control. This has the advantage of controlling specific pests that feed on a crop species producing a defined group of defensive compounds, minimizing effects on nontarget insects.
Insect P450 monooxygenases play a central role in adaptation to plant defense products (Scott et al. 1998; Zeng et al. 2009). The catalytic activities of most eukaryotic P450 monooxygenases rely on cytochrome P450 reductase, cytochrome B5, and cytochrome B5 reductase as partners (Feyereisen 1999; Yang et al. 2010). We noticed that the dsCYP6AE14 cotton plant did have deleterious effects on bollworms, but was not lethal. If multiple genes involved in the P450 complex were targeted by RNAi, the deleterious effects would be magnified. As a recently developed genetic tool, further studies are needed to optimize RNAi-based strategies for crop protection.
References
Bale JS, van Lenteren JC, Bigler F (2008) Biological control and sustainable food production. Philos Trans R Soc Lond B Biol Sci 363:761–776
Baum JA, Bogaert T, Clinton W, Heck GR, Feldmann P, Ilagan O, Johnson S, Plaetinck G, Munyikwa T, Pleau M et al (2007) Control of coleopteran insect pests through RNA interference. Nat Biotechnol 25:1322–1326
Bautista MA, Miyata T, Miura K, Tanaka T (2009) RNA interference-mediated knockdown of a cytochrome P450, CYP6BG1, from the diamondback moth, Plutella xylostella, reduces larval resistance to permethrin. Insect Biochem Mol Biol 39:38–46
Benbouza H, Baudoin J-P, Mergeai G (2006) Improvement of the genomic DNA extraction method with CTAB for cotton leaves. Biotechnologie Agronomie Societe et Environ 10:73–76
Bravo A, Soberon M (2008) How to cope with insect resistance to Bt toxins? Trends Biotechnol 26:573–579
Christou P, Capell T, Kohli A, Gatehouse JA, Gatehouse AM (2006) Recent developments and future prospects in insect pest control in transgenic crops. Trends Plant Sci 11:302–308
Cronin SJ, Nehme NT, Limmer S, Liegeois S, Pospisilik JA, Schramek D, Leibbrandt A, Simoes Rde M, Gruber S, Puc U et al (2009) Genome-wide RNAi screen identifies genes involved in intestinal pathogenic bacterial infection. Science 325:340–343
Du L, Ge F, Zhu S, Parajulee MN (2004) Effect of cotton cultivar on development and reproduction of Aphis gossypii (Homoptera: Aphididae) and its predator Propylaea japonica (Coleoptera: Coccinellidae). J Econ Entomol 97:1278–1283
Dubreuil G, Magliano M, Dubrana MP, Lozano J, Lecomte P, Favery B, Abad P, Rosso MN (2009) Tobacco rattle virus mediates gene silencing in a plant parasitic root-knot nematode. J Exp Bot 60:4041–4050
Ferry N, Edwards MG, Gatehouse J, Capell T, Christou P, Gatehouse AM (2006) Transgenic plants for insect pest control: a forward looking scientific perspective. Transgenic Res 15:13–19
Feyereisen R (1999) Insect P450 enzymes. Annu Rev Entomol 44:507–533
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Gahan LJ, Gould F, Heckel DG (2001) Identification of a gene associated with Bt resistance in Heliothis virescens. Science 293:857–860
Gassmann AJ, Onstad DW, Pittendrigh BR (2009) Evolutionary analysis of herbivorous insects in natural and agricultural environments. Pest Manag Sci 65:1174–1181
Gatehouse JA (2002) Plant resistance towards insect herbivores: a dynamic interaction. New Phytol 156:145–169
Gordon KH, Waterhouse PM (2007) RNAi for insect-proof plants. Nat Biotechnol 25:1231–1232
Huvenne H, Smagghe G (2010) Mechanisms of dsRNA uptake in insects and potential of RNAi for pest control: a review. J Insect Physiol 56:227–235
Johansen LK, Carrington JC (2001) Silencing on the spot. Induction and suppression of RNA silencing in the Agrobacterium-mediated transient expression system. Plant Physiol 126:930–938
Kos M, van Loon JJ, Dicke M, Vet LE (2009) Transgenic plants as vital components of integrated pest management. Trends Biotechnol 27:621–627
Li XB, Cai L, Cheng NH, Liu JW (2002) Molecular characterization of the cotton GhTUB1 gene that is preferentially expressed in fibre. Plant Physiol 130:666–674
Liu CJ, Heinstein P, Chen XY (1999) Expression pattern of genes encoding farnesyl diphosphate synthase and sesquiterpene cyclase in cotton suspension-cultured cells treated with fungal elicitors. Mol Plant Microbe Interact 12:1095–1104
Lu Y, Wu K, Jiang Y, Xia B, Li P, Feng H, Wyckhuys KA, Guo Y (2010) Mirid bug outbreaks in multiple crops correlated with wide-scale adoption of Bt cotton in China. Science 328:1151–1154
Mao YB, Cai WJ, Wang JW, Hong GJ, Tao XY, Wang LJ, Huang YP, Chen XY (2007) Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat Biotechnol 25:1307–1313
Meng YL, Jia JW, Liu CJ, Liang WQ, Heinstein P, Chen XY (1999) Coordinated accumulation of (+)-delta-cadinene synthase mRNAs and gossypol in developing seeds of Gossypium hirsutum and a new member of the cad1 family from G. arboreum. J Nat Prod 62:248–252
Mutti NS, Park Y, Reese JC, Reeck GR (2006) RNAi knockdown of a salivary transcript leading to lethality in the pea aphid, Acyrthosiphon pisum. J Insect Sci 6:1–7
Mutti NS, Louis J, Pappan LK, Pappan K, Begum K, Chen MS, Park Y, Dittmer N, Marshall J, Reese JC, Reeck GR (2008) A protein from the salivary glands of the pea aphid, Acyrthosiphon pisum, is essential in feeding on a host plant. Proc Natl Acad Sci USA 105:9965–9969
Peng JY, Li ZH, Xiang H, Huang JH, Jia SH, Miao XX, Huang YP (2005) Preliminary studies on differential defense responses induced during plant communication. Cell Res 15:187–192
Price DR, Gatehouse JA (2008) RNAi-mediated crop protection against insects. Trends Biotechnol 26:393–400
Qaim M, Zilberman D (2003) Yield effects of genetically modified crops in developing countries. Science 299:900–902
Scott JG, Liu N, Wen Z (1998) Insect cytochromes P450: diversity, insecticide resistance and tolerance to plant toxins. Comp Biochem Physiol C Pharmacol Toxicol Endocrinol 121:147–155
Stipanovic RD, Lopez JD Jr, Dowd MK, Puckhaber LS, Duke SE (2006) Effect of racemic and (+)- and (−)-gossypol on the survival and development of Helicoverpa zea larvae. J Chem Ecol 32:959–968
Tabashnik BE, Gassmann AJ, Crowder DW, Carriere Y (2008) Insect resistance to Bt crops: evidence versus theory. Nat Biotechnol 26:199–202
Tan XP, Liang WQ, Liu CJ, Luo P, Heinstein P, Chen XY (2000) Expression pattern of (+)-delta-cadinene synthase genes and biosynthesis of sesquiterpene aldehydes in plants of Gossypium arboreum L. Planta 210:644–651
Tian H, Peng H, Yao Q, Chen H, Xie Q, Tang B, Zhang W (2009) Developmental control of a lepidopteran pest Spodoptera exigua by ingestion of bacteria expressing dsRNA of a non-midgut gene. PLoS One 4:e6225
Valentine TA, Randall E, Wypijewski K, Chapman S, Jones J, Oparka KJ (2007) Delivery of macromolecules to plant parasitic nematodes using a tobacco rattle virus vector. Plant Biotechnol J 5:827–834
Walshe DP, Lehane SM, Lehane MJ, Haines LR (2009) Prolonged gene knockdown in the tsetse fly Glossina by feeding double stranded RNA. Insect Mol Biol 18:11–19
Wesley SV, Helliwell CA, Smith NA, Wang MB, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA et al (2001) Construct design for efficient, effective and high-throughput gene silencing in plants. Plant J 27:581–590
Whyard S, Singh AD, Wong S (2009) Ingested double-stranded RNAs can act as species-specific insecticides. Insect Biochem Mol Biol 39:824–832
Wittstock U, Agerbirk N, Stauber EJ, Olsen CE, Hippler M, Mitchell-Olds T, Gershenzon J, Vogel H (2004) Successful herbivore attack due to metabolic diversion of a plant chemical defense. Proc Natl Acad Sci USA 101:4859–4864
Wu YR, Llewellyn DJ, Dennis ES (2002) A quick and easy method for isolating good-quality RNA from cotton (Gossypium hirsutum L.) tissues. Plant Molecular Biology Reporter 20:213–218
Wu KM, Lu YH, Feng HQ, Jiang YY, Zhao JZ (2008) Suppression of cotton bollworm in multiple crops in China in areas with Bt toxin-containing cotton. Science 321:1676–1678
Yang CQ, Lu S, Mao YB, Wang LJ, Chen XY (2010) Characterization of two NADPH: cytochrome P450 reductases from cotton (Gossypium hirsutum). Phytochemistry 71:27–35
Zeng RS, Wen Z, Niu G, Schuler MA, Berenbaum MR (2009) Enhanced toxicity and induction of cytochrome P450 s suggest a cost of “eavesdropping” in a multitrophic interaction. J Chem Ecol 35:526–532
Zhou X, Wheeler MM, Oi FM, Scharf ME (2008) RNA interference in the termite Reticulitermes flavipes through ingestion of double-stranded RNA. Insect Biochem Mol Biol 38:805–815
Acknowledgments
This research was supported by the National Science Foundation of China (30870380), the National High-Tech Research Program of China (2009AA10Z112), and the Ministry of Agriculture of China (2008ZX08009-001).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Mao, YB., Tao, XY., Xue, XY. et al. Cotton plants expressing CYP6AE14 double-stranded RNA show enhanced resistance to bollworms. Transgenic Res 20, 665–673 (2011). https://doi.org/10.1007/s11248-010-9450-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-010-9450-1