Abstract
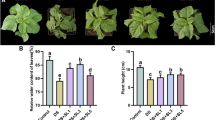
High salinity is one of the most common abiotic stresses that plants have to confront, and selecting salt-tolerant plants is becoming a very important task in breeding. In this study, non-transgenic (NT) and three LbDREB-transgenic Populus ussuriensis Kom. (Dr22, Dr8, and Dr2) lines were treated with 150 mM NaCl. The salt tolerance of NT, Dr2, Dr8, and Dr22 were evaluated and compared. Phenotypic observation, chlorophyll content detection and leaf salt injury index calculation suggested NT exhibited more severe salt injury symptoms and weaker photosynthetic capacity than Dr2, Dr8, and Dr22. LbDREB gene exhibited opposite temporal expression patterns with no apical meristem (NAM), trihelix transcription factor (GT-1) and WRKY transcription factor 70 (WRKY70), and consistent expression patterns with Cu–Zn superoxide dismutase and peroxidase-PO1 genes, in Dr2, Dr8, and Dr22 under salt stress. The superoxide dismutase and peroxidase activities of transgenic lines were higher than NT plants during the salt stress treatment. Malonaldehyde accumulation in NT leaves was more significant than in transgenic leaves, while proline accumulation in NT leaves was much less than that in transgenic leaves under salt stress. The root to shoot ratios were significantly increased in Dr2, Dr8, and Dr22 plants, but were slightly decreased in NT plants, 12 days after salt stress treatment. The relative water content of NT leaves decreased more than transgenic leaves, meanwhile, the relative electrolytic leakage of NT leaves increased more significantly than transgenic leaves after salt stress. In summary, Dr2, Dr8, and Dr22 exhibited stronger salt stress tolerance than NT. The transgenic lines Dr2, Dr8, and Dr22 can serve as alternative salt-tolerant germplasm resources in P. ussuriensis breeding.








Similar content being viewed by others
References
Abogadallah GM (2010) Antioxidative defense under salt stress. Plant Signal Behav 4(5):369–374
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25(12):1263–1274
Agarwal P, Dabi M, Sapara KK, Joshi PS, Agarwal PK (2016) Ectopic expression of JcWRKY transcription factor confers salinity tolerance via salicylic acid signaling. Front Plant Sci 7:1541. doi:10.3389/fpls.2016.01541
Ahmad P, Abdel LA, Hashem A, Abd AE, Gucel S, Tran LS (2016) Nitric oxide mitigates salt stress by regulating levels of osmolytes and antioxidant enzymes in chickpea. Front Plant Sci 7:347. doi:10.3389/fpls.2016.00347
Anjum NA, Sofo A, Scopa A, Roychoudhury A, Gill SS, Iqbal M, Lukatkin AS, Pereira E, Duarte AC, Ahmad I (2015) Lipids and proteins-major targets of oxidative modifications in abiotic stressed plants. Environ Sci Pollut Res 22(6):4099–4121
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL, Gomez-Gomez L, Boller T, Ausubel FM, Sheen J (2002) MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415(6875):977–983
Ban Q, Liu G, Wang Y (2011) A DREB gene from Limonium bicolor mediates molecular and physiological responses to copper stress in transgenic tobacco. J Plant Physiol 168(5):449–458
Benedict C, Skinner JS, Meng R, Chang Y, Bhalerao R, Huner NPA, Finn CE, Chen THH, Hurry V (2006) The CBF1-dependent low temperature signalling pathway, regulon and increase in freeze tolerance are conserved in Populus spp. Plant Cell Environ 29(7):1259–1272
Bouaziz D, Jbir R, Charfeddine S, Saidi MN, Gargouri-Bouzid R (2015) The StDREB1 transcription factor is involved in oxidative stress response and enhances tolerance to salt stress. Plant Cell Tissue Organ Cult 121(1):237–248
Boyce JM, Knight H, Deyholos M, Openshaw MR, Galbraith DW, Warren G, Knight MR (2003) The sfr6 mutant of Arabidopsis is defective in transcriptional activation via CBF/DREB1 and DREB2 and shows sensitivity to osmotic stress. Plant J 34(4):395–406
Chen C, Chen Z (2000) Isolation and characterization of two pathogen- and salicylic acid-induced genes encoding WRKY DNA-binding proteins from tobacco. Plant Mol Biol 42(2):387–396
Chen S, Polle A (2010) Salinity tolerance of Populus. Plant Biol 12(2):317–333
Chen H, Liu L, Wang L, Wang S, Cheng X (2016) VrDREB2A, a DREB-binding transcription factor from Vigna radiata, increased drought and high-salt tolerance in transgenic Arabidopsis thaliana. J Plant Res 129(2):263–273
Chu Y, Huang Q, Zhang B, Ding C, Su X (2014) Expression and molecular evolution of two DREB1 genes in black poplar (Populus nigra). PLoS ONE 9(6):e98334. doi:10.1371/journal.pone.0098334
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33(4):751–763
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179(4):945–963
Gao S, Ouyang C, Wang S, Xu Y, Tang L, Chen F (2008) Effects of salt stress on growth, antioxidant enzyme and phenylalanine ammonia-lyase activities in Jatropha curcas L. seedlings. Plant Soil Environ 54(9):374–381
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48(12):909–930
Golldack D, Luking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30(8):1383–1391
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51(1):463–499
Jin CL (2011) Establishment of genetic transformation system of Populus ussuriensis Kom. and transferred by LbDREB. Northeast Forestry University (Masters Dissertation)
Jube S, Borthakur D (2009) Development of an Agrobacterium-mediated transformation protocol for the tree-legume Leucaena leucocephala using immature zygotic embryos. Plant Cell Tissue Organ Cult 96(3):325–333
Lee SY, Boon NJ, Webb AAR, Tanaka RJ (2016) Synergistic activation of RD29A via integration of salinity stress and abscisic acid in Arabidopsis thaliana. Plant Cell Physiol 57(10):2147–2160
Li C, Yue J, Wu X, Xu C, Yu J (2014) An ABA-responsive DRE-binding protein gene from Setaria italica, SiARDP, the target gene of SiAREB, plays a critical role under drought stress. J Exp Bot 65(18):5415–5427
Li Y, Song Y, Xu B, Xie J, Zhang D (2017) Poplar CBF1 functions specifically in an integrated cold regulatory network. Tree Physiol 37(1):98–115
Liao X, Guo X, Wang Q, Wang Y, Zhao D, Yao L, Wang S, Liu G, Li T (2016) Overexpression of MsDREB6.2 results in cytokinin-deficient developmental phenotypes and enhances drought tolerance in transgenic apple plants. Plant J 89(3):510–526
Liu J, Zhu JK (1997) Proline accumulation and salt-stress-induced gene expression in a salt-hypersensitive mutant of Arabidopsis. Plant Physiol 114(2):591–596
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10(8):1391–1406
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–∆∆C T method. Methods 25(4):402–408
Mittler R, Poulos TL (2007) Ascorbate Peroxidase. In: Smirnoff N (ed) Antioxidants and reactive oxygen species in plants. Blackwell Publishing, Oxford, pp 87–100
Miyazaki Y, Abe H, Takase T, Kobayashi M, Kiyosue T (2015) Overexpression of LOV KELCH protein 2 confers dehydration tolerance and is associated with enhanced expression of dehydration-inducible genes in Arabidopsis thaliana. Plant Cell Rep 34(5):843–852
Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819(2):86–96
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25(2):239–250
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Nakano T (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140(2):411–432
Nietsch J, Brügmann J, Becker D, Fladung M (2017) Old methods rediscovered: application and improvement of two direct transformation methods to hybrid poplar (Populus tremula × P. alba). Plant Cell Tiss Organ Cult 130(1):183–196
Nole-Wilson S, Krizek BA (2000) DNA binding properties of the Arabidopsis floral development protein AINTEGUMENTA. Nucleic Acids Res 28(21):4076–4082
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, Park CY, Jeong JC, Moon BC, Lee JH, Yoon HW, Lee SH, Chung WS, Lim CO, Lee SY, Hong JC, Cho MJ (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135(4):2150–2161
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17(6):369–381
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15(5):247–258
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290(3):998–1009
Sazegari S, Niazi A, Ahmadi FS (2015) A study on the regulatory network with promoter analysis for Arabidopsis DREB-genes. Bioinformation 11(2):101–106
Shabala S, Wu H, Bose J (2015) Salt stress sensing and early signalling events in plant roots: current knowledge and hypothesis. Plant Sci 241:109–119
Shavrukov Y (2013) Salt stress or salt shock: which genes are we studying? J Exp Bot 64(1):119–127
Singh KB, Foley RC, Ate-S ON, A Nchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5(5):430–436
Su XH, Huang QJ, Zhang XH, Zhang QW, Wang B, Yao SZ (2001) Gene resource research on Populus ussuriensis Kom. in China. For Res 14(5):472–478
Sumithra K, Jutur PP, Carmel BD, Reddy AR (2006) Salinity-induced changes in two cultivars of Vigna radiata: responses of antioxidative and proline metabolism. Plant Growth Regul 50(1):11–22
Taulavuori E, Hellstrom EK, Taulavuori K, Laine K (2001) Comparison of two methods used to analyse lipid peroxidation from Vaccinium myrtillus (L.) during snow removal, reacclimation and cold acclimation. J Exp Bot 52(365):2375–2380
Tepfer D (1984) Transformation of several species of higher plants by Agrobacterium rhizogenes: sexual transmission of the transformed genotype and phenotype. Cell 37(3):959–967
Tian QQ, Chen JH, Wang D, Wang HL, Liu C, Wang S, Xia XL, Yin WL (2017) Overexpression of a Populus euphratica CBF4 gene in poplar confers tolerance to multiple stresses. Plant Cell Tissue Organ Cult 128(2):391–407
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97(21):11632–11637
Valvekens D, Montagu MV, Van Lijsebettens M (1988) Agrobacterium tumefaciens-mediated transformation of Arabidopsis thaliana root explants by using kanamycin selection. Proc Natl Acad Sci USA 85(15):5536–5540
Vinocur B, Altman A (2005) Recent advances in engineering plant tolerance to abiotic stress: achievements and limitations. Curr Opin Biotechnol 16(2):123–132
Wang RG, Chen SL, Liu LY, Hao ZY, Wong HJ, Li H et al (2005) Genotypic differences in antioxidative ability and salt tolerance of three poplars under salt stress. J Beijing For Univ 27(3):46–52
Wang HL, Tao JJ, He LG, Zhao YJ, Xu M (2009) cDNA cloning and expression analysis of a Poncirus trifoliata CBF gene. Biol Plant 53(4):625–630
Wang Z, Liu J, Guo H, He X, Wu W, Du J, Zhang Z, An X (2014) Characterization of two highly similar CBF/DREB1-like genes, PhCBF4a and PhCBF4b, in Populus hopeiensis. Plant Physiol Bioch 83:107–116
Wang G, Zhang S, Ma X, Wang Y, Kong F, Meng Q (2016) A stress-associated NAC transcription factor (SlNAC35) from tomato plays a positive role in biotic and abiotic stresses. Physiol Plant 158(1):45–64
Wei T, Deng K, Gao Y, Liu Y, Yang M, Zhang L, Zheng X, Wang C, Song W, Chen C, Zhang Y (2016) Arabidopsis DREB1B in transgenic Salvia miltiorrhiza increased tolerance to drought stress without stunting growth. Plant Physiol Bioch 104:17–28
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yevtushenko DP, Misra S (2010) Efficient Agrobacterium-mediated transformation of commercial hybrid poplar Populus nigra L. × P. maximowiczii A. Henry. Plant Cell Rep 29(3):211–221
Zhang X, Liu X, Wu L, Yu G, Wang X, Ma H (2015) The SsDREB transcription tactor from the succulent halophyte Suaeda salsa enhances abiotic stress tolerance in transgenic tobacco. Int J Genom. doi:10.1155/2015/875497
Zhang YN, Wang Y, Sa G, Zhang YH, Deng JY, Deng SR, Wang MJ, Zhang HL, Yao J, Ma XY, Zhao R, Zhou XY, Lu CF, Lin SZ, Chen SL (2017) Populus euphratica J3 mediates root K+/Na+ homeostasis by activating plasma membrane H+-ATPase in transgenic Arabidopsis under NaCl salinity. Plant Cell Tissue Organ Cult 131(1):75–88
Zhao H, Wang S, Chen S, Jiang J, Liu G (2015) Phylogenetic and stress-responsive expression analysis of 20 WRKY genes in Populus simonii × Populus nigra. Gene 565(1):130–139
Zhao H, Jiang J, Li K, Liu G (2017) Populus simonii × Populus nigra WRKY70 is involved in salt stress and leaf blight disease responses. Tree Physiol 37(6):827–844
Zhou Z, Li YL (2010) Expression pattern of PtCBF5, a CBF homologue gene encoding transcription activator in Populus tomentosa. Sci Silv Sin 46(4):58–63
Zhou M, Ma J, Zhao Y, Wei Y, Tang Y, Wu Y (2012) Improvement of drought and salt tolerance in Arabidopsis and Lotus corniculatus by overexpression of a novel DREB transcription factor from Populus euphratica. Gene 506(1):10–17
Zhu J (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53(1):247–273
Zong J, Li X, Zhou Y, Wang F, Wang N, Dong Y, Yuan Y, Chen H, Liu X, Yao N, Li H (2016) The AaDREB1 transcription factor from the cold-tolerant plant Adonis amurensis enhances abiotic stress tolerance in transgenic plant. Int J Mol Sci 17(4):611
Acknowledgements
This work was supported by the National High-Tech Research and Development Program of China, 863 Program (Key Stress-responsive Genes Identification and Molecular Breeding Technology Research in Woody Plants, Grant No. 2013AA102701).
Author information
Authors and Affiliations
Contributions
HZ and KL conceived and designed the experiments. HZ, ML, YJ, JX and JJ performed the experiments. HZ and XZ analyzed the data. KL contributed reagents/materials/analysis tools. HZ wrote the paper and XZ critical read the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Paula M. Pijut.
Rights and permissions
About this article
Cite this article
Zhao, H., Zhao, X., Li, M. et al. Ectopic expression of Limonium bicolor (Bag.) Kuntze DREB (LbDREB) results in enhanced salt stress tolerance of transgenic Populus ussuriensis Kom. Plant Cell Tiss Organ Cult 132, 123–136 (2018). https://doi.org/10.1007/s11240-017-1317-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-017-1317-1