Abstract
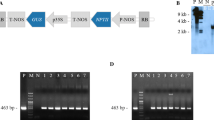
In transgenic plants, multiple T-DNA copies and cytosine methylation usually correlate with a loss of transgene expression while methylation of promoter is associated with transgene silencing. Here, six independent GFP transgenic ‘Bingtang’ sweet orange (Citrus sinensis (L.) Osb.) plants were analyzed with copy number, T-DNA repeat, cytosine methylation and transgene expression. The fluorescence of transgenic plants was normal and bright except that the transgenic B4 plant showed mottled fluorescence. Five of the six transgenic plants had multiple transgene copies and three transgenic plants had direct repeat sequences. The degrees of cytosine methylation within GFP and CaMV 35S promoter region were higher in multiple copies than in single copy transgenic plant B6. Cytosine methylation of nptII in multiple copies transgenic plants except B4 was slightly greater than single copy transgenic plant B6. Furthermore, transgenic B4 plant with mottled GFP expression had the highest degree of methylation of CG cytosine pattern. The transcript level of GFP and nptII was lower in three transgenic plants (B1, B2 and B3) with direct repeat, as revealed by real-time RT–PCR analysis. These results suggest that multiple T-DNA copies and methylation of CaMV 35S promoter affect transgene expression levels but do not cause gene silencing in transgenic sweet orange.







Similar content being viewed by others
Abbreviations
- GFP:
-
Green fluorescent protein
- nptII:
-
Neomycin phosphotransferase II
- PCR:
-
Polymerase chain reaction
- CaMV:
-
Cauliflower mosaic virus
- RB:
-
Right border
- LB:
-
Left border
References
Ahuja MR (2009) Transgene stability and dispersal in forest trees. Trees 23:1125–1135
Brodersen P, Voinnet O (2006) The diversity of RNA silencing pathways in plants. Trends Genet 22:269–278
Brunner AM, Li J, Di Fazio SP, Shevchenko O, Montgomery BE, Mohamed R, Wei H, Ma C, Elias AA, Van Wormer K, Strauss SH (2007) Genetic containment of forest plantations. Tree Genet Genome 3:75–100
Butaye KMJ, Cammue BPA, Delaure SL, De Bolle MFC (2005) Approaches to minimize variation of transgene expression in plants. Mol Breed 16:79–91
Carthew RW, Sontheimer EJ (2009) Origins and mechanisms of miRNAs and siRNAs. Cell 136:642–655
Cervera M, Pina JA, Juarez J, Navarro L, Peña L (2000) A broad exploration of a transgenic population of citrus: stability of gene expression and phenotype. Theor Appl Genet 100:670–677
Chawla R, Ariza Nieto M, Wilson AJ, Moore SK, Srivastava V (2006) Transgene expression produced by biolistic-mediated, site-specific gene integration is consistently inherited by the subsequent generations. Plant Biotechnol J 4:209–218
Cheng YJ, Guo WW, Yi HL, Pang XM, Deng XX (2003) An efficient protocol for genomic DNA extraction from Citrus species. Plant Mol Biol Rep 21:177a–177g
Diéguez MJ, Vaucheret H, Paszkowski J, Mittelsten SO (1998) Cytosine methylation at CG and CNG sites is not a prerequisite for the initiation of transcriptional gene silencing in plants, but it is required for its maintenance. Mol Gen Genet 259:207–215
Domínguez A, Fagoaga C, Navarro L, Moreno P, Peña L (2002) Regeneration of transgenic citrus plants under non selective conditions results in high-frequency recovery of plants with silenced transgenes. Mol Gen Genet 267:544–556
Domínguez A, Cervera M, Pérez RM, Romero J, Fagoaga C, Cubero J, López MM, Juárez JA, Navarro L, Peña L (2004) Characterisation of regenerants obtained under selective conditions after Agrobacterium-mediated transformation of citrus explants reveals production of silenced and chimeric plants at unexpected high frequencies. Mol Breed 14:171–183
Duan YX, Liu X, Fan J, Li DL, Wu RC, Guo WW (2007) Multiple shoot induction from seedling epicotyls and transgenic citrus plant regeneration containing the green fluorescent protein gene. Bot Stud 48:165–171
Duan YX, Fan J, Guo WW (2010) Regeneration and characterization of transgenic kumquat plants containing the Arabidopsis APETALA1 gene. Plant Cell Tiss Organ Cult 100:273–281
Dutt M, Grosser JW (2009) Evaluation of parameters affecting Agrobacterium-mediated transformation of citrus. Plant Cell Tiss Organ Cult 98:331–340
Eamens A, Wang MB, Smith NA, Waterhouse PM (2008) RNA silencing in plants: yesterday, today, and tomorrow. Plant Physiol 147:456–468
Fojtová M, Bleys A, Bedřichová J, Van Houdt H, Křížová K, Depicker A, Kovařík A (2006) The trans-silencing capacity of invertedly repeated transgenes depends on their epigenetic state in tobacco. Nucleic Acids Res 34:2280–2293
Gambino G, Chitarra W, Maghuly F, Laimer M, Boccacci P, Marinoni DT, Gribaudo I (2009) Characterization of T-DNA insertions in transgenic grapevines obtained by Agrobacterium-mediated transformation. Mol Breed 24:305–320
Gambino G, Perrone I, Carra A, ChitarraW Boccacci P, Marinoni DT, Barberis M, Maghuly F, Laimer M, Gribaudo I (2010) Transgene silencing in grapevines transformed with GFLV resistance genes: analysis of variable expression of transgene, siRNAs production and cytosine methylation. Transgenic Res 19:17–27
Goll MG, Anderson R, Stainier DYR, Spradling AC, Halpern ME (2009) Transcriptional silencing and reactivation in transgenic Zebrafish. Genetics 182:747–755
Gruntman E, Qi YJ, Slotkin RK, Roeder T, Martienssen RA, Sachidanandam R (2008) Kismeth: Analyzer of plant methylation states through bisulfite sequencing. BMC Bioinformatics 9:371
Hawkins S, Leple JC, Cornu D, Jouanin L, Pilate G (2003) Stability of transgene expression in poplar: a model forest tree species. Ann For Sci 60:427–438
Hsieh J, Fire A (2000) Recognition and silencing of repeated DNA. Annu Rev Genet 34:187–204
Klimaszewska K, Lachance D, Bernier-Cardou M, Rutledge RG (2003) Transgene integration patterns and expression levels in transgenic tissue lines of Picea mariana, P. glauca and P. abies. Plant Cell Rep 21:1080–1087
Kooter JM, Matzke MA, Meyer P (1999) Listening to the silent genes: transgene silencing, gene regulation and pathogen control. Trends Plant Sci 4:340–347
Krayer von Krauss MP, Kaiser M, Almaas V, van der Sluijs J, Kloprogge P (2008) Diagnosing and prioritizing uncertainties according to their relevance for policy: the case of transgene silencing. Sci Total Environ 390:23–34
Kumar S, Fladung M (2001) Gene stability in transgenic aspen (Populus) II. Molecular characterization of variable expression of transgene in wild and hybrid aspen. Planta 213:731–740
Lechtenberg B, Schubert D, Forsbach A, Gils M, Schmidt R (2003) Neither inverted repeat T-DNA configurations nor arrangements of tandemly repeated transgenes are sufficient to trigger transgene silencing. Plant J 34:507–517
Li JY, Brunner AM, Meilan R, Strauss SH (2009) Stability of transgenes in trees: expression of two reporter genes in poplar over three field seasons. Tree Physiol 29:299–312
Liu Q, Xu J, Liu YZ, Zhao XL, Deng XX, Guo LL, Gu JQ (2007) A novel bud mutation that confers abnormal patterns of lycopene accumulation in sweet orange fruit (Citrus sinensis L. Osbeck). J Exp Bot 58:4161–4171
Mette MF, Aufsatz W, van der Winden J, Matzke MA, Matzke AJ (2000) Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO J 19:5194–5201
Mishiba K, Nishihara M, Nakatsuka T, Abe Y, Hirano H, Yokoi T, Kikuchi A, Yamamura S (2005) Consistent transcriptional silencing of 35S-driven transgenes in gentian. Plant J 44:541–556
Morel JB, Mourrain P, Béclin C, Vaucheret H (2000) DNA methylation and chromatin structure affect transcriptional and post-transcriptional transgene silencing in Arabidopsis. Curr Biol 10:1591–1594
Omar AA, Dekkers MGH, Graham JH, Grosser JW (2008) Estimation of transgene copy number in transformed citrus plants by quantitative multiplex real-time PCR. Biotechnol Prog 24:1241–1248
Sijen T, Vijn I, Rebocho A, van Blokland R, Roelofs D, Mol JNM, Kooter JM (2001) Transcriptional and posttranscriptional gene silencing are mechanistically related. Curr Biol 11:436–440
Skarn M, Eike MC, Meza TJ, Mercy IS, Jakobsen KS, Aalen RB (2006) An inverted repeat transgene with a structure that cannot generate double-stranded RNA, suffers silencing independent of DNA methylation. Transgenic Res 15:489–500
Southern E (2006) Southern blotting. Nat Protoc 1:518–525
Stam M, Mol JNM, Kooter JM (1997) The silence of genes in transgenic plants. Ann Bot 79:3–12
Tan B, Li DL, Xu SX, Fan GE, Fan J, Guo WW (2009) Highly efficient transformation of GFP and KNU genes into precocious trifoliate orange (Poncirus trifoliata [L.] Raf), a potential model genotype for functional genomics studies in Citrus. Tree Genet Genomes 5:529–537
Tang W, Newton RJ, Weidner DA (2007) Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J Exp Bot 58:545–554
Vaucheret H, Fagard M (2001) Transcriptional gene silencing in plants: targets, inducers and regulators. Trends Genet 17:29–35
Wang MB, Waterhouse PM (2000) High-efficiency silencing of a β-glucuronidase gene in rice is correlated with repetitive transgene structure but is independent of DNA methylation. Plant Mol Biol 43:67–82
Xu SX, Cai XD, Tan B, Guo WW (2011) Comparison of expression of three different sub-cellular targeted GFPs in transgenic Valencia sweet orange by confocal laser scanning microscopy. Plant Cell Tiss Organ Cult 104:199–207
Acknowledgments
This research was financially supported by the National 863 program of China (No. 2011AA100205), and the National NSF of China (No. 30921002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fan, J., Liu, X., Xu, SX. et al. T-DNA direct repeat and 35S promoter methylation affect transgene expression but do not cause silencing in transgenic sweet orange. Plant Cell Tiss Organ Cult 107, 225–232 (2011). https://doi.org/10.1007/s11240-011-9973-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-011-9973-z