Abstract
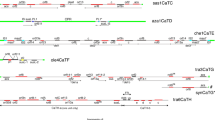
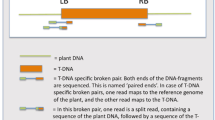
T-DNA integration patterns in 49 transgenic grapevines produced via Agrobacterium-mediated transformation were analyzed. Inverse PCR (iPCR) was performed to identify T-DNA/plant junctions. Sequence comparison revealed several deletions in the T-DNA right border (RB) and left border (LB), and filler DNA and duplications or deletions of grapevine DNA at the T-DNA insertion loci. In 20 T-DNA/grapevine genome junctions microsimilarities were found associated with the joining points and in all grapevine lines microsimilarities were present near the breaking points along the 30 bases of T-DNA adjacent to the two borders. Analysis of target site preferences of T-DNA insertions indicated a non-random distribution of the T-DNA, with a bias toward the intron regions of the grapevine genes. Compositional analysis of grapevine DNA around the T-DNA insertion sites revealed an inverse relationship between the CG and AT-skews and AT rich sequences present at 300–500 bp upstream the insertion points, near the RB of the T-DNA. PCR assays showed that vector backbone sequences were integrated in 28.6% of the transgenic plants analyzed and multiple T-DNAs frequently integrated at the same position in the plant genome, resulting in the formation of tandem and inverted repeats.






Similar content being viewed by others
References
Alonso JM, Stepanova AN, Leisse TJ et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657. doi:10.1126/science.1086391
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
An G, Ebert PR, Mitra A, Ha SB (1988) Binary vectors. In: Gelvin SB, Schilperoort RA (eds) Plant molecular biology manual. Kluwer, Dordrecht, pp A3/1–A3/19
An S, Park S, Jeong D et al (2003) Generation and analysis of end sequence database for T-DNA tagging lines in rice. Plant Physiol 133:2040–2047. doi:10.1104/pp.103.030478
Brukner I, Sanchez R, Suck D, Pongor S (1995) Sequence-dependent bending propensity of DNA as revealed by DNaseI: parameters for trinucleotides. EMBO J 14:1812–1818
Brunaud V, Balzergue S, Dubreucq B et al (2002) T-DNA integration into the Arabidopsis genome depends on sequences of pre-insertion sites. EMBO Rep 3:1152–1157. doi:10.1093/embo-reports/kvf237
Chenna R, Sugawara H, Koike T et al (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31:3497–3500. doi:10.1093/nar/gkg500
De Buck S, De Wilde C, Van Montagu M, Depieker A (2000) T-DNA vector backbone sequences are frequently integrated into the genome of transgenic plants obtained by Agrobacterium-mediated transformation. Mol Breed 6:459–468. doi:10.1023/A:1026575524345
De Neve M, De Buck S, Jacobs A et al (1997) T-DNA integration patterns in co-transformed plant cells suggest that T-DNA repeats originate from co-integration of separate T-DNAs. Plant J 11:15–29. doi:10.1046/j.1365-313X.1997.11010015.x
Durremberger F, Crameri A, Hohn B, Koukolikova-Nicola Z (1989) Covalently bound VirD2 protein of Agrobacterium tumefaciens protects the T-DNA from exonucleolytic degradation. Proc Natl Acad Sci USA 86:9154–9158. doi:10.1073/pnas.86.23.9154
Forsbach A, Schubert D, Lechtenberg B et al (2003) A comprehensive characterization of single-copy T-DNA insertions in the Arabidopsis thaliana genome. Plant Mol Biol 52:161–176. doi:10.1023/A:1023929630687
Fu D, Amand PC, Xiao Y et al (2006) Characterization of T-DNA integration in creeping bentgrass. Plant Sci 170:225–237. doi:10.1016/j.plantsci.2005.08.026
Gambino G, Gribaudo I, Leopold S et al (2005) Molecular characterization of grapevine plants transformed with GFLV resistance genes: I. Plant Cell Rep 24:655–662. doi:10.1007/s00299-005-0006-4
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “Gene-Jockeying” Tool. Microbiol Mol Biol Rev 67:16–37. doi:10.1128/MMBR.67.1.16-37.2003
Gölles R, da Câmara Machado A, Tsolova V et al (1998) Transformation of somatic embryos of Vitis sp. with different constructs containing nucleotide sequences from nepovirus coat protein genes. Acta Hortic 447:265–270
Iglesias VA, Moscone EA, Neuhuberya F et al (1997) Molecular and cytogenetic analyses of stably and unstably expressed transgene loci in tobacco. Plant Cell 9:1251–1264
Jaillon O, Aury JM, Noel B et al (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–468. doi:10.1038/nature06148
Kim S, Lee J, Jun S et al (2003) Transgene structures in T-DNA-inserted rice plants. Plant Mol Biol 52:761–773. doi:10.1023/A:1025093101021
Kim SI, Veena, Gelvin SB (2007) Genome-wide analysis of Agrobacterium T-DNA integration sites in the Arabidopsis genome generated under non-selective conditions. Plant J 51:779–791. doi:10.1111/j.1365-313X.2007.03183.x
Kononov M, Bassuner B, Gelvin SB (1997) Integration of T-DNA binary vector “backbone” sequences into the tobacco genome: evidence for multiple complex patterns of integration. Plant J 11:945–976. doi:10.1046/j.1365-313X.1997.11050945.x
Kumar S, Fladung M (2002) Transgene integration in aspen: structures of integration sites and mechanism of T-DNA integration. Plant J 31:543–551. doi:10.1046/j.1365-313X.2002.01368.x
Lobry JR (1996) Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol 13:660–665
Maghuly F, Leopold S, Machado A et al (2006) Molecular characterization of grapevine plants transformed with GFLV resistance genes: II. Plant Cell Rep 25:546–553. doi:10.1007/s00299-005-0087-0
Maghuly F, Machado A, Leopold S et al (2007) Long-term stability of marker gene expression in Prunus subhirtella: a model fruit tree species. J Biotechnol 127:310–321. doi:10.1016/j.jbiotec.2006.06.016
Matzke MA, Mette MF, Matzke AJM (2000) Transgene silencing by the host genome defense: implications for the evolution of epigenetic control mechanisms in plants and vertebrates. Plant Mol Biol 43:401–415. doi:10.1023/A:1006484806925
Meza TJ, Stangeland B, Mercy IS et al (2002) Analyses of single copy Arabidopsis T-DNA transformed lines show that the presence of vector backbone sequences, short inverted repeats and DNA methylation is not sufficient or necessary for the induction of trandgene silencing. Nucleic Acids Res 30:4556–4566. doi:10.1093/nar/gkf568
Müller AE, Atkinson RG, Sandoval RB, Jorgensen RA (2007) Microhomologies between T-DNA ends and target sites often occur in inverted orientation and may be responsible for the high frequency of T-DNA-associated inversions. Plant Cell Rep 26:617–630. doi:10.1007/s00299-006-0266-7
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications os an inverse polymerase chain reaction. Genetics 120:621–623
Ooms G, Bakker A, Molendijk L et al (1982) T-DNA organization in homogeneus and heterogeneous octopyne-type crown gall tissues of Nicotina tabacum. Cellular 30:589–597. doi:10.1016/0092-8674(82)90255-0
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Schneeberger RG, Zhang K, Tatarinova T et al (2005) Agrobacterium T-DNA integration in Arabidopsis is correlated with DNA sequence compositions that occur frequently in gene promoter regions. Funct Integr Genomics 5:240–253. doi:10.1007/s10142-005-0138-1
Szabados L, Kovács I, Oberschall A et al (2002) Distribution of 1,000 sequenced T-DNA tags in the Arabidopsis genome. Plant J 32:233–242. doi:10.1046/j.1365-313X.2002.01417.x
Thomas MR, Matsumoto S, Cain P, Scott NS (1993) Repetitive DNA of grapevine: classes present and sequences suitable for cultivar identification. Theor Appl Genet 86:173–180
Tinland B (1996) The integration of T-DNA into plant genomes. Trends Plant Sci 1:178–184. doi:10.1016/1360-1385(96)10020-0
Tzfira T, Li J, Lacroix B, Citovsky V (2004) Agrobacterium T-DNA integration: molecules and models. Trends Genet 20:375–383. doi:10.1016/j.tig.2004.06.004
Vain P, Afolabi AS, Worland B, Snape JW (2003) Transgene behaviour in populations of rice plants transformed using a new dual binary vector system: pGreen/pSoup. Theor Appl Genet 107:210–217. doi:10.1007/s00122-003-1255-7
Velasco R, Zharkikh A, Troggio M et al (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2:e1326. doi:10.1371/journal.pone.0001326
Wenck A, Czako M, Kanevsky I, Marton L (1997) Frequent colinear long transfer of DNA inclusive of the whole binary vector during Agrobacterium-mediated transformation. Plant Mol Biol 34:913–922. doi:10.1023/A:1005849303333
Windels P, De Buck S, Van Bockstaele E et al (2003) T-DNA integration in Arabidopsis chromosomes. Presence and origin of filler DNA sequences. Plant Physiol 133:2061–2068. doi:10.1104/pp.103.027532
Wu H, Sparks CA, Jones HD (2006) Characterisation of T-DNA loci and vector backbone sequences in transgenic wheat produced by Agrobacterium-mediated transformation. Mol Breed 18:195–208. doi:10.1007/s11032-006-9027-0
Yin Z, Wang GL (2000) Evidence of multiple complex patterns of T-DNA integration into the rice genome. Theor Appl Genet 100:461–470. doi:10.1007/s001220050060
Yusibov VM, Steck TR, Gupta V, Gelvin SB (1994) Association of single-stranded transferred DNA from Agrobacterium tumefaciens with tobacco cells. Proc Natl Acad Sci USA 91:2994–2998. doi:10.1073/pnas.91.8.2994
Zhai W, Chen C, Zhu X et al (2004) Analysis of T-DNA-Xa21 loci and bacterial blight resistance effects of the transgene Xa21 in transgenic rice. Theor Appl Genet 109:534–542. doi:10.1007/s00122-004-1670-4
Zhang J, Guo D, Chang Y et al (2007) Non-random distribution of T-DNA insertions at various levels of the genome hierarchy as revealed by analyzing 13 804 T-DNA flanking sequences from an enhancer-trap mutant library. Plant J 49:947–959. doi:10.1111/j.1365-313X.2006.03001.x
Zhang J, Cai L, Cheng J et al (2008) Transgene integration and organization in Cotton (Gossypium hirsutum L.) genome. Transgenic Res 17:293–306. doi:10.1007/s11248-007-9101-3
Zhu Q-H, Ramm K, Eamens AL, Dennis ES, Upadhyaya NM (2006) Transgene structures suggest that multiple mechanisms are involved in T-DNA integration in plants. Plant Sci 171:308–322. doi:10.1016/j.plantsci.2006.03.019
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gambino, G., Chitarra, W., Maghuly, F. et al. Characterization of T-DNA insertions in transgenic grapevines obtained by Agrobacterium-mediated transformation. Mol Breeding 24, 305–320 (2009). https://doi.org/10.1007/s11032-009-9293-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-009-9293-8