Abstract
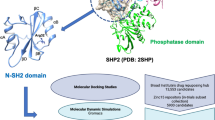
Rheumatoid arthritis (RA) is an autoimmune disorder that causes chronic inflammation with periodic bursts of activity in multiple synovial joints which lead to irreversible damage of cartilage and bone. Although several drugs that reduce inflammation are used for the treatment of RA, they are often associated with side effects. Therefore, the development or identification of a drug with no side effects or reduced side effects is desirable. Protein arginine deiminases (PADs), a set of key enzymes to trigger autoimmune response necessary for the development of RA, can be targeted for the treatment of RA. In the present study, we had developed a pharmacophore model for PAD type 4 (PAD4) protein comprising single aromatic and three hydrogen acceptor groups. Pharmacophore-based virtual screening upon ZINC database mapped several hits which were subsequently reduced by molecular docking with PAD4 protein structure. The best-scoring two ligands (Zinc_00525911 and Zinc_01225171) selected based on docking energy, pharmacophore fitness, and topology among the hits were further validated using molecular dynamics simulation for 10 ns. These two ZINC hits established interactions with key amino acid residues of PAD4 including H-bonds with Arg 372, Arg 374, Asp 350, and His 471 residues. These prioritized hits can be further tested in the in vitro and in vivo models of RA.









Similar content being viewed by others
References
Rheumatoid Arthritis (2018) National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), Maryland. https://www.niams.nih.gov/health-topics/rheumatoid-arthritis. Accessed 10 May 2018
Scott DL, Wolfe F, Huizinga TWJ (2010) Rheumatoid arthritis. Lancet (London, England) 376:1094–1108. https://doi.org/10.1016/S0140-6736(10)60826-4
Gibofsky A (2012) Comparative effectiveness of current treatments for rheumatoid arthritis. Am J Manag Care 18:S303–S314
Venables PJW (2018) Patient education: Rheumatoid arthritis treatment (Beyond the Basics) UpToDate Inc., Waltham. https://www.uptodate.com/contents/rheumatoid-arthritis-treatment-beyond-the-basics Accessed 10 May 2018
Scott DL (2012) Biologics-based therapy for the treatment of rheumatoid arthritis. Clin Pharmacol Ther 91:30–43. https://doi.org/10.1038/clpt.2011.278
Mohanan S, Cherrington BD, Horibata S et al (2012) Potential role of peptidylarginine deiminase enzymes and protein citrullination in cancer pathogenesis. Biochem Res Int 2012:1–11. https://doi.org/10.1155/2012/895343
Mangat P, Wegner N, Venables PJ, Potempa J (2010) Bacterial and human peptidylarginine deiminases: targets for inhibiting the autoimmune response in rheumatoid arthritis? Arthritis Res Ther 12:209. https://doi.org/10.1186/ar3000
Guerrin M, Ishigami A, Méchin MC et al (2003) cDNA cloning, gene organization and expression analysis of human peptidylarginine deiminase type I. Biochem J 370:167–174. https://doi.org/10.1042/BJ20020870
Chirivi RGS, van Rosmalen JWG, Jenniskens GJ et al (2013) Citrullination: a target for disease intervention in multiple sclerosis and other inflammatory diseases? J Clin Cell Immunol 04:1–8. https://doi.org/10.4172/2155-9899.1000146
Ishigami A, Ohsawa T, Asaga H et al (2002) Human peptidylarginine deiminase type II: molecular cloning, gene organization, and expression in human skin. Arch Biochem Biophys 407:25–31
Suzuki A, Yamada R, Chang X et al (2003) Functional haplotypes of PADI4, encoding citrullinating enzyme peptidylarginine deiminase 4, are associated with rheumatoid arthritis. Nat Genet 34:395–402. https://doi.org/10.1038/ng1206
Arita K, Hashimoto H, Shimizu T et al (2004) Structural basis for Ca2+-induced activation of human PAD4. Nat Struct Mol Biol 11:777–783. https://doi.org/10.1038/nsmb799
Teo CY, Shave S, Chor ALT et al (2012) Discovery of a new class of inhibitors for the protein arginine deiminase type 4 (PAD4) by structure-based virtual screening. BMC Bioinformatics 13(Suppl 17):S4. https://doi.org/10.1186/1471-2105-13-S17-S4
Korb O, Stützle T, Exner TE (2009) Empirical scoring functions for advanced protein−ligand docking with PLANTS. J Chem Inf Model 49:84–96. https://doi.org/10.1021/ci800298z
Suzuki A, Yamada R, Yamamoto K (2007) Citrullination by peptidylarginine deiminase in rheumatoid arthritis. Ann N Y Acad Sci 1108:323–339
Baka Z, György B, Géher P et al (2012) Citrullination under physiological and pathological conditions. Joint Bone Spine 79:431–436. https://doi.org/10.1016/j.jbspin.2012.01.008
Eldridge MD, Murray CW, Auton TR et al (1997) Empirical scoring functions: I. the development of a fast empirical scoring function to estimate the binding affinity of ligands in receptor complexes. J Comput Aided Mol Des 11:425–445. https://doi.org/10.1023/A:1007996124545
Nakashima K, Arai S, Suzuki A et al (2013) PAD4 regulates proliferation of multipotent haematopoietic cells by controlling c-myc expression. Nat Commun 4:1836. https://doi.org/10.1038/ncomms2862
Wei Y, Liu R, Liu C et al (2017) Identification of novel PAD4 inhibitors based on a pharmacophore model derived from transition state coordinates. J Mol Graph Model 72:88–95. https://doi.org/10.1016/J.JMGM.2016.11.016
Rahman MB, Chor AL, Salleh AB et al (2013) Ligand-based virtual screening for the discovery of inhibitors for protein arginine deiminase type 4 (PAD4). J Postgenomics Drug Biomark Dev 03. https://doi.org/10.4172/2153-0769.1000118
Sussman JL, Lin D, Jiang J et al (1998) Protein Data Bank (PDB): database of three-dimensional structural information of biological macromolecules. Acta Crystallogr Sect D Biol Crystallogr 54:1078–1084. https://doi.org/10.1107/S0907444998009378
Guex N, Peitsch MC (1997) SWISS - MODEL and the Swiss - PdbViewer : an environment for comparative protein modeling. IS 21:14–2723. https://doi.org/10.1002/elps.1150181505
Kumar SP, Jasrai YT, Pandya HA (2016) Applications of receptor- and ligand-based models in inverse docking experiments: recognition of dihydrofolate reductase using 7,8-Dialkyl- 1,3-Diaminopyrrolo[3,2-f]Quinazolines. Curr Comput Aided Drug Des 12:15–28
Kim S, Thiessen PA, Bolton EE et al (2016) PubChem substance and compound databases. Nucleic Acids Res 44:D1202–D1213. https://doi.org/10.1093/nar/gkv951
Marvin Sketch version 6.3.0 (2014) ChemAxon LLC, Budapest. https://chemaxon.com/company. Accessed 03 Mar 2018
VLife MDS version 4.3 (2008) VLife Sciences Technologies Pvt. Ltd, Pune. http://www.vlifesciences.com/. Accessed 25 Mar 2018
Kumar SP, Rawal RM, Pandya HA, Jasrai YT (2016) Qualitative and quantitative pharmacophore-similarity assessment of anthranilamide-based factor Xa inhibitors: applications on similar molecules with identical biological endpoints. J Recept Signal Transduction 36:189–206
Ingale KB, Bhatia MS (2012) Identification of structural features for 4-Methyl-3-(6-[phenyl methylene] amino} Pyridine-3-yl)-2h Chromen-2-one derivatives as clotting factor XA inhibitors. Med Chem (Los Angeles) 8:299–307
Mareddy J, Suresh N, Kumar CG et al (2017) 1, 2, 3-Triazole-nimesulide hybrid: their design, synthesis and evaluation as potential anticancer agents. Bioorg Med Chem Lett 27:518–523
Deuflhard P (1974) A modified Newton method for the solution of ill-conditioned systems of nonlinear equations with application to multiple shooting. Numer Math 22:289–315
Voet A, Qing X, Lee XY et al (2014) Pharmacophore modeling: advances, limitations, and current utility in drug discovery. J Recept Ligand Channel Res 7:81. https://doi.org/10.2147/JRLCR.S46843
Koes DR, Camacho CJ (2012) ZINCPharmer: pharmacophore search of the ZINC database. Nucleic Acids Res 40:W409–W414. https://doi.org/10.1093/nar/gks378
Discovery Studio Visualizer version 4.0 (2005) Accelrys Software Inc., San Diego. http://www.3dsbiovia.com/. Accessed 25 Mar 2018
Berendsen HJC, Postma JPM, van Gunsteren WF et al (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690. https://doi.org/10.1063/1.448118
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N*log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. https://doi.org/10.1063/1.464397
Liu RH, Meng JL (2003) MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Yi chuan Hered 25:317–321
Ferreira L, dos Santos R, Oliva G, Andricopulo A (2015) Molecular docking and structure-based drug design strategies. Molecules 20:13384–13421. https://doi.org/10.3390/molecules200713384
Kontoyiannis DP, Lewis RE (2004) Toward more effective antifungal therapy: the prospects of combination therapy. Br J Haematol 126:165–175. https://doi.org/10.1111/j.1365-2141.2004.05007.x
Funding
This study is financially supported by the Department of Science and Technology, New Delhi as Innovation in Science Pursuit for Inspired Research (INSPIRE) Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 3858 kb)
Rights and permissions
About this article
Cite this article
Soni, M.N., Kumar, S.P., Johar SR, K. et al. In silico targeting PAD4 for the treatment of rheumatoid arthritis. Struct Chem 30, 1323–1334 (2019). https://doi.org/10.1007/s11224-018-1263-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1263-5