Abstract
While there has been much computational work on the effect of intervention measures, such as vaccination or quarantine, the influence of social distancing on the epidemics’ outbursts is not well understood. We present a realistic, analytically solvable, framework for COVID-19 dynamics in the presence of social distancing measures. The model is a generalization of the compartmental SEIR model that accounts for the effects of these measures. We derive a closed-form mathematical expressions for the time dependence of epidemiological observables, in particular, the detected cases and fatalities. These analytical solutions indicate simple quantitative relations between the model variables and epidemiological observables, which give insights into cause-effect connections that underlie the outburst dynamics but are obscured in more standard (numerical) approaches. While the obtained results and conclusions are based on the study of the COVID-19 pandemic, the presented analysis has general applicability to infection outbursts. Our findings are particularly important in the emergence of new pandemics when effective pharmaceutical treatments are unavailable, and one must rely on well-timed and appropriately chosen social mitigation measures.






Similar content being viewed by others
References
Liu, X., Zheng, X., Balachandran, B.: COVID-19: data-driven dynamics, statistical and distributed delay models, and observations. Nonlinear Dyn. 101, 1527–1543 (2020). https://doi.org/10.1007/s11071-020-05863-5
Mondal, J., Khajanchi, S.: Mathematical modeling and optimal intervention strategies of the COVID-19 outbreak. Nonlinear Dyn. (2022). https://doi.org/10.1007/s11071-022-07235-7
Zhou, B., Jiang, D., Dai, Y., Hayat, T.: Stationary distribution and density function expression for a stochastic SIQRS epidemic model with temporary immunity. Nonlinear Dyn. 105, 931–955 (2021). https://doi.org/10.1007/s11071-020-06151-y
Han, C., Li, M., Haihambo, N., Babuna, P., Liu, Q., Zhao, X., Jaeger, C., Li, Y., Yang, S.: Mechanisms of recurrent outbreak of COVID-19: a model-based study. Nonlinear Dyn. 106, 1169–1185 (2021). https://doi.org/10.1007/s11071-021-06371-w
Al-Darabsah, I.: Threshold dynamics of a time-delayed epidemic model for continuous imperfect-vaccine with a generalized nonmonotone incidence rate. Nonlinear Dyn. 101, 1281–1300 (2020). https://doi.org/10.1007/s11071-020-05825-x
Zhai, S., Luo, G., Huang, T., Wang, X., Tao, J., Zhou, P.: Vaccination control of an epidemic model with time delay and its application to COVID-19. Nonlinear Dyn. 106, 1279–1292 (2021). https://doi.org/10.1007/s11071-021-06533-w
Raza, A., Rafiq, M., Awrejcewicz, J., Ahmed, N., Mohsin, M.: Dynamical analysis of coronavirus disease with crowding effect, and vaccination: a study of third strain. Nonlinear Dyn. (2022). https://doi.org/10.1007/s11071-021-07108-5
Das, P., Upadhyay, R.K., Misra, A.K., Rihan, F.A., Das, P., Ghosh, D.: Mathematical model of COVID-19 with comorbidity and controlling using non-pharmaceutical interventions and vaccination. Nonlinear Dyn. 106, 1213–1227 (2021). https://doi.org/10.1007/s11071-021-06517-w
Guan, G., Guo, Z.: Stability behavior of a two-susceptibility SHIR epidemic model with time delay in complex networks. Nonlinear Dyn. 106, 1083–1110 (2021). https://doi.org/10.1007/s11071-021-06804-6
Das, P., Nadim, S.S., Das, S., Das, P.: Dynamics of COVID-19 transmission with comorbidity: a data driven modelling based approach. Nonlinear Dyn. 106, 1197–1211 (2021). https://doi.org/10.1007/s11071-021-06324-3
Babajanyan, S.G., Cheong, K.H.: Age-structured SIR model and resource growth dynamics: a COVID-19 study. Nonlinear Dyn. 104, 2853–2864 (2021). https://doi.org/10.1007/s11071-021-06384-5
Vilar, J.M.G., Saiz, L.: Reliably quantifying the evolving worldwide dynamic state of the COVID-19 outbreak from death records, clinical parametrization, and demographic data. Sci. Rep. 11, 19952 (2021). https://doi.org/10.1038/s41598-021-99273-1
Vilar, J.M.G., Saiz, L.: Ascertaining the initiation of epidemic resurgences: an application to the COVID-19 second surges in Europe and the Northeast United States. R Soc. Open Sci. 8, 210773 (2021). https://doi.org/10.1098/rsos.210773
Xu, L., Zhang, H., Xu, H., Yang, H., Zhang, L., Zhang, W., Gu, F., Lan, X.: The coSIR model predicts effective strategies to limit the spread of SARS-CoV-2 variants with low severity and high transmissibility. Nonlinear Dyn. 105, 2757–2773 (2021). https://doi.org/10.1007/s11071-021-06705-8
Khyar, O., Allali, K.: Global dynamics of a multi-strain SEIR epidemic model with general incidence rates: application to COVID-19 pandemic. Nonlinear Dyn. 102, 489–509 (2020). https://doi.org/10.1007/s11071-020-05929-4
Ahmed, N., Elsonbaty, A., Raza, A., Rafiq, M., Adel, W.: Numerical simulation and stability analysis of a novel reaction-diffusion COVID-19 model. Nonlinear Dyn. 106, 1293–1310 (2021). https://doi.org/10.1007/s11071-021-06623-9
Di Giamberardino, P., Iacoviello, D., Papa, F., Sinisgalli, C.: A data-driven model of the COVID-19 spread among interconnected populations: epidemiological and mobility aspects following the lockdown in Italy. Nonlinear Dyn. 106, 1239–1266 (2021). https://doi.org/10.1007/s11071-021-06840-2
Jia, Q., Li, J., Lin, H., Tian, F., Zhu, G.: The spatiotemporal transmission dynamics of COVID-19 among multiple regions: a modeling study in Chinese provinces. Nonlinear Dyn. 107, 1313–1327 (2022). https://doi.org/10.1007/s11071-021-07001-1
Sun, G.Q., Wang, S.F., Li, M.T., Li, L., Zhang, J., Zhang, W., Jin, Z., Feng, G.L.: Transmission dynamics of COVID-19 in Wuhan, China: effects of lockdown and medical resources. Nonlinear Dyn. 101, 1981–1993 (2020). https://doi.org/10.1007/s11071-020-05770-9
Zlatić, V., Barjašić, I., Kadović, A., Štefančić, H., Gabrielli, A.: Bi-stability of SUDR+K model of epidemics and test kits applied to COVID-19. Nonlinear Dyn. 101, 1635–1642 (2020). https://doi.org/10.1007/s11071-020-05888-w
Pacheco, P.M.C.L., Savi, M.A., Savi, P.V.: COVID-19 dynamics considering the influence of hospital infrastructure: an investigation into Brazilian scenarios. Nonlinear Dyn. 106, 1325–1346 (2021). https://doi.org/10.1007/s11071-021-06323-4
Li, T., Xiao, Y.: Complex dynamics of an epidemic model with saturated media coverage and recovery. Nonlinear Dyn. (2022). https://doi.org/10.1007/s11071-021-07096-6
Rai, R.K., Khajanchi, S., Tiwari, P.K., Venturino, E., Misra, A.K.: Impact of social media advertisements on the transmission dynamics of COVID-19 pandemic in India. J. Appl. Math. Comput. 68, 19–44 (2022). https://doi.org/10.1007/s12190-021-01507-y
WHO report. URL https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation
Diekmann, O., Heesterbeek, H., Britton, T.: Mathematical tools for understanding infectious disease dynamics. Princeton University Press, Princeton, NJ, USA (2012)
Martcheva, M.: An introduction to mathematical epidemiology. Springer, Berlin, Germany (2015)
Keeling, M.J., Rohani, P.: Modeling infectious diseases in humans and animals. Princeton University Press, Princeton, NJ, USA (2011)
Alagoz, O., Sethi, A.K., Patterson, B.W., Churpek, M., Safdar, N.: Effect of Timing of and Adherence to Social Distancing Measures on COVID-19 Burden in the United States. Ann. Intern. Med. 174(1), 50–57 (2021). https://doi.org/10.7326/m20-4096
Oraby, T., Tyshenko, M.G., Maldonado, J.C., Vatcheva, K., Elsaadany, S., Alali, W.Q., Longenecker, J.C., Al-Zoughool, M.: Modeling the effect of lockdown timing as a COVID-19 control measure in countries with differing social contacts. Sci. Rep. 11, 3354 (2021)
Dressler, D.D.: Social Distancing’s Effects on COVID-19: Roles of Timing and Adherence. NEJM Journal Watch. https://www.jwatch.org/na52712/2020/11/03/social-distancings-effects-covid-19-roles-timing-and (2020)
Nande, A., Adlam, B., Sheen, J., Levy, M.Z., Hill, A.L.: Dynamics of COVID-19 under social distancing measures are driven by transmission network structure. PLoS Comput. Biol. 17(2), e1008684 (2021). https://doi.org/10.1371/journal.pcbi.1008684
Benjamin, M.F., Brockmann, D.: Effective containment explains subexponential growth in recent confirmed COVID-19 cases in China. Science 368, 742–746 (2020). https://doi.org/10.1126/science.abb4557
He, S., Peng, Y., Sun, K.: SEIR modeling of the COVID-19 and its dynamics. Nonlinear Dyn. 101, 1667–1680 (2020). https://doi.org/10.1007/s11071-020-05743-y
Hametner, C., Kozek, M., Boehler, L., Wasserburger, A., Du, Z.P., Kölbl, R., Bergmann, M., Bachleitner-Hofmann, T., Jakubek, S.: Estimation of exogenous drivers to predict COVID-19 pandemic using a method from nonlinear control theory. Nonlinear Dyn. 106, 1111–1125 (2021). https://doi.org/10.1007/s11071-021-06811-7
Carvalho, T., Cristiano, R., Rodrigues, D.S., Tonon, D.J.: Global analysis of a piecewise smooth epidemiological model of COVID-19. Nonlinear Dyn. 105, 3763–3773 (2021). https://doi.org/10.1007/s11071-021-06801-9
Saha, S., Samanta, G.P., Nieto, J.J.: Epidemic model of COVID-19 outbreak by inducing behavioural response in population. Nonlinear Dyn. 102, 455–487 (2020). https://doi.org/10.1007/s11071-020-05896-w
Huang, J., Qi, G.: Effects of control measures on the dynamics of COVID-19 and double-peak behavior in Spain. Nonlinear Dyn. 101, 1889–1899 (2020). https://doi.org/10.1007/s11071-020-05901-2
Saikia, D., Bora, K., Bora, M.P.: COVID-19 outbreak in India: an SEIR model-based analysis. Nonlinear Dyn. 104, 4727–4751 (2021). https://doi.org/10.1007/s11071-021-06536-7
Kwuimy, C.A.K., Nazari, F., Jiao, X., Rohani, P., Nataraj, C.: Nonlinear dynamic analysis of an epidemiological model for COVID-19 including public behavior and government action. Nonlinear Dyn. 101, 1545–1559 (2020). https://doi.org/10.1007/s11071-020-05815-z
Kermack, W.O., McKendrick, A.G.: A contribution to the mathematical theory of epidemics. In: Proceedings of the Royal Society of London. Series A, Mathematical and Physical Sciences 115, 700–721 (1927)
Friedman, A., Kao, C.Y.: Mathematical modeling of biological processes. Springer, Berlin, Germany (2014)
Newman, M.E.J.: The spread of epidemic disease on networks. Phys. Rev. E 66, 016128 (2002). https://doi.org/10.1103/PhysRevE.66.016128
Schutz, G.M., Brandaut, M., Trimper, S.: Exact solution of a stochastic susceptible-infectious-recovered model. Phys. Rev. E 78, 061132 (2008). https://doi.org/10.1103/PhysRevE.78.061132
Awawdeh, F., Adawi, A., Mustafa, Z.: Solutions of the SIR models of epidemics using HAM. Chaos Solitons Fractals 42, 3047–3052 (2009). https://doi.org/10.1016/j.chaos.2009.04.012
Khan, H., Mohapatra, R.N., Vajravelu, K., Liao, S.J.: The explicit series solution of SIR and SIS epidemic models. Appl. Math. Comput. 38, 653–669 (2009). https://doi.org/10.1016/j.amc.2009.05.051
Harko, T., Lobo, F.S.N., Mak, M.K.: Exact analytical solutions of the susceptible-infected-recovered (SIR) epidemic model and of the SIR model with equal death and birth rates. Appl. Math. Comput. 236, 184 (2014). https://doi.org/10.1016/j.amc.2014.03.030
Barlow, N.S., Weinstein, S.J.: Accurate closed-form solution of the SIR epidemic model. Physica D 408, 132540 (2020). https://doi.org/10.1016/j.physd.2020.132540
Carvalho, A.M., Goncalves, S.: An analytical solution for the Kermack–McKendrick model. Physica A 566, 125659 (2021). https://doi.org/10.1016/j.physa.2020.125659
Sadurni, E., Luna-Acosta, G.: Exactly solvable SIR models, their extensions and their application to sensitive pandemic forecasting. Nonlinear Dyn. 103, 2955–2971 (2021). https://doi.org/10.1007/s11071-021-06248-y
Kroeger, M., Schlickeiser, R.: Analytical solution of the SIR-model for the temporal evolution of epidemics. Part A: time-independent reproduction factor. J. Phys. A 53, 505601 (2020). https://doi.org/10.1088/1751-8121/abc65d
Schlickeiser, R., Kroeger, M.: Analytical solution of the SIR-model for the temporal evolution of epidemics. Part B: semi-time case. J. Phys. A 54, 175601 (2021). https://doi.org/10.1088/1751-8121/abed66
Weinstein, S.J., Holland, M.S., Rogers, K.E., Barlow, N.S.: Analytic solution of the SEIR epidemic model via asymptotic approximant. Physica D: Nonlinear Phenomena 411, 132633 (2020). https://doi.org/10.1016/j.physd.2020.132633
Piovella, N.: Analytical solution of SEIR model describing the free spread of the COVID-19 pandemic. Chaos Solitons Fractals 140, 110243 (2020). https://doi.org/10.1016/j.chaos.2020.110243
Miller, J.C.: A note on the derivation of epidemic final sizes. Bull. Math. Biol. 74, 2125–2141 (2012). https://doi.org/10.1007/s11538-012-9749-6
Turkyilmazoglu, M.: Explicit formulae for the peak time of an epidemic from the SIR model. Physica D: Nonlinear Phenomena 422, 132902 (2021). https://doi.org/10.1016/j.physd.2021.132902
Kroeger, M., Turkyilmazoglu, M., Schlickeiser, R.: Explicit formulae for the peak time of an epidemic from the SIR model. Which approximant to use? Physica D: Nonlinear Phenomena 425, 132981 (2021). https://doi.org/10.1016/j.physd.2021.132981
Djordjevic, M., Rodic, A., Salom, I., Zigic, D., Milicevic, O., Ilic, B., Djordjevic, M.: A systems biology approach to COVID-19 progression in population. Adv. Protein. Chem. Struct. Biol. 127, 291–314 (2021). https://doi.org/10.1016/bs.apcsb.2021.03.003
Djordjevic, M., Djordjevic, M., Ilic, B., Stojku, S., Salom, I.: Understanding infection progression under strong control measures through universal COVID-19 growth signatures. Glob. Chall. 5, 2000101 (2021). https://doi.org/10.1002/gch2.202170051
Li, R., Pei, S., Chen, B., Song, Y., Zhang, T., Yang, W., Shaman, J.: Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 368, 489–493 (2020). https://doi.org/10.1126/science.abb3221
He, X., Lau, E.H.Y., Wu, P., Deng, X., Wang, J., Hao, X., Lau, Y.C., Wong, J.Y., Guan, Y., Tan, X., Mo, X., Chen, Y., Liao, B., Chen, W., Hu, F., Zhang, Q., Zhong, M., Wu, Y., Zhao, L., Zhang, F., Cowling, B.J., Li, F., Leung, G.M.: Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat. Med. 26, 672–675 (2020). https://doi.org/10.1038/s41591-020-0869-5
Bar-On, Y.M., Flamholz, A.I., Phillips, R., Milo, R.: Science forum: SARS-CoV-2 (COVID-19) by the numbers. eLife 9, e57309 (2020)
Phillips, R., Kondev, J., Theriot, J., Garcia, H.: Physical biology of the cell. Garland Science, New York, NY, USA (2012)
Worldometer: COVID-19 coronavirus pandemic. https://www.worldometers.info/coronavirus/ (2020)
Bowman, F.: Introduction to bessel functions. Dover, New York NY, USA (1958)
Weisstein, E.W.: CRC concise encyclopedia of mathematics. Chapman & Hall, CRC Press, Boca Raton, FL, USA (2003)
Abramowitz, M., Stegun, I.A.: Handbook of Mathematical Functions. Dover Publications, New York, NY, USA (1972)
Markovic, S., Rodic, A., Salom, I., Milicevic, O., Djordjevic, M., Djordjevic, M.: COVID-19 severity determinants inferred through ecological and epidemiological modeling. One Health 13, 100355 (2021). https://doi.org/10.1016/j.onehlt.2021.100355
Zwillinger, D.: CRC standard mathematical tables and formulae. CRC Press, Boca Raton, FL, USA (1995)
Sarkar, K., Khajanchi, S., Nieto, J.J.: Modeling and forecasting the COVID-19 pandemic in India. Chaos, Solitons Fractals 139, 110049 (2020)
Salom, I., Rodic, A., Milicevic, O., Zigic, D., Djordjevic, M., Djordjevic, M.: Effects of demographic and weather parameters on COVID-19 basic reproduction number. Front. Ecol. Evol. 8, 617841 (2021). https://doi.org/10.3389/fevo.2020.617841
Milicevic, O., Salom, I., Rodic, A., Markovic, S., Tumbas, M., Zigic, D., Djordjevic, M., Djordjevic, M.: PM2.5 as a major predictor of COVID-19 basic reproduction number in the USA. Environ. Res. 201, 111526 (2021). https://doi.org/10.1016/j.envres.2021.111526
Djordjevic, M., Salom, I., Markovic, S., Rodic, A., Milicevic, O., Djordjevic, M.: Inferring the main drivers of SARS-CoV-2 global transmissibility by feature selection methods. GeoHealth 5(9), e2021GH000432 (2021). https://doi.org/10.1029/2021GH000432
Tadic, B., Melnik, R.: Modeling latent infection transmissions through biosocial stochastic dynamics. PloS One 15(10), e0241163 (2020). https://doi.org/10.1371/journal.pone.0241163
Tadic, B., Melnik, R.: Microscopic dynamics modeling unravels the role of asymptomatic virus carriers in SARS-CoV-2 epidemics at the interplay between biological and social factors. Comput. Biol. Med. 133, 104422 (2021). https://doi.org/10.1016/j.compbiomed.2021.104422
Frieden, T.R., Buissonniere, M., McClelland, A.: The world must prepare now for the next pandemic. BMJ Glob. Health 6, e005184 (2021). https://doi.org/10.1136/bmjgh-2021-005184
Boyce, W.E., DiPrima, R.C., Rosatone, L.: Elementary differential equations and boundary value problems. Wiley, Hoboken, NJ, USA (2012)
Kreyszig, E.: Advanced engineering mathematics. Wiley, Hoboken, NJ, USA (2006)
Funding
This work was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia.
Author information
Authors and Affiliations
Contributions
Magdalena Djordjevic and Marko Djordjevic contributed to the study conception and design. Analytical calculations were performed by Magdalena Djordjevic, Bojana Ilic and Igor Salom. Numerical analysis was performed by Bojana Ilic, Magdalena Djordjevic and Marko Djordjevic. The first draft of the manuscript was written by Bojana Ilic and Igor Salom with the help of Magdalena Djordjevic and Marko Djordjevic. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Data Availability
All data used in the study are publically available through Ref. [62].
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A Appendix
A Appendix
1.1 A.1 Analytical derivation of the number of fatalities
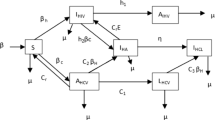
To assess the dynamics of COVID-19 infection, we employed the mechanistic model, defined by equations (1)-(8). In Ref. [58], we have already obtained the number of infectious individuals as a function of time (9).
Throughout the derivations, we will distinguish two-time regions: I) \(t\le t_0\) and II) \(t> t_0\), and will denote the variables in these two regions accordingly.
First, we concentrate on deriving the expression for the time evolution of the number of fatalities. To this end, we start from Eqs. (7)-(8), which lead to a single equation (10). In region I, this second-order inhomogeneous differential equation, after taking into account the first term of Eq. (9) (i.e., \(I_I(t)=I_0 \textrm{e}^{\lambda _+ t}\)), reduces to:
If we assume \(F_I(t=0)=0\) and \(F'_I(t=0)=0\), the time-dependent fatalities for \(t\le t_0\) are given by:
In region II, for simplicity we shift \(t-t_0 \rightarrow t\), and take into account that the expression for infectious now reads \(I_{II}(t) =I_0\textrm{e}^{\lambda _+ t_0} \textrm{e}^{-\frac{\gamma +\epsilon \delta + \sigma }{2}t} \textrm{K}\big (\frac{\gamma +\epsilon \delta - \sigma }{\alpha }, \frac{2\sqrt{\textrm{e}^{-\alpha t} \beta \sigma }}{\alpha } \big )/\) \(\textrm{K}\big (\frac{\gamma +\epsilon \delta - \sigma }{\alpha }, \frac{2\sqrt{\beta \sigma }}{\alpha }\big )\) (the second term of Eq. (9)). Since \(I_{II}(t)\) has this complex form, the further derivations are quite demanding. By substituting \(t\rightarrow y=\frac{2 \sqrt{\textrm{e}^{-\alpha t} \beta \sigma }}{\alpha }\), it can easily be verified that Eq. (10) is in region II reduced to the well-known second-order inhomogeneous Cauchy–Euler equation [76, 77] \(a x^2 y'' +b x y'+cy =g(x)\). After dividing thus obtained differential equation by \(y^2\), we obtain
where \(C=\frac{4 I_0 \epsilon \delta m}{\alpha ^2} \big (2\sqrt{\beta \sigma }/{\alpha } \big )^{-\frac{\gamma +\epsilon \delta +\sigma }{\alpha }} \textrm{e}^{\lambda _+ t_0}/\textrm{K} \big (\frac{\gamma +\epsilon \delta -\sigma }{\alpha }, \frac{2\sqrt{\beta \sigma }}{\alpha } \big )\).
Next, we apply the standard procedure for solving inhomogeneous differential equation given by Eq. (30): the full solution equals homogeneous plus particular solution \(F_{II}(y)=F_{II,h}(y) + F_{II,p}(y)\). It is straightforward to show that auxiliary/characteristic equation yields the following simple form of homogeneous solution \(F_{II,h}(y)=C_1 + C_2 y^{\frac{2 (h+m)}{\alpha }}\) (where linearly independent solutions are \(F_{II,1}(y)=1\) and \(F_{II,2}(y)= y^{\frac{2 (h+m)}{\alpha }}\)), while for obtaining particular solution \(F_{II,p}(y)\) we used the Lagrange’s method of variation of parameters [76]. More precisely, we assume that \(F_{II,p}(y)= C_1(y)+ C_2(y) y^{\frac{2 (h+m)}{\alpha }}\), where again linearly independent solutions of the homogeneous equation are employed. The unknown functions \(C_i\) (\(i=1,2\)) of variable y are sought for via the standard procedure
where f(y) denotes the right-hand side of Eq. (30), while Wronskian [76, 77] is given by \(W=F_{II,1} F'_{II,2} -F_{II,2} F'_{II,1}=\frac{2 (h+m)}{\alpha } y^{\frac{2 (h+m)}{\alpha }-1}\). This leads to:
In spite that the above form of the result will be more useful in what follows, we nevertheless provide also its full-fledged form:
where \(C_0=\frac{ I_0 \epsilon \delta }{\alpha } \frac{m}{h+m} \textrm{e}^{\lambda _+ t_0}/{\textrm{K}\big (\frac{\gamma +\epsilon \delta -\sigma }{\alpha }, \frac{2\sqrt{\beta \sigma }}{\alpha } \big )}\), while
\(_p\tilde{\textrm{F}}_q(a_1,a_2,...a_p;b_1,b_2,...b_q;z)\) denotes regularized generalized hypergeometric function [65]. The general solution of Eq. (30), when returned to variable t, has a form:
where the only unknown parameters are \(C_1\) and \(C_2\). In order to determine these constants we use the following boundary conditions: \(F_{II}(0)=F_I(t_0)\) and \(F'_{II}(0)=F'_I(t_0)\). After some cumbersome calculation steps we obtain the following expressions:
Now, if we use the definition of regularized generalized hypergeometric functions, the above expressions can be significantly simplified and reduced to the form similar to Eq. (32). Namely [65]
where \((a_1)_0=1\), while for \(k\geqslant 1\)
Let us first concentrate on \(C_1\), where from Eq. (35) it follows that \(z= \beta \sigma /{\alpha ^2}\). By denoting \(a_1=(\gamma +\epsilon \delta )/ \alpha \) and \(a_2=\sigma / \alpha \), it is straightforward to infer that the expression in the square brackets of Eq. (35) can be rewritten in a form:
After multiplying the right-hand side by \(z^{\frac{a_1 + a_2}{2}}/z^{\frac{a_1 +a_2}{2}}\) we obtain:
where we made use of the fact that \(\Gamma (n+1)=n \Gamma (n)\) and \(\Gamma (a) (a)_k =\Gamma (a+k)\) (see Eq. (38)). To further simplify Eq. (40), we adopt the following notation \({\mathcal {J}}_1=\sum _{k=0}^{\infty } \frac{z^{k+a_1}}{k! (k+a_1) \Gamma (k+1+a_1-a_2)}\) and differentiate it with respect to z:
As already mentioned, the main idea is to try to reduce Eq. (35) to a form similar to Eq. (32), i.e., to relate the above expression to modified Bessel functions, and the modified Bessel function of the first kind [64] is defined as:
By comparing Eqs. (41) and (42), we observe that their right-hand sides are of a similar form, if \(x/2 \rightarrow \sqrt{z}\), \(n \rightarrow a_1-a_2\), that is:
Proceeding in a similar manner in the case of the remaining term in Eq. (40) \({\mathcal {J}}_2=\sum _{k=0}^{\infty } \frac{z^{k+a_2}}{k! (k+a_2) \Gamma (k+1+a_2-a_1)}\) we arrive at:
Note, from Eq. (40), that \({\mathcal {I}}_1=z^{-\frac{a_1 +a_2}{2}} ({\mathcal {J}}_1 -{\mathcal {J}}_2)\).
By substituting integrated Eqs. (43) and (44) in Eq. (40), and thus obtained expression in Eq. (35), for \(C_1\) we finally obtain:
Note that, in obtaining the above expression the identity [66, 68] relating modified Bessel function of the first and the second kind \(\textrm{I}_{-n}(x) -\textrm{I}_n(x)=\frac{2}{\pi } \sin (n \pi ) \textrm{K}_n(x)\) was used.
The remaining constant \(C_2\) (see Eq. (36)) from fatality counts expression is simplified by applying the same procedure as in the case of \(C_1\). To avoid redundant derivations (i.e., the repetition of the above calculations), we simply outline the final expression:
with the only distinction that, in the process of simplification, parameters \(a_1\) and \(a_2\) now read \((\gamma +\epsilon \delta -h-m)/ \alpha \) and \((\sigma -h-m)/ \alpha \), respectively.
Now that all terms of fatalities count (given by Eq. (34)) are determined, we note that all constants (\(C_1, C_2, C_1(t)\) and \(C_2(t)\)) are still in their integral form. Since, for all countries that we consider it holds \(2 \sqrt{\beta \sigma }/ \alpha \gg 1\), we may further simplify these integrals by utilizing Hankel’s asymptotic expression [66]:
which holds for large \(x=2 \sqrt{\beta \sigma }/ \alpha \).
Along these lines, we rewrite Eqs. (32, 45, 46):
where we also used the same approximation given by Eq. (47) for modified Bessel function of the second kind contained in C and \(C_0\). Note that the second terms of \(C_1\) and \(C_1(t)\) on one side, and the second terms of \(C_2\) and \(C_2(t)\) on the other have the same exponents (keep in mind that, according to Eq. (34), there is still a remaining factor \(\big (2 \sqrt{\beta \sigma }/{{\alpha }} \big )^{\frac{2(h+m)}{\alpha }}\) multiplying \(C_2(t)\)).
In Eq. (48), we encounter the lower incomplete gamma function [66] \(\gamma (s,x)=\int _0^x t^{s-1} \textrm{e}^{-t} \textrm{d}t\), which we go about by utilizing [66]:
where \(\Gamma (s,x)=\int _x^{\infty } t^{s-1} \textrm{e}^{-t} \textrm{d}t\) represents the upper incomplete gamma function [66].
Finally, upon implementing thus calculated constants of Eq. (48) in Eq. (34), and by taking into account Eq. (29), we obtain the expression (16) for the general solution of Eq. (10) at the entire t region.
1.2 A.2 Analytical derivation of detected cases
Next, we concentrate on the time evolution of detected counts. To this end, we make use of Eqs. (6) and (9), i.e.
where \(C_i\) (\(i=3,4\)) stands for the constant of integration. In region I the integration of \(I_I(t)\) (the first term in Eq. (9)) is straightforward and yields \(D_I(t)= \frac{I_0 \epsilon \delta }{\lambda _+} \textrm{e}^{\lambda _+ t} + C_3\), while \(C_3\) is obtained from the initial conditions \(D_I(t=0)=D_0\equiv \frac{I_0 \epsilon \delta }{\lambda _+}\), leading to:
In region II, the integration is more demanding, due to the form of \(I_{II}(t)\) (the second term of Eq. (9)). To address this, we employ the following substitution of variable \(t-t_0 \rightarrow x=\frac{2\sqrt{\beta \sigma }}{\alpha } \textrm{e}^{-\frac{\alpha (t-t_0)}{2}}\). Thus:
Note that, as opposed to our previous notation during the derivation of \(F_{II}(t)\), now we do not make use of the substitution \(t-t_0 \rightarrow t\). Because of this, the boundary condition reads: \(D_{II}(t_0)=D_I(t_0)\), which is used for determining the integration constant \(C_4=\frac{I_0 \epsilon \delta }{\lambda _+} \textrm{e}^{\lambda _+ t_0}\). So, the expression for detected cases reads:
After performing the same algebraic manipulations of regularized generalized hypergeometric functions as in the previous Subsect. A.1, as well as applying Hankel’s approximation (47), we obtain the expression (19) for the general solution of Eq. (11) at the entire t region.
Alternatively, the same expression (19) could be obtained more straightforwardly. Namely, the expression for the number of infectious individuals (Eq. (9)) can be simplified by utilizing Eq. (47):
Now that I(t) has been determined in the desired form, the detected counts can be calculated from Eq. (11). In region I, the derivation is straightforward (and therefore omitted) and leads to Eq. (51). In region II, the integration is more demanding, due to the form of \(I_{II}(t)=I_0 \textrm{e}^{\lambda _+ t_0 +\frac{2\sqrt{\beta \sigma }}{\alpha }} \textrm{e}^{-\frac{\alpha (t-t_0)}{2}(\frac{\gamma +\epsilon \delta +\sigma }{\alpha }-\frac{1}{2})} \textrm{e}^{-\frac{2 \sqrt{\textrm{e}^{-\alpha (t-t_0)} \beta \sigma }}{\alpha }}\). To address this, again we employ the following substitution of variable \(t-t_0 \rightarrow x= \frac{2\sqrt{\beta \sigma }}{\alpha }\) \( \textrm{e}^{-\frac{\alpha (t-t_0)}{2}}\), resulting in the boundary condition \(D_{II}(t_0)=D_I(t_0)\) (which is used for determining the integration constant \(C_4\)). In a similar manner as before, we again encounter the incomplete gamma functions [66]:
The only difference compared to Eq. (48) is that now the lower boundary of integration is not zero but some positive real number. This integral is solved by applying the identity \(\int _a^b x^{s-1} \textrm{e}^{-x}\textrm{d}x =\int _0^b x^{s-1} \textrm{e}^{-x}\textrm{d}x -\int _0^a x^{s-1} e^{-x}\textrm{d}x=\Gamma (s,a)-\Gamma (s,b)\). By combining this result with Eq. (51), we finally arrive at the expression (19) for the number of detected cases.
1.3 A.3 Simplified expressions for fatalities and detected cases
In this section, we show that certain terms in the expressions for fatalities and detected cases can be neglected (when epidemiological parameters are in realistic ranges), without significant loss of predictive precision. First, we start with Eq. (16) for fatality counts. We notice that for \(t>t_0\) first two terms are much smaller than the remaining terms, due to \(2\sqrt{\beta \sigma }/ \alpha \gg 1\), and therefore can be neglected. Additionally, for the same reason all \(\Gamma (s,2\sqrt{\beta \sigma }/ \alpha )\rightarrow \Gamma (s,\infty )\) are approximately equal to zero (i.e., the gamma integral is effectively \(\int _{\infty }^{\infty }\)). Therefore, instead of Eq. (16), the following formula can be safely used in practice:
Continuing in the same manner, the simplified form of the number of detected cases is easily obtained:
We have numerically tested and confirmed that the full-fledged (given by Eq. (16)) and simplified (given by Eq. (56)) fatality curves are practically overlapping (and the same for the detected cases).
1.4 A.4 Expressions for fatalities and detected cases at saturation
We will evaluate the saturation values of fatalities and detected counts, that is, their expressions in the limit of very large t. This means that we can concentrate only on \(t>t_0\), i.e., region II, where we set \(t\rightarrow \infty \). Building on the results of the previous section, from Eq. (56) we observe that, in this limit, the second term in the square brackets can be neglected, due to \(\textrm{e}^{-(h+m)(t-t_0)}\rightarrow 0\). Likewise,
Therefore, we obtain the expressions (17) and (20) for the saturation values.
1.5 A.5 Expressions for the epidemics peak and tipping points
Other important quantities characterizing infection dynamics during the first wave are epidemics peak time and inflection (tipping and turning) points, for which we here provide analytic expressions. Namely, the epidemics peak time is the moment when infected curve reaches its maximal value (i.e., \(\textrm{d}I / \textrm{d}t =0\)), or equivalently \(\textrm{d}^2D / \textrm{d}t^2 =0\). The second derivative of Eq. (19) in region II (or equivalently Eq. (57) in the same region) yields:
In deriving the above expression, we made use of equality \(\textrm{d}\Gamma (\textrm{s},\textrm{x}) / \textrm{d}x =-x^{s-1} \textrm{e}^{-x}\), following from the definition of incomplete gamma functions [66]. Note that, in this subsection, we are interested in region II, where all relevant points lay. After equating the second derivative of \(D_{II}\) with zero, for the epidemics peak time, we obtain:
By evaluating \(\frac{\textrm{d}D}{\textrm{d}t}\) at \(t=t_{max}\), we can straightforwardly obtain the maximum of detected cases per day, given by
Along the same lines, the epidemics inflection points are defined as \(\textrm{d}^2 I / \textrm{d}t^2 =0\), or equivalently \(\textrm{d}^3D / \textrm{d}t^3 =0\):
Eq. (61) has two solutions, which correspond to the infection tipping points
The duration of the epidemic peak can then be defined as a difference between these two tipping points and is equal to:
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ilic, B., Salom, I., Djordjevic, M. et al. An analytical framework for understanding infection progression under social mitigation measures. Nonlinear Dyn 111, 22033–22053 (2023). https://doi.org/10.1007/s11071-023-08692-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11071-023-08692-4